Abstract
Epidemiologic studies have evaluated the association between BRAF mutations and resistance to the treatment of anti-EGFR monoclonal antibodies (MoAb) in patients with metastatic colorectal cancer (mCRC). However, the results are still inconclusive. To derive a more precise estimation of the relationship, we performed this meta-analysis. A total of 11 studies were included in the final meta-analysis. There were seven studies for unselected mCRC patients and four studies for patients with wild type KRAS mCRC. Among unselected mCRC patients, BRAF V600E mutation was detected in 48 of 546 primary tumors (8.8%). The objective response rate (ORR) of patients with mutant BRAF was 29.2% (14/48), whereas the ORR of patients with wild-type BRAF was 33.5% (158/472).The overall RR for ORR of mutant BRAF patients over wild-type BRAF patients was 0.86 (95% CI = 0.57–1.30; P = 0.48). For patients with KRAS wild-type mCRC, BRAF V600E mutation was detected in 40 of 376 primary tumors (10.6%). The ORR of patients with mutant BRAF was 0.0% (0/40), whereas the ORR of patients with wild-type BRAF was 36.3% (122/336). The pooled RR of mutant BRAF patients over wild-type BRAF patients was 0.14 (95% CI = 0.04–0.53; P = 0.004). In conclusion, this meta-analysis provides evidence that BRAF V600E mutation is associated with lack of response in wild-type KRAS mCRC treated with anti-EGFR MoAbs. BRAF mutation may be used as an additional biomarker for the selection of mCRC patients who might benefit from anti-EGFR MoAbs therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Metastatic colorectal cancer (mCRC) is one of the most common human malignant disease and one of the leading causes of cancer-related death worldwide. Despite of recent advances in chemotherapeutic treatment, the 5-year overall survival (OS) is relatively poor, with a median survival of 18–21 months [1].There is a continuous need for more effective therapies. Most recently, two monoclonal antibodies (MoAb) targeting epidermal growth factor receptor (EGFR), cetuximab and panitumumab, have been developed and approved for the treatment of chemotherapy-resistant mCRC [2, 3]. However, resistance to anti-EGFR MoAbs was common. Among unselected mCRC patients, only 10–20% of patients can truly benefit from the treatment [4]. Therefore, there is a clear need for predictive biomarkers to maximize likelihood of response while minimizing toxicities and cost. To data, KRAS mutations have already been identified as clinical predictors of resistance to anti-EGFR MoAb in mCRC [5–7]. However, the occurrence of KRAS mutations only accounts for approximately 30–40% of nonresponsive patients [8–12]. Other mechanisms of resistance to anti-EGFR MoAb thus exist. Mutations in one other key mediator of EGFR signaling, BRAF, have also been shown to be associated with poor clinical outcomes [8, 13–22]. However, the results are still inconclusive. In this article, we performed a meta-analysis of the published studies to summarize the scientific evidence for the association between BRAF mutations and tumor response in mCRC patients treated with anti-EGFR MoAbs.
Materials and methods
Publication search
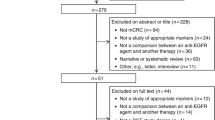
Systematic computerized searches of the PubMed, EMBase, BIOSIS, and SCOPUS (up to September 30, 2009) were performed. Following search terms were used: “metastatic colon cancer”, “metastatic rectal cancer’’, “metastatic colorectal cancer”, “mCRC”, “BRAF”, “monoclonal antibodies”, “MoAb”,“cetuximab”, “panitumumab”. The search was limited to human studies. Eligible studies reported complete response (CR) and partial response (PR) stratified by BRAF mutation status. All eligible studies were retrieved, and their bibliographies were checked for other relevant publications. When the same patient population was used in several publications, only the most recent, largest or complete study was included in the meta-analysis.
Inclusion criteria
The included studies have to meet the following criteria: (1) evaluating the BRAF mutation status and response to anti-EGFR MoAbs in mCRC, (2) studies with full text articles, and (3) sufficient data on CR and PR.
