Abstract
Glaucoma is an ailment causing permanent vision loss but can be prevented through the early detection. Optic disc to cup ratio is one of the key factors for glaucoma diagnosis. But accurate segmentation of disc and cup is still a challenge. To mitigate this challenge, an effective system for optic disc and cup segmentation using deep learning architecture is presented in this paper. Modified Groundtruth is utilized to train the proposed model. It works as fused segmentation marking by multiple experts that helps in improving the performance of the system. Extensive computer simulations are conducted to test the efficiency of the proposed system. For the implementation three standard benchmark datasets such as DRISHTI-GS, DRIONS-DB and RIM-ONE v3 are used. The performance of the proposed system is validated against the state-of-the-art methods. Results indicate an average overlapping score of 96.62%, 96.15% and 98.42% respectively for optic disc segmentation and an average overlapping score of 94.41% is achieved on DRISHTI-GS which is significant for optic cup segmentation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Glaucoma is an ailment which causes vision loss over time and hence is referred to as “silent thief of sight” [37]. The seriousness of the ailment can be estimated from the prediction that the population affected from glaucoma may increase from 76 million in 2020 to 111.8 million in 2040 [56]. Therefore, people between 40 to 64 years of age are advised to get their eye checkup done once in 2 to 4 years, by American Academy of Ophthalmology [20]. However, such a mass screening might raise the chances of human error. Also, the scarcity of ophthalmologists is a major issue. Hence, Computer-Aided Diagnostic (CAD) systems can help to reduce the errors and may assist the healthcare experts in an impartial screening of patients [23]. Glaucoma is recognized by changes in the shape and size of the optic disk and cup. It tends to change the Cup to Disk Ratio (CDR) [42]. The CDR can be calculated by taking the ratio of the vertical length of the optic cup and optic disc [12].
Presently, several imaging techniques are used such as Confocal Scanning Laser Ophthalmoscopy (CSLO), Optical Coherence Tomography (OCT) [57], Scanning Laser Polarimetry (SLP) etc. but the more economical and portable approach is to acquire fundus images using a fundus camera [2]. An add-on advantage of using fundus camera is that images captured from the camera can also be used to detect some other eye diseases such as age-related macular degeneration (AMD) and diabetic retinopathy (DR) [3, 29, 30].
Accurate segmentation of Optic disc and Optic cup is important to obtain the precise Cup-to-Disc Ratio (CDR), which is an essential step for Glaucoma classification, analysis and grading. But the Segmentation boundaries/contours of optic disc and optic cup, usually marked by multiple medical experts, differ by a small amount in every case. An average of these marked boundaries (binary masks) are normally treated as ground truth and are used to train the model.
In particular, the merits and key contributions of the proposed system are highlighted as follows:
-
We present a novel method of training the network with modified ground-truths and build a model, where the boundary pixels have been assigned probabilities instead of binary values of either 1 or 0.
-
We used a modified segmentation measure for accuracy and in representing the subsequent loss function. The intersection/union of two binary maps is now transformed into multiplication of two probability masks, which make more sense.
-
We show the utility of transfer learning, where in the model is trained exclusively on glaucoma image datasets before being evaluated for test performance. This makes the architecture robust and suitable for real-time applications. Also, this approach eliminates the overheads of designing customized features for the segmentation problem.
-
Extensive simulations are conducted to demonstrate the performance of the proposed system.
A broader paper outline is given as follows: Section 2 presents the previous work; Section 3 explains the model architecture that have been utilized in the segmentation work; Section 4 discusses the methodology; Section 5 presents results and analysis; and finally, conclusions are drawn in Section 6.
2 Related work
Numerous methods have been successfully implemented in the past for purpose of automatic diagnosis of the disease [23].There exists various factors such as blood vessels present in the fundus image, affect the classification accuracy of glaucoma present in the eye [19, 22].Chakravarty et al. [9] performed co-training on SVM classifier on image-based and segmentation-based features for the classification of fundus images. In order to find the segmentation-based features, they segmented the optic disc (OD) and optic cup (OC) using coupled sparse dictionary for depth-based segmentation technique. Yanwu-Xu et al. [58] utilized multiple kernel learning framework through the incorporation of class for glaucoma detection. Bock et al. [6] calculated Glaucoma Risk Index (GRI) from the pre-processed image with the use of the probabilistic two-stage scheme. Agarwal et al. [4] implemented classic morphological operations iteratively followed by active contour fitting for the precise segmentation of the optic disc. Issac et al. [26] utilized the concept of super pixels for the removal of false positives followed by the analysis of geometrical features to accurately segment out the optic disc. Raghavendra et al. [43] calculated features using radon transform technique and modified census transform technique to train the SVM classifier, and achieved state of the art results. Feature reduction was done using Locality Sensitive Discriminant Analysis (LSDA).
Literature revealed that high precision results have been achieved on highly challenging datasets using the emerging technique of Convolutional Neural Networks (CNNs) [31]. The depth of these networks is the key factor of its performance hence the effective selection of network depth is task specific [51]. Not only for the classification purpose but also state of the art results have been obtained in the segmentation-oriented tasks, using deep neural networks [24, 38]. Jonathan et al. [49] have achieved state-of-the-art segmentation of PASCAL VOC, NYUDv2 and SIFT Flow, using the learned representations of contemporary classification networks which were transferred after fine-tuning. This highlights the scope in transferring of weights from one problem to another, popularly known as transfer learning. Jason et al. [59] highlight the characteristics attained by the model depending on the layer from which weights are being transferred. They discuss the frozen and non-frozen weights and tried to quantify the generality of the layers. Along with such a wide range of applications, DCNNs is also a very popular approach to computer-aided diagnosis (CAD) of the medical path. Shin et al. [50] discussed the application of deep learning for the detection of thoraco-abdominal lymph node detection and interstitial lung disease, along with under-discussed characteristics of the model including model architecture, dataset characteristics, and transfer learning.
