Abstract
Spot blotch, caused by Cochliobolus sativus, is an important foliar disease of wheat in warmer wheat-growing regions leading to significant reductions in grain yield and quality. Although inoculum levels can be reduced by planting disease-free seed, treatment of plants with fungicides and crop rotation, genetic resistance is likely to be a robust, economical and environmentally friendly tool in the control of spot blotch. The spot blotch resistant synthetic derivative ‘SYN1’ was developed from a cross between two resistance sources, Mayoor and the primary synthetic bread wheat Tksn1081/Ae. squarrosa (222) that are likely to form an important component of resistance in many elite CIMMYT bread wheats. In order to map the loci underlying the resistance of ‘SYN1’, a doubled-haploid population produced from a cross between ‘SYN1’ and the susceptible CIMMYT-derived variety Ocoroni-86 was evaluated in artificially inoculated field nurseries in the 2010–2011 and 2011–2012 crop seasons at CIMMYT’s research station in Agua Fría, Mexico. Disease assessment was performed on three or four occasions and subsequently area under disease progress curve (AUDPC) calculated. Genotyping was with genotyping by sequencing and simple sequence repeat markers. Using inclusive composite interval mapping, three genomic regions were found to have a significant effect on spot blotch AUDPC in each of the 2 years of trials with phenotypic variation explained by QSb.cim-1B of 8.5 %, 17.6 % by QSb.cim-3B and 12.3 % by QSb.cim-5A. The quantitative trait loci (QTL) mapping results showed that the favorable alleles of QSb.cim-1B, QSb.cim-3B and QSb.cim-5A were derived from the synthetic-derived bread wheat SYN1. Genotypes of the parents of SYN1 indicated that the favorable alleles at these three QTLs were all inherited from Mayoor.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Spot blotch caused by Cochliobolus sativus (anamorph: Bipolaris sorokiniana) is an important disease of wheat (Triticum aestivum L.) in warmer wheat-growing regions like eastern India, Southeast Asia, Latin America and sub-Saharan Africa (Saari 1998; Dubin and Duveiller 2000) and can substantially reduce yields (Villareal et al. 1995; Sharma and Duveiller 2006). Diseased plots yielding 43 % less than fungicide protected plots have been reported (Villareal et al. 1995). Spot blotch has also been known as Helminthosporium leaf blight or foliar blight. The increasing threat of this disease with rising global temperatures has increased the urgency for plant geneticists and breeders to identify resistance sources, map the genes and develop spot blotch resistant varieties (Joshi et al. 2007a).
Relatively little work has been reported on resistance to spot blotch. In Nepal, India and Bangladesh, Dubin et al. (1998) reported several wheat genotypes exotic to the region that had better resistance than the local commercial cultivars. More recently, several wheat genotypes with high levels of resistance to spot blotch and favorable agronomic type were reported in south Asia (Duveiller and Sharma 2005; Sharma and Duveiller 2007; Joshi et al. 2007b). The correlations of some morphological and physiological traits with spot blotch resistance have been analyzed (Rosyara et al. 2009, 2010). Results suggested that leaf tip necrosis was associated with resistance to spot blotch (Joshi et al. 2004) and that leaf photochemical efficiency and leaf greenness (measured as SPAD value) as well as the efficiency of photosystem II, which was measured as ratio of variable to maximal chlorophyll fluorescence, F v/F m, could be used as complementary traits in selecting for spot blotch resistance in wheat (Rosyara et al. 2010). Some biparental populations were produced, and the heritability of spot blotch resistance was estimated between 0.32 and 0.70 in different crosses in terms of the area under disease progress curve (AUDPC) using realized heritability procedures (Sharma et al. 2006). Studies of the resistances of Chinese bread wheats Longmai 10, Jinmai and Shanghai by Kumar et al. (2007) indicated the resistance of Jinmai was due to three genes, while the other lines each carried two resistance genes. Intercrosses between these lines indicated one gene was common to all, while transgressive segregation indicated additional diversity that could be combined. Studies on the resistances of ‘Chirya 3’ and ‘Milan/Sanghai #7’ by Neupane et al. (2007) suggested that the resistance was conditioned by a single, dominant gene.
Few reports are available on linkage of spot blotch resistance to molecular markers. Kumar et al. (2009) mapped four quantitative trait loci (QTL) for spot blotch resistance on chromosomes 2AL, 2BS, 5BL and 6DL from Yangmai 6. The leaf rust resistance gene Lr34 was reported as the major locus conferring spot blotch resistance in bread wheat line Saar and was designated as Sb1 (Lillemo et al. 2012). Four QTLs were mapped on chromosomes 2AS, 2BS, 5BL and 7DS in a population from the cross ‘Ning8201’ × ‘Sonalika’, and five QTLs were identified on chromosomes 2BS, 2DS, 3BS, 7BS and 7DS in a population issued from the cross ‘Chirya 3’ × ‘Sonalika’ (Kumar et al. 2010). Aegilops curvifolium is an exotic source of resistance that was crossed to Chinese resistance sources at CIMMYT to produce the Chirya series and Mayoor which have good levels of resistance (Van Ginkel and Rajaram, 1998). It is not clear if genes from Ae. curvifolium contribute to the resistance of these lines. Primary synthetic bread wheats are another exotic source of resistance to spot blotch (Mujeeb-Kazi et al. 2001) that have been widely used in CIMMYT’s breeding activities producing many resistant derivative lines. There are no published reports on mapping of resistance from synthetic wheats. This relatively limited map information on spot blotch resistances complicates selection for resistance as marker-assisted selection cannot be used to complement or partially substitute for phenotypic selection.
Pedigree analysis of a set of synthetic derivatives with good levels of spot blotch resistance pointed to Mayoor and a small number of primary synthetics as likely important resistance sources in CIMMYT bread wheats. In this paper, we report efforts to map the resistance of one of the derivative lines produced from a cross between Mayoor and a resistant primary synthetic bread wheat, Tksn1081/Ae. squarrosa (222). This should contribute to breeding for spot blotch resistance by allowing selection for what are likely to be widespread resistance sources in CIMMYT germplasm and also to begin understanding the diversity for spot blotch resistance currently present in the CIMMYT wheat germplasm pool.
