Abstract
Wheat pre-harvest sprouting (PHS) can cause significant reduction in yield and end-use quality of wheat grains in many wheat-growing areas worldwide. To identify a quantitative trait locus (QTL) for PHS resistance in wheat, seed dormancy and sprouting of matured spikes were investigated in a population of 162 recombinant inbred lines (RILs) derived from a cross between the white PHS-resistant Chinese landrace Totoumai A and the white PHS-susceptible cultivar Siyang 936. Following screening of 1,125 SSR primers, 236 were found to be polymorphic between parents, and were used to screen the mapping population. Both seed dormancy and PHS of matured spikes were evaluated by the percentage of germinated kernels under controlled moist conditions. Twelve SSR markers associated with both PHS and seed dormancy were located on the long arm of chromosome 4A. One QTL for both seed dormancy and PHS resistance was detected on chromosome 4AL. Two SSR markers, Xbarc 170 and Xgwm 397, are 9.14 cM apart, and flanked the QTL that explained 28.3% of the phenotypic variation for seed dormancy and 30.6% for PHS resistance. This QTL most likely contributed to both long seed dormancy period and enhanced PHS resistance. Therefore, this QTL is most likely responsible for both seed dormancy and PHS resistance. The SSR markers linked to the QTL can be used for marker-assisted selection of PHS-resistant white wheat cultivars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pre-harvest sprouting (PHS), the germination of kernels in a ripened wheat (Triticum aestivum) spike before harvesting, can severely reduce yield and end-use quality of wheat grain, and therefore result in great economic losses for wheat farmers (Varughese et al. 1987). Wheat PHS is a major constraint for wheat production worldwide when a long period of wet weather occurs during a harvesting season (Derera 1989; Sharma et al. 1994). Thus, PHS resistance is becoming an attractive research topic and an important breeding objective in wheat-breeding programs worldwide.
Early breakage of seed dormancy has been considered as the major component of PHS (Mares 1987), and the term PHS resistance is often used interchangeably with long seed dormancy (Flintham et al. 2002; Roy et al. 1999; Kulwal et al. 2005; Derera and Bhatt 1980). It is known that PHS and seed dormancy are both complex traits that are determined by genotype together with a large number of other factors including stage of maturity when the plant is harvested; environmental conditions including both biotic and abiotic stresses after seed maturation (Walker-Simmonds 1987); spike and plant morphology (Paterson et al. 1989); and seed color (Groos et al. 2002; Himi et al. 2002; Nilsson-Ehle 1914). Seed dormancy is controlled mainly by the embryo; it is possibly modified by inhibitors endogenous to the bran (Morris and Paulsen 1988) and can be measured by a seed-germination test. However, overall PHS resistance may be affected by additional factors including substances inhibitory to germination residing in chaff tissue (Salmon et al. 1986), physical barriers to water penetration in the spike (Gale 1989), and spike morphology (King 1984). These factors may complicate the procedures for phenotypic evaluation of PHS resistance.
Molecular mapping has been used to locate QTLs for both PHS resistance and seed dormancy in several studies. However, the number and location of QTLs for PHS resistance and seed dormancy are inconsistent among studies. Using restriction fragment length polymorphism (RFLP) markers, Anderson et al. (1993) reported eight genomic regions on chromosomes that might affect PHS resistance (Sorrells and Anderson 1996). Roy et al. (1999) reported that PHS resistance was governed by two genes located on chromosome 6B and 7D, respectively. Groos et al. (2002) reported four QTLs for PHS resistance, and three of them are associated with grain color on the long arms of group 3 chromosomes. Flintham et al. (2002) reported a QTL for PHS resistance on chromosome 4A using a population developed from cross of a white to a red wheat cultivar. More recent studies mapped a major QTL for PHS resistance to chromosome 3A (Kulwal et al. 2005). As for seed dormancy, several studies identified a major QTL on chromosome 4A (Kato et al. 2001; Noda et al. 2002; Mares et al. 2005; Torada et al. 2005) and two other studies identified three QTLs with one major QTL on chromosome 3A and two minor QTLs on chromosome 4A and 4B, respectively (Osa et al. 2003; Mori et al. 2005). Therefore, whether PHS resistance and seed dormancy are controlled by the same or different QTLs is still equivocal.
