Abstract
Introduction
Ependymoma is the third most common malignant pediatric brain tumor. Although the biology that drives ependymoma is slowly being unraveled, the ability to translate these findings to clinical care remains an ongoing challenge. Epigenetic alterations appear to play a central role in the development of molecular classification of ependymoma.
Methods
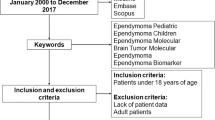
We reviewed the published literature available describing genetic and epigenetic underpinnings of ependymoma that have been reported to date and have summarized the information regarding genetic drivers of ependymoma that may point us toward therapeutic strategies.
Results
Ependymoma is a molecularly heterogeneous disease which has now been divided into at least nine distinct molecular subtypes based on DNA methylation and gene expression profiling. DNA methylation has emerged as an effective tool for classification of brain tumors alongside histopathology and other molecular diagnostics. There have been large retrospective cohorts describing molecular subgroup identity as a powerful independent predictor of outcome. There is limited published data on prospective trials to date however this is forthcoming which will lead to molecular stratification in the next generation of clinical studies.
Conclusion
This is a review of recent advancements in our understanding of the epigenetic basis of ependymoma and discussion of how these findings reveal potential therapeutic opportunities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ependymoma is an aggressive and relentless disease that is one of the most common malignant brain tumors found in children. Brain tumors are the leading cause of cancer related death in pediatrics and despite decades of progress in many other tumor types, the mainstay of treatment for ependymoma patients remains maximal safe surgical resection and radiation therapy. No cytotoxic chemotherapy or targeted agents have been shown to have clear benefits in improving patient outcomes to date [1,2,3,4,5]. Although about 75% of patients with ependymoma survive for over 5 years, most are left with neurologic sequelae of their treatment, which has a significant impact on their quality of life [6]. Ependymoma patients who relapse are often treated with surgery and re-irradiation. Effective and safe targeted therapies are desperately needed for treatment of ependymoma patients. Unfortunately, ependymomas harbor relatively low mutation rates, thus posing a challenge for precision-guided therapeutic strategies. Recurrent alterations that have been consistently detected include C11ORF95-RELA and YAP1 gene fusions which may act as oncogenic transcriptional activators to govern epigenetic and transcriptional programs. Mutation and over-expression of EZHIP (previously named CXORF67) in ependymoma inhibits repressive epigenetic marks (namely H3K27me3), thereby activating oncogenic programs during brain development. Ependymomas are divided into at least nine distinct molecular subtypes based on disparate epigenetic signatures that reflect both unique developmental origins and cancer drivers. These findings suggest that epigenetic mechanisms play a significant role in driving ependymoma development and capturing tumor heterogeneity; this may hold clues for devising desperately needed targeted therapies.
Epigenetic classification of ependymoma informs clinical outcomes
Multiple groups have shown that ependymomas are a molecularly heterogeneous collection of tumors [7, 8]. Ependymoma can be located in the supratentorial (ST) brain, posterior fossa (PF), or spinal cord. In children, 90% of cases arise intracranially, with two thirds in the posterior fossa. Historically, ependymomas have been histologically classified into several groups based on World Health Organization (WHO) Grade I-III. More recently, studies have suggested that given the heterogeneity in biology, distinguishing subgroups based on molecular markers is a more robust, reliable, and informative method [7,8,9,10]. Ependymoma has now been divided into at least nine distinct molecular subtypes based on DNA methylation and gene expression profiling [11].
In the case of PF ependymoma, expansion of DNA methylation profiling has supported the presence of two major subgroups of ependymoma termed PF-EPN-A and PF-EPN-B. In four independent PF ependymoma cohorts of 820 PF tumors, molecular subgroup identity has emerged as a powerful independent predictor of patient outcome [9]. Two examples underscore the clinical ramifications for an understanding of molecular subgroups in ependymoma: (1) sub-totally resected PF-EPN-A tumors have a dismal prognosis and may benefit most from alternative and biologically driven therapies, and (2) PF-EPN-B patients can often be treated with surgery alone, and patients with recurrent PF-EPN-B tumors can be successfully salvaged with delayed external beam radiation.