Data extraction
Information was carefully extracted from all eligible studies. The following data were collected from each study: first author’s name, year of publication, study designs, number of patients screened, number of patients with BRAF mutations, line of treatment, study treatment protocols, response criteria, CR and PR stratified by BRAF mutation status. Data extraction was done independently by two of the authors. Disagreement was resolved by discussion between the two authors. If these two authors could not reach a consensus, another author was consulted to resolve the dispute and a final decision was made by the majority of the votes.
BRAF mutations and clinical endpoints
The primary endpoint was objective response rate (ORR). The ORR was defined as the sum of CR and PR. The correlation between BRAF mutations and ORR was expressed as a relative ratio (RR) for ORR of gene mutant patients over wild-type patients. Thus, a RR equal to 1 indicates a lack of association between BRAF mutations and tumor response to anti-EGFR MoAbs therapy; a RR more than 1 corresponds to a direct correlation between higher ORR and BRAF mutations, and a tendency of mutant BRAF patients to have non-responsiveness is indicated by a RR less than 1.
Statistical methods
The association between BRAF mutations and efficacy of anti-EGFR MoAb therapy was measured by RR with 95% CI. Heterogeneity was checked by a Q-test with degree of freedom equal to the number of analyzed studies minus 1[23]. A P value of more than 0.10 for the Q-test indicates a lack of heterogeneity across studies, so the pooled RR was calculated by the fixed-effects model [24]. Otherwise, the random-effects model was used [25]. Sensitivity analyses were carried out to check if modification of the inclusion criteria of the meta-analysis affected the final results. Begg’s funnel plots and Egger’s linear regression test were used to assess publication bias. Funnel plot asymmetry was assessed by the method of Egger’s linear regression test, a linear regression approach to measure funnel plot asymmetry on the natural logarithm scale of the RR. The significance of the intercept was determined by the t-test as suggested by Egger (P < 0.05 was considered representative of statistically significant publication bias) [26]. All the statistical tests used in our meta-analysis were performed with STATA version 10.0 (Stata Corporation, College Station, TX).
Results
Studies characteristics
Based on our search criteria, a total of 11 studies [8, 13–22] were included in the final meta-analysis. Table 1 lists the studies identified and their main characteristics. Of the 11 studies, sample sizes ranged from 31 to 259. All of these studies were retrospective design. Anti-EGFR MoAb was given as first-line treatment in one studies, as second line or more in ten studies. The patients of two studies received anti-EGFR MoAb monotherapy, while the patients of nine studies received anti-EGFR MoAb-based therapy.
BRAF mutations and ORR in unselected mCRC patients
Totally, seven studies [8, 13, 14, 16, 19, 20, 22] were performed in unselected mCRC patients. BRAF mutational analysis was performed successfully on 546 mCRC. BRAF V600E mutation was detected in 48 (8.8%) primary tumors. Among 546 analyzed patients, 520 patients were assessable for tumor response. The ORR of mCRC patients with mutant BRAF was 29.2% (14/48), whereas the ORR of mCRC patients with wild-type BRAF was 33.5% (158/472). The overall RR for ORR of mutant BRAF patients over wild-type BRAF patients was 0.86 (95% CI = 0.57–1.30; P = 0.48), with no heterogeneity between studies (P = 0.27; I2 = 21.1%; Table 2; Fig. 1). No publication bias was found by Egger’s test (t = 0.18; P = 0.86).
BRAF mutations and ORR in patients with wild-type KRAS mCRC
Totally, four studies [15, 17, 18, 21] were performed in patients with wild-type KRAS mCRC. BRAF mutational analysis was performed successfully on 376 KRAS wild-type mCRC. BRAF V600E mutation was detected in 40 (10.6%) primary tumors. Among 376 analyzed patients, all of patients were assessable for tumor response. The ORR of patients with mutant BRAF was 0.0% (0/40), whereas the ORR of patients with wild-type BRAF was 36.3% (122/336). BRAF V600E mutation had negative effect on tumor response to anti-EGFR MoAbs (pooled RR = 0.14; 95% CI = 0.04-0.53; P = 0.004) in patients with KRAS wild-type mCRC, with no heterogeneity between studies (P = 1.00; I2 = 0.0%; Table 2; Fig. 1). No publication bias was found by Egger’s test (t = 2.10; P = 0.17).
