Abstract
Stripe rust disease is caused by the fungus Puccinia striiformis f. sp. tritici and severely threatens wheat worldwide, repeatedly breaking resistance conferred by resistance genes and evolving more aggressive strains. Wild emmer wheat, Triticum dicoccoides, is an important source for novel stripe rust resistance (Yr) genes. Yr15, a major gene located on chromosome 1BS of T. dicoccoides, was previously reported to confer resistance to a broad spectrum of stripe rust isolates, at both seedling and adult plant stages. Introgressions of Yr15 into cultivated T. aestivum bread wheat and T. durum pasta wheat that began in the 1980s are widely used. In the present study, we aimed to validate SSR markers from the Yr15 region as efficient tools for marker-assisted selection (MAS) for introgression of Yr15 into wheat and to compare the outcome of gene introgression by MAS and by conventional phenotypic selection. Our findings establish the validity of MAS for introgression of Yr15 into wheat. We show that the size of the introgressed segment, defined by flanking markers, varies for both phenotypic selection and MAS. The genetic distance of the MAS marker from Yr15 and the number of backcross steps were the main factors affecting the length of the introgressed donor segments. Markers Xbarc8 and Xgwm493, which are the nearest flanking markers studied, were consistent and polymorphic in all 34 introgressions reported here and are therefore the most recommended markers for the introgression of Yr15 into wheat cultivars. Introgression directed by markers, rather than by phenotype, will facilitate simultaneous selection for multiple stripe rust resistant genes and will help to avoid escapees during the selection process.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stripe rust caused by the fungus Puccinia striiformis f. sp. tritici (Pst) is one of the most destructive diseases of wheat, resulting in yield losses of 10–70 % in susceptible varieties (Chen 2005; Chen et al. 2010). Total losses of harvestable grain have been reported for severe epidemics (McIntosh et al. 1995; Chen 2005). Introgression of resistance (R) genes from wild germplasm has been recognized as an efficient and environmentally safe approach to minimize yield losses due to diseases. For stripe rust, introgression of resistance genes into wheat accessions commenced in the 1980s (Chen 2005), thereby saving hundreds of millions of dollars (Brennan and Murray 1998). Wild emmer wheat, Triticum dicoccoides, discovered in 1906 in Rosh Pina, Israel (Aaronsohn 1910), is an important source for novel stripe rust resistance (Yr) genes (Grama and Gerechter-Amitai 1974; Fahima et al. 1998; Nevo et al. 2002).
New resistance genes for stripe rust are continuously needed due to breakdown of resistance conferred by currently deployed genes as P. striiformis races evolve and migrate (van Silfhout and Grönewegen 1984; Stubbs 1985; Sun et al. 1997; McDonald and Linden 2002). Yr15 is a stripe rust resistance gene discovered in the 1980s in T. dicoccoides accession G25 (Gerechter-Amitai et al. 1989); it was later mapped to chromosome 1BS (Sun et al. 1997; Peng et al. 2000). The resistance conferred by Yr15 was shown to be effective against 24 Pst races from 18 countries around the globe (Gerechter-Amitai et al. 1989) and to 26 international isolates and Chinese races of Pst (58893, 59791, 60105, 61009, 68009, 72107, 74187, 75078, 76088, 76093, 78028, 78080, 80551, 82061, 82517, 85019, 86036, 86094, 86106, 86107, PE92, CYR26, CYR27, CYR29, CYR32, CYR-S) (Li et al. 2006; Niu et al. 2000). More recent studies show that Yr15 is effective against all Pst races identified so far in the USA (Murphy et al. 2009; Chen et al. 2010), Australia (Bariana et al. 2007; Randhawa et al. 2014), and India (H.S. Bariana unpublished results). These findings confirm that Yr15 is a valuable source of resistance to stripe rust. In order to make this resistance gene available to wheat breeders, Yr15 was introgressed into both durum and bread wheat lines by conventional breeding through phenotypic selection at the Agricultural Research Organization, Volcani Center, Bet Dagan, Israel (Grama and Gerechter-Amitai 1974; Grama et al. 1982; van Silfhout and Grönewegen 1984). Following the initial introgression of Yr15 in Israel, breeding programs worldwide have used the resulting lines as a source for introgression of Yr15 into bread and durum wheat cultivars.
Marker-assisted selection (MAS; reviewed in Koebner and Summers 2003; Collard and Mackill 2008; Panigrahi et al. 2013) refers to the use of DNA markers that are tightly linked to target loci as a substitute or assistance for phenotypic selection. By determining the allele of a DNA marker, plants that possess traits linked to that marker may be identified by the marker genotype rather than by their phenotype. MAS is particularly useful for traits with low heritability such as drought tolerance or yield (e.g., Hill et al. 2013), recessive inheritance such as disease resistance (e.g., Tyrka et al. 2008), difficult and costly phenotyping such as baking quality (e.g., Cavanagh et al. 2010), and for pyramiding multiple disease resistance genes (e.g., Tester and Langridge 2010). MAS also provides a rapid means of tracking introgressions (Tester and Langridge 2010). Moreover, it is useful for reducing the length of introgressed chromosome segments in order to minimize the risk of linked genes that can have a negative impact (Holland 2004; Collard and Mackill 2008). MAS has been a successful tool for introgression of genes of interest including disease resistance genes into cultivars in wheat and many other species (Singh et al. 2001; Spielmeyer et al. 2003; Yang et al. 2003; Simko et al. 2009).
Previously, the assignment of Yr15 to chromosome arm 1BS by McIntosh et al. (1996) facilitated the mapping of the gene, which was begun by Sun et al. (1997) and enabled the finer mapping of the Yr15 region with the markers reported here. In the present study, we aimed to validate simple sequence repeat (SSR) markers from the Yr15 region as efficient tools for MAS of Yr15 into wheat and to compare the outcome of gene introgression by MAS and by conventional phenotypic selection. A set of polymorphic SSR markers was used to genotype 34 Yr15 introgression lines (ILs) from different breeding programs obtained both by conventional and by MAS backcrossing, as well as on their susceptible recurrent parents (RPs).
