Abstract
Methionine aminopeptidase (MAP) performs the essential post-translational N-terminal methionine excision (NME) of nascent polypeptides during protein synthesis. To characterize MAP from Mycobacterium tuberculosis, two homolgues, mapA (Rv0734) and mapB (Rv2861c), were over expressed and purified as recombinant proteins in E. coli. In vitro activity assay of apo-MtbMAPs using l-Met-p-nitro anilide as substrate revealed MtbMAP A to be catalytically more efficient compared to MtbMAP B. Ni2+ was the best activator of apo-MtbMAP A, whereas Ni2+ and Co2+ activated apo-MtbMAP B equally. MtbMAP B showed higher thermo-stability, but was feedback inhibited by higher concentrations of l-methionine. Aminopeptidase inhibitors like actinonin and bestatin inhibited both MtbMAPs, more prominently MtbMAP B. Among the site-directed mutants of MtbMAP B, substitution of metal-binding residue D142 completely abolished enzyme activity, whereas substitution of residues forming S1′ pocket, C105S and T94C, had only moderate effects on substrate hydrolysis. Present study identified a specific insertion region in MtbMAP A sequence which differentiates it from other bacterial and eukaryotic MAPs. A deletion mutant lacking amino acids from this insertion region (MtbMAP A-∆164-176) was constructed to probe into their structural and functional role in activity and stability of MtbMAP A. The limited success in soluble expression of this deletion mutant suggests further optimizations of expression conditions or alternative bioinformatics approaches for further characterization of this deletion mutant of MtbMAP A.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Protein synthesis is always initiated with either methionine or N-formyl methionine in prokaryotes and eukaryotes, respectively. The initiator N-terminal methionine is cleaved off from 50 to 70% of nascent polypeptides in a co-translational enzymatic process called N-terminal methionine excision (NME) for their proper folding and functionality. During NME in prokaryotes, peptide deformylase (PDF: E.C. 3.5.1.31) deformylates the N-formyl methionine allowing subsequent methionine cleavage by methionine aminopeptidase (MAP E.C.3.4.11.18) [1]. MAP acts only on polypeptides with non-bulky and uncharged amino acids at their penultimate position [2]. Due to essentiality of NME for metabolism and cell survival, both these enzymes are ideal drug targets for anti-bacterial drug discovery against pathogens. PDF, being prokaryotic specific, attracted initial attention as a drug target. However, the emergence of resistant mutants, bypassing the deformylation step, posed major concerns for developing PDF inhibitors as drugs [3]. The physiological importance of MAP is underscored by the lethality upon deletion or inhibition of MAP genes or gene products in Escherichia coli and Salmonella typhimurium [4, 5]. So there has been a shift of focus towards MAP as drug target in recent years.
Methionine aminopeptidases are ubiquitous metallo-proteases requiring divalent metal ions for catalysis [6]. Many putative MAP coding genes (map) have been identified and characterized across genomes and several of them have been shown to be functional [2, 4, 7]. Based on amino acid sequence alignment, MAPs are divided into two major classes (types I and II) [8, 9]. Eubacteria possess only MAP1; archea have MAP2; while eukaryotes have both the classes [2]. MAP1 enzymes are further subdivided into types 1a, 1b, and 1c depending on the presence and nature of an N-terminal extension [9, 10].
Inhibition of eukaryotic MAP2 (human) and MAP1b (Plasmodium falciparum) has been reported to be effective in treating disorders like cancer, rheumatoid arthritis [11, 12], and malaria [13], respectively. In recent years, many groups are attempting to characterize MAPs from various bacterial pathogens as target for screening and development of MAP-specific antibacterial inhibitors [14, 15]. We hold initial success in purifying and characterizing a putative MAP protein from the cell lysate of Mycobacterium smegmatis mc2155 [16].