In the case of ST ependymoma, approximately 72% of cases are driven by a fusion between C11ORF95 and RELA (termed ST-EPN-RELA) or less frequently involve YAP1 gene fusions (termed ST-EPN-YAP1). DNA methylation based classification has facilitated subgrouping of ST ependymomas alongside gene fusion detection methods (i.e. Break-apart FISH, RNA fusion panels, and RNA-seq). While preliminary, early evidence has demonstrated favorable outcomes in the case of ST-EPN-YAP1 tumors as compared to ST-EPN-RELA ependymomas. One study reporting on European data shows a 5 years progression free survival (PFS) of less than 30% and a 5 years overall survival (OS) of 75% for ST-EPN-RELA compared to a 5 years PFS of 66% and a 5 years OS of 100% for ST-EPN-YAP1 [10, 11]. These findings await further validation in additional and prospective tumor cohorts stratified by molecular subgroup.
DNA methylation has emerged as an effective tool for classification of brain tumors alongside histopathology and other molecular diagnostics. Illumina EPIC Methylation arrays have become the method of choice for several reasons: (1) a reference dataset has been constructed to objectively predict brain tumor types, such as ependymoma, (2) tumor DNA methylation patterns are quite stable in both frozen and FFPE-tissue, (3) methods for identification of some brain tumor subgroups/subtypes are not currently available using other methodologies, (4) low-input DNA requirements, and (5) no batch effects when analyzing cohorts of many samples processed at different times and/or different centers [12]. Illumina EPIC DNA methylation based profiling will provide an increasingly accessible measure of the molecular differences across ependymoma patients in a clinical and trial setting, and should be incorporated in future clinical studies of ependymoma. As an example, the completion of a national US clinical trial evaluating the role of chemotherapy in ependymoma (Children’s Oncology Group: COG:ACNS-0831) would potentially benefit from stratification by molecular subgroups and subsequent comparison of clinical outcomes. Several important insights could be gained from epigenetic characterization of tumors across this cohort such as the relevance of chemotherapy across and within subgroups, outcomes and prevalence of ST-EPN-RELA and ST-EPN-YAP1 tumors, and further validation of PF-EPN-A/B patient outcomes as compared to previous trials (i.e. COG:ACNS-0121) that have accompanying DNA methylation based characterization.
Genetic drivers of supratentorial ependymoma shape tumor epigenomes
Epigenetic marks (such as DNA methylation) or histone marks in ependymoma serve as available tools that could be employed for diagnostic and prognostic purposes [13]. Ependymoma tumor epigenomes also capture somatic alterations that shape oncogenic gene expression programs. Direct examples are ST-EPN-RELA and ST-EPN-YAP1 gene fusions which may act as transcription factors, and are thought to directly impact gene expression programs. The mechanism of gene activation, direct targets, and functional relevance of ependymoma-fusion target genes remain an area of active research.
ST-EPN-RELA accounts for over 70% of supratentorial ependymoma and is driven by a chromothripsis event in chromosome 11, which results in the fusion of the chromosome 11 open reading frame (C11ORF95) to v-rel avian reticuloendotheliosis viral oncogene homolog A (RELA). C11ORF95 has been a historically uncharacterized gene, however, some data suggests that the zinc finger domains of C11ORF95 may be essential for oncogenesis, possibly affecting trafficking, degradation, or target specificity of associated transcription factors [15]. The RELA gene, however, has been well described in the literature; it acts as a transcription factor central to mediating NF-kB pathway activation in processes such as inflammation, cellular metabolism, and chemotaxis [15]. This has led to the notion that C11ORF95-RELA fusion acts directly on DNA/chromatin as an oncogenic transcription factor to drive both canonical NF-kB and neoplastic transcriptional programs. Whether inhibition of canonical NF-kB pathway also disrupts C11ORF95-RELA fusion activity and tumorigenicity is unclear.
Importantly, C11ORF95-RELA fusion alone has been shown to drive ependymoma development in mice, therefore implicating this protein as a bonafide cancer driver [7, 14,15,16]. Proper localization to the nucleus and transcriptional activity of RELA are reliant upon its phosphorylation at serine 276 (S276) and subsequent acetylation at lysine 310 (K310) in response to upstream cytokines such as TNF-α or lipopolysaccharide (LPS) stimulation [14]. Furthermore, mutagenesis of S276 abrogates tumor formation in a native mouse model of C11ORF95-RELA ependymoma [14, 17,18,19]. This indicates that at least some aspects of RELA protein regulation are utilized in the context of the C11ORF95-RELA fusion protein. However, further delineation of the role of the NF-KB pathway in tumorigenesis of ST-EPN-RELA is needed, as this genomic event may pose a lead target for functional and therapeutic investigation.