Materials and methods
Mapping population
The synthetic hexaploid wheat line Tksn1081/Ae. squarrosa (222) and CIMMYT bread wheat ‘Mayoor’ (Pedigree: Chinese-spring/Ae. curvifolium//Glennson-81/3/Alondra/Pavon-76) were characterized as resistant to spot blotch in previous field studies and subsequently crossed to elite bread wheats to develop resistant lines with improved agronomic performance as part of routine breeding and prebreeding activities at CIMMYT. A large number of resistant progenies were developed leading to an interest in genetic mapping of resistance loci that are likely to now be widespread in elite CIMMYT breeding lines. A synthetic derivative SYN1, including both of these resistances sources in its pedigree [Mayoor//Tksn1081/Ae. squarrosa (222)], was found to be resistant to spot blotch and was crossed to a spot blotch susceptible CIMMYT-derived cultivar, Ocoroni-86 (hereafter referred to as Ocoroni), to produce a doubled-haploid (DH) population from which lines could be selected combining the resistances and incorporating them in an improved agronomic background (Table 1). In this study, we screened this population of 161 DH lines and its parents for spot blotch resistance in field trials and with genotyping by sequencing (GBS) markers (Elshire et al. 2011) and simple sequence repeat (SSR, microsatellite) markers to identify linkage groups and map the resistance.
For the initial stage of identifying genomic regions, conferring resistance to spot blotch, resistant and susceptible bulks were made up of the 10 most resistant DH lines and 10 most susceptible DH lines, respectively (Supplementary table 1). Care was taken to ensure similar phenologies in each bulk.
Disease screening
One hundred and sixty-one DH lines from the population ‘SYN1 × Ocoroni’ were evaluated in the field at CIMMYT’s Agua Fría experiment station (state of Puebla, Mexico, altitude 100 m, latitude 20.5°N, average annual rainfall 1,200 mm, temperature range 3–44 °C, wheat-growing season November–March, clay loam soils of pH 7.5–8.5) during the crop seasons 2010–2011 (denoted 2011) and 2011–2012 (denoted 2012) with two repetitions in each season. Parental lines SYN1, Ocoroni and Mayoor were evaluated in the field trial conducted in the 2012 season. Each plot comprised paired rows of 1 m length with 25 cm spacing between the rows and a 50-cm space to the adjacent plot.
Spot blotch inoculum was a mixture of virulent isolates of B. sorokiniana collected from the spot blotch nursery at the Agua Fria station in the previous screening cycle. Isolates showed similar type and color on culture media. Inoculum was increased on autoclaved sorghum grains. These were placed between the paired rows in each plot at the base of the plants. Beginning 4–6 weeks after inoculation, spot blotch severity in each plot was visually scored; four times during the 2011 season and three times during the 2012 season at 7–10 day intervals, using the double-digit scale (00–99) developed as a modification of Saari and Prescott’s severity scale to assess wheat foliar diseases (Saari and Prescott 1975; Eyal et al. 1987). The first digit (D1) indicates how far disease symptoms have progressed up the crop canopy from ground level, and the second digit (D2) is a score of the severity of symptoms based on diseased leaf area. Both D1 and D2 are scored on a scale of 0–9. For each evaluation, percentage disease severity was estimated based on the following formula:
The AUDPC was calculated using data from four disease evaluations in 2011 and three disease evaluations in 2012 using the formula below (Duveiller et al. 1998; Joshi and Chand 2002):
where Y i = spot blotch severity at time t i , t(i+1) − t i = time (days) between two disease scores, n = number of times when spot blotch was recorded.
Final disease index (FDI) was equal to % severity of the last field evaluation.
Plant height (PHT) and days to flowering (DF) were also scored in the field trials.
Genotyping
Parents of the mapping population, SYN1 and Ocoroni, were screened with 790 SSR markers selected as being evenly distributed along each chromosome based on the ITMI reference map (Ganal and Röder 2007). Markers polymorphic between the parents were used to screen resistant and susceptible bulks. Polymorphic markers differing between the contrasting bulks and two parents were used to genotype all the 161 DH lines. Twelve additional SSRs close to putatively resistance-linked loci on chromosomes 1B and 3B were used to genotype the whole population to increase genetic coverage in those regions. Eight of them were assigned onto those two linkage groups. These SSRs were also used to anchor GBS markers.
DNA isolation, PCR and polyacrylamide gel electrophoresis (12 % acrylamide gels) were performed in the Applied Biotechnology Center, CIMMYT following the protocols described in CIMMYT Laboratory protocols (2005). DNA samples of 161 DHs, SYN1 and Ocoroni were sent to Diversity Arrays Technology Pty Ltd., Australia for genotyping with GBS markers.
Statistical analysis and QTL mapping
Analysis of variance (ANOVA) of phenotypic data from the two testing cycles, correlation coefficients among different traits and comparisons of the means of different groups were performed using the statistical analysis software (SAS) (version 9.2). Heritability (h 2) was estimated from the analysis of variance using the following formula (Knapp et al. 1985):
JoinMap 4 (Van Ooijen 2006) was used to construct linkage groups. QTL mapping, QTL by environment analysis and QTL by QTL interaction were conducted by QTL IciMapping version 3.2 (Wang et al. 2011). The inclusive composite interval mapping additive method (ICIM-ADD) was used to map QTLs and analyze QTL by environment with LOD ≥2.5. The inclusive composite interval mapping epistatic method (ICIM-EPI) was used to analyze the epistatic interactions between QTLs with LOD ≥2.5. A LOD of 5 was used as the threshold for significance of interaction effects as suggested by Wang et al. (2011).
