Abstract
Throughout the intestinal epithelium surface there is an intricate polymer network composed by gel-forming mucins, which plays a protective role due to the formation of a physical, chemical and immunological barrier between the organism and the environment. Mucin 2 (MUC2) is the main mucin in the small and large intestine, and it is expressed specifically in the gastrointestinal tract (GIT), which makes its promoter region an important candidate for expression of heterologous genes of biotechnological interest in the GIT of bovine and other ruminants. In order to characterize the bovine MUC2 promoter we designed primers to amplify and isolate a candidate region for this promoter. The amplified sequence was confirmed by sequencing and cloned into a plasmid vector containing the luciferase (LUC) reporter gene. The regulatory sites of the MUC2 promoter already described in the literature were used to find the putative regulatory sites in the bovine MUC2 promoter region. With these data, some deletions were performed in order to find the promoter sequence with greatest expression capacity and specificity. The constructions were tested by transient transfection assays in LoVo cells (human colorectal adenocarcinoma) and bovine fibroblasts. The quantification of the relative expression of the promoter was measured using dual-luciferase assays. Real-time PCR was performed to analyze the expression of endogenous MUC2. The results presented herein prove that the isolated sequence corresponds to the promoter of bovine MUC2 gene, since it was able to induce expression of a reporter gene in an in vitro cell culture experimental platform.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mucins are ubiquitous glycoproteins that cover the ocular surface, the epithelial surface of the respiratory, gastrointestinal, urinary and genital tracts [1, 2]. They play an important role in organism protection since they cover all surfaces with a gel-mucus, forming a physical, chemical and immunological barrier between the organism and the surrounding environment [3]. The gel-forming mucin family includes five genes in mammals: MUC2, MUC5AC, MUC5B, MUC6 and MUC19 that are clustered in human chromosome 11 locus 11p15.5 and in mice chromosome 7 band F5, except MUC 19, which is located in human chromosome 12 locus 12p12 and mouse chromosome 15 band E5 [2, 4,5,6,7]. According to the Bos taurus sequence deposited in GenBank (ARS-UCD1.2), the bovine MUC2 gene is located on chromosome 29.
The MUC2 gene codifies a mucin that is expressed specifically in the intestinal epithelium [8]. It is an o-glycosylated protein of high molecular mass that forms a network of intricate gel-like polymers, responsible for lubrication and protection of the external surface of the internal epithelium tissue [9, 10]. The Muc2 protein is the main constituent of the mucus in the small and large intestines [11], although there is a considerable amount of Muc5AC and Muc6 proteins in the large intestine [12]. These mucins have a protein domain composed of amino acids with repetitive sequences, rich in serine and threonine, where the o-ligated glycosylation occurs. The o-glycans constitute about 80% of the molecular weight of these proteins [13] and are responsible for the generation of aggregates of mucin polymers and the maintenance of the gel structure of these glycoprotein complexes [14]. The Muc2 protein appears to be an important barrier against pathogens [13] and plays many functions in the intestinal homeostasis [15], since its absence leads to an increase in colitis [16] and the development of intestinal cancer with spontaneous progression to metastasis [15]. Furthermore, studies demonstrated that the MUC2 gene is expressed during the beginning of human embryonic development, as soon as nine weeks of gestation [17], which makes it a vital marker to elucidate the mechanisms that regulate the differentiation of secretory cell lines [18].
Regulation of RNA transcription is a critical process for cell growth and differentiation during the development of an organism, and it is vital to keep homeostasis during the organism’s lifetime [19]. Promoters and enhancers are the main regulators of gene expression that integrate information from many signaling pathways, through binding of activators and repressors, named transcription factors (TFs) [19]. The regulation of transcription has been extensively studied in eukaryotes, leading to the identification of conserved DNA sequences that are responsible for the initiation of mRNA synthesis by RNA polymerase II [20]. At least two conserved ubiquitous types of regulatory elements exist in the core promoter, the TATA box and the Inr sequence. The TATA box plays an important role in the identification of transcription initiation site and recruitment of RNA Pol II transcription complex [20]. Promoters that do not have TATA box and have the Inr sequence are less abundant, and this sequence apparently plays the same role as the TATA box. There are also promoters that have both elements in their core region and others that do not have either, but the latter case is less frequent [21]. Therefore, promoters are important regions for gene expression since they are responsible for controlling the beginning of transcription and its intensity [22]. The human and mouse MUC2 promoters are conserved and both present the TATA box as a regulatory element [8, 18].
Characterization of promoter regions is also fundamental in order to express a gene of interest in a specific tissue and in a controlled fashion [23]. The use of transgenic cattle as bioreactors to produce heterologous proteins has the mammary gland as a target tissue, since milk proteins are secreted, which facilitates protein purification [24]. Expression of heterologous proteins in the milk of transgenic animals is induced by promoters of specific milk protein genes, such as α-lactalbumin, β-lactoglobulin and the caseins α, β, γ and κ [24]. Several heterologous proteins have already been expressed in the milk of transgenic cattle, such as human lysozyme, lysine-rich polypeptide, human lactoferrin, human β-defensin-3 and bile salt-stimulated lipase. Commercially available biopharmaceuticals for treating human disease, such as antithrombin (LFB Biotechnologies) and C1-esterase inhibitor (Pharming Group) have also been expressed [24,25,26,27,28,29].
