Abstract
The Bowman–Birk (BBI) protease inhibitors can be used as source of sulfur amino acids, can regulate endogenous protease activity during seed germination and during the defense response of plants to pathogens. In soybean this family has not been fully described. The goal of this work was to characterize in silico and analyze the expression of the members of this family in soybean. We identified 11 potential BBI genes in the soybean genome. In each one of them at least a characteristic BBI conserved domain was detected in addition to a potential signal peptide. The sequences have been positioned in the soybean physical map and the promoter regions were analyzed with respect to known regulatory elements. Elements related to seed-specific expression and also to response to biotic and abiotic stresses have been identified. Based on the in silico analysis and also on quantitative RT-PCR data it was concluded that BBI-A, BBI-CII and BBI-DII are expressed specifically in the seed. The expression profiles of these three genes are similar along seed development. Their expressions reach a maximum in the intermediate stages and decrease as the seed matures. The BBI-DII transcripts are the most abundant ones followed by those of BBI-A and BBI-CII.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Protease inhibitors are essential for regulating protease activity during seed germination, they are involved in plant response to insects and pathogens and in addition may function as storage of sulfur amino acids [1, 2]. In soybeans there are two main families of protease inhibitors: the Kunitz trypsin inhibitors (KTI) and the Bowman–Birk (BBI) trypsin–chymotrypsin inhibitors.
BBI, originally described in soybean by Bowman [3] and Birk [4], consists of a 71 amino acid polypeptide chain containing seven disulfide bonds [5]. These small proteins function as pseudosubstrates for several serine proteases. Their structure typically presents two independent protease ligation sites which are able to ligate and inhibit trypsin, chymotrypsin and elastase [6]. The complex formed by the interaction between the inhibitor and the enzyme substantially limits the proteolytic activity [7].
The genes encoding BBI in Glycine max and G. soja form a multigene family with at least five members: BBI-A, BBI-B, BBI-CII, BBI-DII and BBI-EI. The inhibitory specificity and the primary sequence of BBI-B are very similar to those of BBI-A and supposedly BBI-B is encoded by a gene which is closely related to BBI-A, designated BBI-A2. BBI-EI originates from BBI-DII by proteolysis. As a consequence, the BBI members have been grouped in three classes—A, C and D—with distinct specificities; BBI-A, -C and -D inhibit trypsin/chymotrypsin, elastase/trypsin, and trypsin/trypsin, respectively [8].
Identification, isolation and functional analyses of soybean genes have been speeded up significantly after the soybean genome sequence became available (http://www.phytozome.net/soybean). According to the soybean genomic model (Glyma1), approximately 975 Mb have been organized in 20 chromosomes and 66,153 loci potentially encoding proteins have been predicted [9]. This information will be important to study the organization and regulation of genes related with yield, seed quality, nitrogen fixation, and response to environmental modifications [10].
The main goal of this work was to identify, characterize in silico and analyze the expression of the members of the multigene family encoding BBI in soybean.
Materials and methods
Identification and in silico characterization
To identify the genes encoding BBI the algorithm blastp was used against the modelled proteome data of annotated genes in Glyma1 (the proteome predicted by the soybean genome sequence (Glyma1)) with an E-value threshold ≤1e−5. Fourteen BBI protein sequences (queries) from nine species (G. max, G. microphylla, G. soja, Oryza sativa, Phaseolus vulgaris, Pisum sativum, Triticum aestivum, Vigna unguiculata, and Zea mays) were used.
The search for signal peptide sequences was performed with the software SignalP (http://www.cbs.dtu.dk/services/SignalP/). PREDOTAR (http://urgi.versailles.inra.fr/predotar/predotar.html) and TargetP 1.1 (http://www.cbs.dtu.dk/services/TargetP/) were used to determine the most probable subcellular localization of the predicted proteins. The conserved domains were identified with InterProScan and InterPro DB (www.ebi.ac.uk/interpro). To determine the presence of conserved cysteine residues which are important to maintain the structure of the inhibitor and to determine BBI typical domains the deduced amino acid sequences, not including the possible signal peptides, were aligned using ClustalW (http://www.ebi.ac.uk/Tools/clustalw2) and Superfamily (http://supfam.cs.bris.ac.uk/SUPERFAMILY/).