Discussion
The RAS/RAF/MAPK pathway were major downstream signaling proteins of EGFR, which regulates biological responses such as cell proliferation, cell survival and cell differentiation [27]. Mutations in downstream signaling effectors of EGFR could result in resistance to EGFR inhibitors [28]. In mCRC, KRAS mutations have already been identified as clinical predictors of resistance to anti-EGFR MoAbs [5–7]. However, the occurrence of KRAS mutations only accounts for approximately 30–40% of nonresponsive patients [8–12]. Other mechanisms of resistance to anti-EGFR MoAb thus exist. Recently, emerging data indicate that BRAF mutations were associated with poor clinical outcomes [8, 13–22]. Unfortunately, the results remain conflicting rather than conclusive. To derive a more precise estimation of the predictive significance of BRAF mutations in mCRC patients treated with anti-EGFR MoAbs, we performed this meta-analysis.
Among unselected mCRC patients, data for BRAF mutations and tumor response were available in seven studies, with 520 patients included. The ORR of mCRC patients with mutant BRAF was 29.2% (14/48), whereas the ORR of mCRC patients with wild-type BRAF was 33.5% (158/472). The overall RR indicated that BRAF mutation was not associated with lack of response in unselected mCRC patients treated with anti-EGFR MoAbs.
For patients with KRAS wild-type mCRC, data concerning BRAF mutations and tumor response were available in four studies, with 376 patients included. BRAF V600E mutation was detected in 40 (10.6%) primary tumors. The ORR of patients with mutant BRAF was 0.0% (0/40), whereas the ORR of patients with wild-type BRAF was 36.3% (122/336). The overall RR indicated that BRAF V600E mutation had negative effect on tumor response to anti-EGFR MoAbs in patients with KRAS wild-type mCRC.
Heterogeneity is a potential problem that may affect the interpretation of the results of meta-analyses. Although some diversity in the studies about study designs, inclusion criteria, treatment protocols and response criteria, there was no statistically significant heterogeneity among the studies. This indicated that it may be appropriate to use an overall estimation of the predictive value of BRAF mutations.
Our studies had several limitations that need to be taken into consideration when interpreting the findings. First, inadequate reporting of survival data precludes any meaningful assessment of the effect of BRAF status on survival in mCRC patients treated with anti-EGFR MoAbs. Only three studies presented data on hazard ratio (HR) with 95% CI for progression-free survival (PFS) and overall survival (OS). Therefore, we did not perform pooled analysis for HR. Second, our result was based on unadjusted estimates, while a more precise analysis should be conducted if more detailed individual data were available, which would allow for an adjusted estimate by other prognostic factors such as sex, age, tumor location and previous chemotherapy lines.
Despite of some limitations, this meta-analysis provided evidence that BRAF V600E mutation is associated with lack of response in wild-type KRAS mCRC treated with anti-EGFR MoAbs. The results indicated that BRAF mutation may be used as an additional biomarker for the selection of mCRC patients who might benefit from anti-EGFR MoAbs therapy. However, the number of studies and the number of subjects included in the meta-analysis are relatively small. Large prospective studies using standardized unbiased methods are needed to confirm our results, using homogeneous mCRC patients, with assessors blinded to the clinical data.