Therefore, the identification and characterization of a promoter that can effectively direct the expression of a gene of interest, in a specific manner, in the gastrointestinal tract is very important for the expression of heterologous proteins of biotechnological importance. Furthermore, the characterization of specific intestinal promoters can allow the understanding of the important elements and factors involved in the regulation of intestinal expressed genes, providing a better understanding of the mechanisms responsible for the differentiation patterns of the intestinal epithelium [8]. In addition, a relevant application of this kind of promoter region is the expression of genes that can enhance animal nutrition and increase feed utilization or express genes that could increase disease resistance. By enhancing animal nutrition it is possible to improve the quantity, quality and nutritional composition of livestock [30]. Infections by endoparasites weaken cattle health and lead to economic losses, so the expression of genes that could increase resistance is an interesting approach to this problem [31]. Increasing the efficiency and productivity of livestock has the potential to decrease the environmental footprint generated by animal production [30]. MUC2 is expressed abundantly and specifically in the gastrointestinal tract of mammals, so its promoter becomes an interesting candidate to induce the expression of genes of interest in the gastrointestinal tract. The aim of this study is to characterize the bovine mucin 2 promoter 5′ region.
Materials and methods
All reagents were purchased from Sigma-Aldrich, unless specified.
Cell culture
LoVo cells (human colorectal adenocarcinoma cell line) were obtained from the Cell Bank of Rio de Janeiro (BCRJ) and cultured in F12-K medium with 10% of bovine fetal serum (Invitrogen), 100ui/mL of penicillin and 50 μg/mL of streptomycin. Bovine fibroblasts were harvest from bovine skin and were cultured in D-MEM (Dulbecco′s Modified Eagle′s Medium, Invitrogen) supplemented with 3.7 g/L of sodium bicarbonate, 290 mg/L of l-glutamine 2 mM, 110 mg/L of sodium pyruvate, 10% of bovine fetal serum, 100 μi/mL of penicillin and 50 μg/mL of streptomycin. All cells were maintained in a humidified incubator (Thermo) at 37 °C and 5% carbon dioxide.
MUC2 real-time PCR (qPCR)
Cells were harvested from a cell culture flask with Trypsin–EDTA 0.25% (Invitrogen). TRIzol reagent (Invitrogen) was used to isolate total RNA according to the manufacturer’s protocol. cDNA synthesis was performed using a SuperScript III kit (Invitrogen) according to the manufacturer’s protocol. qPCR was performed using the 7500 Fast Real-time PCR System (Applied Biosystems). β-actin and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) were used as internal housekeeping genes. Each real-time PCR assay contained 1.0 μL cDNA, 12.5 μL of SYBR green reagent (Fermentas), and 0.5 μL of each primer (0.2 μM) in a 25 μL reaction mixture. The PCR reactions were conducted as follows: an initial denaturing step of 10 min at 95 °C, followed by 40 × cycles of 3 s at 95 °C and 30 s at 60 °C, and finally the melt curve stage of 15 s at 95 °C, 1 min at 65 °C, 15 s at 95 °C and 15 s at 60 °C. For each gene analyzed, four technical replicates were used. Since MUC2 is only expressed in LoVo cells, it was not possible to carry out comparative analyses, so expression was analyzed by the melt curve and the fragment visualization through electrophoresis in 2.0% agarose gel. The sequence of the primers used in this study is listed in the supplementary material.
Plasmid construction
Primers (5′-GTACTAGTCAAGGCTGTGGAGGGCTTTAAT and 5′-ATGCGGCCGCGGTGGCCAGAAGGAAGGC) were designed to amplify a region of 3478 bp upstream of the bovine MUC2 gene. Genomic DNA from Bos indicus was used. The PCR was performed using the Kapa HiFi HotStart kit (KAPA Biosystems), and each PCR assay contained 80 ηg of gDNA, 5.0 μL of 5X buffer, 10 mM of KAPA dNTP mix, 0.5 U of KAPA DNA polymerase and 0.3 μM of each primer in a 25 μL reaction mixture. The PCR reactions were conducted as follows: an initial denaturing step of 5 min at 95 °C, followed by 35 × cycles of 20 s at 98 °C, 15 s at 65 °C and 3 min and 20 s at 72 °C, and a final extension of 1 min at 72 °C. To clone the PCR amplicon, a modified pFASTBacI plasmid was used. This plasmid was generously provided by Dr. Bergmann Ribeiro from the University of Brasilia, where the gene of firefly luciferase (LUC) was extracted from the pGEM-LUC (Promega) vector and inserted into the pFASTBacI vector. Smaller fragments (promoter deletions) were amplified using internal primers (supplementary material) from the gDNA or the 3478 bp amplicon, resulting in fragments of 1454 bp, 1035 bp and 101 bp. A larger fragment of 4217 bp was also amplified (primers in supplementary material) to compare with the 3478 bp amplicon for the LUC activity. These constructions were named pMUC2-LUC + size of the fragment. The negative control of the experiments was the promoter sequence cloned into the inverse orientation in the plasmid (pMUC2-LUC CCW), and the positive control was the plasmid with the human EF1α promoter controlling the LUC expression (pEF-LUC). The ligations between the plasmid and the fragments, using T4 DNA Ligase (Promega), were transformed into competent bacteria according to Inoue et al. [32], and the plasmid DNA was extracted using the QIAprep Spin Miniprep Kit (Qiagen) according to the manufacturer’s protocol. All constructions were confirmed by DNA sequencing. The cloned bovine MUC2 promoter sequence is presented in the supplementary material.