Results
Phenotypic variation
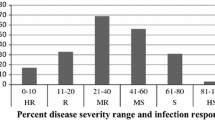
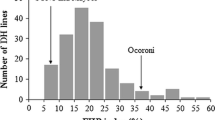
SYN1 was evaluated as highly resistant to spot blotch with an AUDPC of 133.3 in 2012 (Table 1). Mayoor, a parent of SYN1, was similarly resistant to spot blotch with an AUDPC of 182.7 in 2012. The primary synthetic parent Tksn1081/Ae. squarrosa (222) was also highly resistant to spot blotch with an AUDPC of 34.6 in 2012. Ocoroni was highly susceptible to spot blotch based on the evaluation in 2012 with an AUDPC of 1291.4 (Table 1). The means and ranges of spot blotch FDI and AUDPC of the population are listed in Table 1. The population showed large variation in spot blotch reaction among the DHs for FDI and AUDPC (Fig. 1). Spot blotch FDI ranged from 7.4 to 100 % and spot blotch AUDPC ranged from 118.5 to 1,607.4.
The analysis of variance for AUDPC in the mapping population revealed significant variation for genotype and genotype-by-year interaction and highly significant differences between years (Table 2). Variation accounted for by genotype was more than three times greater than genotype by year. The heritability of spot blotch AUDPC in the mapping population was estimated at 0.72.
Trait correlations
A high Pearson correlation coefficient (0.91) was observed between spot blotch AUDPC and FDI in the ‘SYN1 × Ocoroni’ DHs (Table 3). PHT had a low negative correlation with FDI (−0.42**) and AUDPC (−0.44**). DF had low negative correlation with AUDPC (−0.30**), but moderate negative correlation with FDI (−0.53**). No significant correlation between DF and PHT was observed in this population.
Genotypic data and linkage groups
Two hundred and eighty-two SSRs were identified with polymorphism between the two parents of the mapping population. Ninety-two of them were used to genotype the whole population, and 106 loci were identified by these markers. A total of 1,391 GBS markers were reported polymorphic in this population. Of these, 334 GBS markers had missing data in 30 or more DH lines and were not used in map construction. Each of the remaining markers was tested for goodness of fit to the expected 1:1 ratio of SYN1 to Ocoroni allele to identify segregation distortion. The ratios ranged from 0.50 to 4.21:1.0. Chi-squared tests for goodness of fit indicated that 295 of the remaining GBS markers had significant departures from the expected 1:1 ratio. However, these markers were retained in construction of linkage maps. Thirty-four linkage groups were constructed. Among them, 11 groups were anchored with at least two SSRs on chromosomes 1A, 1B, 2A, 2B, 2D, 3B, 4A, 4B, 5A, 5B and 7B.
QTL mapping
In the mapping population ‘SYN1 × Ocoroni’, three QTLs (QSb.cim-3B, QSb.cim-5A and QSb.cim-1A) were detected for spot blotch AUDPC in 2011 (Table 4). The LOD values ranged from 2.6 to 2.8 and the corresponding phenotypic variation explained (PVE) ranged from 5.3 to 8.4 %. Three QTLs QSb.cim-1B, QSb.cim-3B and QSb.cim-5A were detected for spot blotch AUDPC in 2012. The LOD values ranged from 2.9 to 10.2 and the corresponding PVE ranged from 8.0 to 26.0 %. Four QTLs, QSb.cim-1B, QSb.cim-3B, QSb.cim-5A and QSb.cim-14, were detected for average spot blotch AUDPC. The LOD values ranged from 2.8 to 6.7 and the corresponding PVE ranged from 4.9 to 17.6 %. Despite the LOD value for QSb.cim-1B in 2011 of 2.1 being lower than the LOD threshold of 2.5 used in this study, this locus was significant for spot blotch AUDPC in 2012 and the average of spot blotch AUDPC over years. It is most likely that QSb.cim-1B is a QTL with a low but relatively consistent effect.
QSb.cim-3B and QSb.cim-5A showed stronger effects on reducing spot blotch AUDPC. PVE of the 2-year average AUDPC of QSb.cim-3B was 17.6 %, while QSb.cim-5A explained 12.3 %. QSb.cim-1A was only significant in 2011 and QSb.cim-14 only for 2-year average spot blotch AUDPC. These two QTLs with lower PVE were not included in the analyses of QTL by environment and epistasis.
QSb.cim-1B was mapped in the interval between markers Xwmc128 and Xgwm374 (0.4 cM). QSb.cim-3B was mapped in the interval between 990937|F|0 and 1123330|F|0 (2.7 cM). QSb.cim-5A was mapped at the interval 1086218|F|0–982608|F|0 (12.1 cM) (Fig. 2). On chromosome 5A, Vrn-A1 was mapped 30.3 cM away from QSb.cim-5A using a functional maker developed by Yan et al. (2004).
LOD curves for three QTLs on chromosome 1B, 3B and 5A detected for resistance to spot blotch in the DH population ‘SYN1 × Ocoroni’. Genetic positions are shown in cM to the left of vertical axis. The vertical line indicates the LOD significance threshold (2.5) for QTL analysis. Arabic numerals in the brackets following some marker names indicate the number of markers at that locus genotyped in this population. Just one marker at each locus was shown in the figure
No significant epistatic interactions were detected between mapped QTLs in this study which indicated that much of the spot blotch resistance of SYN1 was due to the additive effects of QSb.cim-1B, QSb.cim-3B and QSb.cim-5A. The LOD values of QTL by year interactions for QSb.cim-1B, QSb.cim-3B and QSb.cim-5A were 7.6, 3.5 and 0.6, respectively (Table 4). PVE by QSb.cim-1B by year interaction was 75.5 % of that explained by the additive effect of QSb.cim-1B. By contrast, the QTL by year interactions for QSb.cim-3B and QSb.cim-5A explained only a small proportion of the phenotypic variation compared to the additive effect of these two QTLs with percentages of 5.2 and 5.0 %, respectively (Table 4). This result showed that the QSb.cim-1B and QSb.cim-3B had significant QTL by environment interactions. Meanwhile, QSb.cim-5A showed non-significant QTL by environment interaction.
