Abstract
A library of benzimidazole briged pyrazolo[1,5-a]pyrimidine (6a-q) was designed, synthesized and subjected for evaluation for cytotoxic potential. Antiproliferative activity, ranging from 3.1–51.5 μM, was observed against a panel of cancer cell lines which included MCF-7 (breast cancer), A549 (lung cancer), HeLa (cervical cancer) and SiHa (cervical cancer). Among them, 6k, 6l, 6n and 6o have shown significant cytotoxicity and were investigated further to study their probable mechanism of action against MCF-7 cell line. Accumulation of cells at sub-G1 phase was observed in flow cytometric analysis. The detachment of cells from substratum and membrane blebbing seen under bright field microscopy supports the ability of these conjugates to induce apoptosis. Immunostaining and western blot analysis showed EGFR, p-EGFR, STAT3, and p-STAT3 significant downregulation. Western blot analysis demonstrated an elevated level of apoptotic proteins such as p53, p21, Bax, whereas a decrease in the antiapoptotic protein Bcl-2 and procaspase-9, confirming the ability of these conjugates to trigger cell death by apoptosis. EGFR kinase assay confirms the specific activity of conjugates. Molecular docking simulation study disclosed that these molecules fit well in ATP-binding pocket of EGFR. The analysis of docking poses and the atomic interactions of different conjugates rationalize the structural–activity relationship in this series.
Graphical abstract
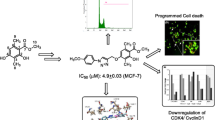
Benzimidazole-linked pyrazolo[1,5-a]pyrimidine conjugates were synthesized and evaluated for their anticancer potential. All the conjugates have significant anticancer potential. Further mechanistic studies revealed that these conjugates arrest cancer cell growth by EGFR/STAT3 inhibition.

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Epidermal growth factor receptor (EGFR) is a member of ErbB family of transmembrane receptor tyrosine kinases [1, 2]. Upon activation of EGFR by growth factors, it forms homodimer or heterodimer with other members of the same family. The dimer subsequently transduces activation signal to the intracellular domain and autophosphorylate intracellular c-terminal tyrosine residues [2,3,4,5]. The phosphorylated receptor relays downstream signals with the help of other signaling molecules in the cascade and regulates important cellular functions such as angiogenesis, invasiveness, proliferation [6, 7], loss of differentiation [8], and decreased apoptosis. Overexpression of EGFR is observed in many solid tumors and is associated with poor prognosis [9,10,11]. It plays a cardinal role in the development of breast cancer [12, 13], and in non-small-cell lung cancer (NSCLC) [14, 15], which is common in lung cancer patients [16].
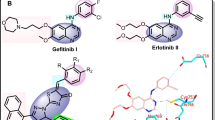
EGFR is known as attractive target for new anticancer drug development and copious inhibitors have been reported. In the past two decades, United States Food and Drug Administration (US FDA) has approved more than ten drugs including Erlotinib (1) and Gefitinib (2), which inhibits EGFR as target and several molecules including BMS-690514 (2) [17] and AEE788 (3) [18], are currently in various stages of clinical trials (Fig. 1) [19, 20]. Thus, EGFR has proven to be an important validated target for the treatment of plentiful of cancers. Hence, exploration of EGFR with newer inhibitors could be a useful approach for the treatment of cancer.
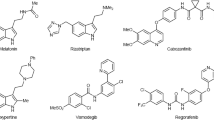
In view of its pharmacological properties, benzimidazole is oldest and one of the most widely used scaffold in the medicinal chemistry. It is a constituent of several drugs for treating different diseases. These include candesartan (antihypertensive), bendamustine (antineoplastic), emedastine (allergic conjunctivitis), omeprazole (proton pump inhibitor), thiabendazole (antiparasitic) and vitamin B12 as well. Anticancer agent nocodazole (5) [21], which is currently undergoing clinical trials, has benzimidazole scaffold in it. This illustrates the potential of benzimidazole for the development of medicinally active conjugates. Similarly, pyrazolo[1,5-a]pyrimidine derivatives are known to exhibit an extended spectrum of biological activities, viz, antitumor [22, 23], anxiolytic [24, 25] and antimicrobial. Comounds bearing pyrazolo[1,5-a]pyrimidine have also been observed in the development of tyrosine kinase inhibitors (CDK) that are involved mitogenic signal tramsmission and other cellular events [22, 26]. A molecule with pyrazolo[1,5-a]pyrimidine scaffold, SCH727965 (4) [27], was ovserved with selective CDK inhibitory activity and is in clinical trials. Besides to this some of the EGFR inhibitors contains pyrazolo[1,5-a]pyrimidine scaffold in it. Our previous attemts towards the synthesis of a variety of hybrid molecules, such as pyrazolo[1,5-a]pyrimidines [26, 28, 29], benzimidazole [30,31,32,33,34] containing hybrids have also led to the development of efficient anticancer agents.
Scaffold hopping is an efficient and commonly known strategy for developing novel and potent molecules [35,36,37,38]. Herein, two or more known pharmacophoric features from the reported molecules were imported into the single-molecule, which may lead to developing a novel molecule with the synergistic effect of two pharmacophores. This strategy is effectively used to minimize the resistance, develop a molecule that can bind to multiple targets, minimize side effects, and improve potency [39]. Thus, it was considered of interest based on the encouraging anticancer profile of pyrazolo[1,5-a]pyrimidines and benzimidazoles; here, we have combined both the pyrazolo[1,5-a]pyrimidine and benzimidazole pharmacophores into a single chemical entity by anticipating an improved biological activity of newly synthesized compounds. The resulting series of benzimidazole-linked pyrazolo[1,5-a]pyrimidines were evaluated for their antiproliferative activity and EGFR inhibitory potential.
Material and methods
Physical measurement and materials
All chemicals and reagents were obtained from Aldrich (Sigma– Aldrich), St. Louis, MO, USA), Lancaster (Alfa Aesar, Johnson Matthey Company, Ward Hill, MA, USA), or Spectrochem Pvt. Ltd (Mumbai, India) and were used without further purification. Reactions were monitored by TLC performed on silica gel glass plate containing 60 GF-254, and visualization was achieved by UV light or iodine indicator. Column chromatography was performed with Merck 60–120 mesh silica gel. 1H and 13C NMR spectra were determined in CDCl3 and DMSO-d6 by using Varian and Avance instruments. Chemical shifts are expressed in parts per million (δ in ppm) downfield from internal TMS and coupling constants are expressed in Hz. 1H NMR spectroscopic data are reported in the following order: multiplicity (s, singlet; br s, broad singlet; d, doublet; dd, a doublet of doublets; t, triplet; m, multiplet), coupling constants in Hz, the number of protons. HRMS analyses were acquired on Agilent Q-TOF-Mass Spectrometer 6540-UHD and carried out in the ESI techniques at 70 eV. Melting points were determined with an Electrothermal melting point apparatus and are uncorrected.
General Procedure for the preparation of 8a-e
Small pieces of metal sodium were added to the ethanol and allowed to react completely. To this solution diethyl oxalate (1 mmol) was added at 0 ºC and the reaction mixture was stirred for 15 min. To this mixture different acetophenones (1 mmol) were added and stirred for 4 h. After reaction completion, the solvent was removed and the mixture was suspended in water extracted with EtOAc. Combined organic phases washed with brine, dried over Na2SO4, concentrated in vacuo. The resulting solid was purified using column chromatography on silica gel to afford pure conjugates 8a-e.
General procedure for preparations of 9a-e
To the solution of compound 8a-e (1 mmol) in ethanol, 3-amino-5-phenylpyrazole (1 mmol) was added followed by 4–5 drops of Conc. HCl. The mixture was refluxed for 4 h, the precipitate was formed. The mixture was cooled to room temperature and precipitated was collected using vacuum filtration, and washed with ethanol to afford pure yellow crystalline compound 9a-e.
Ethyl 7-(4-methoxyphenyl)-2-phenylpyrazolo[1,5-a]pyrimidine-5-carboxylate (9a)
Yellow solid; 90% yield; Mp: 152–154 ºC; 1H NMR (500 MHz, CDCl3): δ 8.31 (d, J = 8.9 H, 2H), 8.01 (d, J = 7.0 2H), 7.68 (s, 1H), 7.47 (t, J = 7.4 Hz, 2H), 7.40 (t, J = 14.6 1H), 7.26 (s, 1H), 7.12 (d, J = 8.5 2H), 4.55 (q, J = 7.1 Hz, 2H), 3.93 (s, 3H), 1.49 (t, J = 7.1 Hz, 3H); MS (ESI): m/z 374 [M + H]+.
Ethyl 7-(3,4-dimethoxyphenyl)-2-phenylpyrazolo[1,5-a]pyrimidine-5-carboxylate (9b)
Yellow solid; 92% yield; Mp: 183–185 ºC; 1H NMR (500 MHz, CDCl3): δ 8.04 (m, 3H), 7.87 (dd, J = 8.5, 2.1 Hz, 1H), 7.71 (s, 1H), 7.47 (t, J = 7.4 Hz, 2H), 7.41 (t, J = 14.5 Hz, 1H), 7.27 (s, 1H), 7.07 (d, J = 8.5 Hz, 1H), 4.56 (q, J = 7.1 Hz, 2H), 4.02 (s, 3H), 4.01 (s, 3H), 1.50 (t, J = 14.3 Hz, 3H); MS (ESI): m/z 404 [M + H]+.