Transient transfection assay and dual-luciferase assay
Transfection assays were performed using Lipofectamine LTX (Invitrogen) for fibroblast and Lipofectamine 2000 (Invitrogen) for LoVo cells, according to the manufacturer’s instructions. For the dual-luciferase assay, a second type of luciferase is needed to normalize the reaction. Renilla luciferase (RNL) was used under the control of the constitutive cytomegalovirus promoter (pRL-CMV). For each plasmid, 3 wells were used as biological replicates. Cells were plated in a 24-well plate and incubated for 24 h to achieve 60–80% of confluence. Transfections were performed using 450 ηg of pMUC-LUC plus 50 ηg of pRL-CMV with 1.5 μL of the transfection reagent in a total volume of 200 μL of serum-free medium. Following 5 h of incubation at 37 °C, the medium was removed from the wells, and 500 μL of standard growth medium was added and the plate incubated for a further 48 h. Cells were harvested following the dual-luciferase kit (Promega) protocol, and the luciferase activity was measured using a GloMax 96 Microplate Luminometer (Promega). LUC activity was normalized by RNL activity. The data were analyzed for normality using the Shapiro–Wilk test. After confirming normality, t test and Anova were used to confirm the statistical significant differences.
Methylation analysis
To investigate methylation modulation of MUC2 promoter on fibroblast cells, a promoter region of 266 bp containing 12 CpG sites was analyzed. The endogenous promoter and the promoter in the pMUC plasmid were evaluated. A transfection assay was performed with fibroblast cells, using 500 ηg of pMUC2-LUC_3478pb. At 48 h after the transfection, the cells were harvested, and the genomic DNA and the plasmid DNA were extracted. The gDNA was treated with sodium bisulfite, using the EZ DNA methylation-lightning kit (Zymo Research), according to the manufacturer’s protocol. PCR was performed to amplify a fragment of 323 bp from the genomic DNA and 387 bp from the plasmid, in which 266 bp were from the MUC2 promoter sequence. The primers were designed using the Bisulfite Primer Seeker tool (https://www.zymoresearch.com/pages/bisulfite-primer-seeker); thus, the forward primer was positioned in the promoter region and the reverse primer for gDNA was located in the MUC2 gene and in the LUC gene for the plasmid, respectively. The fragments were cloned into a pGEM-Teasy (Promega) and transformed into competent bacteria according to Inoue et al. [32]. Ten colonies of each ligation were chosen and the plasmid extracted using the QIAprep Spin Miniprep Kit (Qiagen), according to the manufacturer’s protocol. The 20 clones (gDNA and vector) were sequenced by Sanger sequencing. The methylation pattern was analyzed using QUMA software [33].
In silico analysis of the promoter sequence
Using the online software GPMiner (http://gpminer.mbc.nctu.edu.tw/index.php) and the data from published literature about TF and its target sites, that regulate the human and the mouse MUC2 promoters, it was possible to predict the TF sites on the bovine sequence that could be a potential site for transcription regulation. Using this information, a scheme of the bovine MUC2 promoter sequence with the transcription factor sites was generated using the software SnapGene v5.0.7 (GSL Biotech LLC).
Sequence alignment between mouse, human and bovine MUC2 promoters
Alignment between the promoter sequences of human, mouse and bovine MUC2 gene were performed using the software Geneious v2020.0.5 (Biomatters, New Zealand). The described sequence for the human promoter covers a fragment of 3450 bp [8] and 3582 bp for mouse [18].
Results and discussion
Endogenous expression of bovine MUC2 gene in fibroblasts and LoVo cells
Figure 1 shows the expression of MUC2 in LoVo cells and no expression in bovine fibroblasts, as expected. The endogenous positive controls GAPDH (176 bp) and β-actin (130 bp) can be visualized in both cell types, as expected. In addition, the melt curve confirmed the specificity of the fragment (supplementary material). These data confirm the specificity of MUC2 expression in LoVo cells (intestine origin) and no expression in fibroblast cells.
