Abstract
Introduction
Polymorphisms in the Thiopurine S-Methyltransferase (TPMT) gene are associated with decreased TPMT activity, but little is known about their impact on TPMT protein expression in the liver. This project is to conduct a genome-wide association study (GWAS) to identify single nucleotide polymorphisms (SNPs) associated with altered TPMT protein expression in human livers and to determine if demographics affect hepatic TPMT protein expression.
Methods
Human liver samples (n = 287) were genotyped using a whole genome genotyping panel and quantified for TPMT protein expression using a Data-Independent Acquisition proteomics approach.
Results and Discussion
Thirty-one SNPs were found to be associated with differential expression of TPMT protein in the human livers. Subsequent analysis, conditioning on rs1142345, a SNP associated with the TPMT*3A and TPMT*3C alleles, showed no additional independent signals. Mean TPMT expression is significantly higher in wildtype donors compared to those carrying the known TPMT alleles, including TPMT*3A, TPMT*3C, and TPMT*24 (0.107 ± 0.028 vs. 0.052 ± 0.014 pmol/mg total protein, P = 2.2 × 10−16). After removing samples carrying the known TPMT variants, European ancestry donors exhibited significantly higher expression than African ancestry donors (0.109 ± 0.026 vs. 0.090 ± 0.041 pmol/mg total protein, P = 0.020).
Conclusion
The GWAS identified 31 SNPs associated with TPMT protein expression in human livers. Hepatic TPMT protein expression was significantly lower in subjects carrying the TPMT*3A, TPMT*3C, and TPMT*24 alleles compared to non-carriers. European ancestry was associated with significantly higher hepatic TPMT protein expression than African ancestry, independent of known TPMT variants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thiopurines are used to treat hematologic malignancies and immune-mediated diseases by inhibiting DNA replication. Overexposure to these medications can result in life-threatening toxicities, mainly myelosuppression and hepatotoxicity. Thiopurine medications are metabolized by thiopurine methyltransferase (TPMT), a polymorphic enzyme primarily expressed in the liver. Many polymorphisms within TPMT have been identified that affect enzyme activity, and their frequencies vary by ethnicity [1]. Due to the consequences of overexposure to thiopurines, TPMT activity can be screened prior to initiating treatment by testing the activity in the patient’s red blood cells or testing for TPMT genotypes to determine phenotypic risk [2,3,4].
Current clinical guidelines support the use of preemptive genotyping prior to the initiation of thiopurine treatment [5, 6]. Clinical studies have shown that patients with two loss-of-function alleles of the TPMT gene are at extremely high risk for hematologic toxicities, and patients carrying one loss-of-function allele are classified as the intermediate risk group, having a 30–60% chance of developing toxicity at standard doses [7, 8]. Although genotypes contribute to the interindividual variability in TPMT activity and the toxicity of thiopurines, genotypes do not fully explain observed TPMT activity as some patients carrying wildtype TPMT alleles are classified as having intermediate TPMT activity [9].
Thiopurines are primarily metabolized in the liver. However, few studies have investigated the relationship between the expression of TPMT protein in the liver and genetic polymorphisms at the whole genome level. A meta-analysis conducted by Tamm et al. reported that TPMT activity in the liver was significantly correlated with hepatic TPMT protein levels, while mRNA from whole blood samples did not correlate with TPMT activity or protein expression in the liver [10]. As a result, our study aims to investigate the relationship between the whole genome genotypes and TPMT protein expression in human livers through a genome-wide association study (GWAS) of TPMT protein expression in a large set of human liver samples. Moreover, we determined the effect of the known TPMT alleles TPMT*3A, TPMT*3C, and TPMT*24 and patients’ demographics, including ancestry, sex, and age, on hepatic TPMT protein expression.
Materials and Methods
Materials
A total of 287 human liver samples were obtained from three different sources, including 47 from XenoTech LLC (Lenexa, KS, USA), 79 from the University of Minnesota Liver Tissue Cell Distribution System, and 161 from the Cooperative Human Tissue Network. To reduce confounding factors, all tissue samples were deemed to be healthy. Demographics for these patients are summarized in Table I. Age was reported in 270 of the 287 donors. The sex of the donor was confirmed during the quality control of the whole genome genotyping data.