Emergences of multiple-drug resistant strains of M. tuberculosis have increased the demand for improved TB drugs with alternative targets. In this context, it was decided to focus on characterizing the MAP enzymes from M. tuberculosis. The genome of M. tuberculosis H3Rv has two open reading frames (ORFs), Rv0734 (mapA) and Rv2861c (mapB) encoding MtbMAP A and MtbMAP B proteins (designated as MetAP 1a and MetAP 1c, respectively, in previous studies), respectively. Although, crystal structure of MtbMAP B was reported in 2005 [10], the biochemical properties of both MtbMAPs were largely unknown, except for few recent reports [17–19]. Moreover, the crystal structure of MtbMAP A has not been solved yet, and the most critical amino acids essential for its catalysis and stability are not so well defined. The present study aims to characterize the salient biochemical and biophysical properties of recombinant MtbMAP proteins followed by characterizing their relevant site-directed mutants to explore the role of observed amino acid sequence variations on their enzyme activities.
Materials and methods
Materials
pET28a vector was purchased from Novagen (USA). M. tuberculosis H37Rv genomic DNA was obtained from Colorado State University, USA, based on a research material transfer agreement. All the fine chemicals, PCR reagents, substrates (amino acid derivatives of p-nitro anilide), and inhibitors (bestatin, amastatin, actinonin, and fumagillin) were purchased from Sigma-Aldrich (USA). All the restriction enzymes and DNA modifying enzymes were purchased from Fermentas (USA). Site-directed mutagenesis was performed using Quick-Change Mutagenesis kit (Stratagene, Germany), and purification of his-tagged recombinant enzymes was done using Ni-NTA spin column kit (Qiagen, Germany). Luria–Bertani (LB) media and antibiotics were procured from Hi-Media (India). All the oligonucleotides used in this study were custom synthesized (IDT, USA).
Buffers
Lysis buffer: 20 mM phosphate buffer, pH 7.4, 300 mM NaCl, 5 mM imidazole, 1 mM PMSF, 10 μl protease inhibitor cocktail for his-tagged proteins, and 0.5 mg/ml lysozyme; wash buffer: 20 mM phosphate buffer, pH 7.4, 300 mM NaCl, 20 mM imidazole; elution buffer: 20 mM phosphate buffer, pH 7.4, 300 mM NaCl, 250 mM imidazole. Activity buffer: 50 mM Tris-Cl, pH 8.0, 150 mM KCl; assay buffer: 50 mM Tris-Cl, pH 8.0, 100 mM KCl; blocking buffer; 3% bovine serum albumin (BSA), 10 mM Tris-Cl, pH 7.5, and 150 mM NaCl.
Cloning, expression, and purification of MtbMAPs
All DNA manipulations were performed using standard protocols [20].
PCR amplification and cloning of Rv0734 (mapA) and Rv2861c (mapB)
mapA gene was amplified using Taq DNA polymerase from M. tuberculosis H37Rv genomic DNA using the gene-specific primers; mapA sense 5′TGCGCCCACTGGCACGGCT3′ (designed for blunt end cloning) and mapA antisense: 5′CCCCTCGAGCTAACCGAGCGTCA3′ (XhoI restriction site, underlined). The PCR conditions were as follows: initial denaturation for 5 min at 94°C followed by 30 cycles of 94°C for 45 s, 60°C for 30 s, 72°C for 1 min, and a final extension at 72°C for 10 min. The 801-bp PCR product was ligated as blunt/sticky end insert into pET28a vector, in-frame with N-terminus 6× His-tag. The mapB gene was amplified using mapB sense 5′ACCATAAATGCTAGCCATATGCCTAGTC3′ (NdeI restriction site underlined) and mapB antisense 5′GCCCTCGAGGAATTCT AACTACAGACAGGT 3′ (EcoRI restriction site, underlined) primers. The PCR conditions differed from that of mapA in having 59°C as the annealing temperature. The 858-bp PCR product was ligated into the NdeI/EcoRI site of pET28a, in-frame with N-terminus 6× His-tag of pET28a vector. The recombinant plasmids pET28a: mapA and pET28a: mapB were verified for PCR anomalies by DNA sequencing of both strands using T7 primers (Scigenomics, India). For over-expression and purification of his-tagged recombinant proteins, the confirmed plasmids were transformed into E. coli BL21 (DE3) cells.