Transient expression of LUC reporter gene under control of MUC2 sequences
The diagram of all MUC2 promoter constructions is presented in Fig. 2. LoVo and bovine fibroblasts cells were transfected with pEF-LUC, pMUC2-LUC CCW, pMUC2-LUC_3478 and pMUC2-LUC_1454, and the normalized LUC relative expression is presented in Fig. 3. It is possible to observe that in both cell lines the highest LUC expression occurs with the promoter fragment of 3478 bp. When using a cell line that expresses MUC2 (LoVo), the promoter capability in expressing LUC is superior to a cell line that does not express MUC2 (fibroblasts), p < 0.05. The pEF-LUC expression showed the intrinsic capacity of the cells in response to a strong constitutive promoter with a higher LUC expression in the fibroblast than in LoVo, indicating that the gene expression potential of fibroblasts is stronger than LoVo cells, and the MUC2 promoter fragment showed specificity for LoVo cells, expressing more LUC than fibroblasts. Since the promoter regulation might not be a binary event [34], this difference in expression observed in LoVo and fibroblasts might indicate a considerable specificity of the bovine promoter isolated in this study. Moreover, the epigenetic control of the promoter activity, such as DNA methylation and histone acetylation and methylation, cannot be working properly in the MUC2 promoter sequences cloned in the reporter LUC vectors. This may be due to the lack of time for the epigenetic marks to be established during the transient transfection assays, or the lack of some genomic DNA context (enhancer sequences) that is not present in the MUC2 fragments cloned [35,36,37,38,39]. Furthermore, the human MUC2 promoter modulation through methylation occurs during cell differentiation [40]. These possibilities may explain the LUC expression leak observed in fibroblast cells.
Results of LUC expression in transfection assays in LoVo and bovine fibroblast cells a pMUC2-LUC CCW (C-), pMUC2-LUC_3478 (∆3478) and pMUC2-LUC_1454 (∆1454) b pEF-LUC. Standard deviation of tested samples: C- ± 6,15−5, LoVo ∆3478 ± 0,003, fibroblast ∆3478 ± 0,0002, Lovo ∆1454 ± 0,0009, fibroblast ∆1454 ± 0,0003, LoVo pEF-LUC ± 0,028 and fibroblast pEF-LUC ± 0,017
LUC expression in fibroblast does not invalidate the specificity of the cloned MUC2 promoter, because in vitro cell culture behaves differently from in vivo cells. There is a different environment with different factors and elements that control LUC expression, which may be happening in in vitro cell culture, but not in vivo [41]. The experiments of Woodfint et al. [23] with the transgenic quail expressing GFP under the control of a 2.9 kb chicken MUC2 promoter region clearly demonstrated the specificity of this promoter, since there was GFP expression only in the gastrointestinal tract. However, this result cannot exclude the possibility that the MUC2 promoter might be modulated by factors located outside its core region, or even modulators such as enhancers and silencers that could be present far from the proximal promoter regions, near the MUC2 open reading frame (ORF). Studies have shown the influence and modulation of the human MUC2 promoter by epigenetic markers like DNA methylation in CpG islands [40, 42]. The bovine MUC2 promoter sequence isolated in our study is rich in CpG sites, suggesting that it may also have been modulated by DNA methylation.
In order to refine the core MUC2 promoter sequence capable of modulating the LUC gene, three deletions were made to test if a smaller portion of the promoter could induce a similar expression level of LUC. Besides, a larger fragment of 4217 bp was also tested, but the 3478 bp showed the highest LUC activity. Fragments with 4217 bp, 3478 bp, 1454 bp, 1035 bp and 101 bp were tested by transfection in LoVo cells followed by LUC/RNL measurement (Fig. 4).
Transient LUC expression using the plasmids pMUC2-LUC CCW (C-), pMUC2-LUC_4217 (∆4217), pMUC2-LUC_3478 (∆3478), pMUC2-LUC_1454 (∆1454), pMUC2-LUC_1035 (∆1035) and pMUC2-LUC_101 (∆101) in transfection assays with LoVo cells. * p < 0.0001 when compared to ∆3478 deletion. Standard deviation of tested samples: C- ± 5,47−5, ∆4217 ± 0,0008, ∆3478 ± 0,003, ∆1454 ± 0,0009, ∆1035 ± 0,0013 and ∆101 ± 1,41 × 10−6
When this study started the bovine genome assembly available on GenBank (UMD 3.1.1) did not have the entire MUC2 promoter sequence. The bovine MUC2 gene was at the extremity of the published contig (UMD 3.1.1 version). Since this gene is in the antisense direction (from telomere to centromere), only the fragment of 1454 bp was available downstream of the MUC2 ORF sequence. Therefore, the study started with the fragment of 1454 bp and the deletions (1035 and 101 bp) were cloned based on this fragment. After doing most of the experiments, a new bovine genome assembly (ARS-UCD 1.2) was published. This new assembling was more complete and precise, allowing the positioning of MUC2 gene on chromosome 29 and giving access to a larger downstream MUC2 sequences. Using this new sequence, larger fragments were amplified (4217 and 3478 bp), cloned in the LUC reporter vector and compared to the previous pMUC2 constructions.