Ethyl 2-phenyl-7-(3,4,5-trimethoxyphenyl)pyrazolo[1,5-a]pyrimidine-5-carboxylate (9c)
Yellow solid; 95% yield; Mp: 194–196 ºC; 1H NMR (500 MHz, CDCl3): δ 8.03 (d, J = 7.2 Hz, 2H), 7.72 (s, 1H), 7.57 (s, 2H), 7.53 (t, J = 14.8 Hz, 2H), 7.44 (t, J = 15.7 Hz, 1H), 7.30 (s, 1H), 4.57 (q, J = 7.1 Hz, 1H), 3.99 (s, 9H), 1.51 (t, J = 7.1 Hz, 3H); MS (ESI): m/z 434 [M + H]+.
Ethyl 7-(3,4-difluorophenyl)-2-phenylpyrazolo[1,5-a]pyrimidine-5-carboxylate (9d)
Yellow solid; 81% yield; Mp: 119–121 ºC; 1H NMR (300 MHz, CDCl3): δ 8.31–8.24 (m, 1H), 8.01 (d, J = 6.7 Hz, 3H), 7.68 (s, 1H), 7.53 – 7.34 (m, 4H), 7.30 (s, 1H), 4.56 (q, J = 7.1 Hz, 2H), 1.50 (t, J = 7.1 Hz, 3H); MS (ESI): m/z 380 [M + H]+.
Ethyl 7-(4-chlorophenyl)-2-phenylpyrazolo[1,5-a]pyrimidine-5-carboxylate (9e)
Yellow solid; 80% yield; Mp: 161–163 ºC; 1H NMR (300 MHz, CDCl3): δ 8.27–8.35 (m, 2H), 7.28–7.36 (m, 3H), 8.02 (d, J = 6.7 Hz, 2H), 7.70 (s, 1H), 7.39–7.53 (m, 3H), 4.56 (q, J = 7.1 Hz, 2H), 1.50 (t, J = 7.1 Hz, 3H); MS (ESI): m/z 378 [M + H]+.
General Procedure for preparation of conjugates (8a-e) pyrazolo[1,5-a]pyrimidine-5-carbaldehydes
To the solution of ester 9a-e in dry dichloromethane, DIBAL-H (1.2 mmol) were added at -78 ºC and reaction was stirred for 30 min. The reaction was quenched using diluted HCl and allowed to come to room temperature. The aqueous layer was extracted with dichloromethane, dried over anhydrous Na2SO4 and concentrated in vacuo. The crude was purified by column chromatography using silica gel and eluent EtOAc:Hexane (2:8).
7-(4-Methoxyphenyl)-2-phenylpyrazolo[1,5-a]pyrimidine-5-carbaldehyde (10a)
Yellow solid; 85% yield; Mp: 214–216 ºC; 1H NMR (500 MHz, CDCl3): δ 10.05 (s, 1H), 8.31 (d, J = 8.9 Hz, 2H), 8.06 (d, J = 7.2 Hz, 2H), 7.53 (s, 1H), 7.49 (t, J = 7.5 Hz, 2H), 7.43 (t, J = 7.3 Hz, 1H), 7.30 (s, 1H), 7.13 (d, J = 8.9 Hz, 2H), 3.94 (s, 3H); MS (ESI): m/z 330 [M + H]+.
7-(3,4-Dimethoxyphenyl)-2-phenylpyrazolo[1,5-a]pyrimidine-5-carbaldehyde (10b)
Yellow solid; 75% yield; Mp: 184–187 ºC; 1H NMR (500 MHz, CDCl3): δ 10.06 (s, 1H), 8.06 (d, J = 7.0 Hz, 2H), 8.01 (d, J = 2.1 Hz, 1H), 7.89 (dd, J = 8.5, 2.1 Hz, 1H), 7.55 (s, 1H), 7.50 (t, J = 14.8 Hz, 2H), 7.44 (t, J = 14.6 Hz, 1H), 7.31 (s, 1H), 7.09 (d, J = 8.5 Hz, 1H), 4.03 (s, 3H), 4.02 (s, 3H); MS (ESI): m/z 360 [M + H]+.
2-Phenyl-7-(3,4,5-trimethoxyphenyl)pyrazolo[1,5-a]pyrimidine-5-carbaldehyde (10c)
Yellow solid; 90% yield; Mp: 186–189 ºC; 1H NMR (500 MHz, CDCl3) δ 10.06 (s, 1H), 8.05 (d, J = 7.2 Hz, 2H), 7.56 (s, 2H), 7.55 (s, 1H), 7.50 (t, J = 7.2 Hz, 2H), 7.46–7.42 (m, 1H), 7.33 (s, 1H), 3.99 (s, 3H), 3.99 (s, 6H); MS (ESI): m/z 390 [M + H]+.
7-(3,4-Difluorophenyl)-2-phenylpyrazolo[1,5-a]pyrimidine-5-carbaldehyde (10d)
Yellow solid; 70% yield; Mp: 186–189 ºC; 1H NMR (500 MHz, CDCl3): δ 10.05 (s, 1H), 8.31–8.25 (m, 1H), 8.03 (d, J = 7.2 Hz, 2H), 8.00 – 7.97 (m, 1H), 7.52–7.48 (m, 3H), 7.47–7.38 (m, 2H), 7.34 (s, 1H); MS (ESI): m/z 336 [M + H]+.
7-(4-Chlorophenyl)-2-phenylpyrazolo[1,5-a]pyrimidine-5-carbaldehyde (10e)
Yellow solid; 75% yield; Mp: 214–216 ºC; 1H NMR (400 MHz, CDCl3) δ 10.06 (s, 1H), 8.21 (d, J = 8.6 Hz, 2H), 8.03 (d, J = 6.9 Hz, 2H), 7.60 (d, J = 8.6 Hz, 2H), 7.56–7.41 (m, 4H), 7.33 (s, 1H); MS (ESI): m/z 334 [M + H]+.
General procedure for preparations of conjugates (6a-q)
To the solution of aldehyde in ethanol, aq. sodium metabisulphite and benzene-1,2-diamines 11a-e were added and reflux for overnight. The precipitate were filtered and washed with small amount of ethanol and purified using column chromatography.
5-(1H-Benzo[d]imidazol-2-yl)-7-(4-methoxyphenyl)-2-phenylpyrazolo[1,5-a]pyrimidine (6a)
This conjugate was prepared according to general procedure, by employing 10a (100 mg, 0.30 mmol) and benzene-1,2-diamine (33 mg, 0.30 mmol) to obtain pure product 6a as yellow color solid; 96 mg; 76% yield; Mp: 229–231 ºC; 1H NMR (500 MHz, CDCl3): δ 12.83 (s, 1H), 8.36 (d, J = 8.9 Hz, 2H), 8.11–7.97 (m, 3H), 7.82 (d, J = 7.6 Hz, 1H), 7.65 (d, J = 7.6 Hz, 1H), 7.60 (s, 1H), 7.48 (d, J = 7.4 Hz, 2H), 7.41 (d, J = 7.4 Hz, 1H), 7.33–7.27 (m, 1H), 7.12 (d, J = 8.9 Hz, 2H), 7.05 (s, 1H), 3.94 (s, 3H); 13C NMR (125 MHz, CDCl3 + DMSO): δ 207.0, 162.1, 156.0, 150.6, 147.2, 146.0, 132.8, 131.8, 129.6, 129.3, 126.8, 122.9, 114.5, 104.5, 94.1, 55.9; HRMS calculated for C26H19ON5 [M + H]+ 418.1668, found 418.1666.
7-(4-Methoxyphenyl)-5-(6-methyl-1H-benzo[d]imidazol-2-yl)-2-phenylpyrazolo[1,5-a]pyrimidine (6b)
This conjugate was prepared according to general procedure, by employing 10a (100 mg, 0.30 mmol) and 4-methylbenzene-1,2-diamine (37 mg, 0.30 mmol) to obtain pure product 6b as pale yellow color solid; 108 mg; 82% yield; Mp: 245–247 ºC; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.34 (d, J = 8.8 Hz, 2H), 8.04(s, 1H), 8.01 (s, 2H), 7.60 (s, 1H), 7.49–7.38 (m, 4H), 7.11 (d, J = 8.8 Hz, 3H), 7.03 (s, 1H), 3.93 (s, 3H), 2.49 (s, 3H); 13C NMR (75 MHz, CDCl3 + DMSO): δ 159.7, 153.6, 148.4, 147.1, 145.6, 143.6, 130.6, 129.4, 127.1, 126.8, 124.5, 120.8, 112.1, 102.2, 91.5, 53.5, 19.6; HRMS calculated for C26H19ON5 [M + H]+ 432.1824, found 432.1823.
5-(5,6-Dimethyl-1H-benzo[d]imidazol-2-yl)-7-(4-methoxyphenyl)-2-phenylpyrazolo[1,5-a]pyrimidine (6c)
This conjugate was prepared according to general procedure, by employing 10a (100 mg, 0.30 mmol) and 4,5-dimethylbenzene-1,2-diamine (41 mg, 0.30 mmol) to obtain pure product 6c as yellow color solid. 106 mg; 79% yield; Mp: 251–253 ºC; 1H NMR (500 MHz, CDCl3): δ 8.35 (d, J = 8.9 Hz, 2H), 8.03 (s, 1H), 8.01 (s, 2H), 7.54 (d, J = 8.5 Hz, 1H), 7.47 (t, J = 7.3 Hz, 3H), 7.40 (t, J = 7.3 Hz, 1H), 7.09 (d, J = 8.9 Hz, 2H), 7.00 (s, 1H), 3.93 (s, 3H), 2.39 (s, 6H); 13C NMR (100 MHz, CDCl3): δ 161.9, 150.6, 148.7, 146.5, 132.9, 131.5, 130.2, 129.0, 128.7, 126.6, 125.8, 123.0, 118.9, 117.3, 113.9, 107.9, 103.8, 93.4, 55.5, 19.5; HRMS calculated for C28H23N5O [M + H]+ 445.1981, found 446.19745.
