Abstract
Chlorophyll biosynthesis is catalyzed by two multi subunit enzymes; a light-dependent and a light-independent protochlorophyllide oxidoreductase. The light-independent enzyme consists of three subunits (ChlL, ChlN and ChlB) in photosynthetic bacteria and plastids in which the chlB gene encodes the major subunit that catalyzes the reduction of protochlorophyllide to chlorophyllide. We report here stable integration of the chlB gene from Pinus thunbergii into the chloroplast genome of tobacco. Using helium-driven biolistic gun, transplastomic clones were developed in vitro. The stable integration and homoplasmy for transgenes was confirmed by using PCR and Southern blotting techniques. Nodal cuttings of the homoplasmic transgenic and untransformed wild type shoots were cultured on MS medium in the dark. As expected, shoots developed from the cuttings of the wild type plants in the dark showed etiolated growth with no roots whereas shoots from the cuttings of the transgenic plants developed early and more roots. Upon shifting from dark to light in growth room, leaves of the transgenic shoots showed early development of chlorophyll pigments compared to the wild type shoots. Further, photosynthetically indistinguishable transgenic shoots also showed significant difference in root development from untransformed wild type shoots when cuttings were grown in the light. Therefore, it may be concluded that the chlB gene is involved, directly or indirectly, in the root development of tobacco. Further, the gene promotes early development of chlorophyll pigments, upon illumination from dark, in addition to its role in the light-independent chlorophyll formation when expressed together with subunits L&N in other organisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Photosynthesis is the strategic process in which harvesting and transduction of light energy is carried out by the chlorophyll. Chlorophyll biosynthesis is catalyzed by two multi subunit enzymes; a light dependent protochlorophyllide oxidoreductase, LPOR and a light-independent protochlorophyllide oxidoreductase (dark-operative DPOR). During chlorophyll biosynthesis, protochlorophyllide (PChlide) is reduced to chlorophyllide (Chlide), which is then immediately converted into chlorophyll a. Manipulation of this important compound will therefore have major impact on the capability of plants to carry out photosynthesis [1].
During evolution two genetically and biochemically different strategies of PChlide reduction have been developed [2]. At first PChlide is reduced by nuclear-encoded, plastid-localized oxidoreductase, which requires NADPH and light for its activity. However, in many phototrophic organisms there is another way for chlorophyll synthesis in which PChlide is reduced by a plastid-encoded enzyme complex, independent of light. These two systems can co-exist and play their roles in different ontogenetic stages of plant and in dependence on environmental factors [3]. Genetic and sequence analysis have indicated that DPOR consist of three subunits (ChlL, ChlN and ChlB) in photosynthetic bacteria and plastids [4]. All organisms containing this pathway for PChlide reduction are capable of chlorophyll formation in the dark. The chlB is one of three chloroplast genes shown so far to be required for light-independent chlorophyll synthesis. The chlB is present in the chloroplast genome of the gymnosperms, algae and photosynthetic bacteria [5]. Whereas this gene is absent in chloroplast genomes of tobacco and rice, consistent with the lack of light-independent chlorophyll synthesis in these plants.
The chlB encodes subunit B of light independent protochlorophyllide reductase that catalyzes reduction of PChlide to Chlide in chlorophyll biosynthesis. The complete nucleotide sequence of Pinus thunbergii (Japanese Black Pine) chloroplast genome contains, as in other chloroplast genomes, the chlL and chlN genes in the SSC region, creating an operon; the chlB gene is located separately in LSC region [6]. It encodes a protein of 510 amino acids which was immunochemically detected in membrane fractions of cyanobacteria, showing either cytoplasmic or thylakoid membranes as the site of light-independent reduction of PChlide [7]. However, disruption of the chlB from chloroplast genome, plastome of Chlamydomonas reinhardtii resulted in the development of yellow phenotype of mutants in the dark which was indistinguishable from mutants for chlL or chlN, suggesting that chlB is essential for the activity of DPOR in chloroplasts [8].