With regard to glaucoma analysis, deep convolutional neural networks have been recently used for both classification and segmentation purposes and few ensemble approaches have also been proposed. Fu.et al. [16] proposed Disc-aware Ensemble Network (DENet) for automatic glaucoma screening from fundus image. DENet works consists four deep streams on different levels and modules of fundus image. In these four deep streams, first focus on the classification of fundus image, second detects the disc localization, third works on disc region level and fourth focus on disc region with polar transformation to improve screening performance. The combined streams produce screening performance. Phasuk et al. [41] proposed a glaucoma screening network by utilizing DENet, incorporating Residual deconvolutional neural network for segmentation and artificial neural network for classification network. This approach claimed better accuracy against on DENet ORIGA-650, RIM-ONE and DRISHTI-GS dataset. Bajwa et al. [5] proposed a two-stage framework for classification of glaucoma from fundus images. First stage uses regions with convolutional neural network for localization and second stage uses CNN for classification of glaucoma. Pinto et al. [13] performed assessment of glaucoma on different ImageNet-trained CNN: VGG16, VGG19, InceptionV3, ResNet50 and the performance of Xception architecture was found better compared to other architectures.
Deep Convolutional Neural Networks (DCNNs) have also been a choice for researchers for the glaucoma segmentation tasks previously. Kausu et al. [28] calculated morphological and wavelet features followed by training them on a three-layered artificial neural network, and successfully outperformed state of the art results. Srivastava et al. [55] defined a seven-layered model for the optic disc segmentation in the presence of Parapapillary Atrophy (PPA) and attained state of the art results. An approach of two-folded CNN is given in [8], where they first segmented the OD and then fed that segmented image to the model for classification purpose. Zilly et al. [61] proposed a unique method of CNN with entropy sampling and ensemble learning, and successfully achieved a classification accuracy of 94.1% on RIM-ONE dataset. Lim et al. [33] calculated state-of-art vertical cup to disc error, after the successful segmentation of OD and OC along different orientations. Maninis et al. [34] have implemented a base network architecture with two sets of specialized layers for the segmentation of blood vessels and optic disc. Huazhu et al. [25] have proposed a new network named as M-net architecture for the segmentation of OD and OC, and have successfully attained state-of-the-art results on ORIGA dataset. They also have proposed reasonable glaucoma screening in terms of CDR. JIANG et al. [27] proposed a multi-label DCNN model by utilizing GAN, the better performance of this model is claimed on DRISHTI-GS1 dataset.
U-net architecture is one of the most popular architectures and has performed extraordinarily in various biomedical segmentation-based tasks [44, 46]. It also resolves the problem of the limited number of annotated images. Ronneberger et al. [45] performed segmentation task on three different classes of dental images to achieve dice similarity of 56.4% on a dataset of 39 images provided by the challenge organizers. The number of training images available was only 39, but training was done using data augmentation. Gao et al. [18] utilized U-net architecture along Gaussian matched filter to outperform the existing algorithms of blood vessel segmentation from fundus images. Work proposed in [47] consists of applying a customized loss function on U-net architecture for the purpose of segmentation of OD and OC. They tried to reduce the number of training parameters so as to reduce the computational time and attained state-of-art results. Almost all the papers that we have come across for optic disc or cup segmentation have taken the averaged ground truth labels for the training phase. A better way is perhaps to take all the images with different expert markings to train the data but that is not made available in public datasets. So, there lies an opportunity to build a model which is trained on modified ground truths instead of binarized ones.
3 Model architecture: U-net
Convolutional neural networks are basically mathematical models, which are capable of learning high-level features from the low-level ones [17, 32]. The traditional use of CNNs was for the classification tasks, but various computer vision tasks are segmentation based. Ciresan et al. [11] trained a network on the patches of the image, predicting the class of each pixel. But as visible, the network is too slow and almost impossible to run on huge datasets, as the training dataset is many times larger than the actual dataset. This is where encoder-decoder networks come into the picture. They allow feed an image as a whole and result is the segmented output of the image.
The model proposed in this work is based on the U-net architecture. The name of the architecture is derived from its unique design. It is an encoder-decoder type network which takes the image as an input along with its ground truth and returns the probability map of the ROI as the output. In the network, the feature maps from the convolutional layer of downsampling steps are fed to the convolutional layers of the upsampling part. The U-Net architecture was considered in our experiments because it has proven to achieve outstanding results in the applications of biomedical segmentation tasks [44]. Also, with the help of data augmentation, it has proved to achieve good results even in the absence of a huge dataset. Moreover, the absence of a fully connected layer eliminates the restrictions on the size of the input image. This fact permits the user to process images of different sizes, which is definitely an appealing trait to apply the technique of deep learning on the biomedical images with high precision. The block diagram of the model proposed is depicted in Fig. 1.
Unlike the original U-Net architecture, the proposed network has half the filters in each convolution layer and size of the input image is kept low (128×128).This is done in order to reduce the number of parameters to be trained and to reduce the time during the training phase. However, experiments have also been performed for an image size of 256×256 and 512×512. Total numbers of trainable parameters in the model are 7,760,097. Details about the number of layers and feature maps are mentioned in the following subsections.
