Abstract
The large natural variation existing in Brassica oleracea offers a promising approach to improving B. napus (rapeseed). However, the cytogenetic and genetic characterizations of the interspecific hybridization between B. napus and B. oleracea remain poorly understood. Here, the chromosome behavior of F1 triploid hybrids between B. napus and B. oleracea was observed. Various chromosome pairings in pollen mother cells at diakinesis were found with the predominant configuration of 9II + 10I. The segregation pattern of 9:19 had the highest frequency relative to theoretical distribution estimated at anaphase I. Although the fertility was poor in the F1 generation, it recovered to normal levels in only a few generations. Additionally, B. napus-like individuals in the F3 and F4 generations, referred as new-type rapeseed, showed diverse genetic variation relative to current B. napus and strong heterotic potential. Accordingly, a significantly positive correlation between the introgressed B. oleracea genomic components and heterosis was observed in hybrids made with the new-type rapeseed lines. Our data suggest that the introgression of genetic components of B. oleracea can expand the genetic variation and improve the heterotic potential of rapeseed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Modern agricultural science has significantly improved seed production of crops, such as oil crops with an increase in yield of 125 % between 1985 and 2005 (Foley et al. 2011). However, the genetic variation in crops has been continuously reduced by modern plant breeding (Tanksley and McCouch 1997). For example, Brassica napus (rapeseed, AACC, 2n = 38), one of the main oil crops in the world, possesses a narrower genetic basis than its progenitor species, B. rapa (AA, 2n = 20) and B. oleracea (CC, 2n = 18) (Becker et al. 1995; Seyis et al. 2003). It has been suggested that an approach to broadening the genetic diversity of rapeseed could be the introgression of genetic components from its parental species (Jesske et al. 2013; Seyis et al. 2003).
In the practical breeding program of rapeseed, besides sexual crosses and protoplast fusion between two parental species (Eickermann et al. 2011; Ren et al. 2000; Wen et al. 2008), the cross of rapeseed with B. rapa has been widely adopted due to the high crossability between them and the high frequency of euploids produced from such a cross (Leflon et al. 2006, 2010; Liu 2000; Lu and Kato 2001; Mikkelsen et al. 1996; Olsson 1960; Qian et al. 2005; Shiga 1970; Zhou and Scarth 1995). However, there are few examples of interspecific hybridization between rapeseed and B. oleracea (Ayotte et al. 1988a, b; Chiang et al. 1978; Namai 1987; Quazi 1988; Ripley and Beversdorf 2003). Ultimately, this has left us with a poor understanding of the genetic and cytogenetic characterizations of interspecific hybridization between rapeseed and B. oleracea. Furthermore, the impacts of such activities on yield heterosis require more research, since it has been repeatedly observed that genetic diversity is lower in the C genome than the A genome of rapeseed (Bancroft et al. 2011; Bus et al. 2011; Delourme et al. 2013; Zhao et al. 2013), thus making the C genome as an obvious breeding target (Rahman et al. 2011). In the present study, the chromosomal behavior of an interspecific hybrid created between rapeseed and B. oleracea was characterized, and the B. napus-like progeny, referred as new-type rapeseed, were evaluated for genetic variation and heterotic potential. Our data suggest that it is easy to develop new-type rapeseed from the hybrids between B. napus and B. oleracea, and it possesses diverse genetic variation from current rapeseed and strong heterotic potential.
Materials and methods
Plant materials and field trials
An interspecific hybrid between B. napus and B. oleracea was developed as described in previous research (Li et al. 2013). Briefly, a line of B. oleracea var. acephala, SWU01 was employed as a female to cross with an elite cultivar of B. napus var. Zhongshuang 9, and the immature embryos were rescued 10 days after pollination and reproduced via asexual culture on MS agar medium. Clones of the F1 interspecific hybrid were transplanted in an isolated environment to develop F2 via open pollination, while the successive generations (F3 and F4) were developed via self-pollination.
Fifty-one new-type rapeseed lines (N) in the F4 generation demonstrating good fertility were employed to evaluate genetic diversity relative to a set of current accessions, which were selected randomly from three diverse gene pools of rapeseed (Becker et al. 1995; Bus et al. 2011; Diers and Osborn 1994; Hasan et al. 2006; Qian et al. 2006). This group comprised 15 spring (S), 16 winter (W) and 16 semi-winter (SW) accessions (Supplementary Materials S1).