Materials and methods
Plant material
F2 mapping population
An F2 population of 151 individuals used for the genetic mapping of the SSR markers was developed by crossing the susceptible T. durum accession D447 (LD393/2*Langdon ND58-322) with the resistant BC3F9 and BC3F10 (B9 and B10) introgression lines, which carried the Yr15 gene within a 1BS chromosome segment introgressed from T. dicoccoides accession G25 (Sun et al. 1997; Peng et al. 2000). The F1 plants were tested to verify heterozygosity and then self-crossed to produce a segregating population for linkage analysis.
Yr15 introgression lines
Since its discovery (Gerechter-Amitai et al. 1989), Yr15 has been introgressed into a range of durum and bread wheat genetic backgrounds in a variety of breeding programs around the globe. The 34 Yr15 ILs and the 23 susceptible RPs lacking Yr15, and the Yr15 donor line T. dicoccoides G25 used in the current study, along with their pedigrees, are described in Table 1. The tetraploid (T. turgidum ssp. durum) and hexaploid (T. aestivum) accessions include cultivars and advanced breeding lines from the Agricultural Research Organization, Volcani Center, Bet Dagan, Israel, the University of Sydney, Cobbitty, Australia, and the University of California, Davis, CA, USA.
Stripe rust resistance tests conducted at the seedling stage
Stripe rust resistance tests were conducted at the seedling stage, as described in Cheng et al. (2010). The highly virulent Pst isolate 5006 (that belongs to race 38E134, virulent on Yr2, Yr6, Yr7, Yr9, Yr22, Yr23, YrSD, and YrHVII; Cheng et al. 2010), kindly provided by Dr. Jacob Manisterski (Tel Aviv University, Tel-Aviv, Israel), was used to inoculate the plants. Seedlings in the second-leaf stage were infected with stripe rust by spraying with a mixture of a light mineral oil (Soltrol 170, Chevron-Phillips Chemical Co., Houston) and Pst spores (2 mg ml−1). The sprayed plants were left under non-humid conditions for 20 min in order to allow the mineral oil to dry. The inoculated plants were then transferred to a dew chamber and kept for 24 h at 100 % humidity, first for 12 h at 9 °C in the dark followed by 12 h under light at 15 °C. The plants were then grown at 70 % humidity under the following day/night regime: 12 h at 15 °C with a light intensity of 150 μmol m−2 s−1 followed by 12 h at 9 °C in darkness. Infection types (ITs) were scored 2.5 weeks after inoculation. In case of resistant accessions, the test was repeated once again. The IT produced by plant–pathogen interactions was assessed on a scale of 0–9 (Line and Qayoum 1992). The ITs were summarized by combining them into three classes: 0–3 was considered resistant (R), 4–6 moderately resistant (M), and 7–9 susceptible (S) according to Qayoum and Line (1985). Data regarding susceptibility or resistance to stripe rust were recorded for each accession.
DNA extraction
Leaves of month-old plants were collected, frozen in liquid nitrogen, and stored at −80 °C. Genomic DNA was extracted using a large-scale protocol (Kidwell and Osborn 1992) with the modifications described by Xie et al. (2012).
SSR marker analysis
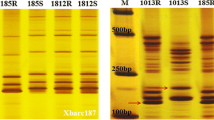
The SSR marker screening of plant DNA samples was performed by PCR in a 15 μl final volume containing 0.2 mM of each dNTP, 250 nM of each primer, 0.6 U of Taq polymerase (DreamTaq, Thermo Fischer Scientific, San Jose, CA), and 100 ng of template DNA. PCR was performed on the Veriti® 96-Well Fast Thermal Cycler (Applied Biosystems Inc. (ABI), Foster City, CA, USA). The PCR program comprised a 5-min initial denaturation at 94 °C followed by 35 cycles of: 94 °C for 30 s; 55–60 °C (depending of annealing temperature of the primer set used) for 30 s; and 72 °C for 30–60 s (depending on the size of the amplicon). Fragment analysis was carried out on a Model 3130xl DNA sequencer (ABI). Allele fragment sizes and genotypes were determined using GeneMapper V3.7 software (ABI). The bread wheat accession Chinese Spring (CS) served as a size reference in each electrophoresis run. The SSR markers that were used in the current study for genetic mapping of Yr15, as well as for screening the IL and RP lines, are listed in Table 2. These markers were selected from a large set of SSR markers mapped to chromosome 1B of bread wheat (Röder et al. 1998; Peng et al. 2000; Song et al. 2002; Somers et al. 2004; Ganal and Röder 2007), based on screening for polymorphism between the parental lines of the Yr15 mapping population, as well as between the IL and RP lines.
Genetic mapping
For each polymorphic SSR marker, a χ 2 analysis was performed to test for deviation from the 1:2:1 expected segregation ratio in the F2 mapping population. Linkage analysis and map construction were performed based on the evolutionary strategy algorithm included in the MultiPoint package (Mester et al. 2003), as described in Peleg et al. (2008).