3.1 Convolutional filters
They form the crux of any deep learning model and can be considered as a cube. Each cell of this cubic filter comprises of some weights which get multiplied to the intensities of the image. The sum of the bias and product terms obtained replaces the value of the center pixel. This model consists 19 convolution layers. Number of filters in each layer varies according to its position in the model and ranges from 32 filters to 512 filters. All the layers except the last one is using relu activation with padding feature kept on.
3.2 Pooling layer or downsampling layer
Pooling layers are generally applied to reduce the redundant features and reduce the image size. Hence, they are also known as downsampling layers. Here we have used max-pooling layers for the purpose. The model is formed with 4 downsampling layers each with a window size of (2,2).
3.3 Upsampling layer and de-convolutional layers
Upsampling layers as the name suggests, are the exact opposite to the pooling layers. An upsampling filter is also a window but this time, one pixel on the input image is copied to all the pixels lying under that window in the output image. Hence a basic window size of 2×2 will result in an image size of 2 N×2 N×d. This layer is used to increase the image size and number of features. Also, it is the most important part of the decoder network. Our model is embedded with de-convolutional layers which are only a bit different. They in place of copying the exact same value, multiply them with certain weights and hence have trainable filters. 4 such layers are present in the proposed model with filter size (2, 2) and number of filters varies as shown in Fig. 1.
4 Methodology
4.1 Preprocessing of fundus images
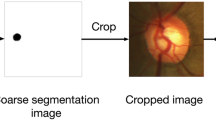
Prior to the segmentation of optic disc and cup, preprocessing is performed in two stages. During the optic disk segmentation phase, contrast of the images was adjusted, using Contrast Limited Adaptive Histogram Equalization (CLAHE) as one of the preprocessing steps. The images were resized to 128×128 pixels and rescaled such that the pixel values are between 0 and 1. Image data is then normalized by mean centering and standard deviation scaling. For the optic cup segmentation phase, the fundus image is cropped to the bounding box of the optic disc obtained as the result of stage 1. As the optic cup is always located inside the optic disc [47], these subimages were used for training the model for optic cup segmentation. A sample image before and after preprocessing of optic disc segmentation step is shown in Fig. 2 while Fig. 3 depicts the complete fundus image and the cropped optic disc region, suitable for the segmentation of optic cup.
4.2 Dataset description
Public datasets like DRISHTI-GS1 [53, 54] provided ground truths for optic disc and optic cup segmentation of fundus images while RIM-ONE [40] and DRIONS-DB [7] cater only to OD segmentation. DRISHTI-GS1 dataset consists of a total of 101 images. These have been divided into 50 training and 51 testing images. All the images have been marked by 4 opthalmologists. The markings by all the four medical experts have been fused to form a soft segmentation map. Rim-one has 169 images (majority of them normal) with 5 manual boundary markings for each one. There were 5 glaucoma domain experts (4 ophthalmologists and 1 optometrist) who developed manual optic disc segmentation in each image. These manual segmentations were subsequently used for establishing the gold standard. Drions-db has 110 images with 2 manual markings for each image. For each image, the gold standard was defined from a contour that was the result of averaging two contours, each of them traced by an expert. Optic disc and cup segmentation are both important parts of ONH segmentation and together they form a base for glaucoma assessment.
4.3 Modification of ground-truth
Optic disc and cup boundaries do not have a steep slope. Instead, the pixel intensities taper off linearly as one moves radially away from the centre of the Optic nerve head (ONH) region. As a result, datasets consist of boundaries marked by more than one expert and this is due to the uncertainty about the exact boundary of optic disc and cup as shown in Fig. 4. It depicts two such images where boundaries were marked by five different experts (Fig. 4a) and by four different experts as shown in Fig. 4b.
There is no one precise boundary but the disc and cup tend to fade over the range of a few pixels. This can be clearly seen in the surface plot of a sample image, which is shown in Fig. 5. The ground truths are formed from averaging these multiple markings and then binarizing the resulting image. In this proposed work, this uncertainty is addressed, and the ground truths have been modified as probability masks instead of binary masks and then are used to train the network.
The provided ground truths are modified in a manner such that, a boundary pixel is assigned a probability instead of a binary value. For this purpose, the circumcircle of the multiple markings is found out and pixels ranging from 0.9R to 1.1R (where ‘R’ is the radius of the circum circle) are assigned certain probabilities. ‘r’ is the radius of the incircle. The circumcircle and the ring created from the circle is illustrated in Fig. 6. It is assumed that all the pixels that are equidistant from the optic disc centre, lie on the circumference of a circle of a specific radius having similar intensity values. The probabilities assigned are such that the value linearly decreases from 0.9R to 1.1R in small steps and varies in the range of (1,0).
The modification of the ground truth (for multiple markings) can be understood through Algorithm-1.

In order to avoid confusion, the modified ground truth will be referred to as GTmod and actual ground truth as GT. The average ground truth GT and the modified ground truth GTmod is depicted in Fig. 7.
4.4 Convolution feature learning
Convolution filter is that part of the network which convolves the features from the input data to extract intrinsic patterns hidden in the image which are then used for further classification. Neurons of these layers are randomly initialized in order to provide adequate distinctions and then are further optimized with the help of the errors over various iterations. Loss functions also referred as cost functions; are used to calculate the error and optimizers are used to improve those weights utilizing the calculated error. Hence the optimum choice of loss function is very important for any model. We have tried to modify the existing evaluation measure (Overlapping Score or Intersection-over-Union) to create a custom loss function [Eq. (3)].