References
Sargent DJ, Wieand HS, Haller DG, Gray R, Benedetti JK, Buyse M, Labianca R, Seitz JF, O’Callaghan CJ, Francini G, Grothey A, O’Connell M, Catalano PJ, Blanke CD, Kerr D, Green E, Wolmark N, Andre T, Goldberg RM, De Gramont A (2005) Disease-free survival versus overall survival as a primary end point for adjuvant colon cancer studies: individual patient data from 20, 898 patients on 18 randomized trials. J Clin Oncol 23:8664–8670
Cunningham D, Humblet Y, Siena S, Khayat D, Bleiberg H, Santoro A, Bets D, Mueser M, Harstrick A, Verslype C, Chau I, Van Cutsem E (2004) Cetuximab monotherapy and cetuximab plus irinotecan in irinotecan-refractory metastatic colorectal cancer. N Engl J Med 351:337–345
Van Cutsem E, Peeters M, Siena S, Humblet Y, Hendlisz A, Neyns B, Canon JL, Van Laethem JL, Maurel J, Richardson G, Wolf M, Amado RG (2007) Open-label phase III trial of panitumumab plus best supportive care compared with best supportive care alone in patients with chemotherapy-refractory metastatic colorectal cancer. J Clin Oncol 25:1658–1664
Meyerhardt JA, Mayer RJ (2005) Systemic therapy for colorectal cancer. N Engl J Med 352:476–487
European Medicines Agency website, European Public Assessment Report. http://www.emea.europa.eu/pdfs/human/opinion/40511307en.pdf. Accessed 31 Mar 2009
European Medicines Agency website, European Public Assessment Report. http://www.emea.europa.eu/pdfs/human/opinion/Erbitux_28040208en.pdf. Accessed 31 March 2009
http://www.fda.gov/AboutFDA/CentersOffices/CDER/ucm172905.htm
Moroni M, Veronese S, Benvenuti S, Marrapese G, Sartore-Bianchi A, Di Nicolantonio F, Gambacorta M, Siena S, Bardelli A (2005) Gene copy number for epidermal growth factor receptor (EGFR) and clinical response to antiEGFR treatment in colorectal cancer: a cohort study. Lancet Oncol 6:279–286
Lièvre A, Bachet JB, Le Corre D, Boige V, Landi B, Emile JF, Côté JF, Tomasic G, Penna C, Ducreux M, Rougier P, Penault-Llorca F, Laurent-Puig P (2006) KRAS mutation status is predictive of response to cetuximab therapy in colorectal cancer. Cancer Res 66:3992–3995
Frattini M, Saletti P, Romagnani E, Martin V, Molinari F, Ghisletta M, Camponovo A, Etienne LL, Cavalli F, Mazzucchelli L (2007) PTEN loss of expression predicts cetuximab efficacy in metastatic colorectal cancer patients. Br J Cancer 97:1139–1145
Di Fiore F, Blanchard F, Charbonnier F, Le Pessot F, Lamy A, Galais MP, Bastit L, Killian A, Sesboüé R, Tuech JJ, Queuniet AM, Paillot B, Sabourin JC, Michot F, Michel P, Frebourg T (2007) Clinical relevance of KRAS mutation detection in metastatic colorectal cancer treated by cetuximab plus chemotherapy. Br J Cancer 96:1166–1169
De Roock W, Piessevaux H, De Schutter J, Janssens M, De Hertogh G, Personeni N, Biesmans B, Van Laethem JL, Peeters M, Humblet Y, Van Cutsem E, Tejpar S (2008) KRAS wild-type state predicts survival and is associated to early radiological response in metastatic colorectal cancer treated with cetuximab. Ann Oncol 19:508–515
Benvenuti S, Sartore-Bianchi A, Di Nicolantonio F, Zanon C, Moroni M, Veronese S, Siena S, Bardelli A (2007) Oncogenic activation of the RAS/RAF signaling pathway impairs the response of metastatic colorectal cancers to anti-epidermal growth factor receptor antibody therapies. Cancer Res 67:2643–2648
Cappuzzo F, Varella-Garcia M, Finocchiaro G, Skokan M, Gajapathy S, Carnaghi C, Rimassa L, Rossi E, Ligorio C, Di Tommaso L, Holmes AJ, Toschi L, Tallini G, Destro A, Roncalli M, Santoro A, Jänne PA (2008) Primary resistance to cetuximab therapy in EGFR FISH-positive colorectal cancer patients. Br J Cancer 99:83–89