5-(6-Fluoro-1H-benzo[d]imidazol-2-yl)-7-(4-methoxyphenyl)-2-phenylpyrazolo[1,5-a]pyrimidine (6d)
This conjugate was prepared according to general procedure, by employing 10a (100 mg, 0.30 mmol) and 4-fluorobenzene-1,2-diamine (38 mg, 0.30 mmol) to obtain pure product 6d as orange color solid. 92 mg; 70% yield; Mp: 270–273 ºC; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.37 (d, J = 8.9 Hz, 2H), 8.03 (d, J = 7.0 Hz, 2H), 7.99 (d, J = 7.1 Hz, 2H), 7.67 (s, 1H), 7.52–7.42 (m, 3H), 7.24 (t, J = 8.9 Hz, 3H), 7.06 (t, J = 8.1 Hz, 1H), 3.95 (s, 3H); 13C NMR (125 MHz, CDCl3 + DMSO): δ 159.8, 153.8, 150.9, 148.4, 145.2, 143.9, 142.2, 138.6, 138.3, 138.1, 130.5, 129.5, 127.3, 126.9, 124.5, 120.7, 112.6, 112.2, 111.9, 91.8, 53.6; HRMS calculated for C26H18FN5O [M + H]+ 436.1574, found 436.1561.
5-(6-Chloro-1H-benzo[d]imidazol-2-yl)-7-(4-methoxyphenyl)-2-phenylpyrazolo[1,5-a]pyrimidine (6e)
This conjugate was prepared according to general procedure, by employing 10a (100 mg, 0.30 mmol) and 4-chlorobenzene-1,2-diamine (43 mg, 0.30 mmol) to obtain pure product 6e as light orange color solid; 110 mg; 80% yield; Mp: 249–251 ºC; 1H NMR (300 MHz, CDCl3 + DMSO): δ 8.36 (d, J = 8.8 Hz, 2H), 8.13 – 7.96 (m, 3H), 7.69–7.64 (m, 2H), 7.50–7.38 (m, 3H), 7.23 (dd, J = 8.6, 1.6 Hz, 1H), 7.13 (d, J = 8.8 Hz, 2H), 7.07 (s, 1H), 3.94 (s, 3H); 13C NMR (75 MHz, CDCl3 + DMSO): δ 161.63, 155.54, 150.47, 150.16, 146.82, 145.61, 132.27, 131.36, 129.19, 128.81, 126.32, 122.38, 114.06, 104.06, 93.68, 55.47; HRMS calculated for C26H18ClN5O [M + H]+ 452.1278, found 452.1258.
5-(1H-Benzo[d]imidazol-2-yl)-7-(3,4-dimethoxyphenyl)-2-phenylpyrazolo[1,5-a]pyrimidine (6f)
This conjugate was prepared according to general procedure, by employing 10b (100 mg, 0.28 mmol) and benzene-1,2-diamine (30 mg, 0.28 mmol) to obtain pure product 6f as yellow color solid; 106 mg; 85% yield; Mp: 180–182 ºC; 1H NMR (300 MHz, CDCl3): δ 10.71 (s, 1H), 8.10 (s, 1H), 8.08–8.05 (m, 2H), 8.03 (s, 1H), 7.95 (dd, J = 8.5, 2.0 Hz, 1H), 7.84 (s, 1H), 7.66 (s, 1H), 7.51–7.41 (m, 3H), 7.32 (d, J = 5.1 Hz, 2H), 7.09 (d, J = 8.6 Hz, 1H), 7.05 (s, 1H), 4.03 (s, 3H),4.01 (s, 3H); 13C NMR (125 MHz, CDCl3 + DMSO): δ 155.8, 151.2, 150.3, 149.3, 148.1, 146.9, 145.9, 132.4, 131.8, 130.6, 128.7, 128.4, 126.1, 124.0, 123.1, 122.7, 112.4, 110.4, 104.0, 93.1, 55.8, 55.7; HRMS calculated for C27H21N5O2 [M + H]+ 448.1773, found 448.1791.
7-(3,4-Dimethoxyphenyl)-5-(6-methyl-1H-benzo[d]imidazol-2-yl)-2-phenylpyrazolo[1,5-a]pyrimidine (6g)
This conjugate was prepared according to general procedure, by employing 10b (100 mg, 0.28 mmol) and 4-methylbenzene-1,2-diamine (34 mg, 0.28 mmol) to obtain pure product 6g as light yellow color solid; 91 mg; 71% yield; Mp: 223–225 ºC;1H NMR (400 MHz, CDCl3): δ 8.05 (s, 2H), 8.01 (d, J = 7.3 Hz, 2H), 7.90 (d, J = 9.7 Hz, 1H), 7.77 (s, 1H), 7.47–7.38 (m, 4H), 7.16 (d, J = 8.3 Hz, 1H), 7.02 (d, J = 8.55 Hz, 1H), 7.00 (s, 1H), 4.00 (s, 3H), 3.98 (s, 3H), 2.50 (s, 3H); 13C NMR (125 MHz, CDCl3 + DMSO): δ 155.4, 150.8, 150.1, 148.6, 147.8, 146.9, 145.5, 132.1, 128.4, 128.1, 125.8, 125.7, 124.5, 122.9, 122.8, 122.5, 112.2, 112.1, 110.2, 103.8, 103.7, 92.7, 55.5, 55.4, 21.2; HRMS calculated for C28H23N5O2 [M + H]+ 462.1930, found 462.1913.
7-(3,4-Dimethoxyphenyl)-5-(5,6-dimethyl-1H-benzo[d]imidazol-2-yl)-2-phenylpyrazolo[1,5-a]pyrimidine (6h)
This conjugate was prepared according to general procedure, by employing 10b (100 mg, 0.28 mmol) and 4,5-dimethylbenzene-1,2-diamine (38 mg, 0.28 mmol) to obtain pure product 6h as light amber color solid; 110 mg; 83% yield; Mp: 230–232 ºC; 1H NMR (500 MHz, CDCl3): δ 8.06 (d, J = 8.2 Hz, 2H), 8.02 (d, J = 7.2 Hz, 2H), 7.96 (d, J = 7.5 Hz, 1H), 7.54–7.38 (m, 5H), 7.06 (d, J = 8.5 Hz, 1H), 7.02 (s, 1H), 4.03 (s, 3H), 4.01 (s, 3H), 2.38 (s, 6H); 13C NMR (75 MHz, CDCl3 + DMSO): δ 155.4, 150.9, 150.1, 147.9, 147.1, 145.5, 135.2, 132.2, 132.1, 128.5, 128.2, 125.8, 122.8, 122.6, 114.9, 112.2, 110.3, 108.6, 103.9, 92.8, 82.5, 55.6, 55.5, 18.9; HRMS calculated for C29H25N5O2 [M + H]+ 476.2087, found 476.2078.
7-(3,4-Dimethoxyphenyl)-5-(6-fluoro-1H-benzo[d]imidazol-2-yl)-2-phenylpyrazolo[1,5-a]pyrimidine (6i)
This conjugate was prepared according to general procedure, by employing 10b (100 mg, 0.28 mmol) and 4-fluorobenzene-1,2-diamine (35 mg, 0.28 mmol) to obtain pure product 6i as yellow color solid; 97 mg; 75% yield; Mp: 224–226 ºC; 1H NMR (300 MHz, CDCl3 + DMSO): δ 12.79 (s, 1H), 8.07–8.04 (m, 4H), 7.96 (d, J = 8.3 Hz, 1H), 7.55–7.32 (m, 5H), 7.15–7.05 (m, 3H), 4.04 (s, 3H), 4.02 (s, 3H); 13C NMR (125 MHz, CDCl3 + DMSO): δ 160.9, 156.3, 155.5, 153.2, 152.1, 151.0, 137.5, 133.9, 133.6, 131.3, 131.1, 128.3, 128.2, 128.2, 127.8, 125.4, 125.4, 117.6, 117.5, 115.7, 109.2, 109.0, 98.2, 60.9, 60.8. HRMS calculated for C27H20FN5O2 [M + H]+ 466.1601, found 466.1661.
5-(6-Chloro-1H-benzo[d]imidazol-2-yl)-7-(3,4-dimethoxyphenyl)-2-phenylpyrazolo[1,5-a]pyrimidine (6j)
This conjugate was prepared according to general procedure, by employing 10b (100 mg, 0.28 mmol) and 4-chlorobenzene-1,2-diamine (40 mg, 0.28 mmol) to obtain pure product 6j as yellow color solid; 102 mg; 76% yield; Mp: 257–260 ºC; 1H NMR (300 MHz, CDCl3): δ 10.73 (s, 1H), 8.04 (d, J = 7.1 Hz, 2H), 8.01 (s, 2H), 7.89 (dd, J = 8.5, 1.8 Hz, 1H), 7.77 (s, 1H), 7.57–7.37 (m, 4H), 7.30 (d, J = 8.6 Hz, 1H), 7.03 (d, J = 9.6 Hz, 2H), 4.01 (s, 3H), 4.00 (s, 3H); 13C NMR (75 MHz, CDCl3): δ 156.5, 151.7, 150.5, 148.5, 146.6, 146.1, 132.6, 129.2, 128.8, 126.5, 124.2, 124.0, 123.9, 123.4, 122.9, 112.7, 112.3, 111.4, 110.7, 103.8, 93.7, 56.1, 56.0; HRMS calculated for C27H20ClN5O2 [M + H]+ 482.1305, found 482.1378.