Preprocessed images discussed in Section 3 are fed to the network with GTmod as the labels; via the data generator which performs the data augmentation. Data augmentation includes width and height shifts, horizontal and vertical flips, zooming within range of 20% and random rotations in the range of 0o to 90o; which is performed in order to avoid the over fitting of the model. 500 images were trained in one epoch with a batch size of 1, due to the hardware constraints and the model is trained for 30 such epochs. For each epoch, the model is validated on 200 test images. The training of the model was stopped after 30 epochs, since the improvement in validation loss of the network was minimal. And moreover after 50 epochs, the nature of the graph of validation loss was far different from the ideally expected one. As mentioned above, the customized loss function is used for the purpose of error calculation. The main idea behind designing the custom loss function was to create a loss function for 2-D labels which considers the probabilities predicted by the network. The original IOU (Intersection-Over-Union) function is designed such that it accepts only 0 s and 1 s as the input values; whereas the modification of the function accepts all values ranging between [0, 1]. This permits the network to calculate the gradient of the function and hence ensures back propagation. The idea was inspired by the work done in [47]. The logarithm of modified IOU function was chosen to be minimized for the purpose of optimization. Eq. (1) and (5) represents the modified IOU function and the loss function, l(A, B) of the network.
Where A is the predicted probability map and B is modified ground truth mask. \( ^{\prime }{a}_i^{\prime}\kern0.5em E\kern0.5em \left[0,1\right] \), represents the pixel values of predicted segmentation map A and ′bi ′ Є [0, 1], represents the pixel values of set B respectively.
Once the result after the first iteration is predicted, it is compared with GTmod and error is calculated with the help of the loss function. This error is then back propagated, and weights are modified in accordance with the optimization function. These weights are then improvised, iteration over iteration for a certain number of epochs. After that, the model starts overfitting and starts memorizing features of the training images and model will not give impressive results on test image datasets. The weights of the network are updated using Adam function as the optimizer with a learning rate of (1e-5) and experiments have also been performed for a learning rate of (1e-3). The features learned by the convolution filters are represented by the output of the filter, given a certain input. The output of the filters shows an interesting pattern as the corresponding layer proceeds towards the final output. Where the initial filters tend to highlight the edges of the foreground object, the contour can easily be seen developing in the filters of final layers. The output of the convolution filter of the first layer and second last layer is shown in Figs. 8, and 9.
4.5 Final output segmentation
After completion of training the network, the fine-tuned model was used for testing on the test datasets. The last layer of the network is a 2D convolution layer with kernel size (1, 1) and only one feature map so that the resulting image is dimensionally equivalent to a probability map. But the resulting image is not actually a probability map, as the result of final convolution could be anything varying in the range (−∞, ∞). For the sake of converting the resulting image into the desired probability map, the sigmoid activation function was applied after the last layer of the network. The sigmoid function limits the input values to a range of [0, 1].
Once the probability map is obtained, the next task was to create the contour of that probability map. To accomplish that, the simple step function was applied to the probability map. The function applied is expressed in eq. (3).
Where ‘a’ is the pixel value of the probability map and ‘u’ is the pixel value of the final segmented contour. Figure 10 depicts the probability map of a sample test image and the contour obtained from the probability map.
5 Results analysis
We have tested the proposed algorithm on three different datasets: DRISHTI-GS dataset [53, 54], RIM-ONE v3 dataset [40] and DRIONS-DB [7] dataset. The former two datasets are tested for optic disc segmentation as well as optic cup segmentation, whereas the third one is only tested for the optic disc segmentation since the ground truth for the optic cup is not provided in the dataset. The DRISHTI-GS dataset is separated into training and testing subsets. The training set consists of 50 images whereas the test set comprises of 51 images. The ground truth for both optic disc and cup are provided within the dataset along with the notch information. The resolution of the images is 2896 × 1944 which is centered at OD and is provided in PNG format. The ground truth available is collected from experts possessing a clinical experience of 3 to 20 years. The third release of the RIM-ONE dataset [Retinal Image database for Optic Nerve Evaluation] comprises 159 fundus images, which are further classified as healthy and glaucoma suspects. All the images are provided along with optic disc and cup notch information from two different expert ophthalmologists. Our experimentation includes all the images provided, out of which 80% are used for the training purpose while 20% is used for testing. The images are shuffled properly before dividing into train and test subsets. DRIONS-DB dataset entails 110 fundus images, for which two expert annotations of the optic disc is provided. The images are centered at the optic disc and have a resolution of 600 × 400. Similar to the RIMONE, this dataset is first shuffled and then divided into training and testing subsets in a ratio of 8:2.
5.1 Evaluation metrics
The evaluation measures for the testing phase are given by Dice Coefficient (DC) and the Intersection over Union (IOU). Dice coefficient of two images P and Q can be seen as the harmonic mean of precision and recall, which are expressed in eq. (4) and (5).
Where ‘tp’ are true positives, ‘fn’ are false positives and ‘fn’ are false negatives of the image P in accordance with Q. The dice coefficient is also referred as F-score and is represented in eq. (6). The value of the dice coefficient varies in the range of [0,1].
Intersection Over Union (IOU) is calculated as the ratio of the intersection of two binary maps of predicted output and ground truths to the union of these images. The IOU function results in the range of [0,1] and is also designated as the Jaccard index or overlapping score. Eq. (7) shows the exact formulation of IOU.
The predicted optic disc contours were originally non-binary masks which have been binarized so that evaluation measures like Intersection-over-Union (IOU) and Dice Coefficient (DC) could be used for comparison with papers in existing literature. However, in recent times, researchers have come up with several new measures to evaluate the accuracy of non-binary foreground maps with respect to binary ground truths. Evaluation Measures such as IOU and DC are based on pixel-wise errors. These ignore the structural similarities of the foreground map. Evaluation measures like weighted Fβ – measure [36], S-measure [15] and E-measure [14] consider the underlying structure of foreground maps and compare it with ground truth masks to evaluate the similarity between both the masks. So, these measures were also considered in evaluation of optic disc and cup contours.
