Abstract
Alzheimer’s disease (AD) is the most common cause of dementia. Increasing evidence shows that mitochondrial DNA (mtDNA) methylation plays an essential role in many diseases related to mitochondrial dysfunction. Since mitochondrial impairment is a key feature of AD, mtDNA methylation may also contribute to AD, but few studies have addressed this issue. Methylation changes of the mitochondrial cytochrome b (CYTB) and cytochrome c oxidase II (COX II) genes in AD have not been reported. We analyzed mtDNA methylation changes of the CYTB and COX II genes in an APP/PS1 transgenic mouse model of AD using pyrosequencing. We examined mtDNA copy numbers and the levels of expression by quantitative real-time PCR. Average methylation levels of different CpG sites were ≤ 4.0%. Methylated mtDNA accounted for only a small part of the total mtDNA. We also observed hypermethylation of mitochondrial CYTB and COX II genes with decreased mtDNA copy numbers and expression in the hippocampi of APP/PS1 transgenic mice. mtDNA methylation may play an important role in AD pathology, which may open a new window for AD therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dementia is characterized by a decline in language, memory, problem solving, and other cognitive indices, which affect the ability to manage daily activities [1]. There were around 50 million cases of dementia globally in 2018, and the total number of cases is projected to reach 152 million by 2050 [2]. Alzheimer’ disease (AD) is the most common cause of dementia; it comprises 60–70% of cases. The neuropathological manifestations of AD are extracellular accumulation of insoluble amyloid-β (Aβ), which accumulates in senile plaques, and intracellular neurofibrillary tangles (NFTs), which consist of hyperphosphorylated microtubule-associated protein tau (hMAPT), along with progressive neuronal loss and cerebral atrophy [3, 4]. The pathogenesis of AD is complex and remains obscure. Increasing evidence shows that mitochondrial impairments, which include the production of reactive oxygen species (ROS), altered mitochondrial dynamics, and apoptosis are prominent features [5]. In several disorders characterized by mitochondrial dysfunction, mitoepigenetics plays an important role [6].
Mitochondria are critical for ATP synthesis [7]. They are intracellular organelles that contain ~ 16.6 kb of mitochondrial DNA (mtDNA), which is separate from the nuclear genome and is inherited in a maternal, non-Mendelian fashion [6]. Human mtDNA is a circular, double-stranded genome encoding 37 genes, including 13 structural genes that encode subunits of the electron transport chain (ETC), 22 tRNA genes, and two rRNA genes [6, 8, 9]. Human mtDNA also includes a non-coding region, the displacement loop (D-loop), which contains an origin of replication and transcriptional promoters [10]. The mtDNA-encoding subunits required for the ETC complexes include: seven subunits of complex I (ND1, ND2, ND3, ND4, ND4L, ND5 and ND6), one subunit of complex III (cytochrome b), three subunits of complex IV (cytochrome c oxidase I to III), and two subunits of complex V (ATPase 6 and 8) [11].
Mitoepigenetics is the study of modulations occurring in the mitochondria that induce heritable phenotype changes without the involvement of alternations in the mtDNA sequence [12]. Mitoepigenetics encompasses four mechanisms: mtDNA methylation/hydroxymethylation, mitochondrial nucleoid modifications, mitochondrial RNA modifications, and non-coding RNA (ncRNA) modulations during mtDNA-encoded gene translation and function [9, 12, 13]. 5-mC/5-hmC and DNA methyltransferase 1 (DNMT 1) have been reported in mitochondria [14, 15]. Human mtDNA represents < 1% of total cellular DNA, but it contains 435 CpG sites and 4747 cytosine residues at non-CpG sites, both of which have cytosines that can be methylated [10]. The methylation of mtDNA is associated with exposure to various environmental agents and endogenous metabolites [10, 16]. MtDNA methylation is linked to many diseases, including cardiovascular diseases, cancer, diabetic retinopathy, nonalcoholic steatohepatitis, obesity, Down’s syndrome, and neurodegenerative diseases [6, 12, 15,16,17,18,19,20]. Although there is a lack of direct evidence, mtDNA methylation may be crucial in many pathophysiological processes through the regulation of mitochondrial gene expression [21,22,23].