Genetic engineering of chloroplast genome was first accomplished in C. reinhardtii, a unicellular green alga [9] followed by chloroplast transformation in Nicotiana tabacum, a flowering plant species [10]. Since then chloroplast transformation has been extended to a diverse group of species due to its very attractive and potential advantages for the gene expression in the plants [11]. These advantages include high level transgene expression [12], absence of gene silencing and position effect variation, the ability to express genes polycistronically from a single promoter [13] and gene containment in most crop plants.
Here we describe the development of stable transgenic plants of tobacco harboring chlB, which have produced vigorous root system, suggesting that the chlB gene is involved, directly or indirectly, in the development of plant roots in addition to its role in the light-independent accumulation of chlorophyll.
Materials and methods
Plant material and growth conditions
Seeds of N. tabacum L. var. Petit Havana were aseptically grown on MS medium solidified with 0.026 % phytagel [14] at pH 5.8 at 27 °C under 100 μmol photons m−2 s−1 (16 h light, 8 h dark conditions). Fully expanded 4–6 weeks old dark green leaves were used in chloroplast transformation experiments.
Genomic analysis of putative transplastomic clones for site specific integration and homoplasmy
Total genomic DNA was extracted from both transformed as well as untransformed wild type tobacco plants by using hexadecyltrimethyl ammonim bromide (CTAB) method with modifications [15]. The isolated DNA was then used as template in polymerase chain reaction performed using a Master Cycler Gradient (Eppendorf, Germany). The integration of transformed cassette was verified by PCR analysis on spectinomycin resistant and wild-type tobacco plants by using gene specific primers to aadA (A19: 5′-GGC TCC GCA GTG GAT GGC GGC CTG-3′; A20: 5′-GGG CTG ATA CTG GGC CGG CAG G-3′) and flanking sequences used for homologous recombination (S19: 5′-GAT ATC AAA ACC CGT CCT CAG TTC GGA TTG C-3′ and S20: 5′-GAT ATC CAC GAG TTG GAG ATA AGC GGA-3′). The PCR program was set for 30 cycles at 95 °C for 2 min. 56 °C for 2 min. and 72 °C for 3 min. with a step of 72 °C for 10 min. The amplified products were separated on 0.8 % agarose gel after staining with ethidium bromide and visualize under UV gel documentation system (VilberLourMat. France).
Site-specific integration and homoplasmy was also investigated through Southern blot analysis. In these experiments, ApaI restricted 10–15 μg genomic DNA was separated on 1 % agarose gel at 60 voltage for 3–4 h. After the washing of the gel with different solutions the DNA was transferred to cellulose nitrate membrane (Schleicher and Schuell, Bioscience Germany) using trans-blotter (Bio-Rad, Hercules, Calif.). The flanking sequence probe (0.81 kb) was prepared by digesting F/TA plasmid with BamHI/BglII, which contains the chloroplast flanking sequences trnI and trnA. The marker gene specific probe (522 bp) of aadA was developed by PCR amplification from FLARE-S by using gene specific primers (A19/A20). Following hybridization of membrane with probe, the detection was carried out using Fermentas Biotin Chromogenic Detection Kit (MBI Fermentas, Italy).
Evaluation of transplastomic plants for photosynthetic efficiency
Physiological parameters as photosynthesis rate, stomatal conductance, transpiration rate and water use efficiency were measured from fully expanded dark grown leaves of both wild type as well as transplastomic plants growing in pots using LCA-UADC portable infrared gas analyzer (Analytical Development Company, Hoddesdon, England). Data was recorded between 11:30 A.M to 12:15 P.M using following parameters: leaf surface area 6.6 cm2, ambient CO2 concentration 1050.5 ppm, temperature of the leaf chamber varied from 35.5 to 42.2 °C, leaf chamber gas flow rate 392.8 ml min−1, Molar flow of the air per leaf surface (Us) 404.84 mol m−2 s−1, Ambient chamber ranged from 20.5 to 23.1 m bar and photosynthetic active radiation (PAR) (Qlaef) at leaf surface was maximum up to 1048 μmol m−2 s−1 [16].