The results showed that the 3478 bp fragment was able to induce the higher expression of LUC, compared to all of the other fragments (p < 0.0001). The fragment of 4217 bp had lower LUC expression than 3478 bp and this may be due to putative negative downstream regulator sequences. The deletions followed a decrease in the reporter gene expression, where the deletion with only 101 bp showed the lowest expression. This result was expected, since the fragment with only 101 bp contains only the TATA box as a regulatory site. The presence of the CACCC box, which is not in the 101 bp fragment, has been demonstrated by Gum et al. [8] to be important in MUC2 promoter efficiency. Other putative transcription factors’ binding sequences were spread alongside the 3478 bp sequence, and its removal in the deletions causes a progressive reduction of the MUC2 promoter capability. These results are expected and corroborated by other equivalent reporter assays using human and mouse MUC2 promoter sequences [8, 18, 43,44,45].
DNA methylation control of the endogenous and reporter MUC2 promoter in fibroblast cells
The endogenous MUC2 promoter from bovine fibroblast cells and the MUC2 promoter of the pMUC2-LUC_3478 vector were analyzed regarding methylation of 12 CpG sites present in the CpG rich 266 bp region upstream of MUC2 transcription start site. A fragment of 323 bp was amplified from the bovine fibroblast gDNA, in which 13 CpG sites were present, one located in the MUC2 gene ORF. For pMUC2-LUC_3478, a fragment of 387 bp was amplified containing 14 CpG sites, in which 2 were present in the LUC ORF. The DNAs treated with bisulfide were amplified, and 10 clones of each amplicon (gDNA and vector) were isolated and sequenced (Fig. 5c). It is clear that the endogenous CpG-rich promoter region is methylated in many sites, and the equivalent region from the transfected plasmid is undermethylated, which can suggest that methylation also plays a role in the control of the bovine endogenous MUC2 promoter. The compiled analysis from all sequenced regions and comparison between the two promoter fragments can be visualized in Fig. 5a, where the overmethylation of CpG sites in the fibroblasts’ endogenous promoter is clear. The endogenous promoter was significantly (p < 0.05) more methylated (61.3% ± 21.6) than the promoter cloned in the reporter plasmid (1.0% ± 2.9) (Fig. 5b).
Endogenous MUC2 promoter of the bovine fibroblast cells (Endogenous) and MUC2 promoter from the pMUC2-LUC_3478 vector (Vector) (a). Comparison between the methylation analysis in the two groups on the 12 CpG sites from the ten clones (each group). The circles represent the CpG sites and the methylated clones are represented by black circles (b). Graphic representing the accumulated methylation values (c). The expanded results of each methylated clone status were shown. The black circles represent a site that is methylated, and white, not methylated
The studies by Yamada et al. [46] showed that only DNA methylation and histone modification were not able to determine MUC2 expression in human cancerous pancreatic cell lines. This was also observed in studies by Vincent et al. [40] that used esophageal, gastric, pancreatic and colon cancerous epithelium cell lines. Therefore, the MUC2 pattern of expression is a result of epigenetic chromatin modifications and the presence of specific transcription factors in the cell types which express MUC2 [40, 46]. This suggests that the same epigenetic mechanisms might direct MUC2 expression in all epithelial cells with different tissue origins [40]. These works on the human MUC2 promoter and our results that showed methylation on the fibroblast promoter suggest that the same mechanisms of modulation may be happening in the bovine promoter. Therefore, the results presented in this study corroborate the MUC2 expression pattern as an outcome of cell specificity (driven by transcription factors), DNA methylation and chromatin modifications. As seen in Fig. 4, the MUC2 promoter of bovine fibroblast cells is under methylation modulation, and the MUC2 gene is not expressed in fibroblasts (Fig. 1). Moreover, when the fibroblasts were transfected with the reporter vector constructions it became possible to see the LUC gene expression, but this expression is lower than the one observed in LoVo cells, which also indicates cell type specificity for the cloned bovine promoter. The inhibitory effect of CpG methylation on the MUC2 endogenous promoter observed in fibroblasts might not have been detected in the reporter assays because of its transient characteristic. In this kind of assay, the LUC activity was analyzed 48 h after transfection, and this period can be insufficient for DNA methyltransferases to act on the promoter region, since the promoter sequence on the transfected plasmid is almost not methylated (Fig. 5). Moreover, the endogenous MUC2 promoter is inserted in a much more complex genome context and it can be regulated by modulator sequences that are kilo or mega-base distant from the MUC2 gene locus.
In silico reconstruction of bovine MUC2 putative regulation sites
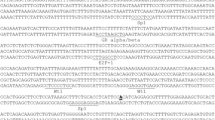
The in silico reconstruction of putative TF binding sites was done (Fig. 6). This reconstruction shows the theoretical locations for consensus sequences of well-established transcription factor binding sites. Due to the scarce literature regarding the characterization of bovine TF binding sites, the sites showed are predictions based on similarities with the human and mouse MUC2 database sequences, which align to the bovine counterpart sequence.