All genes identified were positioned and anchored in the physical map based on the consensus soybean genome sequence available at SoyBase (http://soybeanphysicalmap.org/index.php) [11].
The software SignalScan [12] and database PLACE [13] (http://www.dna.affrc.go.ja/PLACE) were used to identify regulatory cis-elements along the putative promoter regions encompassing 1,500 bp upstream of the theoretical +1 translation site.
To get preliminary information on the expression of the BBI encoding genes, the predicted sequences of their transcripts were used to search (blastn) the GenBank EST data base (http://www.ncbi.nlm.nih.gov/nucest/9205464?dopt=genbank). Alignments with an E-value ≤1e−100 were considered significant matches.
Gene isolation and sequencing
To isolate and sequence the BBI genes, seed genomic DNA of cultivar CAC-1 was extracted according to a procedure previously described [14] and specific primer sets for each BBI putative ORF were designed. PCR reactions were carried out in a thermocycler MasterCycler (Eppendorf, Hamburg, Germany) according to the following program: one initial denaturation step at 94°C for 2 min, followed by 35 cycles at 94°C for 30 s/55°C for 30 s/72°C for 30 s, and a final extension step at 72°C for 4 min. A 10 μL aliquot from each reaction was purified with ExoSAP-IT® (USB, Cleveland, OH, USA) according to the manufacturer’s recommendations and sequenced at Macrogen (Seoul, South Korea).
Plant material and RT-PCR
The expression of the BBI genes was analyzed in roots, stems, leaves and seeds of soybean cultivar CAC-1. The seeds were collected along eight developmental stages based on their fresh weight: 1st—0 to 75 mg; 2nd—76 to 150 mg; 3rd—151 to 225 mg; 4th—226 to 300 mg; 5th—301 to 375 mg; 6th—376 to 450 mg; 7th—451 to 525 mg and 8th—mature seeds.
Total RNA was extracted with phenol:chloroform [15], and treated with RQ1 RNase-Free DNase (PROMEGA, Madison, WI, USA). Synthesis of the first cDNA strand was primed by oligo d(T) using a SuperScript First Strand-Synthesis System (Invitrogen, Grand Island, NY, USA) according to the manufacturer’s recommendations.
The primer-pairs used for isolation and sequencing of the BBI genes were also used in conventional RT-PCR reactions. The amplification reaction conditions were also the same except that the genomic DNA was replaced by cDNA from root, stem, leaves and immature seeds. The soybean actin gene was used as internal control.
The quantitative expression of the BBI genes was determined along the stages of seed development by RT-PCR (qRT-PCR). The specific primer-pairs were designed with the aid of the program Primer Express 3.0 (Applied Biosystems, Foster City, CA, USA). The reactions were carried out in a thermocycler 7500 Real Time PCR System (Applied Biosystems) using SYBR Green PCR Master Mix (Applied Biosystems) according to the manufacturer’s recommendations. A negative control without a cDNA template was run with each analysis to evaluate the overall specificity. The glyceraldehyde 3-phosphate dehydrogenase gene (GAPDH) was used as endogenous reference to normalize the data and for determination of the relative abundance of the transcripts through the 2−ΔCt method [16]. Specificity and reaction efficiency of each primer set were evaluated by a dissociation curve and a standard curve, respectively. All experimental samples were run in triplicate (technical replicates) with three biological replicates for each gene.
Results and discussion
To identify potential BBI encoding genes, 14 BBI protein sequences from different species were blasted (blastp) against the predicted soybean proteome of annotated genes in Glyma1. This analysis revealed the existence of 11 putative BBI encoding genes in the soybean genome, all of them lacking introns: Glyma09g28700, Glyma09g28720, Glyma09g28730, Glyma09g39630, Glyma09g39640, Glyma14g26400, Glyma14g26410, Glyma16g33400, Glyma18g46550, Glyma18g46560 and Glyma18g46580.