There is little research on mtDNA methylation changes in AD. Blanch et al. found that mtDNA methylation increased in the D-loop and decreased in the ND1 gene in the entorhinal cortex in brain samples with AD-related pathology [20]. They also found a dynamic pattern of D-loop methylation changes in the neocortex of amyloid precursor protein/presenilin 1 (APP/PS1) mice that was linked with AD pathology progression (3, 6, and 12 months of age) [20]. A decrease in methylation of the D-loop peripheral blood DNA was found in late-onset AD [24]. We identified a reduction in D-loop methylation and a rise in 12S rRNA gene methylation, while both mtDNA copy number and mitochondrial gene expression decreased in the hippocampi of APP/PS1 transgenic mice [25].The methylation levels of other mtDNA regions have yet to be investigated in AD.
In this study, we analyzed mtDNA methylation changes in the hippocampi of an APP/PS1 transgenic mouse model of AD. We measured the methylation levels of two mitochondrial genes associated with the ETC: cytochrome b (CYTB) and cytochrome c oxidase II (COX II). To the best of our knowledge, methylation changes in these two genes have never been studied using an AD mouse model. We used bisulfite conversion and pyrosequencing to assay DNA methylation [26]. We assessed mtDNA copy numbers and mitochondrial gene expression to identify any association with mtDNA methylation.
Materials and Methods
Animals
Nine-month-old male APP/PS1 transgenic mice and age-matched C57BL/6J mice were purchased from the Beijing HFK Bio-Technology Co. (Beijing, China). The APP/PS1 transgenic mouse expresses a chimeric mouse/human APP (Mo/HuAPP695swe: APP Swedish mutation) and a mutant human PS1 (PS1-dE9) [20]. Mice were housed in a temperature- and humidity-controlled environment on a 12 h light/dark cycle with free access to food and water. Ten mice from each group were anesthetized and sacrificed. The hippocampus of each mouse was promptly isolated, placed on ice, and stored at − 80 ℃. All procedures followed the guidelines of the Animal Ethics Committee of Shandong University (Jinan, China).
Electron Microscopy
The mitochondrial morphology of murine hippocampi were observed by electron microscopy. The isolated hippocampi were cut into 1 mm thick blocks and fixed in glutaraldehyde at 4 ℃ overnight. The blocks were postfixed in osmium for 1 h, embedded in resin, and re-sectioned into 70 nm thick slices using an ultramicrotome (Leica Microsystems, Wetzlar, Germany). The slices were stained with uranyl acetate and lead citrate. Images were captured using a Hitachi HT-7800 electron microscope operating at 80 kV (Hitachi Co., Tokyo, Japan).
DNA Extraction
DNA was extracted using the Rapid Animal Genomic DNA Isolation Kit (Sangon Biotech, Shanghai, China), following the manufacturer’s protocol. The quality of extracted murine DNA was determined by A260/280 ratios on a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). All samples showed ratios between 1.7 and 1.9.
Methylation Analysis of CYTB and COX II Genes
Extracted DNA samples were converted by sodium bisulfite using the EZ DNA Methylation-Gold™ Kit (Zymo Research, Irvine, CA, USA) according to the manufacturer’s instructions. Unmethylated cytosines were converted to uracil, while methylated cytosines remained unchanged.
Primers for PCR amplification designs were based on entries in the Gene Bank: DQ106412.1 mitochondrial genome sequence and synthesized utilizing the PyroMark Assay Design software (Version 2.0, Qiagen, Düsseldorf, Germany) by Sangon Biotech. Forward primers were biotinylated. Table 1 lists the primer sequences. Amplification of bisulfite-treated DNA was performed in a final volume of 50 μl. Each reaction mix comprised 1 μl of forward primer, 1 μl of reverse primer, 1 μl of 10 mM dNTP, 2 μl of bisulfite-converted DNA, 0.5 μl of 5 U/μl Taq DNA polymerase, 5 μl of Taq buffer, 5 μl of 25 mM MgCl2, and 34.5 μl ultrapure water. PCR was performed using the following protocol: (1) 95 ℃ for 3 min; (2) 35 cycles of (a) denaturation at 94 ℃ for 25 s, (b) annealing for 25 s at a specific temperature (60 ℃ for CYTB and 58℃ for COX II); (c) extension at 72℃ for 25 s; (3) one cycle of final extension at 72 ℃ for 5 min; (4) hold at 4 ℃ until use. Each reaction was performed in triplicate. 5 μl of PCR product was run on 2% agarose for gel electrophoresis (Fig. 1).