In order to evaluate heterotic potential, 20 of the 51 new-type rapeseed lines were selected randomly to be crossed with the tester line Zhongshuang 9. The field trial was run in 2011 at two locations in the Yangtze River regions, Chongqing and Nanchang. A split-plot design comprising two blocks, hybrid block and parental line block, was employed with two replications. The plant density was designed according to farming practice, i.e. 10 plants were grown in a 2.5-m row with 0.3-m spacing, and each plot contained 30 plants in three rows. Agronomic traits, including plant height (PH), the height of first branch (HB), the inflorescence length (IL), number of branches (NB), seed yield (SY), seeds per pod (SP) and 1,000-seed weight (TSW), were evaluated at maturity.
Cytological analysis
The ovaries from young buds were collected and treated with 8-hydroxyquinoline for 3–4 h at room temperature, then fixed in Carnoy’s solution (ethanol : acetic acid = 3:1 v/v) and stored at 4 °C for chromosome number counting. The young flower buds were collected and fixed directly in Carnoy’s solution and stored at 4 °C, and the behavior of chromosomes at meiosis in pollen mother cells (PMCs) was studied according to the methods of Li et al. (1995). The fitness of chromosome segregation patterns in the F1 generation was calculated using the ratio of actual frequency to theoretical value according to the methods of Lu and Kato (2001).
The pollen grains from three flowers of each plant were stained with 1 % acetocarmine, and more than 300 pollen grains of each flower were observed under the microscope. The percentage of round and stainable pollen grains was calculated to measure pollen fertility.
Genetic diversity evaluation
The genomic DNA was isolated from young leaves, and the fingerprints of genotypes were developed with 155 primers of simple sequence repeats (SSRs) (Supplementary Material S2). The SSR bands were described by absence (0) or presence (1). The genetic distance (GD) between accessions X and Y was calculated using the formula GDxy = 1 − Nxy/(Nx + Ny), where Nxy is the number of common bands shared by accessions X and Y, and Nx and Ny are the total number of bands in accessions X and Y, respectively (Nei and Li 1979). The data from the GD matrix of 98 genotypes (51 new-type lines and 47 current rapeseed accessions) were subjected to principal component analysis (PCA) using NTSYS-pc software version 2.1 (Rohlf 1997).
The genomic components of B. oleracea in new-type rapeseed were described by the index of subgenomic components (ISG) according to the method of Qian et al. (2005): ISG = n/N × 100, where n represents the number of bands present in both new-type rapeseed and the B. oleracea parent, but absent in the B. napus parent, and N represents the total number of polymorphic SSR bands between the B. napus and B. oleracea parents.
Statistical analysis
Analysis of variance (ANOVA) was performed between hybrids across all environments using the GLM procedure of the Statistical Analysis System (SAS) (SAS Institute 1999). The comparisons between hybrid and control were performed using the F test. Pearson’s simple correlation coefficients were calculated between variables of interest. Mid-parent heterosis (MPH) was calculated as follows: MPH = 100 × (F1 − MP)/MP, where F1 = hybrid performance and MP = mean performance of both parents.
Results
Development of new-type B. napus
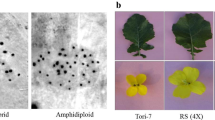
One embryo derived from the cross between B. napus and B. oleracea was rescued and reproduced clones in MS medium. A total of 184 clones were transplanted to the field. The clones exhibited intermediate phenotypes between the two parents (Fig. 1a–c). Seven clones with relatively larger flowers had been identified as hexaploid (AACCCC, 2n = 56) in our previous study (Li et al. 2013), and the remaining 177 clones were identified as triploid (ACC, 2n = 28) in the somatic cells (Fig. 1m). The hexaploid clones were isolated from triploid clones during the flowering stage.