Simultaneous mapping of QTLs for both PHS resistance and seed dormancy may facilitate further understanding of the genetic mechanism of PHS resistance as well as the genetic relationship between the two traits. It may also facilitate identification of new sources of PHS resistance for breeding applications and the development of DNA markers linked to PHS resistance for marker-assisted selection of PHS-resistant wheat cultivars. The objectives of our study were to: (1) identify QTLs for both PHS resistance and seed dormancy in a Chinese landrace, (2) elucidate the genetic relationship of the QTLs for seed dormancy and PHS resistance, and (3) develop molecular markers suitable for marker-assisted selection in wheat breeding programs.
Materials and methods
Plant materials
A mapping population of 152 recombinant inbred lines (RILs) was derived from a cross between the PHS-resistant parent Totoumai A and the PHS-susceptible parent Siyang 936 by single-seed decent. Both parents and their RIL population were evaluated for seed dormancy and PHS tolerance in Kansas State University (Manhattan, KS). The experiment was arranged in a randomized complete block design with three replications at 22 ± 6°C day/15 ± 2°C night temperature with supplemented daylight of 16 h in the greenhouse at Kansas State University. Seed dormancy and spike spouting experiments were both repeated once.
Evaluation of seed dormancy and PHS
Pre-harvest sprouting in an intact spike was evaluated in the spring of 2005 and 2006 by enclosing harvested spikes of RILs in a moist chamber set up in a temperature-controlled growth room at Kansas State University. When wheat spikes reached physiological maturity, five spikes per RIL harvested from each replicate were air-dried for 5 days in the greenhouse, and then stored at −20°C to maintain dormancy for PHS evaluation. After all RILs were collected, the harvested spikes were air-dried again for 2 days in a greenhouse, then surface-sterilized with 20% bleach for 20 min, and immersed in de-ionized water for 5 h. The wet spikes were enclosed in a moist chamber at 22 ± 1°C to facilitate seed germination. High moisture in the chamber was maintained by running a humidifier for 30 min twice daily. After 7 days of incubation, the spikes were individually hand threshed, and the numbers of germinated and non-germinated kernels in each spike were counted. PHS resistance was measured as percentage of germinated kernels in a spike.
Seed dormancy was evaluated in the fall of 2005 and the spring of 2006. To evaluate seed dormancy, wheat spikes were harvested and air-dried as described for PHS evaluation. The harvested spikes were hand threshed. Fifty kernels were surface-sterilized with 25% bleach, placed on a sterilized wet filter paper in a Petri dish, and incubated in a temperature-controlled growth room at 22 ± 1°C. Germinated kernels were counted daily and removed after counting. A weighted germination index (Walker-Simmons 1987) was used to measure seed dormancy based on the following formula:
Germination index (GI%) = 100 × [(x × n 1 + (x−1) × n 2 + ···1 × n i)/total number of grains × total number of days between seed planting and the end of the experiment]
Where x is the total number of days between seed planting and the end of the experiment, n 1, n 2, ...n x are the numbers of kernels that germinated on a specific day i, i is number of days after seed planting ranging from 1 to x.
SSR analysis
Genomic DNA was isolated from 2-week-old wheat leaves of each RIL using a modified CTAB method. Harvested tissue was dried in a freeze dryer (ThermoSavant, Holbrook, NY) for 48 h and ground using a Mixer Mill (MM 300, Rotsch, Germany).