Determination of the chlorophyll contents
In order to check the evidence that the presence of this novel photosynthetic gene will affect the chlorophyll contents, leaves from 3-weeks-old plants of chlB transgenic and wild type tobacco plants were ground in 80 % acetone [17] and the absorbance was determined using spectrophotometer (JENWAY, Labo Med., Inc.) at wavelengths 663, 645 and 480 nm.
Results
Development of chloroplast transformation vector harboring chlB gene
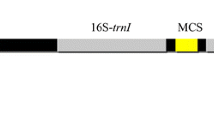
The tobacco chloroplast transformation vector was developed by PCR amplification of inverted repeat regions from tobacco plastome by using primers S19: 5′-GAT ATC AAA ACC CGT CCT CAG TTC GGA TTG C-3′ and S20: 5′-GAT ATC CAC GAG TTG GAG ATA AGC GGA-3′ [18] and the PCR product was cloned into TA cloning vector (MBI Fermentas, Italy). The cloned fragment was then restricted with EcoRV, which was engineered in both primers. The fragment was then ligated into pBluescript II (MBI Fermentas, Italy) cut with PvuII restriction enzyme. A HincII/SmaI fragment carrying MCS (i.e. ClaI, HindIII, EcoRV, EcoRI and PstI) was introduced into the PvuII in tobacco flanking sequences to facilitate the cloning steps [18]. For the screening of transformation events FLARE-S (Fluorescent Antibiotic Resistance Enzyme, Spectinomycin and Streptomycin) marker was used [19]. Chloroplast encoded chlB was PCR amplified using gene specific primers S4/S5 and cloned into TA cloning vector (MBI Fermentas, Italy). A strong constitutive promoter prrn from plastid RNA ribosomal operon was cloned upstream of the chlB by using NheI/KpnI restriction sites. The resultant clone (Fig. 1a) was then confirmed with a series of restriction enzymes and cloned at PstI site in the MCS of tobacco chloroplast transformation vector upstream to the marker gene. The orientation of the cassette in the final tobacco chloroplast transformation vector was verified by a number of enzymes as well as by Polymerase Chain Reaction using different primer combinations.
Confirmation of the integration of chlB transformed cassette with marker gene into inverted repeat region of the chloroplast genome of tobacco and also check homoplasmy level. a Physical map of transgenic plant plastid genomes with primers positions. b PCR amplification of marker gene (aadA) with primer sets A19/A20: lane 1 PCR −ve control, lane 2 +ve control with plasmid DNA, lane 3 untransformed tobacco plant DNA, lanes 4, 5 and 6 are transformed tobacco plant DNA and lane 7 1-kb DNA ladder c Amplification of gene of interest (chlB) by using gene specific primers S4/S5: lane 1, 1-kb DNA ladder, lane 2 PCR −ve control, lane 3 +ve control with plasmid DNA, lane 4 non transformed tobacco plant DNA, lanes 5, 6 and 7 are transformed tobacco plant DNA d, e and f Gels pictures with PCR-amplified left and right border sequence fragments with primer sets A19/S20, S19/A20 and S19/S20 respectively for the confirmation of site specific integration and homoplasmy. Lanes 1 and 8, 1-kb DNA ladder, lane 2 PCR −ve control, lane 3 +ve control with plasmid DNA, lane 4 wild type plant DNA, 5, 6 and 7 are transformed plant DNA
Chloroplast transformation and development of transgenic plants harboring the chlB gene
Chloroplast transformation was performed by using biolistic technology [18–20] with modifications. Green shoots appeared within 4–6 weeks of bombardment on bleached leaf sections on spectinomycin containing RMOP [19]. Multiplying transformed cells and the regenerating shoots on selection medium were regularly inspected with a hand-held long-wave UV lamp for gfp fluorescence. Transformation was confirmed by an Olympus SZX (Olympus SZX9, Japan) stereomicroscope equipped for gfp detection with a CCD camera system. The images produced by gfp fluorescence were viewed on a computer screen attached to the microscope.