TPMT Protein Quantification and GWAS
Human liver microsomes were prepared and subjected to a liquid chromatography-mass spectrometry-based proteomics analysis using a previously published label-free Data-Independent Acquisition (DIA) method [11]. Absolute expression levels of TPMT were calculated using the DIA-based total protein approach (DIA-TPA) algorithm that we developed [12].
DNA was extracted from the human liver samples and genotyped using the Illumina Multi-Ethnic Global Array (Illumina, Miami, USA) platform, which contains 1,779,819 single nucleotide polymorphisms (SNPs). The genotype imputation was conducted using the Michigan Imputation Server (https://imputationserver.sph.umich.edu). Specific thresholds for quality control and genotype imputation were previously reported by Bing et al. [13]
Association of hepatic TPMT protein expression with genome-wide SNPs was performed with PLINK 1.9 [14]. Prior to analysis, quality control was performed on the genomic data as previously outlined [15]. We applied a threshold of 0.02 to remove SNPs and patient samples that had a missing rate higher than 2%. We removed SNPs with a minor allele frequency of less than 0.01. Next, we analyzed the genotyping data for Hardy-Weinberg Equilibrium and removed genetic variants that did not pass the threshold of 10−6. The data was also checked for any individuals who had a heterozygosity rate greater than three standard deviations from the mean and any patients with a PIHAT value greater than 0.2. We performed multidimensional scaling on our dataset and overlayed it with the genotype data from the 1000 genomes database to visually identify and remove ethnic outliers, resulting in a genomic inflation factor of 1, indicating no evidence of inflation. To reduce the effect of population stratification on GWAS analysis, ancestry was imputed using the genotyping data instead of relying on patient self-reported race using a previously published method by Marees et al. [15] We set the GWAS statistically significant P value threshold at 5 × 10−8 to account for multiple comparisons. After quality control and removing ethnic outliers within the dataset, we conducted the GWAS of TPMT protein expression on 243 of the 287 liver samples using 1,685,470 genotyped and imputed markers. Conditional analysis was conducted on SNPs with the lowest P values to determine if there were any other independent signals in the GWAS.
Results
GWAS of TPMT Protein Expression in the Liver
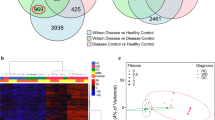
Expression of TPMT protein was quantified in all 287 patient liver samples using the DIA-TPA method. The range of absolute TPMT protein expression levels in the human liver microsomes was between 0.0039–0.1762 pmol/mg total protein, approximately a 45-fold difference between the maximum and minimum quantities. Thirty one SNPs, all located on chromosome 6, passed our genome-wide threshold (P < 5.0 × 10−8) (Fig. 1) (Table II). Further analysis was conducted conditioning on the strongest signal rs1142345, which is associated with the TPMT*3A and TPMT*3C alleles, and no other independent signal was identified.
Manhattan and QQ plots of the GWAS of TPMT protein expression in human liver microsomes. Manhattan plot (1A, left): The y-axis represents the unadjusted p value, while the x-axis represents the genomic coordinates of genotyped and imputed SNPs. QQ plot (1B, right) shows the expected -log10(P) vs. observed -log10(P) for the association of a specific genome location with TPMT expression.
Effect of TPMT Polymorphisms on Hepatic TPMT Protein Expression
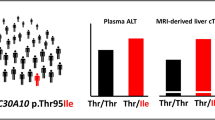
Within the dataset, there were 22 liver sample donors who were heterozygous carriers of one of documented TPMT alleles, including 18 donors with the TPMT*3A allele, 3 donors with the TPMT*3C allele, and 1 donor with the TPMT*24 allele. Using the Welch two-sample t-test, there was a significant difference (P = 6.7 × 10−14) in the mean expression of TPMT protein between the wildtype group (n = 265) and heterozygous carriers of TPMT*3A (n = 18). The mean expression in the wildtype group, 0.107 ± 0.028 pmol/mg total protein, was 1.99-fold higher than the mean expression in the TPMT*3A group, 0.054 ± 0.014 pmol/mg total protein. When combining TPMT*3A, TPMT*3C, and TPMT*24 donors into one group (n = 22), the mean expression was 0.052 ± 0.014 pmol/mg total protein. The mean expression in the wildtype group was 2.07-fold higher than the mean expression in the TPMT variant group (P = 2.2 × 10−16, Welch two-sample t-test) (Fig. 2).