Six of these potential genes encode type A inhibitor (Glyma09g28700, Glyma09g28730 and Glyma14g26410), type C-II (Glyma09g28720 and Glyma14g26400) and type D-II (Glyma16g33400), which have already been described in the literature [8, 17]. Type A BBI genes have been divided into two groups, A1 and A2. These two groups differ by 11 base substitutions which lead to modifications in two amino acid residues. In addition, BBI-A1 genes present a characteristic HindIII restriction site [8]. Based on these differences it was concluded that Glyma09g28700 and Glyma09g28730 encode BBI-A1, and Glyma14g26410 encodes BBI-A2. It is interesting to notice that three genes encoding BBI-A have been originally identified [8]. However, as only two cDNA could be isolated for this gene, the authors concluded that the third one would not be transcribed or that it could be an identical copy of one of the two cDNAs. Indeed, our data allowed us to conclude that the third gene is a copy of BBI-A1.
We also identified two genes encoding BBI-CII (Glyma09g28720 and Glyma14g26410) and one encoding BBI-DII (Glyma16g33400) as previously described [8]. The difference between the two genes encoding BBI-CII resides on three substitutions, two of them are silent but the third one leads to the replacement of an arginine by a histidine. Sequence alignment showed that the transcript encoded by Glyma09g28720 (BBI-CII) is identical to the sequence reported in the literature [17]. There were no reports in the literature about the second transcript. Two transcripts (GmBBI1 and GmBBI2) which are expressed in soybean roots upon interaction with cyst nematodes have been cloned [18]. Alignment of these two cDNAs with transcripts annotated in the soybean genome shows that they are identical to genes Glyma18g46560 and Glyma18g46550. The other sequences we detected (Glyma18g46580, Glyma09g39630 and Glyma09g39640) have not been described in the literature.
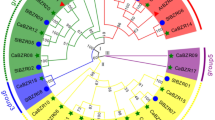
Soybean BBIs typically present two sites which simultaneously inhibit trypsin/chymotrypsin (type A), elastase/trypsin (type C-II) and trypsin/trypsin (type D-II) [8]. In addition, BBI structure is maintained by seven disulfide bonds between conserved cysteine residues [19]. The known BBI sequences present these conserved residues and the two characteristic inhibition sites. However, the novel sequences we detected showed more variability in relation to amino acid composition. As the proteins encoded by these sequences have not been isolated one cannot state that these variations affect their structure and/or function. In relation to the inhibition domains, sequences Glyma09g39640, Glyma18g46550 and Glyma18g46580 presented only one of these typical domains. The second site was not conserved (Fig. 1).
A potential signal peptide was predicted in the N-terminal of all 11 BBI sequences described indicating that either BBI is synthesized and stored in specialized vacuoles (PSVs—protein storage vacuoles), as described for the 11S and 7S soybean seed storage proteins [20], or exported to the intercellular space. Analysis of the soybean embryonary axis and cotyledons demonstrated that BBI may be found in PSVs, in the nucleus and in the cytosol to a minor extent [21]. The KTI was localized in cell walls, protein bodies, the cytoplasm between the lipid-containing spherosomes, and the nucleus of the cotyledon and embryonic axis [22].
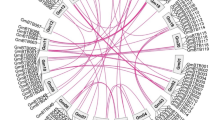
To map the BBI genes we used the soybean consensus map which is available in the USDA database SoyBase. Five of the genes were mapped to chromosome 09 (linkage group (LG) K). Glyma09g28700, Glyma09g28720 and Glyma09g28730 were positioned between SSR markers Sat_043 and Satt 273; and Glyma09g39630 and Glyma09g39640 were flanked by Satt 196 and Satt 588. Glyma16g33400 was mapped to chromosome 16 (LG J) between Satt 441 and Sat_144. Glyma14g26400 and Glyma14g26410 were mapped to chromosome 14 (GL B2) flanked by Satt 318 and Satt474. Glyma18g46550, Glyma18g46560 and Glyma18g46580 were mapped to chromosome 18 (GL G) between Satt 288 and Satt612. Although the molecular confirmation is still necessary, this is the first time the BBI genes are positioned in the soybean physical map (Fig. 2).