A representative agarose gel following PCR amplification of different bisulfite-converted DNA samples. Amplifications of the COX II gene (lanes 1 to 5) and CYTB gene (lanes 7 to 11) with 5 different bisulfite-treated DNA samples were visualized on a 2% agarose gel. The sixth lane is the DNA marker. The product length is 143 bp in COX II gene and 116 bp in CYTB gene
We measured methylation levels of six CpG sites for the CYTB gene and four CpG sites for the COX II gene, using the pyrosequencing technique previously described for the PyroMark Q96 ID machine (Qiagen) [25]. In brief, PCR products were immobilized on Streptavidin Sepharose beads (Streptavidin Sepharose HP, GE Healthcare, Chicago, IL, USA) in binding buffer for 5 min at room temperature. With the pyrosequencing vacuum prep tool (Qiagen), the biotin-labeled template/bead complexes were purified by washing with 70% ethanol for 5 s, then with denaturing buffer for 5 s, and then with washing buffer for 5–10 s. A PSQ HS 96 well plate (Qiagen) was prefilled with 40 μl of annealing buffer that contained 0.5 μM sequencing primer, and the mixture was incubated. DNA strands were denatured for 2 min at 80 °C and reannealed for 10 min at room temperature. Sequencing was carried out with the aid of PyroMark CpG software (Version1.0.11, Qiagen). Data were validated by internal controls. The methylation percentage of each CpG site was determined from the ratio of cytosine to thymidine [26].
Estimation of mtDNA Copy Number
MtDNA copy numbers were assessed by qRT-PCR using a CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad, Hercules, CA, USA). Primers for the mitochondrial CYTB gene and the β-actin gene were designed utilizing Primer Premier (Version 5.0, Premier Biosoft, Palo Alto, CA, USA). The CYTB gene used forward primer 5′-AATCCACTAAACACCCCACCC-3′ (165 bp), reverse primer 5′-GCTTCGTTGCTTTGAGGTATGA-3′; the β-actin gene used forward primer 5′-GGGAAATCGTGCGTGAC-3′, reverse primer 5′-AGGCTGGAAAAGAGCCT-3′. Primers for the mitochondrial COX II gene (forward 5′-ATAACCGAGTCGTTCTGCCAAT-3′, reverse 5′-TTTCAGAGCATTGGCCATAGAA-3′) and the nuclear 18S rRNA gene (forward 5′-CGCGGTTCTATTTTGTTGGT-3′, reverse 5′-AGTCGGCATCGTTTATGGTC-3′) were taken from the literature [27, 28] and provided by Sangon Biotech. qRT-PCR was performed utilizing AceQ qPCR SYBR Green Master Mix (Vazyme, Nanjing, China), following the manufacturer’s protocol. Each 10 μl reaction mixture contained 0.5 μl of 10 μM forward primer, 0.5 μl of 10 μM reverse primer, 5 μl of 2X SYBR Green Mix, 2 μl of DNA template, and 2 μl of ultrapure water. Amplification conditions: (1) initial denaturation at 95 ℃ for 5 min; (2) 40 cycles of (a) denaturation at 95 °C for 10 s and (b) annealing/extension at 60 °C for 30 s. The products were confirmed using the SYBR green single melting curve. Quantification of the mitochondrial CYTB, COX II, and nuclear 18S rRNA gene copies was normalized against a standard curve of the respective amplicons cloned in plasmids. The qRT-PCR data were analyzed by the Bio-Rad CFX Manager software (Bio-Rad). Each PCRreaction was carried out in triplicate to exclude outliers. The copy numbers of relative mtDNA were calculated as the ratio of (CYTB or COX II gene copies)/(18S rRNA gene copies).
RNA Isolation and Reverse Transcription
RNA was isolated using the RNAprep Pure Tissue Kit (Tiagen, Beijing, China) as per the manufacturer’s instructions. The quality of extracted RNA was analyzed using a NanoDrop 2000 spectrophotometer (Thermo Fisher); A260/A280 ratios were between 1.8 and 2.1. RNA was reverse transcribed into cDNA using a HiScript 1st Strand cDNA Synthesis Kit (Vazyme) according to the manufacturer’s protocol.