Over-expression and purification of MtbMAPs
Overnight cultures of E. coli BL21 (DE3) carrying pET28a: mapA or pET28a: mapB were inoculated into 500 ml of LB broth supplemented with 30 μg/ml kanamycin and were incubated at 37°C, 200 rpm until the OD600 nm reached 0.6–0.7. The cultures were then induced with 0.5 mM isopropyl β-d-1-thiogalactopyranoside (IPTG) and allowed to grow for another 12 h at 30°C, 200 rpm. The cells were harvested by centrifugation at 8,590×g (Kubota, Japan) 4°C for 15 min, and cell pellets were stored at −80°C until use. The pellets were resuspended in 25 ml of ice-cold lysis buffer and left on ice for 30 min prior to disruption by sonication (five cycles of 30 s ON 30 s OFF at 42% amplitude) (Sonics, USA), and the resulting lysate was centrifuged at 20,960×g for 30 min at 4°C to recover soluble proteins in the supernatant. Proteins in each fraction were analyzed on 12% SDS-PAGE. Total protein contents in samples were quantified by Bradford’s method using BSA as standard.
Recombinant MtbMAPs were purified from the soluble fractions through Ni-NTA columns under native conditions according to the manufacturer’s protocol (Qiagen). Briefly, the cell lysate was passed through the Ni-NTA columns equilibrated with lysis buffer followed by three washes with wash buffer. The bound recombinant proteins were finally eluted with elution buffer into ethylenediaminetetraacetic acid (EDTA), making the final EDTA concentration as 100 mM, in order to produce purified apo-enzymes of MtbMAPs. EDTA and imidazole were later removed by extensive dialysis for 14 h against the activity buffer at 4°C with two buffer exchanges. The purified apo-enzymes were concentrated using 10 kDa amicon centri-concentrators (Millipore, India) to 1 mg/ml and were stored at −80°C until use. For all assay purposes, enzymes were diluted in the activity buffer.
Western blotting
One microgram soluble protein fractions of MtbMAPs were separated on 12% SDS-PAGE, and the proteins were electrophoretically transferred to a nitrocellulose membrane at 8 mA current for 2.5 h. The blot was then treated with a blocking buffer and incubated with anti-his primary antibodies (Qiagen). Following several washes, the blot was incubated with secondary antibodies (anti-mouse IgG conjugated with alkaline phosphatase, Sigma) and was developed with 5-bromo-4-chloro-3′-indolyl phosphate p-toluidine/nitro-blue tetrazolium chloride solution (BCIP/NBT) (Sigma) until dark brownish-red color was developed.
MAP assay
MAP assay was performed using l-Met-p-nitro anilide (l-Met-pNA) substrate as reported elsewhere [21]. MAP activities of apo-MtbMAP A and apo-MtbMAP B were assayed in the presence of 0.2 mM CoCl2 [17], with 1 mM l-Met-pNA as substrate using 1.6 μg (0.22 μM) of purified apo-MtbMAP A (MW ≈ 29 kDa) and 2 μg (0.275 μM) purified apo-MtbMAP B (MW ≈ 32 kDa), in assay buffer at 37°C in a total reaction volume of 250 μl. The enzyme activities were expressed as μM pNA released/min/mg protein.
Biochemical characterization of MtbMAPs
Substrate specificity
Substrate specificities of MtbMAPs were analyzed using p-NA derivatives of different N-terminal amino acids like methionine, glycine, arginine, proline, leucine, and valine, at 1 mM concentration, in the standard MAP assay in the presence of 0.2 mM CoCl2. Their activities were represented as percentage relative activities of enzymes with l-Met-p-NA.