Results
Identification of highly polymorphic SSR markers for genetic mapping of Yr15
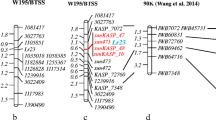
A total of seven SSR markers, among those previously mapped to chromosome 1B, were screened in 58 Yr15 IL and RP lines (Table 2). Among these markers, six polymorphic SSRs showed 7–15 different alleles within these lines, while one marker, Xgwm498, showed a lower polymorphism level with only four alleles (Tables 1, 2). These seven markers were chosen to construct a genetic linkage map of chromosome 1B spanning the Yr15 segment introgressed from the T. dicoccoides donor line G25 (Fig. 1) using an F2 mapping population of 151 plants derived from a cross of the susceptible line D447 with B9 or B10 ILs containing Yr15. The resulting genetic map showed that the seven SSR markers span a region of 31.8 cM on chromosome 1B, with Xgwm911 being the most distal marker (18.2 cM distal to Yr15) and Xgwm762 the most proximal (13.6 cM proximal to Yr15). Sourdille et al. (2004) have used deletion bin mapping to assign marker Xgwm273 to 1BS, whereas Xgwm498 was assigned to 1BL. On this basis, the centromere location was marked in the current study on the Yr15 genetic map between markers Xgwm273 and Xgwm498 (Fig. 1). Therefore, Xgwm273 is the most proximal marker to the Yr15 gene on the short arm of chromosome 1B. The Yr15 gene was mapped to a 6.4 cM interval flanked by marker Xbarc8, located 3.9 cM to the distal side, and by Xgwm413 located 2.5 cM to the proximal side. The obtained genetic map was used as a reference to determine the size of the introgressed chromosome segments harboring Yr15 in the different ILs.
Genetic map of the stripe rust resistance gene Yr15. The map is of chromosome 1BS of wild emmer wheat, developed by genotyping of 155 F2 plants derived from the cross of susceptible D447 line with B9 or B10 ILs that contain Yr15 and genotyped with seven SSR markers. The genetic distance in cM is shown on the left. The approximate position of the centromere, according to deletion bin mapping of markers Xgwm273 to 1BS and Xgwm498 to 1BL (Sourdille et al. 2004), is indicated by an arrowhead
Evaluation of Yr15 donor fragment sizes carried by introgression lines obtained by conventional and MAS backcrossing
The graphical genotypes concept, introduced by Young and Tanksley (1989), was implemented in our study for comparison of the size of Yr15 donor fragments carried by ILs developed by conventional and MAS backcrossing. This method allowed the donor and RP chromosomal segments to be distinguished easily. The seven highly polymorphic SSR markers used for developing the genetic map described above (Fig. 1) were also used to genotype 58 wheat lines (34 ILs and the Yr15 donor line resistant to Pst and 23 susceptible RPs) obtained from three different breeding programs scattered around the globe (Israel, Australia and California). These Yr15 ILs were proved to be resistant at the adult plant stage to a wide range of Pst isolates, prevalent in the respective continents. The spectrum of marker alleles obtained for the seven SSR markers by screening of the 58 wheat lines is presented in Table 1 in a graphical genotype format. The order of the markers was defined by the Yr15 genetic map described above (Fig. 1), and the genetic distances in cM were used to calculate the length of the introgressed donor segment.
In the Israeli breeding program, Yr15 was introgressed from the donor T. dicoccoides line G25 through recurrent backcrossing directed by phenotypic selection. Lines B1, B2, B9, and B10 are ILs derived from the cross between G25 and the susceptible tetraploid T. durum line D447. In these ILs, the Yr15 donor segments expand from the region between Xgwm911 (not present) and Xwmc406 in the short arm to a region beyond Xgwm762 in the long arm. B70 is an additional IL derived from the same cross, which contains a shorter T. dicoccoides donor segment that spans from Yr15 in the short arm to Xgwm762 in the long arm. The B70 alleles for markers Xwmc406 and Xbarc8 are not present in either the donor G25 or the susceptible line D447. ILs 280-1 and 280-2 were found to be resistant to Pst in our survey (Table 1), yet exhibit alleles distinct from those of the Yr15 donor line for all tested SSR markers.
Yr15 was also introgressed into two hexaploid (bread) wheat cultivars, Merav and Mexico 708, resulting in the selection lines designated here as Sel07-97, Sel46, and Sel7. Based on the available markers, these ILs contained the same Yr15 donor segment as the tetraploid lines B1, B2, B9, and B10 described above. The same segment was detected in the hexaploid lines designated as V763 (Table 1). These lines were obtained in the Israeli breeding program by crossing tetraploid durum wheat containing Yr15 (described above) with adapted bread wheat cultivars and then further advancing the progeny with high economic potential (Grama and Gerechter-Amitai 1974). Some of the V763 lines produced in the Israeli program were later used by the Australian Cereal Rust Control Program (ACRCP) at the University of Sydney for introgression of Yr15 into elite Australian cultivars.
The hexaploid line Avocet S, susceptible to stripe rust, was crossed with V763 and subsequently backcrossed to obtain the Yr15-containing line Avocet-Yr15 (Zakari et al. 2003). In a similar way, V763 derivatives (including Avocet-Yr15) were used to introgress Yr15 into five additional Australian bread wheat cultivars (Corrigin, Excalibur, Kulin, Stilleto, and Suncea), which were used as RPs in conventional introgressions by backcrossing. The six ILs derived from these introgressions were screened with the seven SSR markers (Table 1). Among the six ILs, four lines (Avocet-Yr15, Corrigin-Yr15, Kulin-Yr15, and Stilleto-Yr15) contained the long segment including Xwmc406 in the short arm and Xgwm762 in the long arm (likely more than 30 cM) detected in V763. In the other two ILs, shorter donor intervals were found. For the IL Excalibur-Yr15, a donor fragment of 2.5–8.5 cM was identified by markers Xbarc8 distal and Xgwm273 proximal to Yr15. In Excalibur-Yr15, only Yr15 and Xgwm413 were retained from the original G25 donor chromosome. For IL Suncea-Yr15, a donor fragment of less than 31.8 cM was identified by markers Xgwm911 distal and Xgwm762 proximal to Yr15.