Origins of the favorable alleles of detected QTLs
The QTL mapping results showed that the favorable alleles of QSb.cim-14, QSb.cim-1B, QSb.cim-3B and QSb.cim-5A were from SYN1, and only QSb.cim-1A was from the susceptible parent Ocoroni. Both parents of SYN1 [Mayoor and Tksn1081/Ae. squarrosa (222)] are resistant to spot blotch so genotypes of both parents in the region of the QTLs were examined to determine the source of the resistance alleles (Supplementary table 2). The analysis revealed that 1002801|F|0, 1123330|F|0 and 1056249|F|0 in the QSb.cim-3B region, 982608|F|0 and 1019824|F|0 in the QSb.cim-5A region, Xwmc128, Xgwm374, Xwmc31, Xcfd65, 1115814|F|0, 983080|F|0, 1089962|F|0 and 1051996|F|0 near the QSb.cim-1B region as well as 1136240|F|0 and 1057878|F|0 near QSb.cim-14 region were polymorphic between Mayoor and Tksn1081/Ae. squarrosa (222). For all of these polymorphic markers, Mayoor carried the same alleles as SYN1, whereas Tksn1081/Ae. squarrosa (222) always carried a different allele to SYN1. This result indicated that the favorable alleles of QSb.cim-14, QSb.cim-1B, QSb.cim-3B and QSb.cim-5A were inherited from Mayoor.
Phenotypic effects and comparison of QTL/marker alleles and their combinations
The means of spot blotch AUDPC, PHT and DF of DHs carrying different alternative QTL alleles were compared (Supplementary tables 3, 4). All the favorable alleles of QTLs significantly enhanced the resistance to spot blotch. For QSb.cim-3B, QSb.cim-5A and QSb.cim-1B, the spot blotch AUDPC average of DHs with SYN1 alleles were 14.7–35.5 % lower than those with Ocoroni alleles.
For DHs with all the Ocoroni alleles at QSb.cim-3B, QSb.cim-5A and QSb.cim-1B loci, the mean spot blotch AUDPC was 727.7 in 2012 and 975.9 in 2011(Table 5). QTL combinations QSb.cim-3B/QSb.cim-5A and QSb.cim-5A/QSb.cim-1B decreased spot blotch AUDPC by 20.3–43.3 %. DHs carrying all three favorable alleles showed an AUDPC mean of 341.9 in 2012 and 572.2 in 2011, which were 53.0 and 41.4 % lower than the DHs with all three Ocoroni alleles, respectively.
For QSb.cim-3B and QSb.cim-1B, there were no significant differences in PHT and DF between the DHs with SYN1 alleles and DHs with Ocoroni alleles (Supplementary table 4). However, for QSb.cim-5A, the SYN1 allele significantly increased PHT and DF. The SYN1 allele at QSb.cim-5A increased PHT by 5.3 % and DF by 11.4 % compared with Ocoroni allele at this locus.
In the mapping population, the SYN1 Vrn-A1 allele increased DF by 13.7 days. To determine whether the effect on disease response was a pleiotropic effect of the SYN1 Vrn-A1 allele, in line with the negative correlation between DF and AUDPC in the population as a whole (Table 3), mean spot blotch AUDPC of DHs carrying different alleles of QSb.cim-5A within each of the Vrn-A1 allele classes was compared (Supplementary table 5). For DHs with the SYN1 Vrn-A1 allele, the SYN1 allele at QSb.cim-5A decreased spot blotch AUDPC by 26.3 %. Similarly, for DHs with the OCI Vrn-A1 allele, the SYN1 allele of QSb.cim-5A decreased spot blotch AUDPC by 27.9 % indicating that the effect of QSb.cim-5A on disease reaction is distinct from the effect of allelic variation at Vrn-A1. No significant difference in DF was detected between DHs with different alleles at QSb.cim-5A for DHs with the OCI Vrn-A1 allele. However, for DHs with the SYN1 Vrn-A1 allele, the SYN1 allele at QSb.cim-5A significantly increased DF by 8.1. Thus, whether QSb.cim-5A itself has an effect on prolonging the DF cannot be definitively concluded.
Discussion
Phenotypic variation and trait correlations
The CIMMYT research station in Agua Fría has a warm, subtropical climate and high rainfall during the time that spot blotch screens are conducted making it a good site for high and consistent expression of disease symptoms. Lines with resistance in Agua Fria are expected to be resistant in other spot blotch prone areas because there is no defined race structure for spot blotch and differential host-pathogen interactions contribute no more than 1–2 % of the variance (Duveiller 2004). In practice, lines selected for resistance at Agua Fria express high levels of resistance in field tests at a range of locations in south Asia and in glasshouse screens (Duveiller 2004; Mahto et al. 2011).
In previous studies, spot blotch AUDPC and FDI were shown to be strongly correlated although AUDPC was suggested as a more appropriate method for disease evaluation and QTL analysis as it separates genotypes based on disease severity over time (Duveiller et al. 1998; Jeger 2004; Kumar et al. 2009). We confirmed the high correlation between the spot blotch AUDPC and FDI and found that AUDPC showed a lower correlation with DF than FDI in the ‘SYN1 × Ocoroni’ population and two other SYN1 DH populations also tested in both years (data not shown) which supports this suggestion.
Spot blotch AUDPC has been reported to be independent of PHT and days to maturity (Joshi et al. 2002) or to have a negative (−0.29 to −0.43, significant or non-significant) correlation with PHT in three spring wheat populations (Rosyara et al. 2009). Results in this study indicated highly significant negative correlations between spot blotch susceptibility and PHT or DF (−0.30 to −0.53). In genetic studies or in screening of exotic sources of resistance, it is therefore important to ensure that apparent resistance is not a result of increased PHT or slower phenological development because these effects are most likely due to disease escape and are not traits that will be appropriate in varieties targeted at spot blotch prone environments
Differential sowing of material with a large range in phenology is an option to synchronize development of lines between the time of inoculation and maturity to overcome the problem of variation in phenology (Kumar et al. 2009) and may be especially desirable when screening exotic resistance sources. In mapping populations, it is preferable to restrict the range in PHT and phenology to that typical in varieties grown in spot blotch prone areas either by selecting such lines from a larger population or choosing parents with appropriate phenology and similar PHT.