Relevant regulatory sites were found in the bovine sequence for MUC2 promoter, using the characterized sequences from human and mouse, such as TATA box, CACCC box, Sp1 family, CDX domains, AP-1 domains, GATA transcriptional factor and the ligation site of p53. All these similarities between regulatory regions and the similarity observed in the alignment (Fig. 7) corroborate the hypothesis that the sequence isolated in this study corresponds to the promoter sequence of bovine MUC2 gene. The GPMiner software localized sequences that could be a transcriptional factor binding-site, using the human and the mouse database for TFs. Since the literature regarding bovine transcriptional factors is scarce, the presented sites are predictions based on the human and the mouse sequences that have been experimentally confirmed [18, 43,44,45, 47,48,49]. The alignment performed between bovine (3478 bp), human (3450 bp) and mouse (3582 bp) promoters can be visualized in Fig. 7. The similarity between these three sequences is 48.4%, and the similarity between human and mouse promoter is 53.7%, between bovine and mouse promoter 52.4% and between bovine and human promoter 53.3%, which confirms the identity of the isolated bovine sequence as a regulatory region of bovine MUC2 gene.
The work of de Sousa et al. [50] about bovine TFs highlights the scarcity of the literature about DNA regulatory sequences in this species and justifies why they assembled a compendium of possible bovine transcriptional factors derived from the human database. They analyzed family domains, their distribution, evolutionary conservation and specific cell expression patterns. However, experimental evidence is still needed and will be fundamental to validate if the predicted sites are effective and functional TF binding sites, showing that much work still needs to be done in terms of experimental characterization of bovine transcription regulation.
Conclusion
The studies on sequences and regulatory structures of MUC2 provide a better understanding of the precise role of these regulatory sequences in the modulation of expression of the factors involved in intestinal tract physiology [23]. Moreover, a specific promoter of an important target tissue of commercially important livestock has remarkable potential for application in the expression of heterologous proteins of biotechnological interest. The results presented in this research prove that the cloned region of 3478 bp matches the “core promoter" sequence of Bos indicus MUC2 gene, not described previously.
Data availability
The main data generated or analyzed during this study are included in this published article (and its supplementary information files) and the ones not publicly are available from the corresponding author.
References
Corfield AP (2015) Mucins: a biologically relevant glycan barrier in mucosal protection. Biochim Biophys Acta 1850:236–252
Lang T, Klasson S, Larsson E, Johansson ME, Hansson GC, Samuelsson T (2016) Searching the evolutionary origin of epithelial mucus protein components-mucins and FCGBP. Mol Biol Evol 33:1921–1936
Hoorens PR, Rinaldi M, Li RW, Goddeeris B, Claerebout E, Vercruysse J, Geldhof P (2011) Genome wide analysis of the bovine mucin genes and their gastrointestinal transcription profile. BMC Genomics 12:140
Griffiths B, Matthews DJ, West L, Attwood J, Povey S, Swallow DM, Gum JR, Kim YS (1990) Assignment of the polymorphic intestinal mucin gene (MUC2) to chromosome 11p15. Ann Hum Genet 54:277–285
Pigny P, Guyonnet-Duperat V, Hill AS, Pratt WS, Galiegue-Zouitina S, d’Hooge MC, Laine A, Van-Seuningen I, Degand P, Gum JR, Kim YS, Swallow DM, Aubert JP, Porchet N (1996) Human mucin genes assigned to 11p15.5: identification and organization of a cluster of genes. Genomics 38:340–352
Desseyn JL, Laine A (2003) Characterization of mouse muc6 and evidence of conservation of the gel-forming mucin gene cluster between human and mouse. Genomics 81:433–436
Chen Y, Zhao YH, Kalaslavadi TB, Hamati E, Nehrke K, Le AD, Ann DK, Wu R (2004) Genome-wide search and identification of a novel gel-forming mucin MUC19/Muc19 in glandular tissues. Am J Respir Cell Mol Biol 30:155–165
Gum JR, Hicks JW, Kim YS (1997) Identification and characterization of the MUC2 (human intestinal mucin) gene 5’-flanking region: promoter activity in cultured cells. Biochem J 325(Pt 1):259–267