In a different set of crosses produced by the ACRCP, the susceptible Avocet S was crossed with the donor of another stripe rust resistance gene, Yr24, and backcrossed six times to produce the Avocet-Yr24 NIL. This line was then crossed with Avocet-Yr15 to obtain a new Avocet line carrying the two stripe rust resistance genes, designated as Avocet-Yr15Yr24 (Zakari et al. 2003). In these crosses, both MAS and phenotypic selection were applied. The construction of an Avocet IL combining both Yr15 and Yr24 was accomplished by double selection, first for the phenotype of Yr15 and then for the genotype of the marker Xgwm11 [located in 1BS bin 1BS9-0.84-1.06 (Sourdille et al. 2004)] linked to Yr24 (Zakari et al. 2003). We screened five bread wheat RPs (i.e., Baxter, Combat, Sapphire, Shenton, and Ruby) and seven ILs derived from crosses with Avocet-Yr15Yr24. For all IL lines derived from these crosses, the G25 donor fragment did not include the 1B long-arm markers Xgwm498 and Xgwm762. This likely reflects the recombination event between the Yr15 and Yr24 genes, since Yr24 was mapped proximal to Yr15 (Li et al. 2006). The shortest G25 donor fragment in the Yr15Yr24 lines was found in lines HSB-2801 (4.6–12 cM; retained G25 alleles Yr15 and Xgwm273) and HSB-3177 (8.7–20.7 cM; retained G25 alleles Xwmc406 and Yr15). Among the Yr15Yr24 lines, the longest G25 segment was detected in HSB-2955 (<24.2 cM; Table 1).
In the breeding program at the University of California at Davis, accession Sunfield-Yr15 (pedigree WW31/4/W3566/Topo/3/Waite Line/IRN70.511//SUN15B), which harbors the Yr15 donor fragment introgressed from line V763, was used to transfer Yr15 into breeding lines and commercial varieties. In this breeding program, MAS was carried out using markers Xbarc8 and Xgwm273 (http://www.plantsciences.ucdavis.edu/MASWheat/protocols/Yr15/index.htm), which span a 8.5 cM interval (Fig. 1). From this program, we screened seven ILs and their corresponding susceptible RPs with the seven SSR markers. Analysis of ILs UC1037-Yr15, UC1041-Yr15, UC1128-Yr15, UC1107-Yr15, UC1358-Yr15, and UCKern-Yr15, derived from Sunfield-Yr15 crosses, revealed that these lines contain Yr15 donor segments including the short-arm marker Xwmc406 and the long-arm marker Xgwm762. A shorter segment was detected only in IL UC1110-Yr15, where the two long-arm markers include non-G25 alleles. These lines, together with the Avocet-Yr15, have been tested in California for the last 13 years with no virulence detected in any of the lines. This result indicates that Yr15 is still effective for the dozen of Pst races that have been detected during recent years in California (Chen 2005). This is particularly relevant because wheat varieties carrying Yr15 now cover more than 88,000 acres in California (http://smallgrains.ucdavis.edu/CWC_Wheat_Var_Survey_2013.pdf).
Taking these data together, marker analysis with seven SSRs of the 34 Yr15 ILs obtained from three breeding programs revealed that the G25 donor fragment was delimited by SSR marker Xgwm911 on the distal side of the gene; the most distal G25 allele was detected by the Xwmc406 marker. In the majority of ILs (23/34), we found no additional recombination events between Xwmc406 on the short arm and Xgwm762 on the long arm, indicating that the complete centromeric region of G25 1B is present in these lines. Four lines showed shorter G25 1BS segments in the distal region (B70, Excalibur-15, HSB-2408, and HSB2801) and eight have lost the G-25 alleles for the two markers on the 1B long arm (Excalibur-Yr15, HSB-3177, HSB-2408, HSB2801, HSB-2944, HSB-2949, HSB2955, and UC1110-Yr15).
Evaluation of stripe rust resistance in the Yr15 ILs and RPs
Phenotypic tests of the 58 wheat lines listed in Table 1 were made by inoculating with Pst race 38E134, which is avirulent on Yr15 but virulent on Yr2, Yr6, Yr7, and Yr9, Yr22, Yr23, YrSD and YrHVII -containing plants (Cheng et al. 2010). IT types of 0 or 1 (resistant phenotype) were identified in our survey for all accessions that carry the G25 introgressed segment harboring Yr15. No moderate resistance ITs (4–6) were observed in our survey. Plants that were susceptible to stripe rust showed ITs of 7–9. Generally, RPs were found to be susceptible to this race of Pst, whereas ILs that contain the G25 chromosome segment, as identified by the SSR markers used here, showed resistance to Pst, indicating that these lines possess Yr15. Two cases were exceptions: lines 280-1 and 280-2 were found to be resistant, but no alleles of G25 could be identified by SSR genotyping. These lines may possess a stripe rust resistance gene other than Yr15. Alternatively, these two lines may contain a donor chromosomal segment that is shorter than could be detected by the markers used here, i.e., <6.4 cM, as a result of two recombination events, one between Xbarc8 and Yr15 and the other between Yr15 and Xgwm413. In contrast, line HSB-2527 lacked both G25 donor alleles and Pst resistance, indicating that the segment introgressed into this line was probably lost during subsequent generations.
Discussion
Although the wild progenitors of cultivated crops harbor great reservoirs of genes and alleles conferring pathogen resistance, they also carry numerous traits that are undesirable in cultivated crops (Brouwer and St. Clair 2004). Introgression of resistance from wild progenitors is complicated by the need for many backcrossing and selection steps in order to eliminate undesirable donor traits (Niks et al. 2011). Decreasing the size of a donor fragment can remove genes with deleterious effects that are linked to the target gene (Hospital 2001). Therefore, the practical use of a target gene may be dependent on reducing linkage drag during IL development (Jacobsen and Schouten 2007). Molecular markers can be used to accelerate the removal of linked undesirable traits by specifically selecting for recombination between markers linked to the desirable and undesirable traits.
Here, we evaluated the introgression of the stripe rust resistance gene Yr15, which was derived from wild emmer wheat, T. dicoccoides, into commercial wheat cultivars. The ILs investigated here originated from several conventional and MAS breeding programs that differed from each other regarding their experimental design, use of markers, and the number of backcrosses performed. The lines included in our study had not undergone directed selection for limiting the size of the donor fragment surrounding the target gene, except for the lines combining Yr15 and Yr24 where the proximal region of Yr15 was removed to incorporate Yr24. Thus, one of our aims was to evaluate donor fragment sizes in ILs developed by backcrosses under conventional phenotypic selection compared with those developed with MAS.