Comparison of TMPT protein expression in human liver microsomes between wildtype TPMT donors and those with the heterozygous TPMT *3A, *3C, and *24 genotypes. Histogram (2A) shows the distribution of TPMT expression within the 287 HLM 287 samples. The wildtype group is represented in red, with the red dotted line representing the mean expression for this population. Alternatively, the blue bars represent the TPMT*3A/*3C/*24 heterozygous carriers, with the blue dotted line representing the mean expression for this population. The boxplots (2B) visually compare TPMT protein expression between the TPMT*3A/*3C/*24 heterozygous group and the wildtype group. **P < 0.01.
TPMT Expression by Ancestry, Age, and Sex
Exploratory analysis for differential expression of TPMT between demographic groups was conducted by removing all patients who carried a TPMT polymorphism that was known to be associated with TPMT expression (e.g., TPMT*3A, TPMT*3C, TPMT*24). This was done to eliminate the potential confounding effect of those SNPs on the relationship. Of the 34 liver donors of African ancestry, two were removed for being heterozygous carriers of TPMT*3C, and one was removed for being a heterozygous carrier of TPMT*24. Of the 229 liver donors of European ancestry, 15 were removed for being heterozygous carriers of TPMT*3A, and one was removed for being a heterozygous carrier of TPMT*3C. After removal of these donors, the assumed wildtype liver donors of European ancestry exhibited a significantly higher average TPMT expression, 0.109 ± 0.026 pmol/mg total protein, than those of African ancestry, 0.090 ± 0.041 pmol/mg total protein, P = 0.020 (Fig. 3). This difference in expression was maintained when comparing males of European ancestry versus males of African ancestry, 0.111 ± 0.027 (n = 118) vs. 0.092 ± 0.040 (n = 21) pmol/mg total protein, P = 0.046. There was not a significant difference in TPMT expression in females of European ancestry (n = 95) versus females of African ancestry, 0.106 ± 0.024 (n = 95) vs. 0.087 ± 0.046 (n = 10) pmol/mg total protein, P = 0.219. Additional summary statistics are found on Supplementary Table 1. Comparison of other ancestry populations was not possible due to the small sample size. There was no significant association between TPMT expression and age.
Discussion
In the present study, we conducted a GWAS to identify genetic variants associated with TPMT protein expression in the liver. The study revealed that 31 SNPs were significantly associated with differential TPMT expression at the genome-wide level. Further analysis conditioning on rs1142345, a SNP with the lowest P value and associated with the TPMT*3A and TPMT*3C alleles, showed no additional independent signals. TPMT*3A contains two nonsynonymous SNPs, rs1142345 and rs1800460, and is the most common loss-of-function haplotype in our GWAS dataset. The minor allele frequency of TPMT*3A in the Caucasian population is about 5%, [16] which is consistent with the frequency found in our study. TPMT*3A has been characterized by causing rapid TPMT protein degradation by ATP-dependent proteasomes. Tai et al. transfected several mutant TMPT alleles into COS-1 cells and showed that homozygous carriers of TPMT*3A had a 200-fold decrease in TPMT protein expression in comparison to wildtype expressors through Western Blot [17]. These findings have been replicated in COS-1 and COS-7 cell lines, reporting that TPMT*3A homozygotes had no detectable protein levels and enzyme activity [18, 19]. In our dataset, the average TPMT protein levels in TPMT*3A heterozygotes was approximately 50% of that in samples with wildtype TPMT, indicating that the TPMT*3A polymorphism results in almost no detectable protein expression in the human liver.