Previous measures like IOU and DC assumes that the pixels of the foreground map are independent, which is not true. Weighted Fβ – measure tries to capture this interdependency between foreground pixels. It also assigned weights (importance) to false positive pixels based on the distance of these pixels from the foreground maps.
where \( {\mathrm{Precision}}^{\mathrm{w}}=\frac{TP^w}{TP^w+{FP}^w}\kern1em \mathrm{and}\kern0.5em {\mathrm{Recall}}^{\mathrm{w}}=\frac{TP^w}{TP^w+{FN}^{w.}} \)
S-Measure or Structure – measure takes into consideration, the object structure of the predicted foreground mask to evaluate the similarity with the ground truth mask.
where So and Sr are object-aware and region-aware structural similarity measures respectively. Value of α has been kept at 0.5.
E-Measure or the Enhanced-alignment measure accounts for both the pixel level and image level properties and hence is suited for efficient evaluation of foreground maps.
where ϕ is the enhanced alignment matrix.
5.2 Segmentation results
The proposed algorithm utilizes the power of the encoder-decoder network which is trained on three different datasets. The encoding part of the network mainly comprises of the convolutional layers which are used to find different patterns in the image. These features are then fed to the respective deconvolutional layer from the decoding part of the network. The decoder of the network actually differentiates between the foreground and the background objects.
In this section, the model is being trained and tested in the same mode. These results are reported for the image resolution of 128×128 and learning rate (1e-5). The results reported are compared with GT of the image and not GTmod; as the actual ground truth of the image is GT and the purpose of creating GTmod was only to train the model. Finally predicted contours of some images from these three datasets are shown in Fig. 11.
The best and worst segmentation results for the optic disc and cup, in terms of IOU with regard to DRISHTI-GS dataset are shown in Fig. 12.
The proposed method is compared with some of the state-of-the-art results which are reported in Maninis et al. [34] (It will be further referred as DRIU), Artem [47] and Zilly et al. [61]. The proposed work is implemented in Keras framework [10] with Tensorflow [1] backend and for the implementation of adaptive histogram equalization, the CLAHE implementation of skimage library is used. The algorithms are implemented in Python 3.6 on a system with 8 GB RAM and GPU MX 150. Table 1 depicts the segmentation results for the optic disc on all three reported datasets whereas Table 2 depicts the results for the optic cup segmentation.
The segmentation of the optic cup is comparatively more difficult task than the segmentation of the optic disc. This might be due to the low contrast of the optic cup compared to its background (i.e. optic disc region) as the pixel values fade away in a gradual manner with a gentle slope.
Evaluation measures like weighted Fβ – measure [36], S-measure [15] and E-measure [14] were also used on the Drishti-GS test dataset to see the effectiveness of the algorithm for optic disc and cup segmentation (Table 3).
Experiments were conducted on 51 test images of the dataset. For optic disc segmentation, while average weighted Fβ-Measure obtained is 0.9261, average S-Measure obtained is 0.9285 and average E-measure is 0.8361 respectively. For optic cup segmentation, though the E-measure was high, comparable to IOU and DC values, weighted Fβ -Measure and S-measure values were less impressive.
5.3 Comparative analysis
The resolution of the input image can have considerable influence on the computational time and might affect the overall accuracy as well. While a smaller image size can lead to the loss of necessary information; a very large size can heavily increase the computation time. Hence the choice of the appropriate image size has always been important to finalize the model. We have tested our algorithm with three different image sizes: 128×128, 256×256 and 512×512. All the experiments are done for the task of optic disc segmentation to evaluate its importance. An epoch with 500 training images and batch size 1 takes approximately half an hour for a resolution of 128×128. It was seen that in our problem, increasing the image size resulted in increased computation time but it did not lead to increased segmentation accuracy. Hence, image size of 128 × 128 was finalized for the model.
Learning rate is a hyper parameter which is responsible for the amount of change being made to the weights with respect to the loss gradient. And hence a small change in learning rate can sometimes significantly alter the results. While a smaller learning rate can make the model wait forever to reach global minima, a high learning rate may overshoot and skip the minima point. The algorithm is tested for two different learning rates i.e. (1e-5) and (1e-3). The results for optic disc segmentation are tabulated in Table 4 below. It was seen that, a learning rate of (1e-3) gave better results for optic disc segmentation in comparison to a learning rate of (1e-5). Also, learning rate of (1e-5) was better suited for optic cup segmentation on all three datasets.
As part of experimentation, we have also tried to see the effect of inpainting on model accuracy. We have removed the blood vessels from the original images using the inpainting technique described in [39]. Then the inpainted images were fed into the model as input. It was seen that inpainting of images had no mentionable influence on the model accuracy of deep neural networks unlike in traditional methods.