Our results demonstrate that the donor fragment can be dramatically shortened through conventional breeding. Among 19 ILs obtained by conventional breeding, two (10.5 %) possessed donor fragments delimited at both sides by the SSR markers used in the current study. Among the lines containing the Yr15 gene that were investigated, the smallest donor fragment, less than 8.5 cM, was found in Excalibur-Yr15, which was developed by conventional backcross breeding. In comparison, the shortest donor fragment developed by MAS was less than 12 cM (IL HSB-2801). Reduction in the length of the donor fragment by conventional breeding depends not only on the recombination rate in the chromosomal interval surrounding the target gene, but also on the severity of linked negative traits and on the difficulty of recognizing these traits during phenotyping. Hence, although the shortest donor fragment was indeed obtained by conventional breeding, MAS provided a more consistent reduction in linkage drag even when no selection for recombination was applied.
In the Australian program, Avocet-Yr24Yr15 was obtained by MAS using marker Xgwm11 and was then backcrossed conventionally to introgress Yr15 into additional wheat cultivars (Zakari et al. 2003; Bariana unpublished results). All ILs obtained by these crosses contained relatively short donor fragments, bordered at both sides of Yr15 with RP alleles as identified in the current study by SSR markers. In contrast, in the American breeding program, where two markers spanning the target gene served in marker-assisted foreground selection of Yr15 (http://www.plantsciences.ucdavis.edu/MASWheat/protocols/Yr15/index.htm), all but one (UC110-Yr15) IL contained a long donor fragment, which was not delimited on the proximal side of Yr15 (Table 1). The shorter segments in the Yr24Yr15 lines were determined by specific selection for recombination events on the proximal side of Yr15. Since Yr24 is close to Yr15, this resulted in the elimination of most of the original G25 alleles on the proximal side.
The evidence presented here thus highlights the beneficial role of marker choice in MAS programs and the consequent power to both rapidly shorten the segment around a selected gene and even to combine the gene with other desirable traits (e.g., Yr15Yr24). Marker choice in turn depends on the availability of sufficient markers to choose among, which can be limited by lack of polymorphism when transferring markers between populations or affected by variation in genetic positions between different mapping projects (Spielmeyer et al. 2003; Ben-David 2011). The feasibility of using MAS in practical breeding programs is determined by the reproducibility of the marker-gene association across generations and populations (Collard and Mackill 2008). Ideally, markers used for MAS should be diagnostic for traits in a wide range of breeding materials, clearly discriminating between cultivars by their expression of the target trait. SSR markers are widely used in major cereals because these markers are reproducible, codominant, and polymorphic, relatively simple and cheap to use, and generally highly heritable (Gupta et al. 1999; Gupta and Varshney 2000).
Given a particular recombination rate for the region carrying the introgressed gene of interest, the backcross number will affect the length of the donor fragment. It has been calculated that a minimum of six backcrosses are normally required to restore (on average) 99 % of the genome of the recurrent parent when transferring one dominant gene (Stam and Zeven 1981). Because of the phenotype bias of conventional breeding methods, the donor segment can remain very large even with many BC generations (Young and Tanksley 1989). Young and Tanksley (1989) estimated that the size of donor segments may vary from 4 to more than 51 cM in NILs of tomato even after 11 backcross generations. In the current study, lines produced with the shortest donor segments were obtained by the Australian breeding program through five or six backcrosses for both conventional and MAS breeding. The Yr15Yr24 lines, however, were specifically selected to reduce the G25 proximal region of Yr15. Timonova et al. (2013) showed that the length of the At chromosome fragment introgressed into common wheat from Triticum timopheevii was significantly reduced after the third backcross generation. Zhou et al. (2005), who developed 33 wheat NILs for 22 powdery mildew resistance genes, reported that some NILs developed by backcrossing for four generations with MAS were as similar to the RPs as those obtained after six or more conventional backcross generations, therefore demonstrating the high efficiency of MAS.
By screening a large number of Yr15 ILs and RPs, we assessed the utilization of MAS for the practical introgression of Yr15 into various tetraploid and hexaploid wheat accessions. Our results show that markers Xbarc8 and Xgwm413 can be used efficiently to select for Yr15 and, if desired, the further markers Xwmc406 and Xgwm273 can be used in combination with these to select for recombination events in the Yr15 flanking regions (Table 1). However, in some cases, recombination may occur between the marker and the target gene due to loose linkage (Sharp et al. 2001), and therefore, the use of two markers, flanking the gene on both sides, is preferable.
Yr15 confers remarkably broad spectrum resistance against a large collection of Pst isolates from all over the world (Gerechter-Amitai et al. 1989; Niu et al. 2000; Bariana et al. 2007; Murphy et al. 2009; Chen et al. 2010; Randhawa et al. 2014). The genetic mapping of Yr15 indicates that Yr15 behaves as a single dominant gene. Nevertheless, because plant resistance genes tend to cluster (Hammond-Kosack and Jones 1997; McHale et al. 2006; Michelmore and Meyers 1998; Staskawicz et al.1995), it remains formally possible that Yr15 resides within a tightly linked cluster of genes that confers resistance in a more complex fashion. Indeed, wheat chromosome arm 1BS contains additional stripe rust resistance genes including Yr9, Yr10, Yr24, and YrH52 (Li et al. 2006; Lin and Chen 2007; McIntosh et al. 2008). However, the high virulence of isolate 5006 used in mapping (Chen et al. 2010) increases the probability that the resistance conferred by the introgressed segment is due to a monogenic Yr15. Final proof of the single-gene hypothesis will need to await positional cloning of Yr15 and verification of its utility in the absence of flanking regions.