Purification of antibiotic resistant transgenic clones
The initial screening is very important for eliminating the mutants and nuclear transformation events from the transplastomic plant population. Hence two approaches namely; GFP fluorescence and PCR detection were used to screen the transgenic plants form non-transplastomic plants. For the screening and confirmation of the chloroplast transgenic plants from the spontaneous spectinomycin resistant plants as well as escapees by using PCR method, total genomic DNA was isolated from putative transgenic tobacco plants and also from wild type by CTAB method [15, 21] and used this DNA as template in the PCR reactions. The presence of the selection marker gene (aadA) in the putative transgenic tobacco plants was detected by PCR using the primers A19/A20 and only three representative PCR positive clones are shown (Fig. 1b). Primer set S4/S5 specific for chlB (Fig. 1c) and primers S19/S20 representing the flanking region used for site specific integration. The wild type tobacco plant DNA used in the experiment served as negative control in the PCR analysis with no amplification. Site specific chloroplast integration of the transgenic cassette was determined by using a set of primers of which one anneals to the native chloroplast genome and other anneals within the transgenic cassette. Mutants and other nuclear transformants were not expected to produce a PCR product with these primers. The following pair of the primers was used A19/S20 and S19/A20 [18]. The amplification of fragments of 2.4 and 4.1 kb (Fig. 1d, e respectively) with these primers eliminated the possibility of spontaneous mutants, escapes as well as nuclear transformants when expression cassette integrate into the nuclear genome due to illegitimate recombination events from transgenic population. For the confirmation of the homoplasmicity, primer set S19/S20 specific to flanking region was used. All plants resulting in the amplification of 6.1 kb along with the 2.2 kb fragment specific for untransformed wild type control plant as shown in Fig. 1F confirmed the plasmy level of transgenic plants. These results described the heteroplasmic nature of the transplastomic clones, and hence it demands further rounds of selection and regeneration to achieve homoplasmy. Another strategy which was also being used for screening of transformed plants from the untransformed ones was by using the visible reporter genes in addition of selection marker genes. The basic idea was to distinguishing transformed regenerants or shoots in a heterogeneous population. Biolistic delivery of chlB transformation vector containing FLARE-S as a selection marker [19] was carried out. Transplastomic sectors in the chimeric tissues can be identified visually, thus significantly reducing the time and effort required to obtain genetically stable transplastomic lines. The expression of gfp yielded fluorescent shoots at an early age, making the reporter gene useful for early detection of transformation events. Tissue sections of 0.5 by 0.5 cm size when exposed to blue or ultra violet light, gfp was emitting bright green fluorescence (Fig. 2). The sizes of the fluorescent sectors were different in different leaves in heterozygous transgenic clones depending upon the segregation of the transformed cells from wild type non-transformed cells. Thus regeneration of transgenic plants was greatly facilitated by visual identification of the fluorescent sectors. No toxic effect of the gfp was observed and lack of toxicity was supported by the apparently normal phenotype of the plants. The heteroplasmic plants were further subjected to selection and regeneration.
Plasmy level of shoots regenerated from second round of selection and regeneration was confirmed through Southern blotting, using selectable marker gene as well as flanking sequences as a probe (Fig. 3a). Three independent transgenic clones were used to extract genomic DNA. Restriction of genomic DNA with ApaI restriction enzyme results in two fragments, one of about 5.5 kb, carrying the marker gene (aadA) and another of ~2.5 kb having the expression cassette. Wild type tobacco plant produced only one fragment of ~4.2 kb (Fig. 3b). Upon hybridization with aadA probe only the transformed plants showed band whereas no fragment was observed in wild type tobacco plants (Fig. 3c). These results were confirmed the successful integration of transformed cassette into tobacco plants. In order to evaluate the plastome integration as well as for the assessment of homoplasmic state, another strategy was opted for southern blotting. Upon achieving homoplasmy, all of the chloroplast genomes were supposed to contain the integrated transgene cassette hence is identical. When 0.81 kb BamHI/BglII fragment containing chloroplast-flanking sequences was used as probe, the chlB transformed plants produced ~5.5 and 2.5 kb fragments with ApaI restricted plant genomic DNA whereas untransformed chloroplasts showed only one fragment of ~4.2 kb (Fig. 3d). Wild type band was absent in transplastomic lines, indicating that regenerated plants are homoplasmic for transgene integration into the genome of the chloroplasts.