Another important experiment which was conducted was to check the feasibility of using transfer learning. In transfer learning, weights fine-tuned on a pretrained network are used for modeling another task. In most applications, networks trained on Imagenet are often used for transfer learning. This is usually done due to scarcity of data for the present application, in our case, glaucoma images. But if datasets are available for the concerned application, concept of transfer learning may be used to fine tune the weights of the base model [52, 59]. In the present case of glaucoma images, datasets belong to a similar application. The images though differ in image specifications, image acquiring modality and certain conditions. Training the model every single time when it is required to be tested on a new dataset requires a lot of time and effort. Moreover, considering the practical scenarios, the new real-time image is actually not part of any of the dataset and may vary in the specifications. Therefore, transfer learning makes the architecture robust and suitable for real-time applications. Also, it eliminates the overheads of designing customized features for the segmentation problem
Results using transfer learning suggest that optic disc segmentation can achieve appreciable performance. A network trained on few hundred glaucoma images from different datasets can be used to predict the optic disc boundaries for images in a new dataset. For example, optic disc segmentation on DRIONS-DB dataset achieved a model accuracy of 93.76% when the model is trained on DRISHTI-GS dataset. Similarly, the segmentation results for DRISHTI-GS dataset are comparable (95.89%) when trained on DRIONS-DB. The model accuracy may increase if the number of glaucoma images used for training the model is large. The results obtained for optic cup segmentation though, as observed after experimentation via transfer learning, were very poor. This might be attributed to the variation in the size of the optic cup in the datasets. The number of normal cases available in DRISHTI-GS training dataset is less, in other words the number of images with smaller optic cup are less in DRISHTI-GS dataset.
Algorithm results have been compared in particular with four recent methods in literature. Artem [47] have proposed a modified Unet in his paper and further proposed a stack-Unet in a recent paper [48] and reported the findings on these three datasets for optic disc and cup segmentations. Maninis et al. [34], in their paper proposed a Deep retinal image understanding (DRIU) model and reported results on Rim-One and Drions-DB datasets. Zilly on the other hand reported his findings in two papers [61, 62] on Drishti and Rim-One datasets. It can be seen that the proposed algorithm fared well if not significantly better for optic disc segmentation task on all three datasets. With regard to Optic cup segmentation, proposed algorithm gave significantly high overlap accuracy on Drishti dataset [53, 54] (Table 5).
5.4 Threats to validity
Threats to validity play an important role for any experiment. The experimental results should be validated for the population. Threat to external validity bound the results under the experimental settings. To generalize the results and to overcome the threat three benchmarked datasets- DRISHTI-GS dataset [53, 54], RIM-ONE v3 dataset [40] and DRIONS-DB [7] dataset is used, so that generalize, and realistic results can be achieved. Threat to internal validity mainly focuses of independent variables. To overcome this threat in our experiment images is selected in ration of 80:20 for training and testing respectively. The images are shuffled before dividing into training and testing sets. Threat to conclusion validity mainly affects the ability to present the correct conclusion. In our case Dice coefficient and Intersection over union performance parameters is utilized to overcome this threat.
6 Conclusions
In this paper, we have presented a mathematical model to segment the boundaries of optic disc and optic cup in retinal fundus images with improved accuracy. This will ultimately aid in calculating parameters like Cup-to-Disc ratio (CDR), Neuro Retinal Rim Area etc. which are essential for accurate glaucoma analysis, classification and subsequent grading of the disease. The model helped us to improve upon the existing optic disc and cup segmentation results on three different dataset: DRISHTI-GS, DRIONS-DB and RIM-ONE v3. The proposed method aimed at finding accurate boundaries of optic disc and optic cup using deep neural networks. We have also tried to see whether the proposed model works on different datasets formed and acquired in varied conditions. But there were some images where the optic cup was small or on images in which the contrast between optic disc and its background was low, the proposed algorithm produced poor results. This is where salient point detection algorithms [60] could be put to good use along with CNNs to improve upon the model accuracy. Also relevant are context aware saliency algorithms [21, 35] which could be of tremendous value not only to build ground truths but also could provide complementary information in building a robust model.
References
Abadi, M et al. (2015) TensorFlow: Large-scale machine learning on heterogeneous systems. http.tensorflow.org/.
Abramoff MD, Garvin MK, Sonka M (2010) Retinal imaging and image analysis. IEEE Rev Biomed Eng 3:169–208
Acharya UR, Mookiah MRK, Koh JEW, Tan JH, Noronha K, Bhandary SV, Rao AK, Hagiwara Y, Chua KC, Laude A (2016) Novel risk index for the identification of age-related macular degeneration using radon transform and DWT features. Comput Biol Med 73:131–140
Agarwal A, Issac A, Singh A, Dutta MK (2016) Automatic imaging method for optic disc segmentation using morphological techniques and active contour fitting, Ninth International Conference on Contemporary Computing (IC3), Noida, pp. 1–5
Bajwa MN et al. (2019) Correction to: Two-stage framework for optic disc localization and glaucoma classification in retinal fundus images using deep learning (BMC Medical Informatics and Decision Making 19 (136)
Bock R et al (2010) Glaucoma risk index: automated glaucoma detection from color fundus images. Medical image analysis 14.3:471–481
Carmona EJ, Rincón M, García-Feijoo J, Martínez-de-la-Casa JM (2008) Identification of the optic nerve head with genetic algorithms. Artificial Intelligence in Medicine 43(3):243–259
Chai Y, He L, Mei Q, Liu H, Xu L (2017) Deep learning through two-branch convolutional neural network for glaucoma diagnosis, International Conference on Smart Health, pp. 191–201
Chakravarty, Sivaswamy J (2016) Glaucoma classification with a fusion of segmentation and image-based features, IEEE 13th International Symposium on Biomedical Imaging (ISBI), Prague, pp. 689–692
Chollet F (2015) Keras: Deep learning library for theano and tensorflow. https://keras.io/k. Accessed 20 Aug 2019
Ciresan DC, Gambardella LM, Giusti A, Schmidhuber J (2012) Deep neural networks segment neuronal membranes in electron microscopy images, NIPS. pp. 2852–2860
Damms T, Dannheim F (1993) Sensitivity and specificity of optic disc parameters in chronic glaucoma. Investigative Ophthalmology and Visual Science 34(7):2246
Diaz-Pinto A, Morales S, Naranjo V, Köhler T, Mossi JM, Navea A (2019) CNNs for automatic glaucoma assessment using fundus images: an extensive validation. Biomed Eng Online 18(1):1–19
“Enhanced-alignment Measure for Binary Foreground Map Evaluation”, Proceedings of the 27th International Joint Conference on Artificial Intelligence (IJCAI), 2018.