Di Nicolantonio F, Martini M, Molinari F, Sartore-Bianchi A, Arena S, Saletti P, De Dosso S, Mazzucchelli L, Frattini M, Siena S, Bardelli A (2008) Wild-type BRAF is required for response to panitumumab or cetuximab in metastatic colorectal cancer. J Clin Oncol 26:5705–5712
Freeman DJ, Juan T, Reiner M, Hecht JR, Meropol NJ, Berlin J, Mitchell E, Sarosi I, Radinsky R, Amado RG (2008) Association of K-ras mutational status and clinical outcomes in patients with metastatic colorectal cancer receiving panitumumab alone. Clin Colorectal Cancer 7:184–190
Laurent-Puig P, Cayre A, Manceau G, Buc E, Bachet JB, Lecomte T, Rougier P, Lievre A, Landi B, Boige V, Ducreux M, Ychou M, Bibeau F, Bouché O, Reid J, Stone S, Penault-Llorca F (2009) Analysis of PTEN, BRAF, and EGFR status in determining benefit from cetuximab therapy in wild-type KRAS metastatic colon cancer. J Clin Oncol 27:5924–5930
Loupakis F, Ruzzo A, Cremolini C, Vincenzi B, Salvatore L, Santini D, Masi G, Stasi I, Canestrari E, Rulli E, Floriani I, Bencardino K, Galluccio N, Catalano V, Tonini G, Magnani M, Fontanini G, Basolo F, Falcone A, Graziano F (2009) KRAS codon 61, 146 and BRAF mutations predict resistance to cetuximab plus irinotecan in KRAS codon 12 and 13 wild-type metastatic colorectal cancer. Br J Cancer 101:715–721
Molinari F, Martin V, Saletti P, De Dosso S, Spitale A, Camponovo A, Bordoni A, Crippa S, Mazzucchelli L, Frattini M (2009) Differing deregulation of EGFR and downstream proteins in primary colorectal cancer and related metastatic sites may be clinically relevant. Br J Cancer 100:1087–1094
Perrone F, Lampis A, Orsenigo M, Di Bartolomeo M, Gevorgyan A, Losa M, Frattini M, Riva C, Andreola S, Bajetta E, Bertario L, Leo E, Pierotti MA, Pilotti S (2009) PI3KCA/PTEN deregulation contributes to impaired responses to cetuximab in metastatic colorectal cancer patients. Ann Oncol 20:84–90
Sartore-Bianchi A, Di Nicolantonio F, Nichelatti M, Molinari F, De Dosso S, Saletti P, Martini M, Cipani T, Marrapese G, Mazzucchelli L, Lamba S, Veronese S, Frattini M, Bardelli A, Siena S (2009) Multi-determinants analysis of molecular alterations for predicting clinical benefit to EGFR-targeted monoclonal antibodies in colorectal cancer. PLoS One 4:e7287
Tol J, Nagtegaal ID, Punt CJ (2009) BRAF mutation in metastatic colorectal cancer. N Engl J Med 361:98–99
Cochran WG (1954) The combination of estimates from different experiments. Biometrics 10:101–129
Mantel N, Haenszel W (1959) Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst 22:719–748
DerSimonian R, Laird N (1986) Meta-analysis in clinical trials. Control Clin Trials 7:177–188
Egger M, Smith DG, Schneider M, Minder C (1997) Bias in meta-analysis detected by a simple, graphical test. BMJ 315:629–634
Schlessinger J (2002) Ligand-induced, receptor-mediated dimerization and activation of EGF receptor. Cell 110:669–672
Yarden Y, Sliwkowski MX (2001) Untangling the ErbB signaling network. Nat Rev Mol Cell Biol 2:127–137
Acknowledgment
This study was supported by Guangdong Province “211 Project” (Grant No. GW201004).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mao, C., Liao, RY., Qiu, LX. et al. BRAF V600E mutation and resistance to anti-EGFR monoclonal antibodies in patients with metastatic colorectal cancer: a meta-analysis. Mol Biol Rep 38, 2219–2223 (2011). https://doi.org/10.1007/s11033-010-0351-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-010-0351-4