A great number of cis-acting elements have been identified in promoter regions that control a series of biological processes including response to biotic and abiotic stresses and plant development [23]. To characterize the promoter region of the BBI genes we analyzed approximately 1,500 bp upstream of the putative +1 translation site for each of the potential genes in search for known regulatory motifs.
Besides the TATA-box and CAAT-box, cis-elements related to transcriptional activation, seed-specific expression and response to biotic and abiotic stress have been found in the BBI promoters analyzed (Table 1). The following seed-specific cis-elements and elements regularly found in genes encoding seed-storage proteins were found: CANBNNAPA, DPBFCOREDCDC3, EBOXBNNAP, RYPEATBNNAPA, SEF3MOTIFGM, and SEF4MOTIFGM7S.
We also found eight elements—DOFCOREZM, GT1GMSCAM4, MYB1A, MYCATERD1, MYCCONSENSUSAT, T/GBOXATPIN2, WBOXARNPR1, WBOXNTERF3—related to response to biotic and abiotic stress. Among them are the sites for ligation of transcription factors WRKY, MYB and DNA binding with one finger (DOF). WRKY proteins constitute a transcription factor family that has been detected only in plants. Members of this family are induced by pathogen attack, defense signaling and wounding [24]. The MYB family responds to stresses such as UV light, wounding, anaerobiosis and pathogens [25, 26]. The DOF family members are induced by salycylic acid and interact and stimulate the ligation activity of bZIP proteins to DNA which are also responsive to stress conditions [27]. It is noteworthy that in rice the expression of a BBI gene is induced by wounding, jasmonate and ethylene [28], which together with salycylic acid are considered signals that induce the defense response in plants.
Some of the mentioned elements have not been detected in all promoter regions analyzed. The element CANBNNAPA (CNAACAC) involved with seed-specific expression was only present in the genes encoding BBI-A (Glyma09g28700, Glyma28730, and Glyma14g26410), BBI-D (Glyma16g33400) and in a putative BBI gene (Glyma46580) identified in this work. In addition, the promoter regions of one of the genes encoding BBI-C (Glyma14g26400) and the gene Glyma18g46580 encoding a putative BBI do not present a TATA-box and the former does not contain the RY element which is important for seed-specific expression. The absence of these elements suggests that they are not expressed, however, these observations are not enough to show that they are pseudogenes.
To complement the in silico analysis of the promoter region, a virtual expression profile, similar to a virtual northern blotting (VNB), was determined for each gene. The representativity in the GenBank EST data base and the tissue-specific expression of each transcript were evaluated. The transcripts encoding the inhibitors BBI-A, BBI-CII and BBI-DII have been found in expression libraries built from immature cotyledons. The only exception was a hit detected for the BBI-CII transcript in a somatic embryo library. The genes Glyma18g46550 and Glyma18g46560 have been detected mainly in roots submitted to nodulation, pathogen or dehydration. These transcripts correspond to those isolated from roots infected with cyst nematode. In this work the authors determined that the expression of Glyma18g46560 increases 4.5 times in susceptible plants infected with cyst nematode when compared to resistant plants [18]. The last three genes, Glyma09g39630, Glyma09g39640 and Glyma18g39640 have been detected in only one library each: somatic embryo, root submitted to biotic and abiotic stress and seedlings infected with Fusarium solani, the causative agent of Sudden Death Syndrome, respectively. In relation to representativity the most abundant transcript was encoded by Glyma16g33400 (104 hits) followed by those encoded by Glyma09g28700, Glyma09g28730 and Glyma14g26410 (55 hits), Glyma18g46560 (27 hits), Glyma18g46550 (24 hits), Glyma09g28720 and Glyma14g26400 (20 hits).