Southern-blot hybridization analysis to determine integration of transformed cassette into tobacco plants and homoplasmy. a The 522 bp fragment specific to selection marker gene and 0.81 kb flanking sequence fragment that were used as probes for Southern-blot analysis. b Untransformed tobacco plant DNA digested with ApaI (expected fragment size, 4.2 kb) and chlB transformed tobacco plant DNA also digested with ApaI (expected fragments size, 5.3 and 2.5 kb). c Southern blot results by using marker gene as probe; WT, untransformed tobacco plant DNA, TR1, TR2 and TR3 are transformed tobacco plants with chlB gene construct. d Southern blot analysis by using flanking sequences as probe; WT, untransformed tobacco plant DNA, TR1, TR2 and TR3 are transformed tobacco plants with chlB gene construct
Physiological performance of engineered chlB transgenic plants
The chlB gene is reported to play a significant role in photosynthesis and is thought to be of pivotal importance and involved in the formation of chlorophyll precursor i.e. protochlorophyllide [2] thus, chlorophyll contents of chlB transformed tobacco plants were determined. Shoots of homoplasmic transgenic as well as of wild type plants multiplying on MS medium were used in these experiments. Cuttings carrying one node on each segment of same age plants were cultured in both the light or total darkness, and their pigment contents were analyzed by spectrophotometer. Photosynthesis related parameters were investigated by using infra-red gas analyzer (IRGA) on cuttings grown in dark for 14 days and then illuminating for 24 h. Representative data from three independent clones are shown in Table 1. The data show that the chlorophyll a, chlorophyll b and total chlorophyll contents for dark-grown transgenic and wild type shoots were essentially equivalent. Similarly, non-significant difference in the chlorophylls contents of transgenic and wild type shoots growing in the light for 15 days were observed. Nevertheless, the contents were significantly varied under dark and light regimes for both genotypes. Further, early pigment development, within 2–3 h illumination, in transgenic shoots was observed compared to non-transgenic wild type shoots (Fig. 4a, b). In Fig. 4a abaxal and 4B adaxal side of the transgenic leaf is exposed to the camera.
Apart from chlorophyll contents, stomatal conductance, respiration rate; net photosynthesis and water use efficiency of both transgenic and wild type plants were measured and no significant differences between the transgenic lines compared to non-transformed wild type plants were observed. Hence, it was concluded from the experiments that chlB subunit promotes early chlorophyll pigment development rather affects total contents of chlorophyll, which requires integration of all three genes involved in the pathway.
Phenotypic analysis of the engineered chlB transgenic plants
While shoots were growing on MS medium in dark etiolated growth of wild type shoots compared to the transplastomic tobacco plants, expressing chlB gene was observed. Further, the transgenic shoots produced early (within 7 days; Fig. 5a) and vigorous roots (after14 days; Fig. 5b) compared to wild type under dark. Similarly, transgenic shoots produced early and vigorous roots compared to the wild type when grown in the day length (16 h light and 8 h dark) under controlled temperature. Data pertaining to number of primary roots per plant under dark as well as light, number of secondary roots per plant under dark as well as light, root length (cm) under dark as well as light was recorded (Table 2). Data was recorded, after every 7 days of cultivation of cuttings under dark and light, taking two consecutive readings at the same time. In Table 2 means of three observations were statistically analyzed and significant differences were found between chlB transgenic and wild type tobacco plants. This experiment was repeated more than three times and almost similar results were observed. Further, it was observed that root induction in the transgenic shoots was initiated within 3–4 days of culture whereas in wild type after 5–6 days of culture incubated under light conditions. The experiment was repeated several times in the dark and the data (average of three readings) was recorded after 7 and 14 days (Fig. 5c, d). Data recorded from dark- and light grown plants have shown that the ChlB subunit, in addition to PChlide reduction, possibly have some direct or indirect role in root development.