A total of 1,125 SSR primers including BARC, WMC, GWM, and CFD primers were used to screen the parents, and the polymorphic primers were used to screen the RILs. For SSR analysis, a 10-μl PCR mixture contained 40 ng template DNA, 1 mM each of reverse and M13-tailed forward primers, 0.2 mM of each dNTP, 1X PCR buffer, 2.5 mM MgCl2, and 0.6 U Taq polymerase. For PCR detection, 1 pmol fluorescence-labeled M13 primer was added to each PCR reaction (Li-Cor, Lincoln, NE). A touch-town PCR program was used for PCR amplification, in which the reaction mixture was incubated at 95°C for 5 min, then for five cycles of 45 s of denaturing at 95°C, 5 min of annealing at 68°C with a decrease of 2°C in each subsequent cycle, and 1 min of extension at 72°C. For another five cycles, the annealing temperature started at 58°C for 2 min with a decrease of 2°C for each subsequent cycle. PCR continued through an additional 25 cycles of 45 s at 94°C, 2 min at 50°C, and 1 min at 72°C with a final extension at 72°C for 5 min. The amplified PCR fragments were separated in a Li-Cor 4300 DNA analyzer using a 6.5% Gel Matrix (Li-Cor). All marker data were scored by visual inspection and double checked to remove ambiguous data.
Genetic map construction and QTL analysis
A genetic linkage map was constructed with SSR markers using JoinMap version 3.0 (Van Ooijen and Voorrips 2001). Recombination fractions were converted into centiMorgans (cM) using the Kosambi function (Kosambi 1944). The threshold value of LOD (logarithm of odds) score was set at 3.0 to claim linkage between markers with a maximum fraction of recombination at 0.4. For QTL analysis, interval mapping (IM) and composite interval mapping (CIM) were performed with MapQTL Version 5 (Van Ooijen 2004). QTL analysis was done on the basis of line means from individual experiments and from combined phenotypic data from all experiments. Permutation tests were performed to estimate appropriate thresholds for declaring a significant QTL (Doerge and Churchill 1996). On the base of 1,000 permutations, LOD at 2.2 was set as the threshold for declaring a significant QTL. Broad-sense heritability (h 2) was computed as δg 2/(δg 2 + δe 2) based on the estimates of genetic and error variances derived from the analysis of variances (SAS Institute, Cary, NC).
Results
Seed dormancy and PHS in parents and RILs
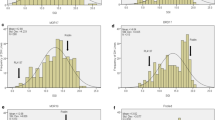
The two parents, Totoumai A and Siyang 936, showed a significant difference in both seed dormancy and PHS resistance. The mean GI% of threshed kernels over 2 years was 57.2% for Totoumai A and 76.8% for Siyang 936, and mean germination rate in an un-threshed spike over 2 years was 33.5% (2006) in Totoumai A and 58.7% in Siyang 936. These results indicated that Totoumai A has not only a lower seed germination rate (longer period of seed dormancy) when threshed kernels were tested, but also a higher PHS resistance than that of Siyang 936 when intact spikes were tested. Therefore, the two parents had significant contrasts in both traits and were appropriate for this study. The frequency of PHS resistance and seed dormancy in the RIL population showed a continuous distribution with a major peak between the two parents (Fig. 1). The broad-sense heritability was about 0.87 for seed dormancy and 0.79 for PHS resistance, indicating that both traits are highly heritable.
Frequency distribution of percentage of germinated kernels to reflect seed dormancy and pre-harvest sprouting (PHS) of wheat recombinant inbred lines (RILs) derived from a cross of Totoumai A and Siyang 936. SD Seed dormancy as measured by germination index (GI%). PHS was measured by seed germination rate (%) of an intact spike after a 7-day incubation in a moist chamber
Genetic map construction and QTL mapping
After screening a total of 1,125 SSR primers, 236 showed polymorphism, and were further used to screen a subset of 94 RILs in the mapping population. A genetic map was constructed with 196 SSR markers distributed in 27 linkage groups covering a genetic distance of 2,139 cM. Genome-wide QTL scanning detected only one significant QTL each for seed dormancy and PHS resistance. Both QTLs co-localized to the same region on the long arm of chromosome 4A.