As a whole, the data obtained from the in silico promoter analysis and the virtual expression profile give support to the idea that BBI protects the plant against herbivores [2] and can function as storage proteins to be used during seed germination [29].
After the in silico analysis, we isolated the 11 putative BBI genes and determined their expression by RT-PCR. As all the genes were devoid of introns we designed primers flanking the ORFs and used soybean genomic DNA to isolate the genes. All amplicons had the predicted sizes and they were sequenced (data not shown). Alignment of the sequences with the soybean genome showed only small differences (identities varying from 96.1 to 99.7%) which might be due to differences between the cultivar we used (cv. CAC-1) and the one used to sequence the whole soybean genome (cv. Williams 82).
The organ-specific expression of each BBI gene was determined by conventional RT-PCR using cDNA from leaf, stem, root and seed. The expression of BBI-A, -CII and -DII genes was seed-specific (Fig. 3), while the expression of the other genes was not detected in these organs (data not shown).
As the genetic material we used was collected under normal conditions, this expression pattern was expected and is in line with the results obtained by the in silico analysis. In dry beans, the expression of two BBI genes has been analyzed in several organs and detected only in developing cotyledons [30]. In addition, Birk [31] reported that 1 week old soybean seedlings presented all types of protease inhibitors in cotyledons and only a small amount in the hypocotyl. In another work, BBI was detected mainly in cotyledons of 12-day-old seedlings [32]. However, a small amount was detected in epicotyls, hypocotyls, and roots. In mature soybean plants, Goldberg [33] observed BBI synthesis only in cotyledons.
The expression of the genes encoding the main BBI types in the seed (BBI-A, -CII and -DII) was evaluated by qRT-PCR during seed development. A specific primer-pair was designed to amplify each type of gene. In the case of BBI-A, a primer-pair was designed to amplify both BBI-A1 and BBI-A2. Only one primer-pair was also used to amplify the two BBI-CII genes. The primer-pairs presented amplification efficiency higher than 90% and each amplified one fragment only. The expression of the three genes was extremely high along seed development, except in the mature seed stage. The transcript levels increase in the initial developmental stages and reach maximum expression at the third stage and reduces thereon. Essentially no transcripts were detected in mature seeds (Fig. 4).
The transcripts for BBI-DII were the most abundant ones in all stages of seed development, followed by BBI-A and finally BBI-CII. These results are in line with those obtained by the in silico expression analysis (Fig. 5). The transcripts for BBI-A, BBI-CII and BBI-DII have been found in seed cDNA libraries, mainly in immature cotyledons (100–300 mg). This size range overlaps the 3rd stage of development (151–225 mg) as defined in the present work. In this stage, we observed the highest BBI transcript levels in vivo, BBI-DII being the most abundant ones.
It is noteworthy that the promoter region of the gene Glyma16g33400 (BBI-DII) possesses three copies of the RY cis-element, which is very important for seed-specific expression. Two of these copies are close to the TATA box. The BBI-A gene presents one copy of this element and BBI-CII harbors two RY copies approximately at 1,200 bp upstream of the TATA box. It is conceivable that the copy number and the position of these elements contribute to determine the expression levels of the BBI genes in the seed. In this sense, a systematic combinatorial in silico analyses of cis-elements and analyses of expression profiles in Arabidopsis indicated that there was a positive correlation between gene response to stimuli and cis-element density in the promoter region [34]. Although BBI-A has been considered the most abundant BBI type in soybean seeds [19], another study reported that BBI-EI is the most abundant type [35]. This last report is in line with our findings considering that BBI-EI derives from a post-translational modification of BBI-DII [8]. BBI-CII seems to be the least abundant BBI type in soybean seeds [17].