Detection of Mitochondrial Gene Expression
Mitochondrial gene expression values were quantified by SYBR green-based qRT-PCR. qRT-PCR of the target genes CYTB and COX II used β-actin gene as an endogenous control. Cycle threshold (CT) values were obtained with the CFX Manager (Version 3.1, Bio-Rad). Absence of non-specific amplicons was confirmed by melting curve analysis. Each sample was run in triplicate. Mitochondrial gene expression values used the 2–ΔΔCT method of relative quantification [29].
Statistical Analysis
Statistical analysis was performed using SPSS software (Version 18.0, SPSS Inc., Chicago, IL, USA). Data obtained are presented as median with interquartile range (IQR). The non-parametric Mann–Whitney U test was used to analyze the data. Graphs were generated using GraphPad Prism 6 software (GraphPad, La Jolla, CA, USA). Statistical significance was set at P < 0.05.
Results
Abnormal Mitochondrial Morphology in APP/PS1 Mice
Electron microscopy analysis revealed mitochondrial damage including mitochondrial swelling and loss of mitochondrial cristae in the hippocampi of APP/PS1 transgenic mice. (Fig. 2, white arrows). In the hippocampi of C57BL/6J mice, most of the mitochondria appeared normal (Fig. 2, black arrows).
Hypermethylation of CYTB and COX II Genes
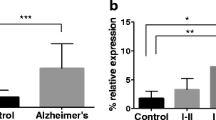
We compared the methylation levels of two mtDNA genes in the hippocampi of APP/PS1 transgenic mice and age-matched C57BL/6J mice (Fig. 3). Methylation levels were quantified by averaging the degree of methylation at six CpG sites of the CYTB gene and four CpG sites of the COX II gene. The average methylation levels of different CpG sites in the two mitochondrial genes were very low (≤ 4.0%); mtDNA was largely unmethylated at CpG sites. In the CYTB gene, cytosine methylation was notably elevated at CpG site 2 (P value = 0.0313, Fig. 4b) and CpG site 6 (P value = 0.0023, Fig. 4f) in the hippocampi of APP/PS1 mice compared with C57BL/6J mice (Fig. 4). The 5-mC of the COX II gene was hypermethylated at CpG site 2 in APP/PS1 mice compared to C57BL/6 J mice (P value = 0.0078, Fig. 4h). No significant differences in methylation were present at the other CpG sites measured.
A representative program in pyrosequencing analysis (COX II gene). The X-axis indicates the dispensation sequence of nucleotides added to the reaction (E enzyme; S substrates), and the Y-axis represents the relative intensity. The wide bars are CpG sites and figures on the top represent the methylation percentage calculated from the ratio of the peak heights of C and T. The narrow bars show a bisulfite treatment control point
Mean methylation values of 6 CpG sites in the CYTB gene (a–f) and 4 CpG sites in the COX II gene (g–j). The X-axis is the CpG sites in the gene. The Y-axis correspond to the mean methylation values of CpG sites. Data was expressed as median with IQR. The statistical significance is represented by the asterisks as follows: *P < 0.05; **P < 0.01
mtDNA Copy Numbers of CYTB and COX II Genes
To determine whether mtDNA methylation might regulate mtDNA copy number [30,31,32], we assayed copy numbers of CYTB and COX II genes by qRT-PCR and calculated the relative copy numbers against the nuclear 18S rRNA gene. We found that the relative mtDNA copy numbers of both genes were significantly lower in APP/PS1 than in age-matched C57BL/6J mice (P value < 0.001, Fig. 5a, b).
a, b Comparative analysis of the relative mtDNA copy numbers of CYTB gene and COX II gene. The results were calculated as the ratio of the CYTB or COX II genes/18S rRNA gene copies. All of the gene copy numbers were determined against reference plasmids containing the amplified sequences. c, d Comparative expression analysis of CYTB gene and COX II gene. Data represented as the amplification levels of CYTB and COX II gene against the β-actin gene using the 2−ΔΔCt method. Data was expressed as median with IQR. *P < 0.05; ***P < 0.001
Reduced Mitochondrial Gene Expression in APP/PS1 Mice
Expression levels of CYTB and COX II genes were significantly lower in APP/PS1 hippocampi than in age-matched C57BL/6J controls (P value < 0.05, Fig. 5c, d).