A major QTL controlling both seed dormancy and PHS tolerance
An additional 58 RILs from the same population were analyzed with the 12 linked SSR markers on chromosome 4A. The data from all 152 RILs of the population were combined for further map construction and QTL analysis. All 12 markers were mapped to the same linkage group, which covered a genetic distance of 46 cM. As expected, interval QTL analysis identified a putative QTL on the same region of the chromosome 4A for both seed dormancy and PHS resistance (Fig. 2). Two SSR markers, Xbarc170 and Xgwm397, flanked this QTL within a marker interval of 9.1 cM. The SSR marker Xbarc 170 alone explained 26.2% of the phenotypic variation for PHS resistance and 19.6% of the phenotypic variation for longer seed dormancy over 2 years. Interval analysis indicated that this QTL explained 28.3% of the phenotypic variation for seed dormancy and 30.6% for PHS resistance based on combined data from 2 years (Table 1).
Interval mapping (IM) of QTL for long seed dormancy and PHS resistance on chromosome 4AL with simple sequence repeat markers using phenotypic data from 2 years. The QTL map is on the top and linkage map is on the bottom. The broken line parallel to the X-axis is the threshold line for the significant LOD value of 2.2 (P < 0.05). Genetic distances are shown in centiMorgans (cM) below the linkage map, and markers are above the linkage map
To evaluate the potential selection efficiency of marker-assistant selection, two flanking markers for the 4AL QTL were used to estimate the selection progress. When a single marker was evaluated, the group of RILs carrying the resistant allele had a 12% and 13% reduction in seed dormancy, and a 12.7% and 14.3% reduction in PHS compared to the group carrying the susceptible marker allele in 2 years. If a single marker was selected, the effect of selection for Xbarc170 was larger than for Xgwm397 (Table 2), about 8% increase for seed dormancy and 12% increase for PHS resistance compared to Xgwm397. When two flanking markers were selected simultaneously, the effect of selection was larger than that of a single marker—about 15% increase for both seed dormancy and PHS resistance compared to selecting for Xbarc170 alone (Table 2).
Discussion
The early breakage of seed dormancy was proposed as a major cause of PHS (Mares 1987). However, the QTL mapping evidence from past studies did not fully support this hypothesis. QTLs for PHS resistance were mapped to chromosomes 2BS (http://maswheat.ucdavis.edu/), 3A (Kulwal et al. 2005), 4AL (Anderson et al. 1993; Flintham et al. 2002; Sorrells and Anderson 1996) 3B, 5A, 6A, 7B (Zanetti et al. 2000), 1AS, 3BL, 4AL, 5DL and 6BL (Anderson et al. 1993; Sorrells and Anderson 1996), 6B and 7D (Roy et al. 1999), 5A and all group 3 chromosomes (Groos et al. 2002), whereas the major QTLs for seed dormancy were mapped to chromosomes 4A (Kato et al. 2001; Noda et al. 2002; Mares et al. 2005; Torada et al. 2005) and 3A (Osa et al. 2003; Mori et al. 2005) in several different studies. In the current study, a major QTL significantly associated with both seed dormancy and PHS tolerance was identified on the long arm of chromosome 4A, which agrees with several previous studies in terms of QTLs for seed dormancy (Kato et al. 2001; Noda et al. 2002; Mares et al. 2005; Torada et al. 2005) and one study in terms of QTLs for PHS resistance (Flintham et al. 2002). This result further confirmed that seed dormancy is the major component of PHS resistance and that the QTL on chromosome 4A controls both PHS resistance and seed dormancy in wheat.