Discussion
Photosynthesis, the most widespread biochemical process on the earth, is the key process affecting crop yield and biomass production [1]. Possible options to improve photosynthesis include; regulating stomatal conductance, reducing respiration rate, engineering the Rubisco enzyme, modification of C3 plants into C4 and/or by increasing photosynthetic pigment contents [12, 22]. Most of these aspects have been well studied and attempted to engineer but no significant achievement has been made. Among all these, photosynthetic pigments are thought to be the pivotal targets in so far as enhancement of photosynthetic productivity is concerned. Chlorophyll a and b are major contributors of light harvesting process along with carotenoids and phycobilins. In chlorophyll biosynthetic pathway, the reduction of PChlide to Chlide is very crucial step and two different enzymes involved in this reduction process are: LPOR and DPOR [2]. DPOR is encoded by three chlorophyll genes (chlL, chlN and chlB) encoded by chloroplast genome of gymnosperms and photosynthetic microbes but absent in angiosperms, the salient cereals and fruit plants. We attempted to increase chlorophyll contents in order to boost up light harvesting ability of tobacco. One of the genes encoding DPOR, chlB was isolated from the plastid genome of Pinus thunbergii, a Japanese Black Pine and then biolistically integrated into the tobacco plastome. The stable integration of the chlB gene along with marker gene/s was confirmed by Polymerase Chain Reaction and Southern blot hybridization techniques. Though transgenic shoots showed early pigment development upon illumination from dark but their photosynthetic performance was not significantly better from wild type plants, suggesting that the gene alone does not influence the photosynthetic response of the plants but requires integration of other relevant genes leading to the completion of dark chlorophyll biosynthetic pathway.
RNA-editing efficiency of chlB gene is developmentally regulated and differs from species to species in larch cotyledons [23], suggesting that the gene may have more secretes in addition to be an essential part of DPOR. It has also been reported that photoautotrophically grown cells of Chlorella protothecoides accumulate less chlB in comparison with heterotrophically and mixotrophically grown cells [24]. In addition, chlB gene participates in the synthesis of molybdopterins (a class of cofactors found in most molybdenum and all tungsten enzymes) and molybdopterin guanine dinucleotide (MGD) in Escherichia coli. The biochemical analysis of the chlB mutants had shown that its product is essential for the addition of the GMP moiety to form MGD, a step which occurs late in the cofactor biosynthetic pathway in E. coli. Wild type cells contained both molybdopterin and MGD, while cells of chlB mutants contained only high levels of molybdopterin but no MGD [25]. These research findings have shown that chlB gene; in addition to be a core part of dark protochlorophyllide oxidoreductase (DPOR) subunit, also regulates certain other developmental processes. Stable integration of chlB gene into the tobacco plastome has revealed that the gene is involved in the root development since more roots appeared on transgenic shoots compared to wild type both under dark and light regimes. Further, both wild type and transgenic plants were acclimatized under normal growth conditions and indistinguishable phenotype was observed (Fig. 6). Hence, the present research findings may be regarded as a step forward to engineer plants to improve root system for better performance.