In addition to TPMT*3A, two other TPMT alleles, TPMT*3C (rs1142345) and TPMT*24 (rs6921269), were also identified in the liver samples. TPMT*3C is a loss-of-function allele that has a low minor allele frequency in the Caucasian population, around 0.2%, but is the most prevalent TPMT polymorphism in Asian and African populations at 2.5% - 5% [19,20,21,22]. TPMT*24 has a minor allele frequency of 2% in Africans and African Americans but is rare in other populations. TPMT*24 is assigned as an uncertain function allele by the Clinical Pharmacogenetics Implementation Consortium (CPIC), although an in vitro study showed that the variant significantly decreased TMPT activity [23]. One TPMT*24 heterozygote with African ancestry was identified in our liver samples. TPMT protein expression in this TPMT*24 heterozygous carrier is 0.043 pmol/mg of total protein versus the mean level of 0.109 ± 0.026 pmol/mg total protein in wildtype subjects, suggesting that, similar to TPMT*3A and *3C, TPMT*24 is also a loss-of-function allele. Of note, there were only three carriers of TPMT*3C and one carrier of TPMT*24 in the study because our samples were primarily of European ancestry. The average TPMT protein expression level in the group of TPMT*3A/*3C/*24 carriers was approximately 50% of the wildtype group.
Differences in TPMT activity and expression between races are typically attributed to the racial differences in the minor allele frequencies of TPMT genetic variants. Few studies have reported a potential difference in TPMT activity in erythrocytes between racial groups. Mcleod et al. reported a significantly lower median erythrocyte TPMT activity in self-reported American black subjects than American white subjects (14.4 versus 16.8 units/ml packed erythrocytes; P < 0.001) [24]. Genetic data was unavailable for this study, but the distribution of participants into normal, intermediate, and poor TPMT activity was similar between the racial groups. Cooper et al. reported that Afro-Caribbean patients had a significantly lower median erythrocyte TPMT activity in comparison to Caucasian and South Asian populations [25]. Similarly, this study did not include genetics analysis but rather used self-reported race. Our study is the first to report a difference in TPMT expression in human livers between European and African ancestry while accounting for TPMT polymorphisms associated with TPMT expression. We found that the average expression of TPMT protein in wildtype samples from European ancestry donors was 1.2-fold of that in wildtype samples from African ancestry donors. This relationship was maintained when comparing male African and European ancestry donors. The racial difference was not statistically significant in female donors, although the average expression for female European ancestry donors was 1.2-fold that of the female African donors. We expect that with a larger sample size, similar to that of the male donors, the difference in TPMT expression would be statistically significant. Since the TPMT polymorphisms known to affect TPMT expression have been removed from the two populations prior to the analysis, it is likely that this difference is due to environmental factors or rare genetic polymorphisms that are not part of the whole genome genotyping panel. Of note, physiologically based pharmacokinetic (PBPK) models have been developed to simulate the concentrations of thiopurines and their metabolites in plasma and tissue based on data from Caucasian populations [26]. Our racial-specific TPMT protein expression data can be used to modify existing PBPK models and more accurately simulate the pharmacokinetics of thiopurines and their metabolites in patients of African ancestry.
A potential limitation of this study is that we did not measure TPMT activity in the liver and, thus, were unable to match the activity data with TPMT expression, genotypes, sex, and age. However, previous studies have shown a strong correlation between hepatic TPMT activity and protein expression and a strong correlation between TPMT activity in erythrocytes and its expression in erythrocyte lysates [10, 17]. Notably, our study showed that some wildtype donors had lower TPMT expression than the TPMT*3A carriers, suggesting that hepatic TPMT expression can also be regulated by other factors in addition to the TPMT*3A allele. Future studies are warranted to determine the genetic and non-genetic factors regulating hepatic TPMT expression in subjects with the wildtype TPMT genotype. Furthermore, future GWAS focusing on other ethnic populations (e.g., African) where TPMT*3A is not the most prevalent loss-of-function allele may identify other genetic polymorphisms affecting TPMT expression within the liver.
Conclusion
For the first time, we quantified absolute TPMT protein expression in the liver using a DIA-TPA approach and conducted a GWAS to identify genetic variants associated with hepatic TPMT protein expression levels. The analysis identified 31 SNPs that were significantly associated with hepatic TPMT protein expression at the genome-wide level. Further analysis conditioning on rs1142345, a SNP associated with the TPMT*3A and *3C haplotypes, did not reveal additional independent signals. Expression of TPMT protein in wildtype donors was 2.07-fold higher than expression in the group of TPMT*3A, TPMT*3C, and TPMT*24 heterozygous carriers. Additionally, liver samples from European ancestry donors had a 1.2-fold higher expression of TPMT protein than the liver samples from African ancestry donors after the exclusion of samples carrying known TPMT variants.