Gum JR Jr, Hicks JW, Toribara NW, Siddiki B, Kim YS (1994) Molecular cloning of human intestinal mucin (MUC2) cDNA. Identification of the amino terminus and overall sequence similarity to prepro-von Willebrand factor. J Biol Chem 269:2440–2446
Bu XD, Li N, Tian XQ, Huang PL (2011) Caco-2 and LS174T cell lines provide different models for studying mucin expression in colon cancer. Tissue Cell 43:201–206
Ambort D, Johansson ME, Gustafsson JK, Nilsson HE, Ermund A, Johansson BR, Koeck PJ, Hebert H, Hansson GC (2012) Calcium and pH-dependent packing and release of the gel-forming MUC2 mucin. Proc Natl Acad Sci USA 109:5645–5650
Jiang Z, Applegate TJ, Lossie AC (2013) Cloning, annotation and developmental expression of the chicken intestinal MUC2 gene. PLoS ONE 8:e53781
Ma J, Rubin BK, Voynow JA (2017) Mucins, mucus, and goblet cells. Chest S0012–3692(17):33080–33085
Johansson ME, Ambort D, Pelaseyed T, Schutte A, Gustafsson JK, Ermund A, Subramani DB, Holmén-Larsson JM, Thomsson KA, Bergstrom JH, van der Post S, Rodriguez-Piñero AM, Sjovall H, Backstrom M, Hansson G (2011) Composition and functional role of the mucus layers in the intestine. Cell Mol Life Sci 68:3635–3641
Velcich A, Yang W, Heyer J, Fragale A, Nicholas C, Viani S, Kucherlapati R, Lipkin M, Yang K, Augenlicht L (2002) Colorectal cancer in mice genetically deficient in the mucin Muc2. Science 295:1726–1729
Van der Sluis M, De Koning BA, De Bruijn AC, Velcich A, Meijerink JP, Van Goudoever JB, Buller HA, Dekker J, Van Seuningen I, Renes IB, Einerhand AW (2006) Muc2-deficient mice spontaneously develop colitis, indicating that MUC2 is critical for colonic protection. Gastroenterology 131:117–129
Buisine MP, Devisme L, Savidge TC, Gespach C, Gosselin B, Porchet N, Aubert JP (1998) Mucin gene expression in human embryonic and fetal intestine. Gut 43:519–524
Aslam F, Palumbo L, Augenlicht LH, Velcich A (2001) The Sp family of transcription factors in the regulation of the human and mouse MUC2 gene promoters. Cancer Res 61:570–576
Core LJ, Martins AL, Danko CG, Waters C, Siepel A, Lis JT (2014) Analysis of nascent RNA identifies a unified architecture of initiation regions at mammalian promoters and enhancers. Nat Genet 46:1311–1320
Carelli FN, Liechti A, Halbert J, Warnefors M, Kaessmann H (2018) Repurposing of promoters and enhancers during mammalian evolution. Nat Commun 9:4066
Quon DVK, Delgadillo MG, Khachi A, Smale ST, Johnson PJ (1994) Similarity between a ubiquitous promoter element in an ancient eukaryote and mammalian initiator elements. Proc Natl Acad Sci USA 91:4579–4583
Mikhaylichenko O, Bondarenko V, Harnett D, Schor IE, Males M, Viales RR, Furlong EEM (2018) The degree of enhancer or promoter activity is reflected by the levels and directionality of eRNA transcription. Genes Dev 32:42–57
Woodfint RM, Chen PR, Ahn J, Suh Y, Hwang S, Lee SS, Lee K (2017) Identification of the MUC2 promoter as a strong promoter for intestinal gene expression through generation of transgenic quail expressing GFP in gut epithelial cells. Int J Mol Sci 18(1):196
Monzani PS, Adona PR, Ohashi OM, Meirelles FV, Wheeler MB (2016) Transgenic bovine as bioreactors: challenges and perspectives. Bioengineered 7:123–131
Yang B, Wang J, Tang B, Liu Y, Guo C, Yang P, Yu T, Li R, Zhao J, Zhang L, Dai Y, Li N (2011) Characterization of bioactive recombinant human lysozyme expressed in milk of cloned transgenic cattle. PLoS ONE 6:e17593
Yu Y, Wang Y, Tong Q, Liu X, Su F, Quan F, Guo Z, Zhang Y (2013) A site-specific recombinase-based method to produce antibiotic selectable marker free transgenic cattle. PLoS ONE 8:e62457
Parc AL, Karav S, Rouquié C, Maga EA, Bunyatratchata A, Barile D (2017) Characterization of recombinant human lactoferrin N-glycans expressed in the milk of transgenic cows. PLoS ONE 12:e0171477
Wang Y, Ding F, Wang T, Liu W, Lindquist S, Hernell O, Wang J, Li J, Li L, Zhao Y, Dai Y, Li N (2017) Purification and characterization of recombinant human bile salt-stimulated lipase expressed in milk of transgenic cloned cows. PLoS ONE 12:e0176864
Zhang S, Ma X, Wang Z, Zhang P, Li Z (2019) Production of transgenic cattle expressing lysine-rich polypeptide in milk by somatic cell nuclear transfer. Transgenic Res 28:317–325
Wheeler MB (2013) Transgenic animals in agriculture. Nat Educ Knowl 4:1
May K, Scheper C, Brügemann K, Yin T, Strube C, Korkuć P, Brockmann GA, König S (2019) Genome-wide associations and functional gene analyses for endoparasite resistance in an endangered population of native German Black Pied cattle. BMC Genomics 20:277