Morphological and cytological characterizations of progenies between B. napus and B. oleracea. Seedling of parental B. napus, var. Zhongshuang 9 (a), parental B. oleracea var. acephala, SWU01 (b), hybrid F1 (c) and F4 (d); pollen fertility of F1 (e), F2 (f), F3 (g) and F4 (h); pods of F1 (i), F2 (j), F3 (k) and F4 (l); m one ovary cell of F1 with 28 chromosomes; n one pollen mother cell (PMC) of F1 at diakinesis with chromosome confugation 9II + 10I (the star symbols indicate bivalents); o one PMC of F1 at anaphase I with chromosome segregation of 9:19; p one ovary cell of a new-type rapeseed with 38 chromosomes. Bar 10 µm
The young buds of triploid clones were analyzed for chromosome behavior at meiosis. Of 234 PMCs observed at diakinesis, 141 (60.26 %) exhibited 9II +10I (Fig. 1n) and 52 exhibited 10II + 8I (22.22 %), accounting for more than 80 % of chromosome pairings. The average chromosomal configuration at diakinesis was 9.06I + 9.12II + 0.10III + 0.08IV + 0.01 V.
In order to calculate the fitness of each chromosome segregation pattern, 366 PMCs were observed at anaphase I. Despite the higher frequency of the 13:15 (45.90 %) and 14:14 (13.39 %) patterns, the fitness of the 9:19 pattern was the highest among the six patterns observed, 16.4-fold more than theoretical expectation (Figs. 1o, 2). This pattern is indicative of a relatively high possibility of producing euploid progeny (C/AC, n = 9/19).
The F1 plants had low pollen fertility (10 %), and produced less than one seed per pod (Fig. 1e, i). Likewise, very few F2 seeds were produced via open pollination. However, there was a dramatic increase in the F2 fertility, where an average of 55.5 % pollen fertility (from 10.8 to 95.6 %) was observed along with an average of 13.4 seeds per pod (from 2.1 to 24.5) among the 485 individuals tested (Fig. 1f, j). The F2 individuals with good seed-set (>15 seeds per pod) were selected for development of the next generations. The seed-set of F3 and F4 plants exhibited normal levels without significant differences from parental B. napus (Zhongshuang 9) and were significantly higher than those of F1 and F2 (Figs. 1e–l, 3). In order to identify chromosome numbers of new-type lines, 14 F3 lines were randomly selected and it was discovered that all of them had the same chromosome numbers as Zhongshuang 9 (2n = 38) (Fig. 1p). It appeared from these results that it was feasible to select new-type B. napus from the progeny of the cross between B. napus and B. oleracea.
Genetic diversity of new-type B. napus
To detect genetic diversity of new-type B. napus, 495 polymorphism bands revealed with 155 SSR primers were employed to calculate genetic distances between 98 accessions, comprising 51 new-type (N), 16 semi-winter (SW), 16 winter (W) and 15 spring (S) ecotype lines. Obvious genetic diversity was detected by PCA (Fig. 4), where the total variation explained by the first and second principal components was 24.57 and 12.10 %, respectively. Using these two principal components, spring types, winter types, semi-winter types and new-type rapeseed lines were assigned to three major clusters (Fig. 4). The average genetic distance between different types of rapeseed was greater than that within them (Table 1). Since the B. napus parent of the new-type lines was a semi-winter ecotype, a comparison of the genetic differentiation between new-type and semi-winter B. napus was performed relative to lines in the winter and spring ecotype groups. The average genetic distance between new-type lines and lines in the winter (N/W, 0.404) and spring (N/S, 0.445) ecotype groups was significantly larger than the same comparison made between lines in the semi-winter group (SW/W, 0.383; SW/S, 0.397) (P < 0.01) (Table 1). This indicated that the introgression of B. oleracea presented a viable option for widening the genetic diversity of current rapeseed.
A significant and positive correlation was detected between the index of subgenomic components (ISG) of B. oleracea in new-type lines and the genetic distance between new-type lines and semi-winter rapeseed (r = 0.79, P < 0.01), spring rapeseed (r = 0.65, P < 0.01) and winter rapeseed (r = 0.46, P < 0.01), although partial genetic components of B. oleracea were transferred into those new-type lines, with an average ISG of 29.9 %, varying from 18.41 to 49.73 %. These findings indicated that new-type rapeseed carrying more genomic components from B. oleracea was more genetically distant from current rapeseed.