For pyramiding genes for one or more traits of interest, MAS is highly useful, especially if these genes cannot be distinguished by phenotypic selection (Stuber 1991; He et al. 2004). MAS had been earlier used to pyramid three bacterial blight resistance genes (xa5, xa13, and Xa21) into indica rice (Singh et al. 2001). Gene pyramiding has a great advantage in the introgression of disease resistance genes because, while individual genes may not confer durable resistance, the combination of four or five genes can provide durable resistance for many years (Joshi and Nayak 2010). Although Yr15 has already succeeded in conferring stripe rust resistance for many years in many introgression lines around the world, its application in a pyramid of resistance genes could extend its lifetime. Indeed, Yr15 has been combined with Yr24 in all HSB lines reported in Table 1. All the commercial varieties including Yr15 in California also include Yr5 specifically for this purpose. These varieties, which currently cover 14 % of the acreage of common wheat in California, have not shown negative traits associated with the introgression of either of these genes.
Conclusion
The molecular markers validated here can serve the wheat breeding community to accelerate the introgression of Yr15 into lines of interest. The nearest flanking markers Xbarc8 and Xgwm413 were consistent in distinguishing between the allele of the segment originating from the G25 donor line and the alleles originating from the susceptible recurrent parents and hence are the current best choice for MAS of Yr15. The introgression of Yr15 has proved to be very useful in controlling the stripe rust epidemics in California and might be of use in the rest of the western USA (Chen 2005, 2007; Chen et al. 2002, 2010). The practical exploitation of wild emmer wheat for genes of interest beneficial to agriculture serves as a strong argument for conservation of the natural habitats in which wild emmer wheat populations grow.
References
Aaronsohn A (1910) Agricultural and botanical explorations in Palestine. Bureau of Plant Industry Bulletin 180, USDA
Bariana H, Brown G, Bansal U, Miah H, Standen G, Lu M (2007) Breeding triple rust resistant wheat cultivars for Australia using conventional and marker-assisted selection technologies. Crop Pasture Sci 58:576–587
Brennan JP, Murray GM (1998) Economic importance of wheat diseases in Australia. Wagga Wagga, NSW Agriculture
Brouwer DJ, St Clair DA (2004) Fine mapping of three quantitative trait loci for late blight resistance in tomato using near isogenic lines (NILs) and sub-NILs. Theor Appl Genet 108:628–638
Cavanagh CR, Taylor J, Larroque O, Coombes N, Verbyla AP, Nath Z, Kutty I, Rampling L, Butow B, Ral JP, Tomoskozi S, Balazs G, Bekes F, Mann G, Quail KJ, Southan M, Morell MK, Newberry M (2010) Sponge and dough bread making: genetic and phenotypic relationships with wheat quality traits. Theor Appl Genet 121:815–828
Chen XM (2005) Epidemiology and control of stripe rust [Puccinia striiformis f. sp tritici] on wheat. Can J Plant Pathol 27:314–337
Chen XM (2007) Challenges and solutions for stripe rust control in the United States. Aust J Agric Res 58:648–655
Chen XM, Moore M, Milus EA, Long DL, Line RF, Marshall D, Jackson L (2002) Wheat stripe rust epidemics and races of Puccinia striiformis f. sp. tritici in the United States in 2000. Plant Dis 86:39–46
Chen XM, Penmanb L, Wanb A, Cheng P (2010) Virulence races of Puccinia striiformis f. sp. tritici in 2006 and 2007 and development of wheat stripe rust and distributions, dynamics, and evolutionary relationships of races from 2000 to 2007 in the United States. Can J Plant Pathol 32:315–333
Cheng J, Yan J, Sela H, Manisterski J, Lewinsohn D, Nevo E, Fahima T (2010) Pathogen race determines the type of resistance response in the stripe rust—Triticum dicoccoides pathosystem. Physiol Plantarum 139:269–279
Collard BC, Mackill DJ (2008) Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Philos Trans R Soc Lond B Biol Sci 363:557–572
Fahima T, Röder M, Grama A, Nevo E (1998) Microsatellite DNA polymorphism divergence in Triticum dicoccoides accessions highly resistant to yellow rust. Theor Appl Genet 96:187–195
Ganal MW, Röder MS (2007) Microsatellite and SNP markers in wheat breeding. In: Varshney RK, Tuberosa R (eds) Genomics-assisted crop improvement, vol 2. Springer Verlag, Heidelberg, pp 1–24. ISBN 978-1-4020-6296-4
Gerechter-Amitai ZK, van Silfhout CH, Grama A, Kleitman F (1989) Yr15—a new gene for resistance to Puccinia striiformis in Triticum dicoccoides sel. G25. Euphytica 43:187–190
Gerechter-Amitai ZK, Grama A, Kleitman F, Daos A (1992) Improvement of cultivated wheat by transfer of high protein potential and resistance to powdery mildew and yellow rust from wild emmer wheat. The Netherlands Ministry of Development Cooperation, The Hague
Grama A, Gerechter-Amitai ZK (1974) Inheritance of resistance to stripe rust (Puccinia striiformis) in crosses between wild emmer (Triticum dicoccoides) and cultivated tetraploid and hexaploid wheats. II. Triticum aestivum. Euphytica 23:393–398
Grama A, Gerechter-Amitai ZK, Blum A, Rubenthaler GL (1982) Breeding bread wheat cultivars for high protein content by transfer of protein genes from Triticum dicoccoides. In: Proceedings of the seed protein improvement by nuclear techniques, Research Coordination Meetings, IAEA, Vienna, December 1982