5-(1H-Benzo[d]imidazol-2-yl)-2-phenyl-7-(3,4,5-trimethoxyphenyl)pyrazolo[1,5-a]pyrimidine (6k)
This conjugate was prepared according to general procedure, by employing 10c (100 mg, 0.26 mmol) and benzene-1,2-diamine (28 mg, 0.26 mmol) to obtain pure product 6k as yellow color solid; 107 mg; 87% yield; Mp: 232–234 ºC; 1H NMR (500 MHz, CDCl3): δ 10.82 (s, 1H), 8.09 (s, 1H), 8.02 (d, J = 7.2 Hz, 2H), 7.90 (d, J = 7.6 Hz, 1H), 7.60 (s, 2H), 7.56 (d, J = 7.5 Hz, 1H), 7.46 (t, J = 7.4 Hz, 2H), 7.43–7.34 (m, 3H), 7.04 (s, 1H), 3.98 (s, 3H), 3.97 (s, 6H); 13C NMR (75 MHz, CDCl3): δ 156.5, 153.0, 150.5, 149.4, 146.6, 134.0, 132.6, 129.2, 128.8, 126.4, 125.6, 124.9, 123.1, 120.4, 111.5, 107.2, 104.4, 93.7, 61.0, 56.4; HRMS calculated for C28H23N5O3 [M + H]+ 478.1879, found 478.1864.
5-(6-Methyl-1H-benzo[d]imidazol-2-yl)-2-phenyl-7-(3,4,5-trimethoxyphenyl)pyrazolo[1,5-a]pyrimidine (6l)
This conjugate was prepared according to general procedure, by employing 10c (100 mg, 0.26 mmol) and 4-methylbenzene-1,2-diamine (32 mg, 0.26 mmol) to obtain pure product 6l as pale yellow color solid; 107 mg; 85% yield; Mp: 210–212 ºC; 1H NMR (400 MHz, CDCl3): δ 8.06 (s, 1H), 8.04 (d, J = 7.09 Hz, 2H), 7.77 (d, J = 9.5 Hz, 1H), 7.60 (s, 2H), 7.50–7.36 (m, 4H), 7.18 (t, J = 8.7 Hz, 1H), 7.03 (s, 1H), 3.98 (s, 3H), 3.97 (s, 6H), 2.51 (s, 3H); 13C NMR (100 MHz, CDCl3): δ 156.5, 153.1, 150.6, 148.9, 146.9, 146.6, 142.6, 140.7, 135.2, 134.3, 132.7, 129.2, 128.8, 126.4, 125.7, 125.0, 119.9, 111.2, 107.3, 104.3, 93.7, 61.0, 56.5, 21.9; HRMS calculated for C29H25N5O3 [M + H]+ 492.2036, found 492.2021.
5-(5,6-Dimethyl-1H-benzo[d]imidazol-2-yl)-2-phenyl-7-(3,4,5-trimethoxyphenyl)pyrazolo[1,5-a]pyrimidine (6m)
This conjugate was prepared according to general procedure, by employing 10c (100 mg, 0.26 mmol) and 4,5-dimethylbenzene-1,2-diamine (35 mg, 0.26 mmol) to obtain pure product 6m as amber color solid; 95 mg; 73% yield; Mp: 218–221 ºC; 1H NMR (500 MHz, CDCl3): δ 8.08 (s, 1H), 8.02 (d, J = 7.3 Hz, 2H), 7.65 (s, 2H), 7.47 (t, J = 7.4 Hz, 3H), 7.41 (t, J = 7.3 Hz, 2H), 7.05 (s, 1H), 4.01 (s, 6H), 3.99 (s, 3H), 2.39 (s, 6H); 13C NMR (125 MHz, CDCl3): δ 155.9, 152.7, 150.3, 148.4, 147.2, 145.8, 140.1, 132.4, 132.4, 128.8, 128.5, 126.1, 126.0, 125.5, 107.0, 104.5, 93.2, 60.6, 56.1, 19.1; HRMS calculated for C30H27N5O3 [M + H]+ 506.2192, found 506.2169.
5-(6-Fluoro-1H-benzo[d]imidazol-2-yl)-2-phenyl-7-(3,4,5-trimethoxyphenyl)pyrazolo[1,5-a]pyrimidine (6n)
This conjugate was prepared according to general procedure, by employing 10c (100 mg, 0.26 mmol) and 4-fluorobenzene-1,2-diamine (25 mg, 0.26 mmol) to obtain pure product 6m as light orange color solid; 95 mg; 75% yield; Mp: 243–246 ºC; 1H NMR (400 MHz, CDCl3): δ 10.67 (s, 1H), 8.03 (d, J = 6.8 Hz, 3H), 7.61 (s, 2H), 7.51–7.33 (m, 4H), 7.27 (s, 1H), 7.13 (dd, J = 15.9, 8.1 Hz, 1H), 7.07 (m, 1H), 3.99 (s, 9H); 13C NMR (125 MHz, CDCl3 + DMSO): δ 160.8, 157.8, 155.5, 155.4, 152.2, 150.8, 145.3, 137.5, 137.0, 136.3, 136.2, 134.2, 133.9, 131.4, 131.3, 130.7, 112.4, 112.3, 109.9, 109.7, 98.8, 65.4, 61.3; HRMS calculated for C28H22FN5O3 [M + H]+ 496.1785, found 496.1771.
5-(6-Chloro-1H-benzo[d]imidazol-2-yl)-2-phenyl-7-(3,4,5-trimethoxyphenyl)pyrazolo[1,5-a]pyrimidine (6o)
This conjugate was prepared according to general procedure, by employing 10c (100 mg, 0.26 mmol) and 4-chlorobenzene-1,2-diamine (37 mg, 0.26 mmol) to obtain pure product 6o as orange color solid; 93 mg; 71% yield; Mp: 219–221 ºC; 1H NMR (400 MHz, CDCl3): δ 8.04 (s, 2H), 8.02 (s, 1H), 7.62 (s, 2H), 7.50–7.41 (m, 4H), 7.32 (d, J = 8.7 Hz, 1H), 7.27 (s, 1H), 7.08 (s, 1H), 4.00 (s, 6H), 4.00 (s, 3H); 13C NMR (125 MHz, CDCl3 + DMSO): δ 156.0, 152.6, 150.2, 150.1, 146.6, 145.9, 140.1, 132.2, 130.6, 130.6, 128.8, 128.4, 126.0, 125.3, 123.6, 113.8, 107.6, 106.9, 104.8, 104.4, 93.3, 60.5, 56.0; HRMS calculated for C28H22ClN5O3 [M + H]+ 512.1489, found 512.14850.
5-(1H-Benzo[d]imidazol-2-yl)-7-(3,4-difluorophenyl)-2-phenylpyrazolo[1,5-a]pyrimidine (6p)
This conjugate was prepared according to general procedure, by employing 10d (100 mg, 0.30 mmol) and benzene-1,2-diamine (32 mg, 0.30 mmol) to obtain pure product 6p as pale yellow color solid; 101 mg; 80% yield; Mp: 206–208 ºC; 1H NMR (300 MHz, CDCl3): δ 12.14 (s, 1H), 8.42–8.27 (m, 1H), 8.15–7.97 (m, 3H), 7.75 (d, J = 54.0 Hz, 2H), 7.52–7.39 (m, 4H), 7.36 (s, 1H), 7.34 (d, J = 5.7 Hz, 2H), 7.07 (s, 1H); 13C NMR (125 MHz, CDCl3 + DMSO): δ 154.3, 148.6, 147.4, 145.8, 141.9, 130.6, 127.5, 127.1, 126.0, 125.2, 124.8, 124.7, 121.6, 117.5, 117.3, 116.2, 116.1, 103.7, 103.6, 92.3; HRMS calculated for C28H22ClN5O3 [M + H]+ 424.1374, found 424.1386.
5-(1H-Benzo[d]imidazol-2-yl)-7-(4-chlorophenyl)-2-phenylpyrazolo[1,5-a]pyrimidine (6q)
This conjugate was prepared according to general procedure, by employing 10e (100 mg, 0.30 mmol) and benzene-1,2-diamine (32 mg, 0.30 mmol) to obtain pure product 6q as yellow color solid; 100 mg; 79% yield; Mp: 268–270 ºC; 1H NMR (300 MHz, CDCl3): δ 13.51 (s, 1H), 8.36 (d, J = 8.6 Hz, 2H), 8.09 (d, J = 7.0 Hz, 2H), 8.01 (s, 1H), 7.76 (t, J = 7.3 Hz, 3H), 7.60 (d, J = 7.7 Hz, 1H), 7.52–7.42 (m, 3H), 7.40 (s, 1H), 7.34–7.23 (m, 2H); 13C NMR (126 MHz, DMSO): δ 156.14, 150.54, 149.54, 147.99, 145.22, 144.27, 136.50, 135.77, 132.66, 131.93, 129.78, 129.35, 129.23, 126.87, 124.64, 122.89, 120.20, 112.84, 105.73, 94.46; HRMS calculated for C25H16ClN5 [M + H]+.; 422.1172 found: 422.1155.