Much effort has been made to elucidate the number and location of QTLs for either PHS resistance or seed dormancy. The results from different studies have been inconsistent. Some studies indicated multigenic control of PHS resistance (Anderson et al. 1993; Sorrells and Anderson 1996; Zanetti et al. 2000), whereas others demonstrated evidence of oligogenic control (Mares et al. 2005; Torada et al. 2005). Zanetti et al. (2000) detected several QTLs with a major effect on PHS resistance on chromosomes 3B, 5A, 6A, 7B, whereas Anderson et al. (1993) reported that eight genomic regions on chromosomes 1AS, 3BL, 4AL, 5DL and 6BL were associated with PHS tolerance (Sorrells and Anderson 1996). Roy et al. (1999) reported that two genes on chromosomes 6B and 7D were responsible for PHS resistance. Using a population from a cross between white and red wheat, Groos et al. (2002) detected QTLs for PHS resistance on chromosome 5A and all group 3 chromosomes where the kernel color gene R and taVp1 were previously mapped. An SSR marker, Xbarc055 on 2BS, has been associated with a major QTL in Cayuga (http://maswheat.ucdavis.edu/), which explained at least 32% of the phenotypic variation for PHS over 3 years. For seed dormancy, only one major QTL on 4A (Kato et al. 2001; Noda et al. 2002; Mares et al. 2005) or 3A (Osa et al. 2003; Mori et al. 2005) has been reported. In this study, we identified one major QTL that only partly explained the genetic variation for both seed dormancy and PHS resistance. Obviously, PHS resistance is most likely controlled by several QTLs, with at least one of them having a major effect. The remaining undetected QTLs in this study are due mainly to the lack of polymorphism in the markers used, or the lack of available markers closely linked to other QTLs. Alternatively, the QTL effects were too small to be detected relative to environmental effects in this study. Further studies to add other markers may improve the map resolution to facilitate identification of other QTLs.
PHS resistance in wheat is a complex trait. Many factors may affect PHS resistance. Phenotyping methods and many environmental factors can affect accurate identification of QTLs for PHS resistance. A suitable method for phenotypic evaluation is critical to the successful mapping of QTLs. An improved method for phenotype evaluation of PHS was employed in this study. In this study, the numbers of germinated and non-germinated grains were hand-threshed and counted for each spike, instead of the visual scoring on a scale of 1 to 9 used in previous studies (McMaster and Derera 1976; Roy et al. 1999; Kulwal et al. 2004, 2005). Germinability, measured as the percentage germination, was explored to estimate the degree of PHS. Although this method is time-consuming, it provides a more objective measure of sprouting damage in a spike and more accurate phenotypic data for QTL mapping.
In Asia, white wheat grain is preferred over the red grain due to consumers’ preference for the white color of steam bread, noodles and other wheat products. Therefore, developing white-grained wheat cultivars in the United States is an attractive measure to expand marketing potential. However, most white wheat cultivars are more susceptible to PHS than red wheat cultivars (Nilsson-Ehle 1914). The association between PHS and grain color in red wheat may be due to either a pleiotropic effect of the genes controlling grain color or to genetic linkage between seed color genes and the genes affecting PHS (Gale 1989; Watanabe and Ikebata 2000). In this study, two white parents were used to develop the mapping population. Totoumai A has a similar level of PHS as some red PHS-resistant wheat cultivars, and it has significantly higher PHS resistance and longer seed dormancy than the sensitive parent Siyang 936. Therefore, the difference in PHS resistance between Totoumai A and Siyang 936 excludes the seed color effect, and the QTL identified in this study can be used to improve PHS resistance of white wheat cultivars.
Since PHS is affected by many environmental factors, phenotyping of PHS is time-consuming and labor intensive. Marker-assisted selection is a desirable approach to developing PHS-resistant cultivars. The QTL detected in the present study will prove valuable in marker-assisted selection aimed at improving the grain quality of wheat in terms of increased resistance to PHS. The genetic distance between the QTL identified in this study and its closest SSR marker, Xbarc170, was about 3.0 cM (Fig. 2). Xbarc170 should be a good marker for marker-assisted selection in early breeding generation to reduce the cost of marker-analysis. However, selecting for flanking markers may increase selection accuracy in late generations when the population size becomes smaller. The markers are also useful for pyramiding PHS resistance with QTLs from other sources and as a starting point for fine mapping and map-based cloning of the major QTL for PHS resistance.