Field performance of new-type rapeseed
To evaluate the potential of new-type rapeseed for seed production, 20 F4 new-type lines were randomly selected and evaluated for seed yield and yield-related traits, together with the hybrids between them and Zhongshuang 9 at two locations, Chongqing and Nanchang. Heavy rain in Nanchang during flowering significantly reduced the seed yield at Nanchang (av. 1.41 T/ha) relative to Chongqing (av. 1.96 T/ha) (P < 0.01). Nevertheless, a positive correlation was detected for the yield of lines grown in the two environments (r = 0.35, P = 0.035). There was no significant difference between the seed yield of the new-type line and Zhongshuang 9. However, the new-type line was higher than Zhongshuang 9 by 17.09 % for number of branches (P = 0.018), 4.62 % for seeds per pod, 3.99 % for plant height (P = 0.015) and 5.16 % for main inflorescence length. Furthermore, the hybrids made between new-type lines and Zhongshuang 9 were significantly better than Zhongshuang 9 for seed yield (31.63 %, P = 0.05), seeds per pod (11.61 %, P = 0.002), number of branches (16.11 %, P = 0.01), plant height (10.61 %, P < 0.01), height of first branch (6.17 %, P = 0.01) and main inflorescence length (12.90 %, P < 0.01) (Fig. 5). A significant positive correlation was noted between ISG and mid-parental heterosis between the new-type line and Zhongshuang 9 for seed yield (r = 0.54, P = 0.02).
Discussion
Rapeseed is an important but young oil crop, which was domesticated no more than 400 years ago (Gomez-Campo and Prakash 1999; Toxopeus 1979). The short history of domestication and the reproductive isolation from the progenitor species might be critical causes for the narrow genetic diversity in rapeseed, particularly in the C subgenome of rapeseed (Bus et al. 2011; Jesske et al. 2013; Li et al. 2013). In comparison, B. oleracea possesses wide genetic variation, and differs substantially from the C subgenome of rapeseed (Mei et al. 2010, 2011a, b). In order to take advantage of this variation, the genomic components of B. oleracea were introgressed during the creation of new-type rapeseed developed in this study. Our data indicated that this new-type rapeseed exhibited diverse genetic variation from current rapeseed. This is likely due to the introgression of B. oleracea. In the F1 triploid derived from crossing B. oleracea with B. napus, variable chromosome pairings and segregation were observed in this and previous studies (Namai 1987), which indicated a strong possibility of homologous and homoeologous recombination and rearrangement. Subsequently, this could have induced genomic changes in the progeny, such as deletion, duplication and translocation as reported in previous studies (Cifuentes et al. 2010; Gaeta and Pires 2010).
The transfer of genetic variation from the progenitor species into rapeseed has traditionally occurred by resynthesizing B. napus from crosses between B. rapa and B. oleracea. However, those synthetic lines have limited use in breeding programs due to wild or disadvantageous characteristics from both parental species (Fujii and Ohmido 2011; Gaeta et al. 2007; Girke et al. 2012a, b; Jesske et al. 2013; Szadkowski et al. 2010, 2011; Xiong et al. 2011). In practical breeding programs, the synthetic lines are normally backcrossed with current rapeseed for several generations to improve selection against the negative traits originating from the parental species (Becker et al. 1995), resulting in a dilution in the genetic variance gained from parental species. In comparison, interspecific hybridization between rapeseed and the progenitor species can often minimize the negative impacts in lines derived from crosses between the diploid parental species. For example, interspecific hybridization between B. napus and B. rapa has been widely employed and resulted in the release of some cultivars in China and Japan (Liu 2000). However, few examples of interspecific hybridization between B. napus and B. oleracea exist in practical breeding programs, possibly due to the low crossability between B. napus and B. oleracea. Our data suggested that F1 lines could be developed between B. napus and B. oleracea via embryo rescue and new-type rapeseed could easily be developed from the progenies. It was interesting that those new-type lines possessed the strong potential to widen the genetic diversity and improve the heterosis of current rapeseed. Similar observations have been reported in other introgression lines of rapeseed carrying genetic components of B. rapa and B. carinata (BBCC) (Fu et al. 2012; Li et al. 2006; Qian et al. 2005; Xiao et al. 2010; Zou et al. 2010). Thus, it seems that harvesting genetic diversity from B. oleracea via interspecific hybridization between B. oleracea and B. napus is an important breeding strategy in rapeseed.