In summary, the soybean genome presents 11 loci that potentially encode BBI. Among them six encode type A, -CII and -DII inhibitors which are seed-specific. The other sequences are present in cDNA libraries from tissues that have been submitted to stress. Their expression has not been detected under normal cultivation conditions. The expression profiles of the genes encoding seed BBI follow a similar pattern along seed development, however, transcripts for BBI-DII are the most abundant followed by those of BBI-A and BBI-CII.
References
Mayer AM, Shain Y (1974) Control of seed germination. Annu Rev Plant Physiol 25:167–193
Ryan CA (1990) Proteinase inhibitors in plants: genes for improving defenses against insects and pathogens. Annu Rev Phytopathol 28:425–429
Bowman DE (1946) Differentiation of soybean anti-tryptic factors. Proc Soc Exp Biol Med 63:547–550
Birk Y (1961) Purification and some properties of a highly active inhibitor of trypsin and α-chymotrypsin from soybean. Biochim Biophys Acta 54:378–380
Chen J, Mao SJ, Xie Y (2006) Expression and inhibitory activity analysis of a 25-kD Bowman–Birk protease inhibitor in rice. Chin Sci Bull 51:54–62. doi:10.1007/s11434-005-0937-8
Fernadez JH, Mello MO, Galgaro L, Tanaka AS, Silva-Filho MC, Neshich G (2007) Proteinase inhibition using small Bowman–Birk-type structures. Genet Mol Res 6:846–858
Tiffin P, Gaut BS (2001) Molecular evolution of the wound-induced serine protease inhibitor wip1 in Zea and related genera. Mol Biol Evol 18:2092–2101
Deshimaru M, Yoshimi S, Shioi S, Terada S (2004) Multigene family for Bowman–Birk type proteinase inhibitors of wild soja and soybeans: the presence of two BBI-A genes and pseudogenes. Biosci Biotechnol Biochem 68:1279–1286. doi:10.1271/bbb.68.1279
Schmutz J, Cannon SB, Schlueter J et al (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183. doi:10.1038/nature08670
Mochida K, Yoshida T, Sakurai T, Yamaguchi-Shinozaki K, Shinozaki K, Tran L-SP (2009) In silico analysis of transcription factor repertoire and prediction of stress responsive transcription factors in soybean. DNA Res 16:353–369. doi:10.1093/dnares/dsp023
Grant D, Nelson RT, Cannon SB, Shoemaker RC (2010) SoyBase, the USDA-ARS soybean genetics and genomics database. Nucleic Acids Res 38:D843–D846. doi:10.1093/nar/gkp798
Pestridge DS (1991) SIGNAL SCAN: a computer program that scans DNA sequences for eukaryotic transcriptional elements. Comput Appl Biosci 7:203–206. doi:10.1093/bioinformatics/7.2.203
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res 27:297–300. doi:10.1093/nar/27.1.297
MacDonald MB, Elliot IJ, Sweeney PM (1999) DNA extraction from dry seeds for RAPD analyses in varietal identification studies. Seed Sci Technol 22:171–176
Sambrook J, Frietsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor, New York
Livak KJ, Schmittgen TD (2001) Analysis of relative expression data using real-time quantitative PCR and the 2−ΔΔCt. Method 25:402–408. doi:10.1006/meth.2001.1262
Baek J-M, Song J-C, Choi YD, Kim SI (1994) Nucleotide sequence homology of cDNAs encoding soybean Bowman–Birk type proteinase inhibitor and its isoinhibitors. Biosci Biotechnol Biochem 58:843–846
Rashed NA, MacDonald MH, Matthews BF (2008) Protease inhibitor expression in soybean roots exhibiting susceptible and resistant interactions with soybean cyst nematode. J Nematol 40:138–146
Odani S, Ikenaka T (1977) Studies on soybean trypsin inhibitors. X. Isolation and partial characterization of four soybean double-headed proteinase inhibitors. J Biochem 82:1513–1522