In this study, a single major QTL for controlling both PHS resistance and seed dormancy in the Chinese landrace Totoumai A was detected on chromosome 4AL. This result provides direct evidence that seed dormancy is the most important component of PHS resistance in wheat. Markers linked to the QTL should be valuable for marker-assisted selection of PHS-resistant wheat cultivars.
References
Anderson JA, Sorrells ME, Tanksley SD (1993) RFLP analysis of genomic regions associated with resistance to preharvest sprouting in wheat. Crop Sci 33:453–459
Derera NF (1989) Breeding for preharvast sprouting tolerance. In: Derera NF (ed) Preharvast field sprouting in cereals. CRL, Boca Raton, pp 111–128
Derera NF, Bhatt GM (1980) Germination inhibition of the bracts in relation to pre-harvest sprouting tolerance in wheat. Cereal Res Commun 8:199–201
Doerge RW, Churchill GA (1996) Permutation tests for multiple loci affecting a quantitative character. Genetics 142:285–294
Flintham JE, Adlam R, Bassoi M, Holdsworth M, Gale M (2002) Mapping genes for resistance to sprouting damage in wheat. Euphytica 126:39–45
Gale MD (1989) The genetics of pre-harvest sprouting in cereals, particularly in wheat. In: Derera NF (ed) Pre-harvest field sprouting in cereals. CRC, Boca Raton, pp 85–110
Groos C, Gay G, Perretant M-R, Gervais L, Bernard M, Dedryver F, Charmet G (2002) Study of the relationship between pre-harvest sprouting and grain color by quantitative trait loci analysis in a white × red grain bread-wheat cross. Theor Appl Genet 104:39–47
Himi E, Mares DJ, Yanagisawa A, Noda K (2002) Effect of grain colour gene (R) on grain dormancy and sensitivity of the embryo to abscisic acid (ABA) in wheat. J Exp Bot 53:1569–1574
Kato K, Nakamura W, Tabiki T, Miura H, Sawada S (2001) Detection of loci controlling seed dormancy on group 4 chromosomes of wheat and comparative mapping with rice and barley genomes. Theor Appl Genet 102:980–985
King RW (1984) Water uptake in relation to pre-harvest sprouting damage in wheat: grain characteristics. Aust J Agric Res 36:337–345
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Kulwal PL, Singh R, Balyan HS, Gupta PK (2004) Genetic basis of pre-harvest sprouting tolerance using single-locus and two-locus QTL analyses in bread wheat. Funct Integr Genomics 4:94–101
Kulwal PL, Kumar N, Gaur A, Khurana P, Khurana JP, Tyagi AK, Balyan HS, Gupta PK (2005) Mapping of a major QTL for pre-harvest sprouting tolerance on chromosome 3A in bread wheat. Theor Appl Genet 111:1052–1059
Mares DJ (1987) Pre-harvest sprouting tolerance in white-grained wheat. In: Mares SJ (ed) Fourth international symposium on pre-harvest sprouting in cereals. Westview, Boulder, pp 64–74
Mares DJ, Mrva K, Cheong J, Williams K, Watson B, Storlie E, Sutherland M, Zou Y (2005) A QTL located on chromosome 4A associated with dormancy in white- and red-grained wheats of diverse origin. Theor Appl Genet 111:1357–1364
McMaster GJ, Derera NF (1976) Methodology and sample preparation when screening for sprouting damage in cereals. Cereal Res Commun 4:251–254
Mori M, Uchino N, Chono M, Kato K, Miura H (2005) Mapping QTLs for grain dormancy on wheat chromosome 3A and group 4 chromosomes, and their combined effect. Theor Appl Genet 110:1315–1323
Morris CF, Paulsen GM (1988) Localization and physical properties of endogenous germination inhibitors in white wheat grain. Cereal Chem 65:404–408