The remaining subgroup of supratentorial ependymoma is characterized by YAP1 (YES-associated protein) fusions, termed the ST-EPN-YAP1 subgroup. The majority (~ 80%) of YAP1 fusion ependymomas are characterized by fusions between YAP1 and MAMLD1 (mastermind like domain containing 1). MAMLD1 is an important regulator of Notch signaling transcription and p53 tumor suppressor pathways [11, 20, 21]. A less frequent fusion has also been described between YAP1 and FAM118B [11].
YAP is a well characterized oncoprotein that is one of the major downstream effectors of the Hippo signaling pathway, which is a cancer pathway that has been shown to be dysregulated in multiple cancers such as ovarian carcinoma, non-small cell lung carcinoma, and hepatocellular carcinoma [20]. Additionally, YAP1 has been shown to co-activate alternative Wnt signaling as well as the canonical Wnt/B-catenin pathway in cancers such as colon cancer [1]. When the YAP pathway is activated, YAP and its co-activator tafazzin (TAZ) are phosphorylated and unable to translocate to the nucleus. Without phosphorylation, however, YAP-TAZ can translocate to the nucleus and associate with transcription factors such as TEAD1-4. This interaction results in upregulation of genes involved in cell proliferation and down-regulation of those involved in apoptosis and differentiation [22,23,24,25]. When the interaction between YAP and TEAD transcription factors is prohibited, tumor formation in mice is abolished, thus demonstrating that elements of canonical YAP1 signaling are required for ependymoma development [21].
Together these findings point to a role of C11ORF95-RELA and YAP1 fusion in activating oncogenic gene expression programs. Whether this is a direct or indirect interaction with DNA or chromatin is unclear. RELA and YAP1 canonical pathway activation is likely to contribute to tumor formation and yield potential therapeutic insights. These fusion proteins are likely to confer novel gene activation. Supporting this concept is that hyperactivation mutants of RELA are unable to induce ependymomas in mice, and that RELA or YAP1 gene amplifications are rarely ever observed in human tumors [8, 14, 15]. Also important to consider is the biology of non-RELA and non-YAP fusion driven tumors and mechanistic insights that can be gained from studying these less-common fusion proteins together in mouse models. Understanding the transcriptional impact of ependymoma fusion proteins will be important to developing specific and effective therapeutic approaches.
Posterior fossa ependymoma : a purely epigenetic disease?
In line with several other pediatric brain tumors, ependymomas exhibit low mutation rates and low-frequency of recurrent somatic mutations [8, 15]. Of somatic gene mutations identified, EZHIP (or CXORF67) or H3F3A-K27M alterations have been reported in PF-EPN-A ependymoma (never in PF-EPN-B) [10, 26,27,28]. In nearly all cases, except when H3F3A-K27M mutations are present, EZHIP expression is elevated. EZHIP functions as a natural inhibitor of the polycomb-repressive complex 2, a protein complex that contains the EZH2 enzyme that catalyzes the repressive chromatin mark, H3K27me3. EZHIP expression or H3K27M mutation is sufficient to repress H3K27me3 modification. The direct transcriptional effects and contribution to ependymoma development of EZHIP are unclear. Importantly, in many cases EZHIP over-expression is the only reported alteration in PF-EPN-A ependymomas, without any clear additional cancer gene mutation or alteration.