Activation by divalent metal ions
Apo-enzymes of MtbMAPs were assayed in the presence of varying concentrations (1–1,000 μM) of divalent metal chlorides (Co2+, Ni2+, Mn2+, Mg2+ and Fe2+) while performing standard MAP assay with 1 mM l-Met-pNA. Appropriate metal ion blanks were performed to cancel out any background absorbance. The optimum concentrations of the best activating metal ions for each of these enzymes were selected for their further characterizations.
Temperature and pH optimum
Enzyme assays of MtbMAPs were carried out at different temperatures ranging from 10 to 65°C in different buffers of pH ranging from 3.5 to 10.5 in order to determine the optimum temperature and pH for their enzyme activities. For temperature optimization, the standard MAP assay was performed at pH 8.0, and at 37°C for pH optimization. The buffers used to set different pH were 50 mM of citrate buffer (pH 4.8), sodium acetate buffer (pH 3.5–5.0), phosphate buffer (pH 5.5–7.5), Tris-Cl buffer (pH 8–9.5), and sodium-glycine buffer (pH 9.5–10.5), all of them containing 100 mM KCl.
Thermo-stability
Thermo-stability was assessed at 37 and 55°C by pre-incubating 10 μg of purified apo-MtbMAPs for 1 h. The enzyme activities were monitored at 37°C by adding aliquots of pre-incubated MtbMAP A and MtbMAP B into the standard MAP assay in the presence of optimized concentrations of Ni2+, at every 10 min intervals. The residual activities at each time intervals were represented as their percentage relative activities at time 0.
Enzyme kinetics
Continuous MAP assay was performed against various concentrations of l-Met-p-NA (0–1 mM) at room temperature at pH 8.0 using 0.22 μΜ MtbMAP A and 0.275 μΜ MtbMAP B in the presence of optimized concentrations of best activating divalent metal ion (100 μΜ Ni2+ and 40 μΜ Co2+ for MtbMAP A; 100 μΜ Νi2+ and Co2+ for MtbMAP B). Reaction rates of MtbMAPs at various concentrations of substrate were calculated from linear cure fits of μΜ p-NA released versus time data. K m and V max were determined from slopes of various concentrations of substrate by applying a non-linear curve fit. Kinetic analysis was performed using Graph Pad Prism version 5.0 (Graph pad software, San Diego, CA, USA). The turnover number, K cat value, was determined directly from the software by applying the formula K cat = V max/[ET], where V max is the maximum velocity and E T is the total enzyme concentration.
Effect of inhibitors
The effects of potential aminopeptidase inhibitors such as bestatin, amastatin [22], actinonin [23], and fumagillin [24] were tested at three different inhibitor concentrations (0.1, 1, and 10 μM) on Ni2+-loaded MtbMAPs in the standard MAP assay. The inhibitors were dissolved in 100% DMSO. Ni2+-loaded MtbMAPs were incubated with the indicated concentrations of inhibitors for 15 min at room temperature prior to starting the standard MAP assay by addition of 1 mM l-Met-p-NA. Appropriate inhibitor blanks were also performed. The uninhibited reaction in each case was taken as control, and the results were expressed as percentage relative activity of the control.
Feedback inhibition by l-methionine
In order to determine the product inhibition on MtbMAPs, standard MAP assay was performed using Ni2+-loaded MtbMAPs in the presence of varying concentrations (0–100 μM) of l-methionine. Corresponding blank reactions using l-methionine were performed, and values were corrected by subtracting appropriate blanks.
Design, construction, and characterization of site-directed mutants of MtbMAPs
The amino acid sequences of MtbMAP A and MtbMAP B were aligned with E. coli and human type I MAPs using the ClustalW program (http://www.ebi.ac.uk/clustalW/index.html), and sites were selected for creating substitution or deletion mutants of these enzymes, based on the sequence alignment.