The results of QTL mapping in the ‘SYN1 × Ocoroni’ DH population suggested that some spot blotch resistances are associated with PHT and DF, while others that are not associated with these traits were also present. Correlation analysis between spot blotch resistance and PHT or DF is necessary before conducting QTL mapping. Furthermore, the effects of mapped QTLs on PHT and DF should be analyzed if the correlation between spot blotch resistance and PHT or DF is significant.
Genotypic data and linkage groups
This study used a combination of SSR and GBS markers to develop linkage maps. For the GBS markers, we observed an average 13.0 % missing data and distorted inheritance was observed at some loci. The results showed that 295 markers had significant departures from the expected 1:1 ratio. Missing marker data and segregation distortion are commonly encountered in QTL mapping populations. The effect of missing markers can be quantified by a population with a reduced size similar to the rate of missing markers (Zhang et al. 2010). Markers with segregation distortion were included in construction of linkage maps considering that segregation distortion loci are common in several crop species making the adjacent markers show distortion from the expected Mendelian ratio (Hedrick and Muona 1990; Lorieux et al. 1995a, b; Xu 2008; Tai et al. 2000). Generally, distorted markers will not have a great effect on the position and estimated QTL effects (Xu 2008). Nonetheless, although high-density linkage maps were constructed with the more than seven hundred GBS and SSR makers, there are still gaps of more than 20 cM on D genome chromosomes, and the D genome linkage maps had a lower density than those of the A and B genomes. Of the 34 linkage groups identified, only 11 groups were anchored with at least two SSRs on chromosomes 1A, 1B, 2A, 2B, 2D, 3B, 4A, 4B, 5A, 5B and 7B.
QTL mapping
Three consistent QTLs for spot blotch, QSb.cim-1B, QSb.cim-3B and QSb.cim-5A, were detected from SYN1. These QTLs explained a large part of the phenotypic variance (46.1 % in 2012 and 43.3 % for the mean spot blotch AUDPC over years). No significant epistatic interactions were found between these QTLs indicating the resistance of SYN1 to spot blotch is mainly due to additive effects of these QTLs. Analysis of interactions between QTL and year showed that QSb.cim-1B and QSb.cim-3B had significant QTL by year interactions although similar effects in each year indicate the interaction is associated with a change in scale, not a crossover interaction.
QSb.cim-1B maps to the same chromosome as a minor QTL for spot blotch resistance that was mapped by Lillemo et al. (2012) on chromosome 1BL corresponding to Lr46 between SSR markers barc80 and wmc719. The map distance between this locus and QSb.cim-1B was about 60 cM based on the consensus map (Somers et al. 2004) indicating that they are distinct QTLs. The LOD curves on chromosome 1B indicated that QSb.cim-1B may be located beyond the end of the linkage map (Fig. 2). As no other loci for spot blotch resistance have been reported on chromosome 1B, we suggest that QSb.cim-1B is a new QTL for spot blotch resistance. The linked SSR markers Xwmc128, Xgwm374, Xwmc31 and Xcfd65 could be used to select for this QTL in wheat breeding. Further screening to identify more closely linked, higher throughput markers may be desirable for use in marker-assisted selection of this QTL.
QSb.cim-3B maps close to QSb.bhu-3B from Chirya 3 reported by Kumar et al. (2010), flanked by Xgwm533 and Xgwm1037 and genes Sr2 and Fhb1, according to the consensus map (Somers et al. 2004). Pedigree analysis shows that Chirya 3 (Pedigree: Chinese-Spring/Ae.cu//Glennson-81/3/Alondra/Pavon-76/4/Ningmai-4/Olesen//Alondra/Yangmai-4) and Mayoor (Pedigree: Chinese-Spring/Ae.cu//Glennson-81/3/Alondra/Pavon-76) are closely related indicating a reasonable probability that this QTL is the same as QSb.cim-3B. Phenotypic variation accounted for by QSb.bhu-3B was 9.5 % of the total in one environment and was not significant in the other environment (Kumar et al. 2010). In our studies, the effect of QSb.cim-3B appeared greater and was significant in both years, explaining 17.6 % of the phenotypic variation across the 2 years. In the SYN1 × Ocoroni population and the Chirya × Sonalika population in which QSb.bhu-3B was identified (Kumar et al. 2010), the spot blotch susceptible parent contributing the susceptible allele on 3BS carries Sr2 (Spielmeyer et al. 2003, McIntosh et al. 1995) indicating that QSb.cim-3B and QSb.bhu-3B are linked in repulsion with Sr2. While Fhb1 is also linked in repulsion with Sr2, phenotyping of the SYN1 × Ocoroni population for Fusarium head blight (FHB) reaction indicates QSb.cim-3B has no effect on FHB and is distinct from Fhb1 (CIMMYT unpublished).
The QSb.cim-5A QTL is the first spot blotch resistance reported on chromosome 5A. Phenotypic effect analysis showed that the SYN1 allele of QSb.cim-5A increased DF by 11.4 days compared to the Ocoroni allele at this locus. Interestingly, QSb.cim-5A was mapped 30.3 cM away from Vrn-A1. As the SYN1 Vrn-A1 allele increases DF compared to the OCI Vrn-A1 allele, the linkage with SYN1 Vrn-A1 might be responsible for the observed effect of QSb.cim-5A on prolonging DF. For DHs with the OCI Vrn-A1 allele, no significant difference in DF was detected between DHs carrying different alleles of QSb.cim-5A suggesting that it does not have an effect on DF per se. However, for DHs with the SYN1 Vrn-A1 allele, the SYN1 allele at QSb.cim-5A increased DF by 8.1. It is therefore not possible from the current study to definitively state if QSb.cim-5A has an effect on phenology or not, and this would need to be clarified in a future study. However, our results indicated it is possible to select a spot blotch resistant QSb.cim-5A allele without increasing DF.