Indeed, global loss of H3K27me3 represents a hallmark feature of PF-EPN-A ependymoma; analogous to H3K27M driven midline high-grade glioma (mHGG) [29, 30]. A major difference however is the lack of accompanying alterations such as TP53, ATRX, PDGFRA that are seen in mHGG [31, 32]. This difference raises the question as to how loss of H3K27me3 in ependymoma alone contributes to tumor formation and the cellular and mechanistic differences that distinguish PF-EPN-A ependymoma and H3K27M glioma. Can exclusive EZHIP over-expression, accompanied by loss of H3K27me3, drive tumorigenesis? Also unclear are the role of EZHIP mutations that are observed in PF-EPN-A ependymomas, and the mechanisms of tumor initiation that lead to EZHIP over-expression. One commonality between PF-EPN-A ependymoma and H3K27M glioma is the global depletion of DNA methylation, and in the case of PF-EPN-A tumors retention of DNA hypermethylation at CpG islands. Genes regulated by DNA hypermethylation in PF-EPN-A ependymoma are enriched in known targets of PRC2. This may create a potential therapeutic vulnerability to EZH2 inhibitors in H3K27me3 depleted tumors, which are thought to require minimal levels of H3K27me3 for tumor suppressor gene silencing and cell survival.
The potential result of repressive gene silencing mechanisms is the subsequent activation of repeat element transcription [including endogenous retroviruses (ERVs)] that comprise over two-thirds the human genome [33]. As shown in ATRT, aberrant transposable element expression leads to mobilization of transposons and disruption of the INI-1 tumor suppressor gene [34, 35]. Although the mechanism is unclear, repeat elements and ERVs are transcribed in PF-EPN-A ependymoma and H3K27M glioma, which activates intrinsic ellular defense responses and a therapeutic vulnerability known as viral mimicry [36]. Pathways of repeat element expression and their contribution to ependymoma-genesis continue to be explored as an active area of research.
In contrast, and counter-intuitively, PF-EPN-B tumors have frequent and recurrent large-scale copy number alterations, yet patients tend to have better clinical outcomes [8]. One hypothesis is that PF-EPN-B tumors arise from a more differentiated cell type and require additional ‘hits’ for neoplastic transformation. In the case of PF-EPN-A ependymoma, evidence supports that these tumors arise during early embryonic brain development (within the radial glial cell lineage) and that cellular identity program together with EZHIP over-expression could be a driver of tumorigenesis. Furthermore, are there cell-extrinsic factors such as metabolism and cell–cell communication, that aberrantly lead to EZHIP over-expression?
The end of the beginning
Over the last decade, ependymomas have undergone exhaustive genomic characterization. While there are likely to be additional genomic insights into the disease that may be uncovered, clear drivers particularly in ST ependymoma have been discovered, in the form of gene fusions. The next step is likely deep characterization and mouse modeling of these gene fusions to learn more about their molecular functions with the goal of identifying new therapeutic strategies. ‘Drugging’ the fusion proteins themselves is likely to be challenging, however, new methods are being developed to assist with this goal such as the application of protein-degrader methods and advanced protein structural methods (i.e. CRYO-EM). There is also the potential of identifying proteins that are required for ependymoma-gene fusion activity, such as transcriptional co-activators and epigenetic modifiers, which may have small molecules already available. In the case of the most common ependymoma in children, PF-EPN-A, there are similar (if not more difficult) challenges in terms of pre-clinical mouse modeling, which will be vital to the investigation of EZHIP biology. There may also be benefit in comparing and contrasting PF-EPN-A against H3K27M driven gliomas which also harbor a similar (although not identical) epigenetic phenotype (i.e. H3K27me3 depletion). A major future direction of this field will be understanding the fundamental epigenetic basis of ependymoma subtypes, each as different diseases, and bridging these findings to new therapies and/or molecular diagnostics.