Based on the multiple sequence alignment, six substitution mutants of MtbMAP B (T94C, C105S, W255L, H114A, D142A, and E269V) and one deletion mutant of MtbMAP A (MtbMAP A-∆164-176) were designed to characterize the role of these substituted/deleted amino acids in enzyme activities of MtbMAPs. Internal primers having corresponding mutations were designed (Table S1). pET28a: mapA or pET28a: mapB constructs served as template for constructing site-directed mutants. Site-directed mutagenesis was performed essentially according to the manufacturer’s protocol. Briefly, the whole plasmid was PCR amplified with Pfu-turbo DNA polymerase using internal mutagenic primers, and the amplicons were digested with DpnI in order to remove parent plasmids. After transformation, plasmids were isolated from E. coli DH5α cells, and each mutation was confirmed by DNA sequencing (Scigenomics, India). The confirmed mutants were transformed into E. coli BL21 (DE3) cells for expression and purification.
Protein expressions from all the confirmed mutants were performed similar to their wild-type enzymes. The expressed mutant proteins were purified, processed, and stored similarly. The enzyme activities of purified variants were determined relative to the wild-type enzymes by performing MAP assay at optimized conditions.
Results
Cloning, recombinant expression, and purification of MtbMAPs
mapA (801 bp) and mapB (858 bp) genes were successfully cloned into pET28a vector with N-teminal his-tag fusion. Even though both MtbMAPs were sufficiently expressed in soluble fractions of E. coli BL21 (DE3) cell lysate, MtbMAP A was expressed mainly in the insoluble fraction compared to MtbMAP B (Fig. S1a). Both recombinant proteins were purified from the soluble fractions using native conditions. Recombinant MtbMAPs got eluted from Ni-NTA column at 250 mM imidazole concentration, providing purity to near homogeneity. Purified MtbMAP A exhibited a single purified band at 34 ± 2 kDa, whereas purified MtbMAP B was seen as 37 ± 2 kDa band at 12% SDS-PAGE (Fig. S1b). The final yield of purified apo-MtbMAP A and MtbMAP B were 1–2 and 6–8 mg/l, respectively. The authentic recombinant nature of MtbMAPs and their difference in soluble expression were further confirmed by immuno-blotting with anti-His antibodies (Fig. S1c).
Enzyme activities and substrate specifcities of MtbMAPs
In the absence of any added metal ion, both apo-MtbMAPs showed negliglible hydrolysis of l-Met-p-NA, proving the efficiency in the preparation of apo-enzymes. But, in the presence of 0.2 mM CoCl2, at 37°C, MtbMAP B showed almost twice the activity of MtbMAP A (Fig. 1a). Both MtbMAPs showed their highest specificity toward l-Met-p-NA, even though MtbMAP A could hydrolyse p-NA substrates with other uncharged amino acid to a very small extent (Fig. 1b).
Enzyme activity and substrate specificity of MtbMAPs. a Enzyme activity of purified apo-MtbMAPs with 1 mM l-Met-pNA performed in assay buffer, pH 8.0, at 37°C in the presence of 0.2 mM CoCl2. b Substrate preference of MtbMAPs determined in enzyme assay using 1 mM of the indicated amino acyl-pNAs in the presence of 0.2 mM CoCl2, at pH 8.0 and 37°C. The results are represented as percentage relative activities of each enzyme against l-Met-pNA
Metal ion preferences of MtbMAPs
During titration of enzyme activity with different metal ions, 0.22 μM MtbMAP A was activated best at 100 μM of Ni2+ followed by 40 μM of Co2+. Slight activation of MtbMAP A activity was also noted in the entire range by Mn2+ and in limited range by Fe2+ (Fig. 2a). In the case of MtbMAP B, both Ni2+ and Co2+ showed similar trends in activation of enzyme activity. Both these metal ions activated substrate hydrolysis best at 100 μM. Significant MtbMAP B enzyme activity was triggered by Mn2+ at 400 μM concentration, but both Fe2+ and Mg2+ were inefficient activators (Fig. 2b). In both cases, metal ions had inhibitory effect on enzyme activities beyond a threshold concentration.