Discussion
Increasing evidence indicates that DNA methylation plays a role in the pathogenesis of AD. DNA methylation is an inheritable but reversible biochemical process, and has been considered to be an epigenetic biomarker that can be used in the diagnosis, prognosis, and treatment of AD [33]. Genome-wide and locus-specific epigenetic alterations have been reported in AD [34], but the majority of studies have focused on nuclear DNA methylation. mtDNA methylation is a recent and promising field [16]. It plays a crucial role in many diseases related to mitochondrial dysfunction [10, 12, 16, 35]. Mitochondrial impairments, including ROS production, impaired mitochondrial dynamics, and apoptosis, are prominent features of AD [5]. mtDNA methylation is a potential mechanism that might lead to mitochondrial dysfunction in AD, but few studies have been performed to address this issue [16]. mtDNA methylation levels of mitochondrial CYTB and COX II genes had not previously been investigated in AD.
We aimed to explore this by estimating the methylation levels of the CYTB and COX II genes. APP/PS1 transgenic mice, a standard AD model, were used to simulate the pathological changes and progressive course of AD [20]. We also quantified mtDNA copy numbers and expression levels of the two genes. The average methylation levels of different CpG sites were low and the methylated mtDNA accounted for only a small part of total mtDNA, consistent with previous studies [16, 24, 36, 37]. There is consensus that the D-loop region is one of the most methylated sites in mtDNA [16]. We found hypermethylation of mitochondrial CYTB and COX II genes with lower mtDNA copy numbers and expressions in the hippocampi of APP/PS1 mice. To the best of our knowledge, this is the first study to study methylation changes in these genes in an AD mouse model.
Complex III of the mitochondrial respiratory chain accepts electrons from reduced coenzyme Q and transfers them on to cytochrome c [3]. Complex III comprises 11 subunits; CYTB is one of the three conserved core subunits, and the only mtDNA-encoded subunit of Complex III [3, 38]. No previous study has indicated that mitochondrial CYTB gene methylation is related to any disease [39]. We observed that methylation of the CYTB gene decreased in the hippocampi of APP/PS1 mice at 9 months of age. Our study is the first to find hypermethylation of the CYTB gene in an AD mouse model.
Complex IV is the terminal oxidase of the mitochondrial respiratory chain and catalyzes electron transfer from reduced cytochrome c to O2, coupling it to the pumping of protons into the intermembrane space [3]. This complex is a copper-heme A hetero-multimeric complex formed by three functional core subunits (COX-III) encoded in mtDNA and 11 subunits encoded in the nuclear genome [40]. Impaired COX function is related to cellular energy metabolism and increased ROS production [3]; one study showed that the methylation of platelet mitochondrial COX II was higher in cardiovascular disease (CVD) [26]. Platelet COX II gene hypermethylation, which could serve as easy-to-obtain and non-invasive marker, may be implicated in the etiology of CVD Exposure to low perinatal doses of flame retardant polybrominated diphenyl ethers (PBDEs) may result in COX II gene hypomethylation in the frontal lobes of rat offspring [41]. To the best of our knowledge, COX II gene methylation has not been studied in other diseases. We found that mitochondrial COX II gene methylation was significantly higher in the hippocampi of 9-month-old APP/PS1 transgenic mice. This is the first study to show methylation changes in the COX II gene in an AD mouse model.
The pathophysiological mechanism and function of mtDNA remain unclear. To further explore the connections between abnormal mtDNA methylation, mtDNA content, and gene expression in an AD mouse model, we assayed the copy numbers and transcription levels of two genes. Altered mtDNA copy number is an indicator of abnormal mitochondrial biogenesis and has been previously detected in several diseases [31]. Some research indicates that demethylation of mtDNA could cause an elevation of mtDNA copy number [30, 32] and that mtDNA methylation of the D-loop is inversely proportional to mtDNA copy number in some diseases [17, 23]. Methylation changes in mitochondrial CYTB and COX II genes haves not been previously reported in association with mtDNA copy number in any disease, including AD. We report hypermethylation of CYTB and COX II genes with lower mtDNA copy number in the hippocampi of APP/PS1 mice than in C57BL/6J mice.
The effect of 5-mC is considered to interfere with transcriptional initiation, which thereby silences gene expression [33]. Although there is a lack of direct evidence, it appears that mtDNA methylation may play a key role in many pathophysiological processes through the regulation of mitochondrial gene expression [21,22,23]. One study in AD showed lower mitochondrial ND1 gene methylation and higher levels of ND1 mRNA in entorhinal cortex of eight AD patients. The authors proposed that hypomethylation of the ND1 gene could promote a rise in levels of mRNA expression [20]. We found that APP/PS1 mice with hypermethylation of CYTB and COX II genes exhibited lower mitochondrial gene expression. We speculate that hypermethylation of these genes might reduce gene expression. This warrants further investigation.