Inoue H, Nojima H, Okayama H (1990) High efficiency transformation of Escherichia coli with plasmids. Gene 96:23–28
Kumaki Y, Oda M, Okano M (2008) QUMA: quantification tool for methylation analysis. Nucleic Acids Res 36(Web Server):W170–W175
Louis M, Becskei A (2002) Binary and graded responses in gene network. Sci Signal. https://doi.org/10.1126/stke.2002.143.pe33
Carter D, Chakalova L, Osborne CS, Dai Y, Fraser P (2002) Long-range chromatin regulatory interactions in vivo. Nat Genet 32:623–626
Turker M (2002) Gene silencing in mammalian cells and the spread of DNA methylation. Oncogene 21:5388–5393
Zhao C, Guo X, Chen S, Li C, Yang Y, Zhang J, Chen S, Jia Y, Wang T (2017) Matrix attachment region combinations increase transgene expression in transfected Chinese hamster ovary cells. Sci Rep 7:42805
Chen F, Zhang Q, Deng X, Zhang X, Chen C, Lv D, Li Y, Li D, Zhang Y, Li P, Diao Y, Kang L, Owen GI, Chen J, Li Z (2018) Conflicts of CpG density and DNA methylation are proximally and distally involved in gene regulation in human and mouse tissues. Epigenetics 13:721–741
Thormann V, Rothkegel MC, Schöpflin R, Glaser LV, Djuric P, Li N, Chung HR, Schwahn K, Vingron M, Meijsing SH (2018) Genomic dissection of enhancers uncovers principles of combinatorial regulation and cell type-specific wiring of enhancer-promoter contacts. Nucleic Acids Res 46:2868–2882
Vincent A, Perrais M, Desseyn JL, Aubert JP, Pigny P, Van Seuningen I (2007) Epigenetic regulation (DNA methylation, histone modifications) of the 11p15 mucin genes (MUC2, MUC5AC, MUC5B, MUC6) in epithelial cancer cells. Oncogene 26:6566–6576
Duval K, Grover H, Han LH, Mou Y, Pegoraro AF, Fredberg J, Chen Z (2017) Modeling physiological events in 2D vs. 3D cell culture. Physiology 32:266–277
Hamada T, Goto M, Tsutsumida H, Nomoto M, Higashi M, Sugai T, Nakamura S, Yonezawa S (2005) Mapping of the methylation pattern of the MUC2 promoter in pancreatic cancer cell lines, using bisulfite genomic sequencing. Cancer Lett 227:175–184
Mesquita P, Jonckheere N, Almeida R, Ducourouble MP, Serpa J, Silva E, Pigny P, Silva FS, Reis C, Silberg D, Van Seuningen I, David L (2003) Human MUC2 mucin gene is transcriptionally regulated by Cdx homeodomain proteins in gastrointestinal carcinoma cell lines. J Biol Chem 278:51549–51556
Van der Sluis M, Melis MH, Jonckheere N, Ducourouble MP, Buller HA, Renes I, Einerhand AW, Van Seuningen I (2004) The murine Muc2 mucin gene is transcriptionally regulated by the zinc-finger GATA-4 transcription factor in intestinal cells. Biochem Biophys Res Commun 325:952–960
Burger-van Paassen N, Vincent A, Puiman PJ, van der Sluis M, Bouma J, Boehm G, van Goudoever JB, van Seuningen I, Renes IB (2009) The regulation of intestinal mucin MUC2 expression by short-chain fatty acids: implications for epithelial protection. Biochem J 420:211–219
Yamada N, Hamada T, Goto M, Tsutsumida H, Higashi M, Nomoto M, Yonezawa S (2006) MUC2 expression is regulated by histone H3 modification and DNA methylation in pancreatic cancer. Int J Cancer 119:1850–1857
Hagen G, Muller S, Beato M, Suske G (1992) Cloning by recognition site screening of two novel GT box binding proteins: a family of Sp1 related genes. Nucleic Acids Res 20:5519–5525
Ookawa K, Kudo T, Aizawa S, Saito H, Tsuchida S (2002) Transcriptional activation of the MUC2 gene by p53. J Biol Chem 277:48270–48275
Almeida R, Silva E, Santos-Silva F, Silberg DG, Wang J, De Bolos C, David L (2003) Expression of intestine-specific transcription factors, CDX1 and CDX2, in intestinal metaplasia and gastric carcinomas. J Pathol 199:36–40
de Sousa MM, Zerlotini A, Geistlinger L, Tizioto PC, Taylor JF, Rocha MIP, Diniz WJS, Coutinho LL, Regitano LCA (2018) A comprehensive manually-curated compendium of bovine transcription factors. Sci Rep 8:13747
Funding
This research was financed by Embrapa (Grant 01.13.06.001.06), and Yamashita MSA was financed by a scholarship from the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brasil (CAPES).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yamashita, M.S.A., Vargas, L.N. & Melo, E.O. Characterization of the regulatory 5′-flanking region of bovine mucin 2 (MUC2) gene. Mol Cell Biochem 476, 2847–2856 (2021). https://doi.org/10.1007/s11010-021-04133-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-021-04133-1