References
Appell ML, Berg J, Duley J, Evans WE, Kennedy MA, Lennard L, Marinaki T, McLeod HL, Relling MV, Schaeffeler E, Schwab M, Weinshilboum R, Yeoh AEJ, McDonagh EM, Hebert JM, Klein TE, Coulthard SA. Nomenclature for alleles of the thiopurine methyltransferase gene. Pharmacogenet Genomics. 2013;23(4):242–8.
Dervieux T, Meyer G, Barham R, Matsutani M, Barry M, Boulieu R, Neri B, Seidman E. Liquid chromatography-tandem mass spectrometry analysis of erythrocyte thiopurine nucleotides and effect of thiopurine methyltransferase gene variants on these metabolites in patients receiving azathioprine/6-mercaptopurine therapy. Clin Chem. 2005;51(11):2074–84.
Chrzanowska M, Kuehn M, Januszkiewicz-Lewandowska D, Kurzawski M, Droździk M. Thiopurine S-methyltransferase phenotype-genotype correlation in children with acute lymphoblastic leukemia. Acta Pol Pharm. 2012;69(3):405–10.
Lennard L. Implementation of TPMT testing. Br J Clin Pharmacol. 2014;77(4):704–14.
Weitzel KW, Smith DM, Elsey AR, Duong BQ, Burkley B, Clare-Salzler M, Gong Y, Higgins TA, Kong B, Langaee T, McDonough CW, Staley BJ, Vo TT, Wake DT, Cavallari LH, Johnson JA. Implementation of standardized clinical processes for TPMT testing in a diverse multidisciplinary population: challenges and lessons learned. Clin Transl Sci. 2018;11(2):175–81.
Relling MV, Schwab M, Whirl-Carrillo M, Suarez-Kurtz G, Pui CH, Stein CM, Moyer AM, Evans WE, Klein TE, Antillon-Klussmann FG, Caudle KE, Kato M, Yeoh AEJ, Schmiegelow K, Yang JJ. Clinical Pharmacogenetics Implementation Consortium Guideline for Thiopurine Dosing Based on TPMT and NUDT15 Genotypes: 2018 Update. Clin Pharmacol Ther. 2019;2019 105(5):1095–105.
Stocco G, Cheok M, Crews K, Dervieux T, French D, Pei D, Yang W, Cheng C, Pui CH, Relling M, Evans W. Genetic polymorphism of inosine triphosphate pyrophosphatase is a determinant of Mercaptopurine metabolism and toxicity during treatment for acute lymphoblastic leukemia. Clinical Clin Pharmacol Ther. 2009;85(2):164–72.
Benkov K, Lu Y, Patel A, Rahhal R, Russell G, Teitelbaum J, f.t.N.C.o.I.B. Disease, role of Thiopurine metabolite testing and Thiopurine methyltransferase determination in pediatric IBD. J Pediatr Gastroenterol Nutr. 2013;56(3):333–40.
Coelho T, Andreoletti G, Ashton JJ, Batra A, Afzal NA, Gao Y, Williams AP, Beattie RM, Ennis S. Genes implicated in thiopurine-induced toxicity: comparing TPMT enzyme activity with clinical phenotype and exome data in a paediatric IBD cohort. Sci Rep. 2016;6(1):1–9.
Tamm R, Mägi R, Tremmel R, Winter S, Mihailov E, Smid A, Möricke A, Klein K, Schrappe M, Stanulla M, Houlston R, Weinshilboum R, Mlinarič Raščan I, Metspalu A, Milani L, Schwab M, Schaeffeler E. Polymorphic variation in TPMT is the principal determinant of TPMT phenotype: a Meta-analysis of three genome-wide association studies. Clin Pharmacol Ther. 2017;101(5):684–95.