Fan D-P, Cheng M-M, Liu Y, Li T, Borji A (2017) Structure-measure: a new way to evaluate foreground maps, ICCV, pp. 4550–4557
Fu H, Cheng J, Xu Y, Zhang C, Wong DWK, Liu J, Cao X (2018) Disc-aware ensemble network for Glaucoma screening from fundus image. IEEE Trans Med Imaging 37(11):2493–2501
Fukushima K (1980) Neocognitron: A self-organizing neural network model for a mechanism of pattern recognition unaffected by shift in position. Biol. Cyb. 36:193–202
Gao X, Cai Y, Qiu C, Cui Y (2017) Retinal blood vessel segmentation based on the Gaussian matched filter and U-net" ,10th International Congress on Image and Signal Processing, BioMedical Engineering and Informatics(CISP-BMEI),Shanghai,pp.1–5
Ghoshal R, Saha A, Das S (2019) An improved vessel extraction scheme from retinal fundus images. Multimed Tools Appl 78(18):25221–25239
Glaucoma Research Foundation, “Five common glaucoma tests”, https://www.glaucoma.org/glaucoma/diagnostic-tests.php, 2017.
Goferman S, Zelnik-Manor L, Tal A (2010) Context-aware saliency detection, Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit, pp. 2376–2383
Guo S, Wang K, Kang H, Zhang Y, Gao Y, Li T (2019) BTS-DSN: Deeply supervised neural network with short connections for retinal vessel segmentation, Int. J. Med. Inform., vol. 126, no. January, pp. 105–113
Hagiwara Y, Koh JEW , Tan JH, Bhandary SV, Laude A, Ciaccio EJ, Tong L, Rajendra Acharya U (2018) Computer-aided diagnosis of Glaucoma using fundus images: a review, Computer Methods and Programs in Biomedicine
Hsu HY, Srivastava G, Wu HT, Chen MY (2020) Remaining useful life prediction based on state assessment using edge computing on deep learning. Comput Commun 160:91–100
Huazhu F, Cheng J, Xu Y, Wong DWK, Liu J, Cao X (2018) Joint optic disc and cup segmentation based on multi-label deep network and polar transformation. IEEE Transactions on Medical Imaging (TMI) 37(7):1597–1605
Issac A, Sengar N, Singh A, Sarathi MP, Dutta MK (2016) Automated computer vision method for optic disc detection from non-uniform illuminated digital fundus images, 2nd International Conference on Communication Control and Intelligent Systems (CCIS), Mathura,pp.76–80
Jiang Y, Tan N, Peng T (2019) Optic disc and cup segmentation based on deep convolutional generative adversarial networks. IEEE Access 7:64483–64493
Kausu TR, Gopi VP, Wahid KA, Doma W, Niwas SI (2018) Combination of clinical and multiresolution features for glaucoma detection and its classification using fundus images, Biocybernetics and Biomedical Engineering
Koh JEW, Acharya UR, Hagiwara Y, Raghavendra U, Tan JH, Sree SV, Bhandary SV, Rao AK, Sivaprasad S, Chua KC, Laude A, Tong L (2017) Diagnosis of retinal health in digital fundus images using continuous wavelet transform (CWT) and entropies. Comput Biol Med 84:89–97
Koh JEW, Ng EYK, Bhandary SV, Laude A, Acharya UR (2017) Automated detection of retinal health using PHOG and SURF features extracted from fundus images, Applied Intelligence, Springer US, pp. 1–15
Krizhevsky A, Sutskever I, Hinton GE (2012) ImageNet Classification with Deep Convolutional Neural Networks, Neural Information Processing Systems. 25. https://doi.org/10.1145/3065386
LeCun Y, Bottou L, Bengio Y, Haffner P (1998) Gradient-based learning applied to document recognition, IEEE 86 (11), pp. 2278–2324
Lim G, Cheng Y, Hsu W, Lee ML (2015) Integrated Optic Disc and Cup Segmentation with Deep Learning, IEEE 27th International Conference on Tools with Artificial Intelligence (ICTAI), Vietrisul Mare,pp.162–169
Maninis K-K, Pont-Tuset J, Arbel’aez P, Van Gool L (2016) Deep retinal image understanding, International Conference on Medical Image Computing and Computer-Assisted Intervention, pp. 140–148, Springer
Margolin R, Tal A, Zelnik-Manor L (2013) What makes a patch distinct?, Proc. IEEE Comput. Soc. Conf. Comput. Vis. Pattern Recognit., pp. 1139–1146
Margolin R, Zelnik-Manor L, Tal A (2014) How to evaluate foreground maps?, CVPR, pp. 248–255
Marsden J Glaucoma: The “silent thief of sight”, Nursing Times, 110, 42
Medeiros FA (2019) Deep learning in glaucoma: progress, but still lots to do, The Lancet Digital Health, vol. 1, no. 4. Elsevier Ltd, pp. e151–e152, 01-Aug
Partha Sarathi M, Dutta MK, Singh A, Travieso CM (2016) Blood vessel inpainting based technique for efficient localization and segmentation of optic disc in digital fundus images. Biomedical Signal Processing and Control 25:108–117
Pena-Betancor C, Gonzalez-Hernandez M, Fumero-Batista F, Sigut J, Mesa E, Alayon S, de la Rosa MG (2015) Estimation of the relative amount of hemoglobin in the cup and neuro-retinal rim using stereoscopic color fundus images, IOVS, pp. IOVS–14–15592