Cell cultures, maintenance and antiproliferative activity evaluation
All cell lines used in this study were purchased from the American Type Culture Collection (ATCC, United States). MCF-7, A549, HeLa, SiHa, HEK293 and MRC5 were grown in Dulbecco’s modified Eagle’s medium (containing 10% FBS under a humidified atmosphere of 5% CO2 at 37 °C). Cells were trypsinized when sub-confluent from T25 flasks/60 mm dishes and seeded in 96-well plates. The synthesized test conjugates were tested for their in vitro antiproliferative activities in four different human cancer cell lines. A protocol of 24 h continuous drug exposure was used and an MTT cell proliferation assay was used to estimate cell viability. Individual cell lines were seeded into 96-well microtiter plates in 200 μL aliquots at plating densities depending on the doubling time of individual cell lines. The microtiter plates were incubated at 37 °C, 5% CO2, 95% air and 100% relative humidity for 24 h prior to the addition of experimental drugs. Aliquots of 2 μL of the test conjugates were added to the wells already containing 198 μL of cells, resulting in the required final drug concentrations. For each conjugate, five concentrations (0.01, 0.1, 1, 10 and 100 μM) were evaluated and each was done in triplicate wells. Plates were incubated further for24 h and the assay was terminated by the addition of 10 μL of 5% MTT and incubated for 60 min at 37 °C. Later, the plates were air-dried. The bound stain was subsequently eluted with100 μL of DMSO and the absorbance was read on a multimode plate reader (Varioscan Flash) at a wavelength of 560 nm. Percent growth was calculated on a plate by plate basis for test wells relative to control wells. The above determinations were repeated thrice. The growth inhibitory effects of the conjugates were analyzed by generating dose–response curves as a plot of the percentage of surviving cells versus the conjugate concentration. The sensitivity of the cancer cells to the test conjugate was expressed in terms of IC50, a value defined as the concentration of the conjugate that produced a 50%reduction as compared to the control absorbance. IC50values are indicated as means ± SD of three independent experiments.
Analysis of cell cycle
MCF-7 cells were grown in 60 mm dishes and were incubated for 24 h in the presence or absence of test conjugates 6k, 6l, 6n, 6o and erlotinib at 5 μM concentration. Cells were harvested using Trypsin–EDTA, fixed with ice-cold 70% ethanol at 4 °C for 30 min, ethanol was removed by centrifugation and cells were stained with 1 mL of DNA staining solution [0.05 mg of propidium iodide (PI) and 100 μg RNase A for 30 min in the dark at 37 °C. The DNA contents of 20,000 events were measured using a flow cytometer (BD MoFlo Legacy). Histograms were analyzed using SummitV4.3.39 [40].
Immunocytochemistry
MCF-7 cells were seeded on a glass coverslip and incubated for 24 h in the presence or absence of test conjugates 6k, 6l, 6n, 6o and erlotinib at a concentration of 5 μM. The cells grown on the coverslips were fixed in 4% formaldehyde in phosphate-buffered saline (PBS), pH 7.4, for 10 min at room temperature. Cells were permeabilized for 6 min in PBS containing 0.5% Triton X-100 (Sigma) and 0.05% Tween-20 (Sigma). The permeabilized cells were blocked with 2% BSA (Sigma) in PBS for 1 h. Later, the cells were incubated with a primary antibody for Rabbit Anti p-EGFR (Cat No: 3777S, Cell Signaling Technology, Inc. USA) at 1: 200 and Rabbit Anti STAT3 (Cat No: 9132L, Cell Signaling Technology, Inc. USA) at 1: 200 diluted in blocking solution for 2 h at room temperature. Subsequently, the antibodies were removed and the cells were washed thrice with PBS. Cells were then incubated with a Cy-3 labeled anti-rabbit secondary antibody (1: 500) for 1 h at room temperature. Cells were washed thrice with PBS and mounted in a medium containing DAPI (Vecta Shield). Images were captured using the Olympus confocal microscope FLOW VIEW FV 1000series and analyzed using FV10ASW 1.7 series software [28].
Western blot analysis
Cells were lysed and the lysate was extracted using RIPA lysis buffer after treatment with conjugates for 24 h at 5uM final drug concentration. Protein was quantitated using the Bradford assay and 20 μg of total protein was loaded per well, and resolved on 8.0%, 10.0%, or 12% SDS–polyacrylamide gels. The gels were then transferred to Immoblion-P, PVDF (Millipore, Billerica, Massachusetts) using semidry transfer technique and probed with the primary and secondary antibodies. ECL (GE, Pittsburgh, Pennsylvania) was used as the chemiluminescent substrate [41].
Plasmid DNA constructs and transfection procedure
EGFR-GFP plasmid was a gift from Alexander Sorkin (Addgene plasmid #32,751) [42]. HEK293 cells were seeded in 60 mm cell culture dishes at the semi-confluent level 2 µg of plasmid was transfected using Lipofectamine2000 under serum depletion for 6 h. Media was changed after 6 h to normal growth medium supplemented with 10% FBS. 48 h post transfection cells were treated with conjugates at 5 µM final concentration for 24 h and total protein was isolated using RIPA cell lysis buffer.
EGFR kinase assay
EGFR kinase assay was performed using EGFR Kinase Enzyme System (Promega Cat#V3831) and activity was detected using ADP-Glo™ Kinase Assay (Promega Cat#V9101) according to the manufacturer protocol provided. To determine the IC50 value of the compounds 6k, 6l, 6n, 6o and Erlotinib. Compounds were serially diluted to get the final concentrations in between 10,000 nM to 0.01 nM and enzyme was taken 200 ng to 0.1953 ng by serial dilution technique, the essay is performed employing dose response curve method according to protocol mentioned by the manufacturer.
Molecular modeling studies
Coordinates of the protein were obtained from Protein Data Bank (PDB ID 1M17) [43]. Necessary corrections to the protein were done using Protein Preparation Wizard from Schrodinger package [44]. The protein preparation was accomplished by assigning bond orders, deleting water molecules from the crystal structure, hydrogens were added for pH 7.0 using Epik and termini were capped. Finally, the protein structure was optimized and minimized using the OPLS3e force field to avoid steric clashes between atoms. The geometry of the ligand molecules was optimized in Gaussian 09 using the PM3 semi-empirical method [45]. AutoGrid4 was used to create grid maps. The receptor grid centered on the cocrystal ligand erlotinib and the grid box size was 40 X 40 X 40 Å. AutoDock4 (version 4.2) with the Lamarckian genetic algorithm was used to perform the docking studies [46]. Docking parameters selected for AutoDock4 runs were as follows: 10 docking runs, the population size of 150, random starting position and conformation, 2.5 million energy evaluations, translation step ranges of 2.0 Å, the mutation rate of 0.02 and crossover rate of 0.8. Docked conformations were clustered using a tolerance of 2.0 Å RMSD. Visualization of docking results and image generation was done in PyMol software [47].
Results and discussion
Chemistry
Benzimidazole-linked pyrazolo[1,5-a]pyrimidines (6a-q) were synthesized as in Scheme 1. Different acetophenones (7a-e) were reacted with diethyl oxalate and sodium ethoxide in a solvent ethanol to give 1,3-diketones (8a-e). 8a-e that were subsequently cyclized with 3-amino-5-phenyl-1H-pyrazole in ethanol to produce pyrazolo[1,5-a]pyrimidine esters (9a-e). The pyrazolo[1,5-a]pyrimidine esters (9a-e) were reduced using DIBAL-H to yield corresponding pyrazolo[1,5-a]pyrimidine-5-carbaldehydes (10a-e). Different benzene-1,2-diamines were racted with pyrazolo[1,5-a]pyrimidine-5-carbaldehydes to offer benzimidazole-linked pyrazolo[1,5-a]pyrimidines (6a-q).
Biology
Antiproliferative activity
The compounds (6a-q) were assess for in vitro antiproliferative potential against a selected of cancer cell lines such as A549 (lung cancer), MCF-7 (breast cancer), HeLa (cervical cancer), SiHa (cervical cancer) and MRC5 (normal lung fibroblasts) by employing 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay. Cytotoxic activity is expressed as IC50 values in μM using roscovitine as a positive control (Table 1). All the conjugates demonstrated significant antiproliferative activity against the aforementioned cell lines however no significant effect was observed on non- cancerous cell line MRC5. Moreover, it was very interesting to study the effect of substitution pattern on the antiproliferative activity. The introduction of different substituents on C-7 phenyl ring and C-5 benzimidazole ring were exemplified by electron withdrawing and electron donating groups. With the exception of 6m, conjugates with 3,4,5-trimethoxyphenyl on C-7 showed superior activity with IC50 ranging from 3.2–6.6 μM on MCF-7. Among these conjugates, 6n with 3,4,5-trimethoxyphenyl on the C-7 and 5-F on C-5 benzimidazole ring had the highest potency (IC50 3.2 μM) on MCF-7 cell line. The same conjugate showed IC50 values of 4.2 μM, 8.9 μM and 7.9 μM against the A549, HeLa and SiHa cell lines, respectively. Conjugates with electron donating groups, such as 3,4,5-trimethoxy, 3,4-dimethoxy, and 4-methoxy on C-7 phenyl ring (6a-o) exhibited superior activity as compared to the conjugates with electron withdrawing groups, such as 3,4-diF and 4-Cl (6p, 6q). The activity order for C-7 phenyl substituents was 3,4,5-trimethoxy > 3,4-dimethoxy ≥ 4-methoxy > 3,4-F ≥ 4-Cl. Specifically, conjugates with hydrogen bond acceptor on the meta- and para- positions of the C-7 phenyl ring had superior activity than other conjugates in the same series. In contrast, electron withdrawing substituents on the C-5 benzimidazole ring enhanced the activity, while electron donating groups reduced it. Thus the activity order for the C-5 benzimidazole substitutions was 5-F > 5-Cl > 5-Me > 5,6-diMe. Electron donating groups at C-7 phenyl ring enhanced the activity while electron withdrawing groups diminished the activity. This was quite significantly in contrast to SAR observed from substituents on the C-5 benzimidazole ring. In the literature, many benzimidazole pharmacophores containing anticancer molecules were reported [48, 49]. Sharma et al. reported the benzimidazole-thiazolidinedione hybrids as anticancer agents [50]. The most active molecule 11p exhibited cytotoxicity 11.46 µM on the A549 cancer cell line. Gopal et al. synthesized and characterized N-benzimidazole linked pyrimidine for their anticancer potential [51]. Compound 8b possesses the 39.6 µM activity. Our synthesized compounds possess higher cytotoxic potential compared to the above compounds. Four of the most potent conjugates (6k, 6l, 6n and 6o) were selected for the mechanistic studies based on the cytotoxic activity results.