Wang X, He B, Shi J, Li Q, Zhu H-J. Comparative proteomics analysis of human liver Microsomes and S9 fractions. Drug Metab Dispos. 2020;48(1):31–40.
He B, Shi J, Wang X, Jiang H, Zhu H-J. Label-free absolute protein quantification with data-independent acquisition. J Proteome. 2019;200:51–9.
He B, Shi J, Wang X, Jiang H, Zhu H-J. Genome-wide pQTL analysis of protein expression regulatory networks in the human liver. BMC Biol. 2020;18(1):97.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, De Bakker PIW, Daly MJ, Sham PC. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81(3):559–75.
Marees AT, De Kluiver H, Stringer S, Vorspan F, Curis E, Marie-Claire C, Derks EM. A tutorial on conducting genome-wide association studies: quality control and statistical analysis. Int J Methods Psychiatr Res. 2018;27(2):e1608.
Gurwitz D, Rodríguez-Antona C, Payne K, Newman W, Gisbert JP, De Mesa EG, Ibarreta D. Improving pharmacovigilance in Europe: TPMT genotyping and phenotyping in the UK and Spain. Eur J Hum Genet. 2009;17(8):991–8.
Tai H-L, Krynetski EY, Schuetz EG, Yanishevski Y, Evans WE. Enhanced proteolysis of thiopurine S-methyltransferase (TPMT) encoded by mutant alleles in humans (TPMT*3A, TPMT*2): mechanisms for the genetic polymorphism of TPMT activity. Proc Natl Acad Sci. 1997;94(12):6444–9.
Ujiie S, Sasaki T, Mizugaki M, Ishikawa M, Hiratsuka M. Functional characterization of 23 allelic variants of thiopurine S-methyltransferase gene (TPMT*2 – *24). Pharmacogenet Genomics. 2008;18(10):887–93.
Salavaggione OE, Wang L, Wiepert M, Yee VC, Weinshilboum RM. Thiopurine S-methyltransferase pharmacogenetics: variant allele functional and comparative genomics. Pharmacogenet Genomics. 2005;15(11):801–15.
Hon YY, Fessing MY, Pui C-H, Relling MV, Krynetski EY, Evans WE. Polymorphism of the Thiopurine S-methyltransferase gene in African-Americans. Hum Mol Genet. 1999;8(2):371–6.
Srimartpirom S, Tassaneeyakul W, Kukongviriyapan V, Tassaneeyakul W. Thiopurine S-methyltransferase genetic polymorphism in the Thai population. Br J Clin Pharmacol. 2004;58(1):66–70.
Ando M, Ando Y, Hasegawa Y, Sekido Y, Shimokata K, Horibe K. Genetic polymorphisms of thiopurine S-methyltransferase and 6-mercaptopurine toxicity in Japanese children with acute lymphoblastic leukaemia. Pharmacogenet Genomics. 2001;11(3):269–73.
Garat A, Cauffiez C, Renault N, Lo-Guidice JM, Allorge D, Chevalier D, Houdret N, Chavatte P, Loriot MA, Gala JL, Broly F. Characterisation of novel defective thiopurine S-methyltransferase allelic variants. Biochem Pharmacol. 2008;76(3):404–15.
McLeod HL, Lin J-S, Scott EP, Pui C-H, Evans WE. Thiopurine methyltransferase activity in American white subjects and black subjects. Clin Pharmacol Ther. 1994;55(1):15–20.
Cooper SC, Ford LT, Berg JD, Lewis MJ. Ethnic variation of thiopurine S-methyltransferase activity: a large, prospective population study. Pharmacogenomics. 2008;9(3):303–9.
Ogungbenro K, Aarons L. Physiologically based pharmacokinetic model for 6-mercpatopurine: exploring the role of genetic polymorphism in TPMT enzyme activity. Br J Clin Pharmacol. 2015;80(1):86–100.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 14 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Smith, L.S., Wang, X., Shi, J. et al. Genome-Wide Association Study for the Genetic Determinants of Thiopurine Methyltransferase Protein Expression in Human Livers and Racial Differences. Pharm Res 40, 2525–2531 (2023). https://doi.org/10.1007/s11095-023-03558-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11095-023-03558-1