Gupta PK, Varshney RK (2000) The development and use of microsatellite markers for genetic analysis and plant breeding with emphasis on bread wheat. Euphytica 113:163–185
Gupta PK, Varshney RK, Sharma PC, Ramesh B (1999) Molecular markers and their applications in wheat breeding. Plant Breed 118:369–390
Hammond-Kosack KE, Jones JD (1997) Plant disease resistance genes. Annu Rev Plant Physiol Plant Mol Biol 48:575–607
He Y, Li X, Zhang J, Jiang G, Liu S, Chen S, Tu J, Xu C, Zhang Q (2004) Gene pyramiding to improve hybrid rice by molecular marker techniques. In: Proceedings of the 4th international crop science congress, Brisbane, Australia
Hill CB, Taylor JD, Edwards J, Mather D, Bacic A, Langridge P, Roessner U (2013) Whole-genome mapping of agronomic and metabolic traits to identify novel quantitative trait loci in bread wheat grown in a water-limited environment. Plant Physiol 162:1266–1281s
Holland JB (2004) Implementation of molecular markers for quantitative traits in breeding programs—challenges and opportunities. In: Proceedings of the 4th international crop science congress, Brisbane, Australia
Hospital F (2001) Size of donor chromosome segments around introgressed loci and reduction of linkage drag in marker-assisted backcross programs. Genetics 158:1363–1380
Jacobsen E, Schouten HJ (2007) Cisgenesis strongly improves introgression breeding and induced translocation breeding of plants. Trends Biotechnol 25:219–223
Joshi RK, Nayak S (2010) Gene pyramiding-A broad spectrum technique for developing durable stress resistance in crops. Biotechnol Mol Biol Rev 5:51–60
Kidwell KK, Osborn TC (1992) Simple plant DNA isolation procedures. In: Beckmann JS, Osborn TC (eds) Plant genomes: methods for genetic and physical mapping. Kluwer Academic Publishers, Dordrecht, pp 1–13
Koebner R, Summers RW (2003) 21st century wheat breeding: plot selection or plate detection? Trends Biotechnol 21:59–63
Li GQ, Li ZF, Yang WY, Zhang Y, He ZH, Xu SC, Singh RP, Qu YY, Xia XC (2006) Molecular mapping of stripe rust resistance gene YrCH42 in Chinese wheat cultivar Chuanmai 42 and its allelism with Yr24 and Yr26. Theor Appl 112:1434–1440
Lin F, Chen XM (2007) Genetics and molecular mapping of genes for race-specific all-stage resistance and non-race-specific high-temperature adult-plant resistance to stripe rust in spring wheat cultivar Alpowa. Theor Appl Genet 114:1277–1287
Line RF, Qayoum A (1992) Virulence, aggressiveness, evolution, and distribution of races of Puccinia striiformis (the cause of stripe rust of wheat) in North America, 1968–87. US Dep Agric Agric Res Serv Tech Bull 1788
McDonald BA, Linden C (2002) The population genetics of plant pathogens and breeding strategies for durable resistance. Euphytica 124:163–180
McHale L, Tan X, Koehl P, Michelmore RW (2006) Plant NBS-LRR proteins: adaptable guards. Genome Biol 7:212–222
McIntosh RA, Wellings CR, Park RF (1995) Wheat rusts: an atlas of resistance genes. CSIRO, Melbourne, Australia
McIntosh RA, Silk J, The TT (1996) Cytogentic studies in wheat. XVII. Monosomic analysis and linkage relationships of the gene Yr15 for resistance to stripe rust. Euphytica 89:395–399
McIntosh RA, Yamazaki Y, Dubcovsky J, Rogers J, Morris C, Somers DJ, Appels R, Devos KM (2008) Catalogue of gene symbols for wheat. In: Proceedings of the 11th international wheat genetics symposium, pp 1–59
Mester D, Ronin Y, Minkov D, Nevo E, Korol AB (2003) Constructing large-scale genetic maps using an evolutionary strategy algorithm. Genetics 165:2269–2282
Michelmore RW, Meyers BC (1998) Clusters of resistance genes in plants evolve by divergent selection and a birth-and-death process. Genome Res 8:1113–1130
Murphy LR, Santra D, Kidwell K, Yan G, Chen X, Campbell KG (2009) Linkage maps of wheat stripe rust resistance genes Yr5 and Yr15 for use in marker-assisted selection. Crop Sci 49:1786–1790
Nevo E, Korol AB, Beiles A, Fahima T (2002) Evolution of wild emmer and wheat improvement. Springer, Heidelberg. ISBN 978-1-4020-6297-1
Niks RE, Parlevliet JE, Lindhout P, Bai Y (2011) Breeding crops with resistance to diseases and pests. Wageningen Academic Publishers, Wageningen, p 198
Niu YC, Qiao Q, Wu LR (2000) Postulation of resistance genes to stripe rust in commercial wheat cultivars from Henan, Shandong and Anhui provinces. Acta Phytopathol Sin 30:122–128
Panigrahi J, Mishra RR, Sahu AR, Rath SC, Seth S, Mishra SP (2013) Marker-assisted breeding for simple inherited traits conferring stress resistance in crop plants. The Ecoscan 02/2013; III(Special):217–233
Peleg Z, Saranga Y, Suprunova T, Ronin Y, Röder MS, Kilian A, Korol AB, Fahima T (2008) High-density genetic map of durum wheat × wild emmer wheat based on SSR and DArT markers. Theor Appl Genet 117:103–115
Peng JH, Fahima T, Röder MS, Huang OY, Dahan A, Li YC, Grama A, Nevo E (2000) High-density molecular map of chromosome region harboring stripe-rust resistance gene YrH52 and Yr15 derived from wild emmer wheat, Triticum dicoccoides. Genetica 109:199–210
Qayoum A, Line RF (1985) High-temperature, adult-plant resistance to stripe rust of wheat. Phytopathology 75:1121–1125
Randhawa M, Bansal U, Valárik M, Klocová B, Doležel J, Bariana H (2014) Molecular mapping of stripe rust resistance gene Yr51 in chromosome 4AL of wheat. Theor Appl Genet 127:317–324
Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Sharp PJ, Johnston S, Brown G, McIntosh RA, Pallotta M, Carter M, Jones MGK (2001) Validation of molecular markers for wheat breeding. Crop Pasture Sci 52:1357–1366