There are some limitations in our work. We studied the methylation of several CpG sites in CYTB and COX II genes in a mouse model of AD. Further studies are required to assess mtDNA methylation changes in other mtDNA regions, non-CpG sites, and AD patient samples. An unresolved question is whether mtDNA methylation in an AD mouse model is a consequence or cause of mitochondrial dysfunction. The epigenetic regulatory mechanism of mtDNA methylation remains elusive. With advances in techniques for targeted DNA methylation editing these issues should be resolved in the near future [42].
In summary, methylated mtDNA accounted for only a small part of the total mtDNA. Our data showed hypermethylation of the mitochondrial CYTB and COX II genes with reduced mtDNA copy numbers and lower gene expressions in an APP/PS1 mouse model of AD. We hypothesize that mtDNA methylation, one type of mitoepigenetic modification, might participate in the pathogenesis of AD by affecting mtDNA copy number and modulating gene expression, but further research is required to validate this hypothesis. mtDNA methylation may play a role in AD pathology, which opens a new window for AD therapy.
References
Alzheimer’s Association (2016) 2016 Alzheimer’s disease facts and figures. Alzheimer’s Dement 12(4):459–509
World Alzheimer Report 2018, Alzheimer’s Disease International, 2018: pp 1–48
Rigotto G, Basso E (2019) Mitochondrial dysfunctions: a thread sewing together alzheimer’s disease, diabetes, and obesity. Oxid Med Cell Longev 2019:7210892
Joe E, Ringman JM (2019) Cognitive symptoms of Alzheimer’s disease: clinical management and prevention. BMJ 367:l6217
Hroudová J, Singh N, Fišar Z (2014) Mitochondrial dysfunctions in neurodegenerative diseases: relevance to Alzheimer’s disease. Biomed Res Int 2014:175062
Devall M, Mill J, Lunnon K (2014) The mitochondrial epigenome: a role in Alzheimer’s disease? Epigenomics 6(6):665–675
Devall M et al (2016) Epigenetic regulation of mitochondrial function in neurodegenerative disease: New insights from advances in genomic technologies. Neurosci Lett 625:47–55
Kelly RD et al (2012) Mitochondrial DNA copy number is regulated in a tissue specific manner by DNA methylation of the nuclear-encoded DNA polymerase gamma A. Nucleic Acids Res 40(20):10124–10138
Ghosh S et al (2015) Mitoepigenetics: The different shades of grey. Mitochondrion 25:60–66
Iacobazzi V et al (2013) Mitochondrial DNA methylation as a next-generation biomarker and diagnostic tool. Mol Genet Metab 110:25–34
Phillips NR, Simpkins JW, Roby RK (2014) Mitochondrial DNA deletions in Alzheimer’s brains: a review. Alzheimer’s Dementia 10(3):393–400
Dong Z, Pu L, Cui H (2020) Mitoepigenetics and its emerging roles in cancer. Front Cell Dev Biol 8:4
Romano G et al (2018) RNA methylation in ncRNA: classes, detection, and molecular associations. Front Genet 9:243
Shock LS et al (2011) DNA methyltransferase 1, cytosine methylation, and cytosine hydroxymethylation in mammalian mitochondria. Proc Natl Acad Sci USA 108(9):3630–3635
Infantino V et al (2011) Impairment of methyl cycle affects mitochondrial methyl availability and glutathione level in Down’s syndrome. Mol Genet Metab 102(3):378–382
Coppedè F, Stoccoro A (2019) Mitoepigenetics and neurodegenerative diseases. Front Endocrinol 10:86
Stoccoro A et al (2018) Mitochondrial DNA copy number and D-loop region methylation in carriers of amyotrophic lateral sclerosis gene mutations. Epigenomics 10(11):1431–1443
Mishra M, Kowluru RA (2015) Epigenetic modification of mitochondrial DNA in the development of diabetic retinopathy. Invest Ophthalmol Vis Sci 56(9):5133–5142
Pirola CJ et al (2013) Epigenetic modification of liver mitochondrial DNA is associated with histological severity of nonalcoholic fatty liver disease. Gut 62(9):1356–1363
Blanch M et al (2016) Altered mitochondrial DNA methylation pattern in alzheimer disease-related pathology and in Parkinson disease. Am J Pathol 186(2):385–397