Cell cycle analysis
Effect of four compounds 6k, 6l, 6n and 6o on cell cycle progression consisting of MCF-7 cells was studied at 5 μM concentration for 24 h. Cell cycle analysis performed with PI staining showed accumulation of cells at sub-G1 phase (Fig. 2). Untreated cells showed 7.9% cell distribution in sub-G1 phase. In contrast, compounds 6k, 6l, 6n and 6o treated cells showed accumulation of 41.5%, 20.1%, 67.5% and 73.0%, respectively. Erlotinib was used as positive control, and it showed 9.3% sub-G1 population. Thus, molecules under investigation showed a more significant effect in restricting MCF-7 cells to sub-G1 region, which in itself is a strong evidence of apoptotic cell death.
Effect of 6k, 6l, 6n and 6o on Cell Cycle Distribution. FACS cell cycle studies showing increased sub-G1 population upon treatment with compounds 6k, 6l, 6n and 6o at 5 μM concentration for 24 h. erlotinib served as positive control at 5 μM, Sub-G1 region indicated with red lines and the percentage of population in the region is mentioned above it. Compounds showed percentage of sub-G1 population as 6n > 6o > 6k > 6l > erlotinib > UT
Cell morphology study
In order to further substantiate the cell cycle arrest which resulted in cell death by means of apoptosis, cell morphological changes were also studied upon treatment of conjugate. MCF-7 cells were seeded in a 6-well plate and then incubated with the conjugates at 5 μM concentration for 24 h. Cells were captured under bright field microscope with a 10 × objective. Highly detached cells and cell membrane bleb were observed with conjugates 6k, 6o, and 6n, whereas molecule 6l showed cytosolic vacuoles and cell detachment (Fig. 3). Untreated and erlotinib-treated cells did not show significant morphological changes. This observation hinted towards apoptotic cellular death is indeed the preferred mode of cytotoxicity. The loss of cellular structural integrity and detachment with substratum and membrane bleb indicated the ability of the conjugates to induce apoptosis in MCF-7 cells.
Effect on EGFR/STAT3 pathway
The present study was also intended to elucidate the pathway responsible for the cell death through apoptosis. MCF-7 cells were treated with test compounds under investigation at 5 μM concentration for 24 h, followed by total protein lysate extraction. Protein lysate was subjected to western blot analysis using specific antibodies to determine their cellular levels and their implications on cell proliferation and apoptosis. Interestingly EGFR and phosphorylated EGFR levels dropped significantly upon conjugate treatment as regulation of STAT3 and its phosphorylated form was also observed. STAT3 is a well-known transcription factor responsible for cell survival and proliferation and is commonly compared to the control. These results highly correlated with erlotinib-treated cells (Fig. 4). Further, a significant down overexpressed in most cancer types. Any mode of therapy that lowers the levels of STAT3 in cancer patients will thus become an efficient strategy to control the cancerous growth. Among the conjugates studied, 6l, 6n and 6o also demonstrated a significant effect in the modulation of the EGFR/STAT3 axis as compared to the untreated control (Fig. 4). They also showed a high degree of correlation with the erlotinib-treated cells used as a positive control.
MCF-7 cells were treated with 6k, 6l, 6n and 6o at 5 μM concentration and erlotinib as positive control at 5 μM for 24 h. Protein lysate was obtained and subjected to western blotting, Data showing the significant down regulation of EGFR, p-EGFR and its downstream transcription factor STAT3, p-STAT3 by 6l, 6n and 6o followed by positive control erlotinib. β-Actin served as loading control
Immunostaining
Further the study was pondered to confirm the deregulation of the EGRF/STAT axis in MCF-7 cells. Cellular protein levels and the localization of p-EGFR and STAT3 proteins were identified by immunofluorescence method. The confocal images of p-EGFR and panSTAT3 treated with the test compounds along with the untreated and erlotinib-treated cells are depicted in Fig. 5. Cells treated with 6k, 6n and 6o showed a significant reduction of red fluorescence resulting from the Cy3-conjugated secondary antibody against anti p-EGFR and STAT3, respectively. In contrast, only a moderate effect was seen with 6l and erlotinib when compared to the control image.
Immunofluorescence of p-EGFR and STAT3 in MCF-7 cells: MCF-7 cells were seeded on cover glass and treated with 6k, 6l, 6n, 6o and erlotinib at 5 μM at sub confluent level followed by incubation for 24 h. After incubation samples were formalin fixed and subjected to immunofluorescence study using confocal microscopy. Data showing DAPI (blue), p-EGFR and STAT3 (Cy3 red) and merge picture showing cellular distribution of the respective proteins
Effect on proapoptotic and antiapoptotic proteins
Synthesized compounds were further evaluated for the role in regulating proapoptotic and antiapoptotic proteins in MCF-7 cells and facilitates apoptotic death. MCF-7 cells were treated with test conjugates at 5 μM concentration for 24 h followed by total protein lysate extraction. Lysate obtained from the treated cells was subjected to western blotting, and probed with different antibodies against proapoptotic proteins like p53, p21, BAX, procaspase-9 and antiapoptotic protein Bcl-2. The test conjugates significantly increased the levels of p53, p21 and BAX, and decreased that of procaspase-9 as portrayed in Fig. 6. The increase in the levels of many of these proapoptotic proteins also showed a concomitant decrease in the levels of antiapoptotic protein Bcl-2 upon conjugate treatment. Only moderate effects were seen in the erlotinib-treated cells, which served as a positive control.
Western Data of antiapoptotic and proapoptotic proteins in MCF-7 cells: MCF-7 cells were treated with 6k, 6l, 6n, 6o and erlotinib at 5 μM for 24 h and protein lysate obtained was subjected to western blotting. Data showing protein levels of various proapoptotic and antiapoptotic proteins, β-Actin served as gel loading control
EGFR kinase assay
Synthesized compounds were subjected for specific EGFR inhibitory activity, EGFR kinase assay was performed according to the manufacturer protocol employing 25 ng EGFR kinase and conjugates 6k, 6l, 6n, 6o and erlotinib at 5 µM, 2.5 µM, 1.25 µM and 0.625 µM concentration individually. IC50 values were calculated from the % enzyme inhibition at various concentrations of hybrids. 6k, 6l, 6n, 6o and erlotinib showed IC50 values as 0.82 ± 0.21 µM, 0.31 ± 0.01 µM, 0.37 ± 0.01 µM, 0.29 ± 0.04 µM and 0.45 ± 0.01 µM respectively (Table 2).
Effect on wild type EGFR over expressing vector was transfected in to HEK392 cells
To verify activity of these hybrids, the specific, wtEGFR overexpressing plasmid was transfected in to HEK293 cells and 48 h post transfection cells were treated with test conjugates at 5 µM for 24 h. Total protein isolated from the cells was exposed to different conjugates and subjected to western blotting analysis to check the EGFR and STAT3 levels. Conjugate 6k, 6l, 6n, 6o and erlotinib showed significant down regulation when compared with untreated cells harbouring wtEGFR overexpression (Fig. 7). This shows the conjugates specificity is towards EGFR.
Molecular Docking studies
In order to understand the binding mode for the benzimidazole conjugates, molecular docking studies were performed. Docking results showed that all conjugates (6a-q) bind well within the active site of EGFR. All the conjugates had a better docking score than the cocrystal ligand erlotinib (The docking score and binding interactions are given in Table 3). The docking score for erlotinib was 8.14 and the docking score for all conjugates was above 8.75. The conjugates 6e, 6g, 6i and 6l had docking scores above 9. The docking studies suggest that 6g was the most efficient binder, with a docking score of 9.45. It was forming the two hydrogen bonding interactions with the target molecule in that one hydrogen-bonding interaction with Met769 and other hydrogen-bonding interaction with Asp831. Whereas erlotinib was forming one hydrogen-bonding interactions with Met769. The above docking results indicate that synthesized conjugates and erlotinib had similar binding poses. However, synthesized conjugates had a better binding score and extra hydrogen bonding interactions, indicting the superior inhibitory potential of synthesized conjugates.