For many years, authors have suggested that B. rapa and B. oleracea are the parents of B. napus. UN (1935) proposed a process of origination via a cross between 2× B. rapa and 2× B. oleracea, followed by genome duplication. This scenario remains possible since it is available for direct synthesis of rapeseed via crossing between B. rapa and B. oleracea (Girke et al. 2012a, b; Jesske et al. 2013; Ren et al. 2000; Seyis et al. 2003; Wen et al. 2008). The relatively high frequency of euploid progency in ACC triploid hybrids observed in this study, as well as in AAC triploid hybrids detected in a previous study (Lu and Kato 2001), suggest that an alternative pathway, a cross between 2× B. rapa and 4× B. oleracea or between 4× B. rapa and 2× B. oleracea, followed by successive self-pollination, may be a more likely possibility for the origination of B. napus, although we did not perform those crosses.
References
Ayotte R, Harney PM, Souza MV (1988a) The transfer of triazine resistance from Brassica napus L. to B. oleracea L. II. Morphology, fertility and cytology of the F1 hybrid. Euphytica 37:189–197
Ayotte R, Harney PM, Souza MV (1988b) The transfer of triazine resistance from Brassica napus L. to B. oleracea L. III. First backcross to parental species. Euphytica 38:137–142
Bancroft I, Morgan C, Fraser F, Higgins J, Wells R, Clissold L, Baker D, Long Y, Meng J, Wang X, Liu S, Trick M (2011) Dissecting the genome of the polyploid crop oilseed rape by transcriptome sequencing. Nat Biotechnol 29:762–766
Becker HC, Engqvist GM, Karlsson B (1995) Comparison of rapeseed cultivars and resynthesized lines based on allozyme and RFLP markers. Theor Appl Genet 91:62–67
Bus A, Körber N, Snowdon RJ, Stich B (2011) Patterns of molecular variation in a species-wide germplasm set of Brassica napus. Theor Appl Genet 123:1413–1423
Chiang BY, Grant WF, Chiang MS (1978) Transfer of resistance to race 2 of plasmodiophora Brassica from Brassica napus to cabbage (B. oleracea var. capitata). II. Meiosis in the interspecific hybrids between B. napus and 2x and 4x cabbage. Euphytica 27:81–93
Choi SR, Teakle GR, Plaha P, Kim JH, Allender CJ, Beynon E, Piao ZY, Soengas P, Han TH, King GJ, Barker GC, Hand P, Lydiate DJ, Batley J, Edwards D, Koo DH, Bang JW, Park BS, Lim YP (2007) The reference genetic linkage map for the multinational Brassica rapa genome sequencing project. Theor Appl Genet 115:777–792
Cifuentes M, Eber F, Lucas MO, Lode M, Chèvre AM, Jenczewski E (2010) Repeated polyploidy drove different levels of crossover suppression between homoeologous chromosomes in Brassica napus allohaploids. Plant Cell 22:2265–2276
Delourme R, Falentin C, Fomeiu BF, Boillot M, Lassalle G, André I, Duarte J, Gauthier V, Lucante N, Marty A, Pauchon M, Pichon JP, Ribière N, Trotoux G, Blanchard P, Ribière N, Martinant JP, Pauquet J (2013) High-density SNP-based genetic map development and linkage disequilibrium assessment in Brassica napus L. BMC Genom 14:120
Diers BW, Osborn TC (1994) Genetic diversity of oilseed Brassica napus germplasm based on restriction fragment length polymorphisms. Theor Appl Genet 88:662–668
Eickermann M, Ulber B, Vidal S (2011) Resynthesized lines and cultivars of Brassica napus L. provide sources of resistance to the cabbage stem weevil (Ceutorhynchus pallidactylus (Mrsh.)). Bull Entomol Res 101:287–294
Foley JA, Ramankutty N, Brauman KA, Cassidy ES, Gerber JS, Johnston M, Mueller ND, O’Connell C, Ray DK, West PC, Balzer C, Bennett EM, Carpenter SR, Hill J, Monfreda C, Polasky S, Rockström J, Sheehan J, Siebert S, Tilman D, Zaks DP (2011) Solutions for a cultivated planet. Nature 478(7369):337–342