References
Lee SH, Chung CK, Kim CH, Yoon SH, Hyun SJ, Kim KJ et al (2013) Long-term outcomes of surgical resection with or without adjuvant radiation therapy for treatment of spinal ependymoma: a retrospective multicenter study by the Korea Spinal Oncology Research Group. Neuro-Oncology 15(7):921–929
Metellus P, Barrie M, Figarella-Branger D, Chinot O, Giorgi R, Gouvernet J et al (2007) Multicentric French study on adult intracranial ependymomas: prognostic factors analysis and therapeutic considerations from a cohort of 152 patients. Brain : J Neurol 130(Pt 5):1338–1349
Ostrom QT, Gittleman H, Liao P, Rouse C, Chen Y, Dowling J et al (2014) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2007–2011. Neuro-Oncology. https://doi.org/10.1093/neuonc/nou223
Taylor MD, Poppleton H, Fuller C, Su X, Liu Y, Jensen P et al (2005) Radial glia cells are candidate stem cells of ependymoma. Cancer Cell 8(4):323–335
Wu J, Armstrong TS, Gilbert MR (2016) Biology and management of ependymomas. Neuro-Oncology 18(7):902–913
Merchant TE, Li C, Xiong X, Kun LE, Boop FA, Sanford RA (2009) Conformal radiotherapy after surgery for paediatric ependymoma: a prospective study. Lancet Oncol 10(3):258–266
Pajtler KW, Mack SC, Ramaswamy V, Smith CA, Witt H, Smith A et al (2017) The current consensus on the clinical management of intracranial ependymoma and its distinct molecular variants. Acta Neuropathol 133(1):5–12
Mack SC, Witt H, Piro RM, Gu L, Zuyderduyn S, Stutz AM et al (2014) Epigenomic alterations define lethal CIMP-positive ependymomas of infancy. Nature 506(7489):445–450
Ramaswamy V, Hielscher T, Mack SC, Lassaletta A, Lin T, Pajtler KW et al (2016) Therapeutic impact of cytoreductive surgery and irradiation of posterior fossa ependymoma in the molecular era: a retrospective multicohort analysis. J Clin Oncol 34(21):2468–2477
Pajtler KW, Wen J, Sill M, Lin T, Orisme W, Tang B et al (2018) Molecular heterogeneity and CXorf67 alterations in posterior fossa group A (PFA) ependymomas. Acta Neuropathol 136(2):211–226
Pajtler KW, Witt H, Sill M, Jones DT, Hovestadt V, Kratochwil F et al (2015) Molecular classification of ependymal tumors across all CNS compartments, histopathological grades, and age groups. Cancer Cell 27(5):728–743
Capper D, Jones DTW, Sill M, Hovestadt V, Schrimpf D, Sturm D et al (2018) DNA methylation-based classification of central nervous system tumours. Nature 555(7697):469–474
Mack SC, Pajtler KW, Chavez L, Okonechnikov K, Bertrand KC, Wang X et al (2018) Therapeutic targeting of ependymoma as informed by oncogenic enhancer profiling. Nature 553(7686):101–105
Ozawa T, Arora S, Szulzewsky F, Juric-Sekhar G, Miyajima Y, Bolouri H et al (2018) A de novo mouse model of C11orf95-RELA fusion-driven ependymoma identifies driver functions in addition to NF-kappaB. Cell Rep 23(13):3787–3797
Parker M, Mohankumar KM, Punchihewa C, Weinlich R, Dalton JD, Li Y et al (2014) C11orf95-RELA fusions drive oncogenic NF-kappaB signalling in ependymoma. Nature 506(7489):451–455
Griesinger AM, Witt DA, Grob ST, Georgio Westover SR, Donson AM, Sanford B et al (2017) NF-κB upregulation through epigenetic silencing of LDOC1 drives tumor biology and specific immunophenotype in Group A ependymoma. Neuro-Oncology 19(10):1350–1360
Chen LF, Mu Y, Greene WC (2002) Acetylation of RelA at discrete sites regulates distinct nuclear functions of NF-kappaB. EMBO J 21(23):6539–6548
Hoesel B, Schmid JA (2013) The complexity of NF-κB signaling in inflammation and cancer. Molecular Cancer 12:86
Huang B, Yang XD, Lamb A, Chen LF (2010) Posttranslational modifications of NF-kappaB: another layer of regulation for NF-kappaB signaling pathway. Cell Signal 22(9):1282–1290
Andreiuolo F, Varlet P, Tauziède-Espariat A, Jünger ST, Dörner E, Dreschmann V et al (2019) Childhood supratentorial ependymomas with YAP1-MAMLD1 fusion: an entity with characteristic clinical, radiological, cytogenetic and histopathological features. Brain Pathol (Zurich, Switzerland) 29(2):205–216
Pajtler KW, Wei Y, Okonechnikov K, Silva PBG, Vouri M, Zhang L et al (2019) YAP1 subgroup supratentorial ependymoma requires TEAD and nuclear factor I-mediated transcriptional programmes for tumorigenesis. Nature Commun 10(1):3914