References
Willows RD (2003) Biosynthesis of chlorophylls from protporphyrin IX. Nat Prod Rep 20:1–16
Armstrong GA (1998) Greening in the dark: light-independent chlorophyll biosynthesis from anoxygenic photosynthetic bacteria to gymnosperms. J Photochem Photobiol B Biol 43:87–100
Fujita Y, Bauer CE (2000) Reconstitution of light-independent protochlorophyllide reductase from purified BchL and BchN–BchB subunits. J Biol Chem 275:23583–23588
Fujita Y, Takagi H, Hase T (1996) Identification of the ChlB gene product essential for light-independent chlorophyll biosynthesis in the cyanobacterium Plecronema boyanum. Plant Cell Physiol 37:313–323
Li J, Goldschmidt-Clermont M, Timko P (1993) Chloroplast-encoded ChlB is required for light-independent protochlorophyllide reductase activity in Chlamydomonas reinhardtii. Plant Cell 5:1817–1829
Wakasugi T, Tsudsuki J, Ito S, Nakashima K, Tsudsuki T, Sugiura M (1994) Loss of all ndh genes as determined by sequencing the entire chloroplast genome of the black pine Pinus thunbergii. Proc Natl Acad Sci USA 91:9794–9798
Nomata J, Ogawa T, Kitashima M, Inou K, Fujita Y (2008) NB-protein (BchN-BchB) of dark-operative protochlorophyllide reductase is the catalytic component containing oxygen-tolerant Fe–S clusters. FEBS Lett 582:1346–1350
Liu X-Q, Xu H, Huang C (1993) Chloroplast ChlB is required for light independent chlorophyll accumulation in Chlamydomonas reinhordhi. Plant Mol Biol 23:297–308
Boynton JE, Gillham NW, Harns EH, Hosler JP, Johanson AM, Jones AR, Randolph-Anderson BL, Robertson O, Klein TM, Shark KB, Sanford JC (1988) Chloroplast transformation in Chlamydomonas with high-velocity microprojectiles. Science 240:1534–1538
Svab Z, Hajdukiewlcz P, Maliga P (1990) Stable transformation of plastids in higher plants. Proc Natl Acad Sci USA 87:8526–8530
Wang H, Yin W, Hu Z (2009) Advances in chloroplast engineering. J Genet Genomics 36:387–398
Bock R, Khan MS (2004) Taming plastids for a green future. Trends Biotechnol 22:311–318
Daniell H, Khan MS, Allison L (2002) Milestones in chloroplast genetic engineering: an environmentally friendly era in biotechnology. Trends Plant Sci 7:84–91
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Rogers SO, Bendich AJ (1985) Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Biol 5:69–76
Wheatherley PE (1950) Studies in water relations of the cotton plant. I: the field measurement of water deficit in leaves. New Phytol 49:81–97
Arnon DI (1949) Copper enzymes in isolated chloroplasts. Polyphenoloxidase in Beta vulgaris. Plant Physiol 24:1–15
Khan MS, Hameed W, Nozoe M, Shiina T (2007) Disruption of the psbA gene by the copy correction mechanism reveals that the expression of plastid-encoded genes is regulated by photosynthesis activity. J Plant Res 120:421–430
Khan MS, Maliga P (1999) Fluorescent antibiotic resistance marker for tracking plastid transformation in higher plants. Nat Biotechnol 17:910–915
Svab Z, Maliga P (1993) High-frequency plastid transformation in tobacco by selection for a chimeric aadA gene. Proc Natl Acad Sci USA 90:913–917
Spychalla J, Bevan MW (1993) Agrobacterium mediated transformation of potato stem and tuber tissues, regeneration and PCR screening for transformation. Plant Tissue Cult Man 11:1–18
Khan MS (2007) Engineering photorespiration in chloroplasts: a novel strategy for increasing biomass production. Trends Biotechnol 25:437–440
Demko V, Pavlovib A, Valkova D, Slovakova L, Grimm B, Hudak J (2009) A novel insight into the regulation of light-independent chlorophyll biosynthesis in Larix decidua and Picea abies seedlings. Planta 230:165–176
Shi C, Shi X (2006) Expression switching of three genes encoding light-independent protochlorophyllide oxidoreductase in Chlorella protothecoides. Biotechnol Lett 28:261–265
Johnson JL, Indermaur W, Rajagopalan KV (1991) Molybdenum cofactor biosynthesis in Escherichia coli. Requirement of the ChlB gene product for the formation of molybdopterin guanine dinucleotide. J Biol Chem 266:12140–12145
Acknowledgments
This research work was supported by the Higher Education Commission (HEC) Islamabad, Pakistan. Authors also thank Ministry of Science and Technology (MoST) for awarding projects to MSK at NIBGE.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nazir, S., Khan, M.S. Chloroplast-encoded chlB gene from Pinus thunbergii promotes root and early chlorophyll pigment development in Nicotiana tabaccum . Mol Biol Rep 39, 10637–10646 (2012). https://doi.org/10.1007/s11033-012-1953-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-1953-9