Fu D, Qian W, Zou J, Meng J (2012) Genetic dissection of intersubgenomic heterosis in Brassica napus carrying genomic components of B. rapa. Euphytica 184:151–164
Fujii K, Ohmido N (2011) Stable progeny production of the amphidiploid resynthesized Brassica napus cv. Hanakkori, a newly bred vegetable. Theor Appl Genet 123:1433–1443
Gaeta RT, Pires JC (2010) Homoeologous recombination in allopolyploids: the polyploid ratchet. New Phytol 186:18–28
Gaeta RT, Pires JC, Iniguez-Luy F, Leon E, Osborn TC (2007) Genomic changes in resynthesized Brassica napus and their effect on gene expression and phenotype. Plant Cell 19:3403–3417
Girke A, Schierholt A, Becker HC (2012a) Extending the rapeseed gene pool with resynthesized Brassica napus I: genetic diversity. Genet Resour Crop Evol 59:1441–1447
Girke A, Schierholt A, Becker HC (2012b) Extending the rapeseed gene pool with resynthesized Brassica napus II: heterosis. Theor Appl Genet 124:1017–1026
Gomez-Campo C, Prakash S (1999) Origin and domestication. In: Gomez-Campo C (ed) Biology of Brassica coenospecies. Elsevier Science B.V, The Netherlands, pp 33–58
Hasan M, Seyis F, Badani AG, Pons-Kühnemann J, Friedt W, Lühs W, Snowdon RJ (2006) Analysis of genetic diversity in the Brassica napus L. gene pool using SSR markers. Genet Resour Crop Evol 53:793–802
Jesske T, Olberg B, Schierholt A, Becker HC (2013) Resynthesized lines from domesticated and wild Brassica taxa and their hybrids with B. napus L.: genetic diversity and hybrid yield. Theor Appl Genet 126:1053–1065
Leflon M, Eber F, Letanneur JC, Chelysheva L, Coriton O, Huteau V, Ryder CD, Barker G, Jenczewski E, Chèvre AM (2006) Pairing and recombination at meiosis of Brassica rapa (AA) × Brassica napus (AACC) hybrids. Theor Appl Genet 113:1467–1480
Leflon M, Grandont L, Eber F, Huteau V, Coriton O, Chelysheva L, Jenczewski E, Chèvre AM (2010) Crossovers get a boost in Brassica allotriploid and allotetraploid hybrids. Plant Cell 22:2253–2264
Li Z, Liu HL, Luo P (1995) Production and cytogenetics of intergeneric hybrids between Brassica napus and Orychophragmus violaceus. Theor Appl Genet 91:131–136
Li M, Chen X, Meng J (2006) Intersubgenomic heterosis in rapeseed production with a partial new-type Brassica containing subgenome Ar from B. rapa and Cc from Brassica carinata. Crop Sci 46:234–242
Li Q, Mei J, Zhang Y, Li J, Ge X, Li Z, Qian W (2013) A large-scale introgression of genomic components of Brassica rapa into B. napus by bridge of hexaploid derived from hybridization between B. napus and B. oleracea. Theor Appl Genet 126:2073–2080
Liu HL (2000) Genetics and breeding in rapeseed. Chinese Agricultural Universities Press, Beijing
Long Y, Shi J, Qiu D, Li R, Zhang C, Wang J, Hou J, Zhao J, Shi L, Park B-S, Choi SR, Lim YP, Meng J (2007) Flowering time quantitative trait loci analysis of oilseed Brassica in multiple environments and genomewide alignment with Arabidopsis. Genetics 177:2433–2444
Lowe AJ, Moule C, Trich M, Edwards KJ (2003) Efficient large-scale development of microsatellites for marker and mapping applications in Brassica crop species. Theor Appl Genet 108:1103–1112
Lu C, Kato M (2001) Fertilization fitness and relation to chromosome number in interspecific progeny between Brassica napus and B. rapa: a comparative study using natural and resynthesized B. napus. Breed Sci 51:73–81
Mei J, Li Q, Yang X, Qian L, Liu L, Yin J, Frauen M, Li J, Qian W (2010) Genomic relationships between wild and cultivated Brassica oleracea L. with emphasis on the origination of cultivated crops. Genet Resour Crop Evol 57:687–692