Simko I, Pechenick DA, McHale LK, Truco MJ, Ochoa OE, Michelmore RW, Scheffler BE (2009) Association mapping and marker-assisted selection of the lettuce dieback resistance gene Tvr1. BMC Plant Biol 9:135
Singh S, Sidhu JS, Huang N, Vikal Y, Li Z, Brar DS, Dhaliwal HS, Khush GS (2001) Pyramiding three bacterial blight resistance genes (xa5, xa13 and Xa21) using marker-assisted selection into indica rice cultivar PR106. Theor Appl Genet 102:1011–1015
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Song QJ, Fickus EW, Cregan PB (2002) Characterization of trinucleotide SSR motifs in wheat. Theor Appl Genet 104:286–293
Sourdille P, Singh S, Cadalen T, Brown-Guedira G, Gay G, Qi L, Qi LL, Dufour P, Murigneux A, Bernard M (2004) Microsatellite-based deletion bin system for the establishment of genetic-physical map relationships in wheat (Triticum aestivum L.). Funct Integr Genomics 4:12–25
Spielmeyer W, Sharpb PJ, Lagudaha ES (2003) Identification and validation of markers linked to broad-spectrum stem rust resistance gene Sr2 in wheat (Triticum aestivum L.). Crop Sci 43:333–336
Stam P, Zeven AC (1981) The theoretical proportion of the donor genome in near-isogenic lines of self-fertilizers bred by backcrossing. Euphytica 30:227–238
Staskawicz BJ, Ausubel FM, Baker BJ, Ellis JG, Jones JDG (1995) Molecular genetics of plant disease resistance. Science 268:661–667
Stubbs RW (1985) Stripe rust. In: Roelfs AP, Bushnell WR (eds) The cereal rusts, vol 2., Diseases, distribution, epidemiology, and controlAcademic Press, Orlando, FL, USA, pp 61–101
Stuber CW (1991) Biochemical and molecular markers in plant breeding. Plant Breed Rev 9:37–61
Sun GL, Fahima T, Korol AB, Turpeinen T, Grama A, Ronin YI, Nevo E (1997) Identification of molecular markers linked to the Yr15 stripe rust resistance gene of wheat originated in wild emmer wheat, Triticum dicoccoides. Theor Appl Genet 95:622–628
Tester M, Langridge P (2010) Breeding technologies to increase crop production in a changing world. Science 327:818–822
Timonova EM, Leonova IN, Röder MS, Salina EA (2013) Marker-assisted development and characterization of a set of Triticum aestivum lines carrying different introgressions from the T. timopheevii genome. Mol Breed 31:123–136
Tyrka M, Perovic D, Wardynska A, Ordon F (2008) A new diagnostic SSR marker for selection of the Rym4/Rym5 locus in barley breeding. J Appl Genet 49:127–134
van Silfhout CH, Grönewegen LJM (1984) The use of T. dicoccoides in winter wheat breeding. Vle conf. eur. et méditerr. Sur les rouilles des cereals. Grignon, 4–7 Septembre 1984. Ed. INRA Publ., 1984 (Les colloques de l’INRA, nr 25). Dissertation
Xie W, Ben-David R, Zeng B, Dinoor A, Xie C, Sun Q, Röder MS, Fahoum A, Fahima T (2012) Suppressed recombination rate on 6VS/6AL translocation region carrying the Pm21 locus introgressed from Haynaldia villosa into hexaploid wheat. Mol Breed 29:399–412
Yang ZP, Gilbert J, Somers DJ, Fedak G, Procunier JD, McKenzie IH (2003) Marker assisted selection of fusarium head blight resistance genes in two doubled- haploid populations of wheat. Mol Breed 12:309–317
Young ND, Tanksley SD (1989) RFLP analysis of the size of chromosomal segments retained around the Tm-2 locus of tomato during backcross breeding. Theor Appl Genet 77:353–359
Zakari A, McIntosh RA, Hovmoller MS, Wellings CR, Shariflou MR, Hayden M, Bariana HS (2003) Recombination of Yr15 and Yr24 in chromosome 1BS. In: Pogna NE, Romano N, Pogna EA, Galterio G (eds) Proc 10th Int Wheat Genet Symp, vol 1. Instituto Sperimentale per la Cerealcoltura, Rome, pp 417–420
Zhou R, Zhu Z, Kong X, Huo N, Tian Q, Li P, Jia J (2005) Development of wheat near-isogenic lines for powdery mildew resistance. Theor Appl Genet 110:640–648
Acknowledgments
This work was supported by grants from Sixth Framework Programme (FP6) of the European Union through the BioExploit project (CT-2005-513949), the United States—Israel Binational Agricultural Research and Development Fund (BARD), Grant IS-4628-13, and the Israel Science Foundation equipment Grant 1719/08. Dina Raats is grateful for the Eshkol Fellowship awarded by the Israeli Ministry of Science. Dr. J. Dubcovsky acknowledges support from HHMI, GBMF, and USDA-NIFA Grant 2011-68002-30029. The authors would like to acknowledge the help of the late Dr. Z. Gerechter-Amitai from the Agricultural Research Organization, Beit Dagan, Israel, Dr. X. Zhang from the University of California, and Drs. R. McIntosh and T. The from the University of Sydney, Australia, in providing us with the Yr15 introgression lines and the corresponding recipient lines, together with their pedigree information. The authors thank Dr. J. Manisterski and P. Ben-Yehuda from Tel Aviv University for providing spores of Puccinia striiformis isolate 5006 (race 38E134). The authors would like to thank O. Gurevich and T. Kis-Papo from the University of Haifa, Israel for their skillful technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Elitsur Yaniv and Dina Raats have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Yaniv, E., Raats, D., Ronin, Y. et al. Evaluation of marker-assisted selection for the stripe rust resistance gene Yr15, introgressed from wild emmer wheat. Mol Breeding 35, 43 (2015). https://doi.org/10.1007/s11032-015-0238-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-015-0238-0