Herman EM, Larkins BA (1999) Protein storage bodies and vacuoles. Plant Cell 11:601–613
Horisberger M, Tacchini-Vonlanthen M (1983) Ultrastructural localization of Bowman–Birk inhibitor on thin sections of Glycine max (soybean) cv Maple Arrow by the gold method. Histochem Cell Biol 3:313–321
Horisberger M, Tacchini-Vonlanthen M (1983) Ultrastructural localization of Kunitz Inhibitor on thin sections of Glycine max (soybean) cv Maple Arrow by the gold method. Histochemistry 77:37–50. doi:10.1007/BF00496634
Vandepoele K, Quimbaya M, Casneuf T, De Veylder L, Van de Peer Y (2009) Unraveling transcriptional control in Arabidopsis using cis-regulatory elements and coexpression networks. Plant Physiol 50:535–546. doi:/10.1104/pp.109.136028
Eulgem T, Rushton PJ, Robatzek S, Somssich IE (2000) The WRKY superfamily of plant transcription. Trends Plant Sci 5:199–206. doi:10.1016/S1360-1385(00)01600-9
Jin H, Martin C (1999) Multifunctionality and diversity within the plant MYB-gene family. Plant Mol Biol 41:577–585. doi:10.1023/A:1006319732410
Stracke R, Werber M, Weisshaar B (2001) The R2R3-MYB gene family in Arabidopsis thaliana. Curr Opin Plant Biol 4:447–456. doi:10.1016/S1369-5266(00)00199-0
Kang HG, Singh K (2000) Characterization of salicylic acid responsive, Arabidopsis Dof domain proteins: overexpression of OBP3 leads to growth defects. Plant J 21:329–339. doi:10.1046/j.1365-313x.2000.00678.x
Rakwal R, Agrawal GK, Jwa N-S (2001) Characterization of a rice (Oryza sativa L.) Bowman–Birk proteinase inhibitor: tightly regulated induction in response to cut, jasmonic acid, ethylene and protein phosphatase 2A inhibitors. Gene 263:189–198. doi:10.1016/S0378-1119(00)00573-4
Tan-Wilson AL, Cosgriff SE, Duggan MC, Obach RS, Wilson KA (1985) Bowman–Birk protease isoinhibitor complements of soybean strains. J Agric Food Chem 33:389–393. doi:10.1021/jf00063a017
Galasso I, Piergiovanni AR, Lioi L, Campion B, Bollini R, Sparvoli F (2009) Genome organization of Bowman–Birk inhibitor in common bean (Phaseolus vulgaris L.). Mol Breed 23:617–624. doi:10.1007/s11032-009-9260-4
Birk Y (1967) Chemistry and nutritional significance of proteinase inhibitors from plant sources. Ann NY Acad Sci 146:388–399. doi:10.1111/j.1749-6632.1968.tb20299.x
Hwang DL, Yang W-K, Foard DE (1978) Rapid release of protease inhibitors from soybeans. Plant Physiol 61:30–34
Goldberg RB, Crouch ML, Walling L (1983) Regulation of soybean seed protein gene expression. In: Nagley P, Linnane AW, Peacok WJ, Pateman JA (eds) Manipulation and expression of genes in eukaryotes. Academic Press, New York, pp 193–201
Walther D, Brunnemann R, Selbig J (2007) The regulatory code for transcriptional response diversity and its relation to genome structural properties in A. thaliana. PLoS Genet 3:e11. doi:10.1371/journal.pgen.0030011
Tan-Wilson AL, Rightmire BR, Wilson KA (1982) Different rates of metabolism of soybean proteinase inhibitors during germination. Plant Physiol 70:493–497
Acknowledgments
This work was funded by CNPq, FAPEMIG and FINEP (Brazilian Government). B.A. Barros was the recipient of a PhD fellowship from CNPq.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
de Almeida Barros, B., da Silva, W.G., Moreira, M.A. et al. In silico characterization and expression analysis of the multigene family encoding the Bowman–Birk protease inhibitor in soybean. Mol Biol Rep 39, 327–334 (2012). https://doi.org/10.1007/s11033-011-0742-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-011-0742-1