Phasuk S et al. (2019) Automated Glaucoma screening from retinal fundus image using deep learning, Proc Annu Int Conf IEEE Eng Med Biol Soc EMBS, pp. 904–907
Quigley HA, Green WR (1979) The histology of human glaucoma cupping and optic nerve damage: clinical correlation in 21 eyes. Ophthalmology 86(10):1803–1830
Raghavendra U, Bhandary SV, Gudigar A, Acharya UR (2017) Novel expert system for glaucoma identification using non-parametric spatial envelope energy spectrum with fundus images, Biocybernetics and Biomedical Engineering
Ronneberger O, Fischer P, Brox T (2015) U-Net: Convolutional Networks for Biomedical Image Segmentation
Ronneberger O, Fischer P, Brox T Dental X-ray Image Segmenation using a U-shaped Deep Convolutional Network
Sadhukhan S, Ghorai GK, Maiti S, Karale VA, Sarkar G, Dhara AK (2018) Optic disc segmentation in retinal fundus images using fully convolutional network and removal of false-positives based on shape features, Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support, pp. 369–376
Sevastopolsky A (2017) Optic Disc and Cup Segmentation Methods for Glaucoma Detection with Modification of U-Net Convolutional Neural Network, Pattern Recognition and Image Analysis. 27. https://doi.org/10.1134/S1054661817030269
Sevastopolsky A , Drapak S , Kiselev K, Snyder BM , Keenan JD, Georgievskaya A (2019) Stack-U-net: refinement network for improved optic disc and cup image segmentation, Medical Imaging: Image Processing
Shelhamer E, Long J, Darrell T (2017) Fully Convolutional Networks for Semantic Segmentation, IEEE Transactions on Pattern Analysis and Machine Intelligence, vol. 39, no. 4, pp. 640–651, 1 April
Shin H-c, Roth H, Gao M, Lu L, Xu Z, Nogues I , Yao J, Mollura D , Summers R (2016) Deep Convolutional Neural Networks for Computer-Aided Detection: CNN Architectures, Dataset Characteristics and Transfer Learning. IEEE Transactions on Medical Imaging. 35. 10.1109/TMI.2016.2528162
Simonyan K, Zisserman A (2014) Very Deep Convolutional Networks for Large-Scale Image Recognition, arXiv 1409.1556
Singh A, Sengupta S, Lakshminarayanan V (2019) Glaucoma diagnosis using transfer learning methods, in Applications of Mach Learn vol. 11139, p. 27.
Sivaswamy J, Krishnadas SR, Chakravarty A, Dutt G, Joshi U, Syed TA (2015) A Comprehensive Retinal Image Dataset for the Assessment of Glaucoma from the Optic Nerve Head Analysis. JSM Biomedical Imaging Data Papers 2(1):1004
Sivaswamy J, Krishnadas KR, Joshi GD, Madhulika J, Ujjwal and Syed Abbas T. Drishti-GS: Retinal Image Dataset for Optic Nerve Head(ONH) Segmentation, IEEE ISBI, Beijing
Srivastava R, Cheng J, Wong DWK, Liu J (2015) Using deep learning for robustness to parapapillary atrophy in optic disc segmentation, IEEE 12th International Symposium on Biomedical Imaging (ISBI), New York,NY,pp.768–771
Tham YC, Li X, Wong TY, Quigley HA, Aung T, Cheng CY (2014) Global prevalence of glaucoma and projections of glaucoma burden through 2040: a systematic review and meta-analysis. Ophthalmology 121(11):2081–2091
Xu Y, Duan L, Wong D, Kee W, Wong TY, Liu J (2015) Glaucoma detection by learning from multiple informatics domains., Proceedings of the Ophthalmic Medical Image Analysis Second International Workshop, OMIA
Xu Y; Duan L; Wong D, Kee W, Wong, Yin T, Liu J (2015) Glaucoma detection by learning from multiple informatics domains, Proceedings of the Ophthalmic Medical Image Analysis Second International Workshop, OMIA
Yosinski J, Clune J, Bengio Y, Lipson H (2014) How transferable are features in deep neural networks?, Advances in Neural Information Processing Systems 27 (NIPS ‘14), NIPS Foundation
Zhao J-X, Liu J-J, Fan D-P, Cao Y, Yang J-F, Cheng M-M (2019) EGNet: Edge Guidance Network for Salient Object Detection, In Proceedings of the IEEE International Conference on Computer Vision, pp. 8779–8788
Zilly J, Buhmann JM, Mahapatra D (2015) Glaucoma detection using entropy sampling and ensemble learning for automatic optic cup and disc segmentation. Computerized Medical Imaging and Graphics 55:28–41
Zilly JG, Buhmann JM, Mahapatra D (2015) Boosting convolutional filters with entropy sampling for optic cup and disc image segmentation from fundus images, in International Workshop on Machine Learning in Medical Imaging, pp. 136–143, Springer
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mangipudi, P.S., Pandey, H.M. & Choudhary, A. Improved optic disc and cup segmentation in Glaucomatic images using deep learning architecture. Multimed Tools Appl 80, 30143–30163 (2021). https://doi.org/10.1007/s11042-020-10430-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11042-020-10430-6