Inconsistent QTLs with minor effects were also detected in this study, but they only explain a small additional proportion of the phenotypic variance. Although heritability over the 2 years of the study was high (0.72), around 43 % of the genotypic variance was not accounted for. This additional genotypic variation could be due to QTLs of moderate effect that were not detected due to gaps in the map, to a large number of QTLs of small effect or due to interactions with other loci or the environment over the 2 years of the study. A further possibility is that some of the additional variation is due to variation in phenology or linkage of resistance QTLs with genes affecting phenology. Gaps in the map and exclusion of phenology-related loci probably reflect selection of markers based on polymorphism between resistant and susceptible bulks and the manner in which the range in phenology was restricted in those bulks. QTLs of smaller effect are less likely to be clearly separated and identified in a bulked segregant approach.
Origins of the favorable alleles of detected QTLs and future efforts mapping spot blotch resistance of CIMMYT wheats
The favorable allele of a minor QTL, Qsb.cim-1A, was inherited from the spot blotch susceptible parent Ocoroni, whereas the favorable alleles of other QTLs, QSb.cim-14, QSb.cim-1B, QSb.cim-3B and QSb.cim-5A, were inherited from the spot blotch resistant parent SYN1. Analysis of the alleles of markers linked to these QTLs in the parents of SYN1, Mayoor and Tksn1081/Ae. squarrosa (222), showed that the favorable alleles of QSb.cim-14, QSb.cim-1B, QSb.cim-3B and QSb.cim-5A were likely inherited from Mayoor. This study therefore provides useful information on the basis of resistance to spot blotch of Mayoor that may be relevant to its derivatives. As both parents of SYN1 were highly resistant to spot blotch, there is a possibility that the spot blotch resistance of Tksn1081/Ae. squarrosa (222), or even components of the resistance of Mayoor, was lost during the process of selection. Therefore, this study provides no information on the basis of resistance in Tksn1081/Ae. squarrosa (222) and may not have identified all of the important resistance QTLs from Mayoor. Failure to capture resistance QTLs from Tksn1081/Ae. squarrosa (222) in SYN1 may be due to its poor agronomic type including tall PHT and late maturity. If the spot blotch resistance of Tksn1081/Ae. squarrosa (222) is associated with these or other undesirable traits, there is a greater chance that they were lost along with the undesirable traits during selection. Although the approach of crossing two resistant lines together as occurred in the development of SYN1 offers the attractive possibility of combining the resistances if they are genetically distinct, it runs the risk of losing genetic variation from both of the resistance sources due to unfavorable linkages with poor phenotypes or simply due to small sample sizes and the high level of resistance observed across a range of combinations of QTLs. The result of this study shows it is preferable to cross a resistant line directly to a susceptible parent when developing a mapping population. It is possible that although all of the important alleles in the SYN1 × Ocoroni DH population are from the Mayoor parent of SYN1 that additional Mayoor resistance QTLs may not have been captured in SYN1 and remain to be identified. The population used in the research reported here was developed long before mapping efforts were initiated. The research reported represented an attempt to map resistance QTLs in two widely used resistance sources. Other widely used resistance sources from primary synthetics have been identified, and appropriately targeted mapping populations are under development to characterize their genetic basis and determine the level of diversity they represent for spot blotch resistance.
Phenotypic effects and comparison of QTL/marker alleles and their combinations
In both years, the reductions in spot blotch AUDPC by the favorable alleles of QSb.cim-3B or QSb.cim-5A were much greater than those of QSb.cim-1B. Neither of the SYN1 alleles of QSb.cim-1B or QSb.cim-3B was associated with PHT or DF. As a result, it is possible to introduce these two QTLs into breeding lines without increased PHT and delayed flowering time. By contrast, the favorable allele of QSb.cim-5A for spot blotch significantly increased PHT and DF in our study population. Our data suggest that the effect of QSb.cim-5A on PHT and DF may be due to linkage with SYN1 Vrn-A1 which was mapped nearby rather than a per se effect (Supplementary table 5). SYN1 Vrn-A1 prolongs the vegetative development stage of the plants and delays flowering time allowing plants to escape the highest levels of spot blotch disease pressure. The prolonged vegetative development stage also increases the PHT which reduces the rate of movement of disease up the canopy due to splashing of spores. Our results indicate that combining the three favorable alleles at QSb.cim-1B, QSb.cim-3B and QSb.cim-5A reduces disease levels by around 50 % under the high disease pressure of Agua Fria. It should be possible to recover lines carrying the desirable alleles at all three loci, and with appropriate flowering time based on the earlier flowering OCI Vrn-A1 allele. This level of disease reduction will probably be greater in more typical spot blotch environments where disease pressure is lower.
Strategies for use of QTLs in breeding for spot blotch resistance
The QTLs of larger effect mapped in this study, QSb.cim-1B, QSb.cim-3B and QSb.cim-5A, are likely to be present in many CIMMYT wheat lines with Mayoor in their pedigrees, and the linked markers identified in this study could be used initially to characterize parent lines. If the loci are present in a high proportion of parent lines, it may be warranted to convert markers to high-throughput forms amenable to use in routine screening. The reduction in spot blotch disease development in typical spot blotch prone environments is likely to be sufficient with only these three QTLs combined assuming they have equivalent effects in different environments and with different strains of the spot blotch fungus. These QTLs may be particularly useful to breeding programs targeting spot blotch resistance that do not have a reliable phenotyping environment. In the CIMMYT program, they may be useful for selection in segregating material that cannot be effectively screened for spot blotch resistance.
The repulsion phase linkage between QSb.cim-3B and Sr2 indicated in this study may reduce the frequency of this resistance QTL in CIMMYT breeding lines due to stronger and earlier stage selection for the presence of Sr2 and should be investigated further.
The QTLs of smaller effect are less likely to offer sufficient additional benefit to justify investment in developing high-throughput markers.
In the CIMMYT program, markers for the key QTLs could be used in selection in early generations to ensure fixation or at least a higher frequency of these resistances prior to phenotypic screening of which is restricted to more advanced, inbred material due to insufficient space at the Agua Fria station, the cost of increasing the amount of material screened and the heritability not being high enough on a single plant basis. Inbred lines could then be screened in plots like the ones used in this study to identify additional variation, possibly from QTLs of smaller effect, and select the most resistant material.