Lester A, McDonald KL (2020) Intracranial ependymomas: molecular insights and translation to treatment. Brain Pathol (Zurich, Switzerland) 30(1):3–12
Totaro A, Panciera T, Piccolo S (2018) YAP/TAZ upstream signals and downstream responses. Nat Cell Biol 20(8):888–899
Yu FX, Zhang Y, Park HW, Jewell JL, Chen Q, Deng Y et al (2013) Protein kinase a activates the hippo pathway to modulate cell proliferation and differentiation. Genes Dev 27(11):1223–1232
Zhao B, Tumaneng K, Guan KL (2011) The hippo pathway in organ size control, tissue regeneration and stem cell self-renewal. Nat Cell Biol 13(8):877–883
Hübner JM, Müller T, Papageorgiou DN, Mauermann M, Krijgsveld J, Russell RB et al (2019) EZHIP/CXorf67 mimics K27M mutated oncohistones and functions as an intrinsic inhibitor of PRC2 function in aggressive posterior fossa ependymoma. Neuro-Oncology 21(7):878–889
Jain SU, Do TJ, Lund PJ, Rashoff AQ, Diehl KL, Cieslik M et al (2019) PFA ependymoma-associated protein EZHIP inhibits PRC2 activity through a H3 K27M-like mechanism. Nature Commun 10(1):2146
Piunti A, Smith ER, Morgan MAJ, Ugarenko M, Khaltyan N, Helmin KA et al (2019) CATACOMB: An endogenous inducible gene that antagonizes H3K27 methylation activity of Polycomb repressive complex 2 via an H3K27M-like mechanism. Sci Adv. https://doi.org/10.1126/sciadv.aax2887
Panwalkar P, Clark J, Ramaswamy V, Hawes D, Yang F, Dunham C et al (2017) Immunohistochemical analysis of H3K27me3 demonstrates global reduction in group-A childhood posterior fossa ependymoma and is a powerful predictor of outcome. Acta Neuropathol 134(5):705–714
Pratt D, Quezado M, Abdullaev Z, Hawes D, Yang F, Garton HJL et al (2020) Diffuse intrinsic pontine glioma-like tumor with EZHIP expression and molecular features of PFA ependymoma. Acta Neuropathologica Commun 8(1):37
Wu G, Broniscer A, McEachron TA, Lu C, Paugh BS, Becksfort J et al (2012) Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat Genet 44(3):251–253
Schwartzentruber J, Korshunov A, Liu XY, Jones DT, Pfaff E, Jacob K et al (2012) Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482(7384):226–231
de Koning AP, Gu W, Castoe TA, Batzer MA, Pollock DD (2011) Repetitive elements may comprise over two-thirds of the human genome. PLoS Genet 7(12):e1002384
Henssen AG, Koche R, Zhuang J, Jiang E, Reed C, Eisenberg A et al (2017) PGBD5 promotes site-specific oncogenic mutations in human tumors. Nat Genet 49(7):1005–1014
Henssen AG, Kentsis A (2018) Emerging functions of DNA transposases and oncogenic mutators in childhood cancer development. JCI Insight 3(20):e123–172
Krug B, De Jay N, Harutyunyan AS, Deshmukh S, Marchione DM, Guilhamon P et al (2019) Pervasive H3K27 acetylation leads to ERV expression and a therapeutic vulnerability in H3K27M gliomas. Cancer Cell 35(5):782–97.e8
Acknowledgements
S.C.M. is supported by a Cancer Prevention Research Institute of Texas (CPRIT) scholar award (RR170023), an Alex s Lemonade Stand Foundation (ALSF) A award and Young Investigator award, a Pediatric Brain Tumor Foundation award, a Chad Tough Young Investigator award, a Cookies for Cancer research grant, a RALLY research grant, a BEAR Necessities Pediatric Cancer Foundation grant, a Children's Cancer Research Fund award, a Children's Brain Tumor Foundation award, and a Baylor College of Medicine Junior Faculty award.
Author information
Authors and Affiliations
Contributions
All authors contributed to the review conception and design. Material preparation, literature search, and data analysis was performed by AS, KB, PW, AS, and SM. The first draft of the manuscript was written by AS, KB, and PW and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors of this manuscript have no conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Stuckert, A., Bertrand, K.C., Wang, P. et al. Weighing ependymoma as an epigenetic disease. J Neurooncol 150, 57–61 (2020). https://doi.org/10.1007/s11060-020-03562-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-020-03562-0