Mei JQ, Fu Y, Qian LW, Xu XF, Li JN, Qian W (2011a) Effectively widening the gene pool of oilseed rape (Brassica napus L.) by using Chinese B. rapa in a ‘virtual allopolyploid’ approach. Plant Breed 130:333–337
Mei J, Li Q, Qian L, Fu Y, Li J, Frauen M, Qian W (2011b) Genetic investigation of the origination of allopolyploid with virtually synthesized lines: application to the C subgenome of Brassica napus. Heredity 106:955–961
Mikkelsen TR, Jensen J, Jørgensen RB (1996) Inheritance of oilseed rape (Brassica napus) RAPD markers in a backcross progeny with Brassica campestris. Theor Appl Genet 92:492–497
Namai H (1987) Inducing cytogenetical alteration by means of interspecific and intergeneric hybridization in Brassica crops. Gamma Field Symp 26:41–87
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endouncleases. Proc Natl Acad Sci USA 76:5269–5273
Olsson G (1960) Species crossing within the genus Brassica napus. II. Artificial Brassica napus L. Hereditas 46:351–396
Qian W, Chen X, Fu D, Zou J, Meng J (2005) Intersubgenomic heterosis in seed yield potential observed in a new-type of Brassica napus introgressed with partial Brassica rapa genome. Theor Appl Genet 110:1187–1194
Qian W, Meng J, Li M, Frauen M, Sass O, Noack J, Jung C (2006) Introgression of genomic components from Chinese Brassica rapa contributes to widening the genetic diversity in rapeseed (B. napus L.), with emphasis on the evolution of Chinese rapeseed. Theor Appl Genet 113:49–54
Qiu D, Morgan C, Shi J, Long Y, Liu J, Li R, Zhuang X, Wang Y, Tan X, Dietrich E, Weihmann T, Everett C, Vanstraelen S, Beckett P, Fraser F, Trick M, Barnes S, Wilmer J, Schmidt R, Li J, Li D, Meng J, Bancroft I (2006) A comparative linkage map of oilseed rape and its use for QTL analysis of seed oil and erucic acid content. Theor Appl Genet 114:67–80
Quazi MH (1988) Interspecific hybrids between Brassica napus L. and B. oleracea L. developped by embryo culture. Theor Appl Genet 75:309–318
Rahman MH, Bennett RA, Yang RC, Thiagarajah MR (2011) Exploitation of the late flowering species Brassica oleracea L. for the improvement of earliness in B. napus L.—an untraditional approach. Euphytica 177:365–374
Ren JP, Dickson MH, Earle ED (2000) Improved resistance to bacterial soft root by protoplast fusion between Brassica rapa and B. oleracea. Theor Appl Genet 100:810–819
Ripley VL, Beversdorf WD (2003) Development of self-incompatible Brassica napus (I) introgression of S-alleles from Brassica oleracea through interspecific hybridization. Plant Breed 122:1–5
Rohlf F (1997) NTSYS-PC 2.1. Numerical taxonomy and multivariate analysis system. Exeter Software. In, Setauket, NY
SAS Institute (1999) SAS Online Doc (R), version 8.0. Cary, NC, USA
Seyis F, Snowdon RJ, Luhs W, Friedt W (2003) Molecular characterization of novel resynthesized rapeseed (Brassica napus) lines and analysis of their genetic diversity in comparison with spring rapeseed cultivars. Plant Breed 122:473–478
Shiga T (1970) Rapa breeding by interspecific crossing between Brassica napus and Brassica campestris in Japan. Jpn Agric Res Quart 5:5–10
Szadkowski E, Eber F, Huteau V, Lodé M, Huneau C, Belcram H, Coriton O, Manzanares-Dauleux MJ, Delourme R, King GJ, Chalhoub B, Jenczewski E, Chèvre AM (2010) The first meiosis of resynthesized Brassica napus, a genome blender. New Phytol 186:102–112
Szadkowski E, Eber F, Huteau V, Lodé M, Coriton O, Jenczewski E, Chèvre AM (2011) Polyploid formation pathways have an impact on genetic rearrangements in resynthesized Brassica napus. New Phytol 191:884–894
Szewc-McFadden AK, Kresovich S, Bliek SM, Mitchell SE, McFerson JR (1996) Identification of polymorphic, conserved simple sequence repeats (SSRs) in cultivated Brassica species. Theor Appl Genet 93:534–538