While there is no clear evidence of race specificity in the spot blotch pathogen, it seems prudent to continue genetic characterization of diverse resistance sources to increase the levels of resistance, identify more QTLs of large effect that may simplify resistance breeding and to ensure sufficient genetic diversity in resistance to guard against possible pathogenic changes. The large number of spot blotch resistant primary synthetic bread wheats capturing variation from a range of Ae. tauschii accessions and the widespread use of these primary synthetics in the CIMMYT program make these an attractive target for studying the genetics of resistance. Mapping populations for many of these resistance sources are already in the process of being generated and should add greatly to the understanding of the diversity and genetic architecture of resistance to spot blotch from sources that are likely to be relatively common in advanced CIMMYT breeding lines and derived cultivars. Together this knowledge will allow development of improved strategies for breeding spot blotch resistant wheats at CIMMYT and in other breeding programs.
It is possible that, as suggested in this study, a good proportion of the useful genetic variation for spot blotch resistance will be due to many genes of small effect. In that case, it may be possible that phenotypic selection in good environments like Agua Fria and selection for QTLs with significant effects could be complemented with a genomic selection approach (Meuwissen et al. 2001). Such an approach assumes many genes of small effect and determines values of alleles across the genome without the restriction of those allele effects attaining a specific level of statistical significance. This is becoming more realistic as increasing numbers of CIMMYT advanced lines are being routinely genotyped with high-density, whole-genome markers (Bonnett unpublished). This genotypic data in combination with the routine spot blotch phenotyping of advanced lines may also be effectively used to identify significant QTLs through association mapping approaches.
References
CIMMYT (2005) Laboratory protocols, 3rd edn. CIMMYT Applied Molecular Genetics Laboratory, Mexico
Dubin HJ, Duveiller E (2000) Helminthosporium leaf blights of wheat: integrated control and prospects for the future. In: Mitra DK (ed) Proceeding of international conference on integrated plant disease management for sustainable agriculture, New Delhi, India, pp 575–579
Dubin HJ, Arun B, Begum SN, Bhatta M, Dhari R, Goel LB, Joshi AK, Khanna BN, Malaker PK, Pokhrel DR, Rahman MM, Saha NK, Shaheed MA, Sharma RC, Singh AK, Singh RM, Singh RV, Vargas M, Verma PC (1998) Results of the South Asia regional helminthosporium leaf blight and yield experiment, 1993–94. In: Duveiller E, Dubin HJ, Reeves J, McNab A (eds) Helminthosporium blights of wheat: spot blotch and tan spot. CIMMYT, Mexico, pp 182–187
Duveiller E (2004) Controlling foliar blights of wheat in the rice–wheat systems of Asia. Plant Dis 88:552–556
Duveiller E, Sharma RC (2005) Results of the 13th helminthosporium monitoring nursery 2004–2005. CIMMYT South Asia, Kathmandu
Duveiller E, Garcia I, Franco J, Toledo J, Crossa J, Lopez F (1998) Evaluating spot blotch resistance of wheat: improving disease assessment under controlled condition and in the field. In: Duveiller E, Dubin HJ, Reeves J, McNab A (eds) Helminthosporium blights of wheat: spot blotch and tan spot. CIMMYT, Mexico, pp 171–181
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES, Mitchell SE (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS One 6(5):e19379. doi:10.1371/journal.pone.0019379
Eyal Z, Scharen AL, Prescott JM, van Ginkel M (1987) The Septoria disease of wheat: concepts and methods of disease management. CIMMYT, Mexico
Ganal MW, Röder MS (2007) Microsatellite and SNP markers in wheat breeding. In: Varshney RK, Tuberosa R (eds) Genomics assisted crop improvement. Genomics applications in crops, vol 2. Springer, The Netherlands, pp 1–24
Hedrick PW, Muona O (1990) Linkage of viability genes to marker loci in selfing organisms. Heredity 64:67–72
Jeger MJ (2004) Analysis of disease progress as a basis for evaluating disease management practices. Annu Rev Phytopathol 42:61–82
Joshi AK, Chand R (2002) Variation and inheritance of leaf angle, and its association with spot blotch (Bipolaris sorokiniana) severity in wheat (Triticum aestivum). Euphytica 124:283–291
Joshi AK, Chand R, Arun B (2002) Relationship of plant height and days to maturity with resistance to spot blotch in wheat. Euphytica 123:221–228
Joshi AK, Chand R, Kumar S, Singh RP (2004) Leaf tip necrosis: a phenotypic marker associated with resistance to spot blotch diseases in wheat. Crop Sci 44:792–796
Joshi AK, Ortiz-Ferrara G, Crossa J, Singh G, Alvarado G, Bhatta MR, Duveiller E, Sharma RC, Pandit DB, Siddique AB, Das SY, Sharma RN, Chand R (2007a) Associations of environments in South Asia based on spot blotch disease of wheat caused by Cochliobolus sativus. Crop Sci 47(3):1071–1081
Joshi AK, Ortiz-Ferrara G, Crossa J, Singh G, Sharma RC, Chand R, Parsad R (2007b) Combining superior agronomic performance and terminal heat tolerance with resistance to spot blotch (Bipolaris sorokiniana) of wheat in the warm humid Gangetic Plains of South Asia. Field Crop Res 103(1):53–61
Knapp SJ, Stroup WW, Ross WM (1985) Exact confidence intervals for heritability on a progeny mean basis. Crop Sci 25:192–194
Kumar S, Prasad LC, Kumar U, Tyagi K, Arun B, Joshi AK (2007) Inheritance and allelic relationship of resistance genes to spot blotch of wheat caused by Bipolaris sorokiniana. In: Buck HT, Nisi JE, Salomon N (eds) Wheat production in stressed environments. Proceedings of the international wheat conference, pp 113–118