Nilsson-Ehle H (1914) Zur Kenntnis der mit Keinumgsphysiologie des Weizens in Zusammenhang stehenden inneren Faktoren. Z Pflanzenzuecht 2:153–157
Noda K, Matsuura T, Maekawa M, Taketa S (2002) Chromosomes responsible for sensitivity of embryo to abscisic acid and dormancy in wheat. Euphytica 123:203–209
Osa M, Kato K, Mori M, Shindo C, Torada A, Miura H (2003) Mapping QTLs for seed dormancy and the Vp1 homologue on chromosome 3A in wheat. Theor Appl Genet 106:1491–1496
Paterson AH, Sorrells ME, Obendorf RL (1989) Methods of evaluation for preharvest sproutiong resistance in wheat breeding programs. Can J Plant Sci 69:681–689
Roy JK, Prasad M, Varshney RK, Balyan HS, Blake TK, Dhaliwal HS, Singh H, Edwards KJ, Gupta PK (1999) Identification of a microsatellite on chromosomes 6B and a STS on 7D of bread wheat showing an association with preharvest sprouting tolerance. Theor Appl Genet 99:336–340
Sorrells ME, Anderson JA (1996) Quantitative trait loci associated with preharvest sprouting in white wheat. In: Noda K, Mares DJ (eds) Preharvest sprouting in cereals 1995. Center for Academic Societies, Japan, pp 137–142
Salmon DF, Helm JH, Duggan TR, Lakeman DM (1986) The influence of chaff extracts on the germination of spring triticale. Agron J 78:863–867
Sharma SK, Dhaliwal HS, Multani DS, Bains SS (1994) Inheritance of preharvast sprouting tolerance in Triticum aestivum and its transfer to an amber-gained cultivar. J Hered 85:312–314
Torada A, Ikeguchi S, Koike M (2005) Mapping and validation of PCR-based markers associated with a major QTL for seed dormancy in wheat. Euphytica 143:251–255
Van Ooijen JW, Voorrips RE (2001) JionMap version 3.0: Software for the calculation of genetic linkage maps
Van Ooijen JW (2004) MapQTL 5: software for the mapping of quantitative trait loci in experimental populations
Varughese G, Barker T, Saari E (1987) Triticale. CIMMYT, Mexico
Walker-Simmonds MK (1987) ABA levels and sensitivity in developing embryos of sprouting resistant and susceptible cultivars. Plant Physiol 84:61–66
Watanabe N, Ikebata N (2000) The effects of homoeologous group 3 chromosomes on grain color dependent seed dormancy and brittle rachis in tetraploid wheat. Euphytica 115:215–220
Zanetti S, Winzeler M, Keller M, Keller B, Messmer M (2000) Genetic analysis of pre-harvest sprouting resistance in a wheat × spelt cross. Crop Sci 40:1406–1417
Acknowledgments
This research is partly supported by the National Research Initiative of the United States Department of Agriculture-The Cooperative State Research, Education, and Extension Service, CAP Grant Number 2006-55606-16629. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture. This is contribution No. 07-188-J from the Kansas Agricultural Experiment Station, Manhattan, Kansas, USA.
Author information
Authors and Affiliations
Corresponding author
Additional information
Shi-Bin Cai and Cui-Xia Chen contributed equally to this work.
Rights and permissions
About this article
Cite this article
Chen, CX., Cai, SB. & Bai, GH. A major QTL controlling seed dormancy and pre-harvest sprouting resistance on chromosome 4A in a Chinese wheat landrace. Mol Breeding 21, 351–358 (2008). https://doi.org/10.1007/s11032-007-9135-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-007-9135-5