Binding pose for 6k (Fig. 8b) has shown that the molecule binds well in the ATP binding site. The C-2 phenyl ring was buried in the hydrophobic specificity pocket enclosed by Ala719 (1.87 Å), Ile720 (2.99 Å), Lys721 (2.72 Å), Glu738 (4.23 Å), Leu764 (2.25 Å), Ile765 (2.93 Å) and Thr766 (2.16 Å) amino acids. These interactions in the hydrophobic specificity pocket are important for attaining the specificity among the other kinases. The superimposed pose of 6k with cocrystal ligand erlotinib (Fig. 8d) showed that the C-2 phenyl ring of 6k overlapped with the phenylacetylene group of erlotinib. The C-7 phenyl ring was found oriented towards the hinge region, where 3-methoxy (2.07 Å) and 4-methoxy (3.63 Å) formed hydrogen bonds with the backbone NH of Met769. This hydrogen bond is vital for EGFR inhibitors and is present in all the drug molecules and ATP as well [52, 53]. However, the C-5 benzimidazole ring is oriented towards the A-loop. The NH of benzimidazole formed a hydrogen bond with the side chain of Asp831 (2.18 Å) of A-loop.
The conjugates with halogen-substituted at C-5 of benzimidazole showed slightly enhanced activity over the methyl and dimethyl substituents. The additional polar interactions of the halogens on the benzimidazole ring with the NH of Arg817 (3.60 Å) could perhaps be attributed for the slight increase in activity of these conjugates. Comparison of activity of C-7 substituents, it was quite evident that the conjugates with methoxy substituents (6a-o) have superior activity over the molecules with 3,4-difluoro (6p) and 4-chloro (6q). From the binding pose of 6k, it was obvious that these molecules (6p, 6q) were devoid of hydrogen bond donor to form hydrogen bonding with the Met769. Hence, these molecules exhibited a completely different binding pose as compared to the methoxy-substituted conjugates. While for 6p, the C-7 substitution diverted towards the A-loop and the C-5 benzimidazole substitution towards the hinge region. N4 of pyrazolo[1,5-a]pyrimidine formed a weak hydrogen bond with Met769 (Fig. 8f 2.26 Å). Moreover, molecules with only 4-methoxy substitution on the C-7-phenyl (6a-e) have slightly different binding pose than the 3,4-dimethoxy and 3,4,5-trimethoxy substituted molecules (Fig. 8e). Whereas only 4-methoxy group forms a hydrogen bond with the Met769 (2.41 Å). In 6a, both hydrogen bonds have longer bond distance than the 6k. This could be reason for higher activity of 4-methoxy substituted conjugates than 3,4-dimethoxy and 3,4,5-trimethoxy substituted conjugates. The aforementioned detailed docking results complement the cytotoxic activity.
Conclusions
A library of benzimidazole-linked pyrazolo[1,5-a]pyrimidines was synthesized and evaluated for its cytotoxic activity. These conjugates exhibited significant anticancer activity against a panel of cell lines such as MCF-7, A549 and HeLa with IC50 values ranging from 3.2–41.1 μM and were found less cytotoxic to normal lung fibroblasts MRC5 cells. Structure–activity relationship for the conjugates was also elucidated. Some of the active conjugates (6k, 6l, 6n and 6o) showed significant cytotoxicity against MCF-7 cell lines. Cell cycle analysis indicated a significant accumulation of cells at a sub-G1 phase when treated with 6k, 6l, 6n and 6o at 5 μM for 24 h. Morphological studies revealed an altered structural integrity of cells like detachment from the substratum and membrane blebbing and gave a hint of apoptotic cell death. Western blot and immunostaining analyses suggested the downregulation of EGFR, p-EGFR, STAT3, and p-STAT3, implying that these conjugates strongly modulate the EGFR/STAT3 axis which is important in cancer cell survival and proliferation. Western blot analysis also showed the up-regulation of proteins like p53, p21 and BAX; all well-known tumor suppressor and proapoptotic proteins which facilitate cancer cell death via apoptosis. The down-regulation of a very important antiapoptotic protein, Bcl-2, also correlates the facilitated apoptotic cell death by these conjugates. Hybrids 6k, 6n and 6o significantly reduced the procaspase-9 compared to untreated, which is the effector's caspase in apoptosis that leaves the signature of facilitated apoptosis by these conjugates. Moreover, EGFR kinase assay and wtEGFR transfection study showed the EGFR inhibition. Molecular docking studies were able to rationalize the structure activity relationship and the binding mode of these molecules at the EGFR-binding site. Our investigations suggest that these newer conjugates may have a potential to be developed as chemotherapeutic agents to treat cancer.
References
Schlessinger J (2000) Cell signaling by receptor tyrosine kinases. Cell 103:211–225. https://doi.org/10.1016/S0092-8674(00)00114-8
Yarden Y, Sliwkowski MX (2001) Untangling the ErbB signalling network. Nat Rev Mol Cell Biol 2:127–137. https://doi.org/10.1038/35052073
Schlessinger J (2004) Common and distinct elements in cellular signaling via EGF and FGF receptors. Science 306:1506–1507. https://doi.org/10.1126/science.1105396
Thiel KW, Carpenter G (2007) Epidermal growth factor receptor juxtamembrane region regulates allosteric tyrosine kinase activation. Proc Natl Acad Sci 104:19238–19243. https://doi.org/10.1073/pnas.0703854104
Jura N, Endres NF, Engel K et al (2009) Mechanism for activation of the EGF receptor catalytic domain by the juxtamembrane segment. Cell 137:1293–1307. https://doi.org/10.1016/j.cell.2009.04.025
Holbro T, Hynes NE (2004) ErbB receptors: directing key signaling networks throughout life. Annu Rev Pharmacol Toxicol 44:195–217. https://doi.org/10.1146/annurev.pharmtox.44.101802.121440
Speake G, Holloway B, Costello G (2005) Recent developments related to the EGFR as a target for cancer chemotherapy. Curr Opin Pharmacol 5:343–349. https://doi.org/10.1016/j.coph.2005.02.007
Olayioye MA, Neve RM, Lane HA, Hynes NE (2000) The ErbB signaling network: receptor heterodimerization in development and cancer. EMBO J 19:3159–3167. https://doi.org/10.1093/emboj/19.13.3159
Temam S, Kawaguchi H, El-Naggar AK et al (2007) Epidermal growth factor receptor copy number alterations correlate with poor clinical outcome in patients with head and neck squamous cancer. J Clin Oncol 25:2164–2170. https://doi.org/10.1200/JCO.2006.06.6605
Tanaka E, Hashimoto Y, Ito T et al (2005) The clinical significance of Aurora-A/STK15/BTAK expression in human esophageal squamous cell carcinoma. Clin Cancer Res 11:1827–1834. https://doi.org/10.1158/1078-0432.CCR-04-1627
Kurai M, Shiozawa T, Shih H-C et al (2005) Expression of Aurora kinases A and B in normal, hyperplastic, and malignant human endometrium: Aurora B as a predictor for poor prognosis in endometrial carcinoma. Hum Pathol 36:1281–1288. https://doi.org/10.1016/j.humpath.2005.09.014
Rimawi MF, Shetty PB, Weiss HL et al (2010) Epidermal growth factor receptor expression in breast cancer association with biologic phenotype and clinical outcomes. Cancer 116:1234–1242. https://doi.org/10.1002/cncr.24816
Salomon DS, Brandt R, Ciardiello F, Normanno N (1995) Epidermal growth factor-related peptides and their receptors in human malignancies. Crit Rev Oncol/Hematol 19:183–232. https://doi.org/10.1016/1040-8428(94)00144-I
Jemal A, Siegel R, Ward E, Murray T, Xu J, Thun MJ (2007) Cancer statistics, 2007. Ca-Cancer J Clin 57:43–66. https://doi.org/10.3322/canjclin.57.1.43
Sharma SV, Bell DW, Settleman J, Haber DA (2007) Epidermal growth factor receptor mutations in lung cancer. Nat Rev Cancer 7:169–181. https://doi.org/10.1038/nrc2088
Molina JR, Yang P, Cassivi SD, Schild SE, Adjei AA (2008) Non-small cell lung cancer: epidemiology, risk factors, treatment, and survivorship. Mayo Clin Proc 83:584–594. https://doi.org/10.4065/83.5.584
Wong TW, Lee FY, Yu C et al (2006) Preclinical antitumor activity of BMS-599626, a pan-HER kinase inhibitor that inhibits HER1/HER2 homodimer and heterodimer signaling. Clin Cancer Res 12:6186–6193. https://doi.org/10.1158/1078-0432.CCR-06-0642
Traxler P, Allegrini PR, Brandt R et al (2004) AEE788 A dual family epidermal growth factor receptor/ErbB2 and vascular endothelial growth factor receptor tyrosine kinase inhibitor with antitumor and antiangiogenic activity. Cancer Res 64:4931–4941. https://doi.org/10.1158/0008-5472.CAN-03-3681
Kris MG, Natale RB, Herbst RS et al (2003) Efficacy of gefitinib, an inhibitor of the epidermal growth factor receptor tyrosine kinase, in symptomatic patients with non–small cell lung cancer: a randomized trial. JAMA 290:2149–2158. https://doi.org/10.1001/jama.290.16.2149