Kumar U, Joshi AK, Kumar S, Chand R, Röder MS (2009) Mapping of resistance to spot blotch disease caused by Bipolaris sorokiniana in spring wheat. Theor Appl Genet 118:783–792
Kumar U, Joshi AK, Kumar S, Chand R, Röder MS (2010) Quantitative trait loci for resistance to spot blotch caused by Bipolaris sorokiniana in wheat (T. aestivum L.) lines ‘Ning 8201’ and ‘Chirya 3’. Mol Breed 26(3):477–491
Lillemo M, Joshi AK, Prasad R, Chand R, Singh RP (2012) QTL for spot blotch resistance in bread wheat line Saar co-locate to the biotrophic disease resistance loci Lr34 and Lr46. Theor Appl Genet. doi:10.1007/s00122-012-2012-6
Lorieux M, Goffinet B, Perrier X, González de León D, Lanaud C (1995a) Maximum likelihood models for mapping genetic markers showing segregation distortion. 1. Backcross population. Theor Appl Genet 90:73–80
Lorieux M, Goffinet B, Perrier X, González de León D, Lanaud C (1995b) Maximum likelihood models for mapping genetic markers showing segregation distortion. 2. F2 population. Theor Appl Genet 90:81–89
Mahto BN, Gurung S, Adhikari TB (2011) Assessing genetic resistance to spot blotch, Stagonospora nodorum blotch and tan spot in wheat from Nepal. Eur J Plant Pathol 131:249–260
McIntosh RA, Wellings CR, Park RF (1995) Wheat rusts. An atlas of resistance genes. CSIRO, Melbourne
Meuwissen THE, Hayes BJ, Goddard ME (2001) Prediction of total genetic value using genome-wide dense marker maps. Genetics 157:1819–1829
Mujeeb-Kazi A, Fuentes Dávila G, Villareal RL, Cortés A, Roasas V, Delgado R (2001) Registration of five synthetic hexaploid wheat and seven bread wheat lines resistant to wheat spot blotch. Crop Sci 41(5):1653–1654
Neupane RB, Sharma RC, Duveiller E, Ortiz-Ferrara G, Ojha BR, Rosyara UR, Bhandari D, Bhatta MR (2007) Major gene controls of field resistance to spot blotch in wheat genotypes Milan/Shanghai #7 and Chirya. 3. Plant Dis 91(6):692–697
Rosyara UR, Khadka K, Subedi S, Sharma RC, Duveiller E (2009) Field resistance to spot blotch is not associated with undesirable physio-morphological traits in three spring wheat populations. J Plant Pathol 91(1):113–122
Rosyara UR, Subedi S, Duveiller E, Sharma RC (2010) Photochemical efficiency and SPAD value as indirect selection criteria for combined selection of spot blotch and terminal heat stress in wheat. J Phytopathol 158(11–12):813–821
Saari EE (1998) Leaf blight disease and associated soil borne fungal pathogens of wheat in South and Southeast Asia. In: Duveiller E, Dubin HJ, Reeves J, McNab A (eds) Helminthosporium blights of wheat: spot blotch and tan spot. CIMMYT, Mexico, pp 37–51
Saari EE, Prescott JM (1975) A scale for appraising the foliar intensity of wheat disease. Plant Dis Rep 59:377–380
Sharma RC, Duveiller E (2006) Spot blotch continues to cause substantial grain yield reductions under resource-limited farming conditions. J Phytopathol 154(7–8):482–488
Sharma RC, Duveiller E (2007) Advancement toward new spot blotch resistant wheats in South Asia. Crop Sci 47(3):961–968
Sharma RC, PandEy-Chhetri B, Duveiller E (2006) Heritability estimates of spot blotch resistance and its association with other traits in spring wheat crosses. Euphytica 147(3):317–327
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Spielmeyer W, Sharp PJ, Lagudah ES (2003) Identification and validation of markers linked to broad-spectrum stem rust resistance gene Sr2 in wheat (Triticum aestivum L.). Crop Sci 43(1):333–336
Tai GCC, Seabrook JEA, Aziz AN (2000) Linkage analysis of anther-derived monoploids showing distorted segregation of molecular markers. Theor Appl Genet 101:126–130
van Ginkel M, Rajaram S (1998) Breeding for resistance to spot blotch in wheat: global perspective. In: Duveiller E, Dubin HJ, Reeves J, McNab A (eds) Helminthosporium blights of wheat: spot blotch and tan spot. CIMMYT, Mexico, pp 162–170
Van Ooijen JW (2006) JoinMap® 4, software for the calculation of genetic linkage maps in experimental populations. Kyazma B.V., Wageningen
Villareal RL, Mujeeb-Kazi A, Gilchrist L, Del Toro E (1995) Yield loss to spot blotch in spring bread wheat in warm nontraditional wheat production areas. Plant Dis 79(9):893–897
Wang J, Li H, Zhang L, Meng L (2011) Users’ manual of QTL IciMapping version 3.2. Chinese Academy of Agricultural Sciences (CAAS), Beijing
Xu S (2008) Quantitative trait locus mapping can benefit from segregation distortion. Genetics 180:2201–2208
Yan L, Helguera M, Kato K, Fukuyama S, Sherman J, Dubcovsky J (2004) Allelic variation at the VRN-1 promoter region in polyploidy wheat. Theor Appl Genet 109(8):1677–1686
Zhang L, Wang S, Li H, Deng Q, Zheng A, Li S, Li P, Li Z, Wang J (2010) Effects of missing marker and segregation distortion on QTL mapping in F2 populations. Theor Appl Genet 121(6):1071–1082
Acknowledgments
The senior author was financially support by the CIMMYT Global Wheat Program, the China Scholarship Council and the China Agriculture Research System. GBS genotyping was supported by the Seeds of Discovery (SeeD) project, one of the four components of the MasAgro program funded by the Mexican Ministry of Agriculture, Livestock, Rural Development, Fisheries and Food (SAGARPA). The authors would like to thank Dr. Etienne Duveiller for his advice on data collection and analysis, Drs. Xiang Wang and Caixia Lan for providing SSR primers to this study, Roman Delgado, Hugo Lopez, and Jesus Sanchez for their technical assistance.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhu, Z., Bonnett, D., Ellis, M. et al. Mapping resistance to spot blotch in a CIMMYT synthetic-derived bread wheat. Mol Breeding 34, 1215–1228 (2014). https://doi.org/10.1007/s11032-014-0111-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-014-0111-6