van der Wijst MG, Rots MG (2015) Mitochondrial epigenetics: an overlooked layer of regulation? Trends Genet 31(7):353–356
Feng S et al (2012) Correlation between increased ND2 expression and demethylated displacement loop of mtDNA in colorectal cancer. Mol Med Rep 6(1):125–130
Sanyal T et al (2018) Hypomethylation of mitochondrial D-loop and ND6 with increased mitochondrial DNA copy number in the arsenic-exposed population. Toxicology 408:54–61
Stoccoro A et al (2017) Decreased methylation of the mitochondrial D-loop region in late-onset Alzheimer’s disease. J Alzheimer’s Dis 59(2):559–564
Xu Y et al (2019) Altered mitochondrial DNA methylation and mitochondrial DNA copy number in an APP/PS1 transgenic mouse model of Alzheimer disease. Biochem Biophys Res Commun 520(1):41–46
Baccarelli AA, Byun HM (2015) Platelet mitochondrial DNA methylation: a potential new marker of cardiovascular disease. Clin Epigenet 7:44
Baek JH et al (2019) Chronological aging standard curves of telomere length and mitochondrial DNA copy number in twelve tissues of C57BL/6 male mouse. Cells 8(3):247
Podlesniy P et al (2013) Low cerebrospinal fluid concentration of mitochondrial DNA in preclinical Alzheimer disease. Ann Neurol 74:655–668
Schefe JH et al (2006) Quantitative real-time RT-PCR data analysis: current concepts and the novel “gene expression’s CT difference” formula. J Mol Med 84(11):901–910
Tong H et al (2017) Methylation of mitochondrial DNA displacement loop region regulates mitochondrial copy number in colorectal cancer. Mol Med Rep 16(4):5347–5353
Wen SL, Zhang F, Feng S (2013) Decreased copy number of mitochondrial DNA: a potential diagnostic criterion for gastric cancer. Oncol Lett 6(4):1098–1102
Gao J et al (2015) De-methylation of displacement loop of mitochondrial DNA is associated with increased mitochondrial copy number and nicotinamide adenine dinucleotide subunit 2 expression in colorectal cancer. Mol Med Rep 12(5):7033–7038
Cui D, Xu X (2018) DNA methyltransferases, DNA methylation, and age-associated cognitive function. Int J Mol Sci 19(5):1315
Sanchez-Mut JV, Gräff J (2015) Epigenetic alterations in Alzheimer’s disease. Front Behav Neurosci 9:347
Breton CV et al (2019) Effects of air pollution on mitochondrial function, mitochondrial DNA methylation, and mitochondrial peptide expression. Mitochondrion 46:22–29
Byun HM, Barrow TM (2015) Analysis of pollutant-induced changes in mitochondrial DNA methylation. Methods Mol Biol (Clifton, N.J.) 1265:271–283
D’Aquila P et al (2015) Age-and gender-related pattern of methylation in the MT-RNR1 gene. Epigenomics 7(5):707–716
Yang XH, Trumpower BL (1986) Purification of a three-subunit ubiquinol-cytochrome c oxidoreductase complex from Paracoccus denitrificans. J Biol Chem 261(26):12282–12289
D’Aquila P et al (2017) Mitochondrial genome and epigenome: two sides of the same coin. Front Biosci 22:888–908
Timón-Gómez A et al (2018) Mitochondrial cytochrome c oxidase biogenesis: recent developments. Semin Cell Dev Biol 76:163–178
Byun HM et al (2015) Epigenetic effects of low perinatal doses of flame retardant BDE-47 on mitochondrial and nuclear genes in rat offspring. Toxicology 328:152–159
Pflueger C, Swain T, Lister R (2019) Harnessing targeted DNA methylation and demethylation using dCas9. Essays Biochem 63(6):813–825
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 81870848) and the Key Research and Development Program of Shandong Province (Grant No. 2017GSF218046).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Xu, Y., Cheng, L., Sun, J. et al. Hypermethylation of Mitochondrial Cytochrome b and Cytochrome c Oxidase II Genes with Decreased Mitochondrial DNA Copy Numbers in the APP/PS1 Transgenic Mouse Model of Alzheimer’s Disease. Neurochem Res 46, 564–572 (2021). https://doi.org/10.1007/s11064-020-03192-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-020-03192-y