Tanksley SD, McCouch SR (1997) Seed banks and molecular maps: unlocking genetic potential from the wild. Science 277:1063–1066
Toxopeus H (1979) The domestication of Brassica crops. In: Proceedings of Eucarpia conference on the breeding of cruciferous crops, Wageningen, Netherlands, pp 47–56
UN (1935) Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jpn J Bot 7:389–452
Uzunova MI, Ecke W (1999) Abundance, polymorphism and genetic mapping of microsatellites in oilseed rape (Brassica napus L.). Plant Breed 118:323–326
Wen J, Tu JX, Li ZY, Fu TD, Ma CZ, Shen JX (2008) Improving ovary and embryo culture techniques for efficient resynthesis of Brassica napus from reciprocal crosses between yellow-seeded diploids B. rapa and B. oleracea. Euphytica 162:81–89
Xiao Y, Chen L, Zou J, Tian E, Xia W, Meng J (2010) Development of a population for substantial new type Brassica napus diversified at both A/C genomes. Theor Appl Genet 121:1141–1150
Xiong Z, Gaeta RT, Pires JC (2011) Homoeologous shuffling and chromosome compensation maintain genome balance in resynthesized allopolyploid Brassica napus. Proc Natl Acad Sci USA 108:7908–7913
Zhao M, Du J, Lin F, Tong C, Yu J, Huang S, Wang X, Liu S, Ma J (2013) Shifts in the evolutionary rate and intensity of purifying selection between two Brassica genomes revealed by analyses of orthologous transposons and relics of a whole genome triplication. Plant J 76:211–222
Zhou YM, Scarth R (1995) Microspore culture of hybrids between Brassica napus and B. campestris. Acta Bot Sin 37:848–855
Zou J, Zhu J, Huang S, Tian E, Xiao Y, Fu D, Tu J, Fu T, Meng J (2010) Broadening the avenue of intersubgenomic heterosis in oilseed Brassica. Theor Appl Genet 120:283–290
Acknowledgments
This study is partly supported by the Fundamental Research Funds for the Central Universities (XDJK2013A013, XDJK2014C148), the Key Projects in the National Science & Technology (2010BAD01B02), 111 project (B12006), NSFC (31171585), and the open funds of the National Key Laboratory of Crop Genetic Improvement, China.
Author information
Authors and Affiliations
Corresponding author
Additional information
Qinfei Li and Qinghong Zhou have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11032_2014_153_MOESM1_ESM.xlsx
Supplementary materials S1. List of current Brassica napus lines to compare with new type lines for genetic variation revealed with simple sequence repeat markers (XLSX 11 kb)
11032_2014_153_MOESM2_ESM.xlsx
Supplementary materials S2. Distribution of loci amplified with simple sequence repeat primers. The primers were prefixed with BRAS/CB from Celera AgGen Brassica Consortium; SWUC, AG, Pod, YD, Br, h, Bn, CEN from Southwest University; Na, Ra, Ol from BBSRC (Lowe et al. 2003); NIAB from The National Institute of Agricultural Biotechnology; PUT from http://www.plantgdb.org; sR, sN, sS from Agriculture and Agri-Food Canada; BRMS from Suwabe et al. (2002); CNU from Chungnam National University; FITO from http://www.asbornlab.agronomy.wisc.edu/research/maps/ssrs.html; BN were from Szewc-McFadden et al. (1996); MR from Uzanova and Ecke (1999); sOR from Qiu et al.(2006); with ENA and GOL from Choi et al.(2007); CN from Long et al.(2007) (XLSX 17 kb)
Rights and permissions
About this article
Cite this article
Li, Q., Zhou, Q., Mei, J. et al. Improvement of Brassica napus via interspecific hybridization between B. napus and B. oleracea . Mol Breeding 34, 1955–1963 (2014). https://doi.org/10.1007/s11032-014-0153-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-014-0153-9