Dowell J, Minna JD, Kirkpatrick P (2005) Erlotinib hydrochloride. Nat Rev Drug Discov 4:13–14. https://doi.org/10.1038/nrd1612
Vasquez RJ, Howell B, Yvon A, Wadsworth P, Cassimeris L (1997) Nanomolar concentrations of nocodazole alter microtubule dynamic instability in vivo and in vitro. Mol Biol Cell 8:973–985. https://doi.org/10.1091/mbc.8.6.973
Heathcote DA, Patel H, Kroll SH et al (2010) A novel pyrazolo [1, 5-a] pyrimidine is a potent inhibitor of cyclin-dependent protein kinases 1, 2, and 9, which demonstrates antitumor effects in human tumor xenografts following oral administration. J Med Chem 53:8508–8522. https://doi.org/10.1016/j.bmcl.2020.127432
Gao M, Duan L, Luo J et al (2013) Discovery and optimization of 3-(2-(Pyrazolo [1, 5-a] pyrimidin-6-yl) ethynyl) benzamides as novel selective and orally bioavailable discoidin domain receptor 1 (DDR1) inhibitors. J Med Chem 56:3281–3295. https://doi.org/10.1021/jm301824k
Selleri S, Bruni F, Costagli C et al (2005) A novel selective GABAA α1 receptor agonist displaying sedative and anxiolytic-like properties in rodents. J Med Chem 48:6756–6760. https://doi.org/10.1021/jm058002n
Popik P, Kostakis E, Krawczyk M et al (2006) The anxioselective agent 7-(2-chloropyridin-4-yl) pyrazolo-[1, 5-a]-pyrimidin-3-yl](pyridin-2-yl) methanone (DOV 51892) is more efficacious than diazepam at enhancing GABA-gated currents at α1 subunit-containing GABAA receptors. J Pharmacol Exp Ther 319:1244–1252. https://doi.org/10.1124/jpet.106.107201
Kamal A, Tamboli JR, Nayak VL, Adil S, Vishnuvardhan M, Ramakrishna S (2013) Synthesis of pyrazolo [1, 5-a] pyrimidine linked aminobenzothiazole conjugates as potential anticancer agents. Bioorg Med Chem Lett 23:3208–3215. https://doi.org/10.1016/j.bmcl.2013.03.129
McClue SJ, Blake D, Clarke R et al (2002) In vitro and in vivo antitumor properties of the cyclin dependent kinase inhibitor CYC202 (R-roscovitine). Int J Cancer 102:463–468. https://doi.org/10.1002/ijc.10738
Kamal A, Tamboli JR, Ramaiah MJ et al (2012) Anthranilamide-Pyrazolo [1, 5-a] pyrimidine Conjugates as p53 Activators in Cervical Cancer Cells. ChemMedChem 7:1453–1464. https://doi.org/10.1002/cmdc.201200205
Bagul C, Rao GK, Makani VKK, Tamboli JR, Pal-Bhadra M, Kamal A (2017) Synthesis and biological evaluation of chalcone-linked pyrazolo [1, 5-a] pyrimidines as potential anticancer agents. MedChemComm 8:1810–1816. https://doi.org/10.1039/C7MD00193B
Mullagiri K, Nayak VL, Sunkari S et al (2018) New (3-(1 H-benzo [d] imidazol-2-yl))/(3-(3 H-imidazo [4, 5-b] pyridin-2-yl))-(1 H-indol-5-yl)(3, 4, 5-trimethoxyphenyl) methanone conjugates as tubulin polymerization inhibitors. MedChemComm 9:275–281. https://doi.org/10.1039/C7MD00450H
Donthiboina K, Anchi P, Gurram S et al (2020) Synthesis and biological evaluation of substituted N-(2-(1H-benzo [d] imidazol-2-yl) phenyl) cinnamides as tubulin polymerization inhibitors. Bioorg Chem 103:104191. https://doi.org/10.1016/j.bioorg.2020.104191
Mani GS, Anchi P, Sunkari S et al (2020) Synthesis of (Z)-3-(arylamino)-1-(3-phenylimidazo [1, 5-a] pyridin-1-yl) prop-2-en-1-ones as potential cytotoxic agents. Bioorg Med Chem Lett 30:127432. https://doi.org/10.1016/j.bmcl.2020.127432
Ramya PS, Angapelly S, Rani RS et al (2020) Hypervalent iodine (III) catalyzed rapid and efficient access to benzimidazoles, benzothiazoles and quinoxalines: Biological evaluation of some new benzimidazole-imidazo [1, 2-a] pyridine conjugates. Arab J Chem 13:120–133. https://doi.org/10.1016/j.arabjc.2017.02.007
Korrapati SB, Yedla P, Pillai GG et al (2021) In-silico driven design and development of spirobenzimidazo-quinazolines as potential DNA gyrase inhibitors. Biomed Pharmacother 134:111132. https://doi.org/10.1016/j.biopha.2020.111132
Kamal A, Reddy KS, Ahmed SK et al (2006) Anti-tubercular agents. Part 3. Benzothiadiazine as a novel scaffold for anti-Mycobacterium activity. Bioorg Med Chem 14:650–658. https://doi.org/10.1016/j.bmc.2005.08.063
Pancholia S, Dhameliya TM, Shah P et al (2016) Benzo[d]thiazol-2-yl(piperazin-1-yl) methanones as new anti-mycobacterial chemotypes: Design, synthesis, biological evaluation and 3D-QSAR studies. Eur J Med Chem 116:187–199. https://doi.org/10.1016/j.ejmech.2016.03.060
Mahesh R, Nayak VL, Babu KS et al (2017) Design, Synthesis, and in vitro and in vivo Evaluations of (Z)-3, 4, 5-Trimethoxystyrylbenzenesulfonamides/sulfonates as Highly Potent Tubulin Polymerization Inhibitors. ChemMedChem 12:678–700. https://doi.org/10.1002/cmdc.201600643
Bhagat S, Supriya M, Pathak S, Sriram D, Chakraborti AK (2019) α-Sulfonamidophosphonates as new anti-mycobacterial chemotypes: Design, development of synthetic methodology, and biological evaluation. Bioorg Chem 82:246–252. https://doi.org/10.1016/j.bioorg.2018.09.023
Schneider G, Schneider P, Renner S (2006) Scaffold-hopping: how far can you jump? Qsar Comb Sci 25:1162–1171. https://doi.org/10.1002/qsar.200610091
Padmaja P, Rao GK, Indrasena A et al (2015) Synthesis and biological evaluation of novel pyrano [3, 2-c] carbazole derivatives as anti-tumor agents inducing apoptosis via tubulin polymerization inhibition. Org Biomol Chem 13:1404–1414. https://doi.org/10.1039/C4OB02015D
Srinivas C, Ramaiah MJ, Lavanya A et al (2015) Novel etoposide analogue modulates expression of angiogenesis associated micrornas and regulates cell proliferation by targeting STAT3 in breast cancer. PLoS ONE 10:e0142006. https://doi.org/10.1371/journal.pone.0142006
Carter RE, Sorkin A (1998) Endocytosis of functional epidermal growth factor receptor-green fluorescent protein chimera. J Biol Chem 273:35000–35007. https://doi.org/10.1074/jbc.273.52.35000
Stamos J, Sliwkowski MX, Eigenbrot C (2002) Structure of the epidermal growth factor receptor kinase domain alone and in complex with a 4-anilinoquinazoline inhibitor. J Biol Chem 277:46265–46272. https://doi.org/10.1074/jbc.M207135200
Madhavi Sastry G, Adzhigirey M, Day T, Annabhimoju R, Sherman W (2013) Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput Aided Mol Des 27:221–234. https://doi.org/10.1007/s10822-013-9644-8
Frisch MJ, Trucks GW, Schlegel HB, et al. Gaussian 09. Wallingford, CT; 2016.
Morris GM, Goodsell DS, Halliday RS et al (1998) Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J Comput Chem 19:1639–1662. https://doi.org/10.1002/(SICI)1096-987X(19981115)19:14
he PyMOL Molecular Graphics System, Version 1.2r3pre, Schrödinger, LLC.
Shrivastava N, Naim MJ, Alam MJ, Nawaz F, Ahmed S, Alam O (2017) Benzimidazole scaffold as anticancer agent: synthetic approaches and structure–activity relationship. Arch Pharm 350:e201700040. https://doi.org/10.1002/ardp.201700040
Kamal A, Shaik AB, Polepalli S et al (2015) Synthesis of arylpyrazole linked benzimidazole conjugates as potential microtubule disruptors. Bioorg Med Chem 23:1082–1095. https://doi.org/10.1016/j.bmc.2015.01.004
Sharma P, Reddy TS, Thummuri D et al (2016) Synthesis and biological evaluation of new benzimidazole-thiazolidinedione hybrids as potential cytotoxic and apoptosis inducing agents. Eur J Med Chem 124:608–621. https://doi.org/10.1016/j.ejmech.2016.08.029
Padhy GK, Panda J, Behera AK (2019) Synthesis and Characterization of Novel N-Benzylbenzimidazole Linked Pyrimidine Derivatives as Anticancer Agents. Indian J Pharm Educ Res 53:S129–S134. https://doi.org/10.5530/ijper.53.2s.57
Johnson LN (2009) Protein kinase inhibitors: contributions from structure to clinical compounds. Q Rev Biophys 42:1–40. https://doi.org/10.1017/S0033583508004745
Warnault P, Yasri A, Coisy-Quivy M et al (2013) Recent advances in drug design of epidermal growth factor receptor inhibitors. Curr Med Chem 20:2043–2067. https://doi.org/10.2174/0929867311320160001
Acknowledgements
The author C.B acknowledges the Department of Pharmaceuticals (Ministry of Chemicals and Fertilizers, Govt. of India) for the financial support & the CSIR-IICT, Hyderabad for providing facilities. K.R.G thanks UGC for his fellowship. This work was financially supported by the CSIR 12th FYP CSC0111 (SMiLE). The authors thank Y. Suresh for the Flow Cytometry and Confocal Microscopy studies.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bagul, C., Rao, G.K., Veena, I. et al. Benzimidazole-linked pyrazolo[1,5-a]pyrimidine conjugates: synthesis and detail evaluation as potential anticancer agents. Mol Divers 27, 1185–1202 (2023). https://doi.org/10.1007/s11030-022-10481-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-022-10481-x