Optimum pH and temperature for MtbMAP activity
The optimum temperature for enzyme activity of MtbMAP A was 37°C and for that of MtbMAP B was 55°C (Fig. 3a). MtbMAP B activity increased only in very small increments from 37 to 55°C, reaching the maximum activity at 55°C. Both the enzymes were not active beyond 65°C. Both MtbMAP A and MtbMAP B had maximum activity at pH 8.0, with very low activity in acidic and basic range (Fig. 3b). Although MtbMAP B had 10% relative increase in activity at 55°C, for sake of comparison and for ease of handling, their further activity assays were also performed at 37°C at pH 8.0, similar to MtbMAP A.
MtbMAP B was found to be moderately stable at 37°C with a half-life (t 1/2) more than 1 h. At 55°C, this enzyme retained only 15% of its original activity after 1 h with t 1/2 close to 40 min (Fig. 4a). MtbMAP A was found to be less stable compared to MtbMAP B and had a t 1/2 of 50 min at 37°C. The enzyme lost all its activity after incubating at 55°C for 1 h with a t 1/2 close to 20 min at 55°C (Fig. 4b).
Thermo-stability of MtbMAPs. a MtbMAP B. b MtbMAP A. 10 μg of each apo-enzyme was incubated at the indicated temperatures, and their residual activities were determined in standard MAP assay at every 10 min in the presence of 100 μM Ni2+. The residual activities are expressed as percentage relative activities of each enzyme at time 0
Kinetic properties of MtbMAPs
MtbMAPs followed typical Michealis–Menten kinetics for catalysis against l-Met-pNA. The kinetic parameters of MtbMAPs are summarized in Table 1. Enzyme kinetic parameters suggested Ni2+-loaded MtbMAP A to be nearly six-fold catalytically more efficient than MtbMAP B, with higher substrate affinity and higher enzyme turnover.
Inhibition of MtbMAP enzyme activities
Among the four metallo-protease inhibitors studied, only bestatin and actinonin had inhibitory activities on MtbMAPs. At 10 μM concentration of bestatin, MtbMAP A and MtbMAP B lost nearly 25 and 50% of their activities, respectively. Actinonin showed remarkable inhibition of MtbMAPs activity at 10 μM concentration, leaving behind only 60 and 30% of original activities of MtbMAP A and MtbMAP B, respectively. In general, there was better inhibition of MtbMAP B compared to MtbMAP A. Not surprisingly, fumagillin, a known inhibitor of type II MAPs did not show any inhibitory effects on MtbMAPs at the tested concentration (Table 2).
Methionine as a feed back inhibitor had no effects on the in vitro activity of MtbMAP A, but inhibited MtbMAP B to a greater extent. At the highest concentration tested (100 μM), methionine inhibited 70% of original MtbMAP B activity (Fig. S2).
Site-directed mutagenesis of MtbMAPs
Multiple sequence alignment of MtbMAPs with representatives of bacterial and human type I MAPs is shown in Fig. 5. Both MtbMAPs share only 33% of sequence identity between themselves and have less than 45% of similarity with E. coli MAP 1 and 48% similarity with human MAP.
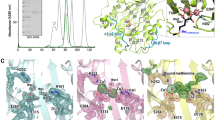
Multiple sequence alignment of MtbMAPs with type I MAP sequences from E. coli and H. sapiens . The conserved residues involved in metal-binding in MAP asterisk (*). The residues involved in the formation of substrate binding S1′ pockets in E. coli filled circle. The sites selected for creating substitution mutants in MtbMAP B arrow heads. The residues deleted from the insertion region of MtbMAP A in creating deletion mutant triangles. The multiple sequence alignment was constructed using ClustalW program (http://www.ebi.acuk/clustalW/index.html)
Within these variations, there was a high level of conservation of all the five metal-binding residues in all these MAPs. The metal-binding residues in case of MtbMAP A are D106, D117, H186, E219, and E250. The corresponding residues in MtbMAP B are D131, D142, H205, E238, and E269 (Fig. 5). Some residues involved in architecture of S1′ pocket in case of MtbMAPs showed variations from those of E. coli and human type I MAPs [25, 26].
Upon over-expression, only three mutants of MtbMAP B (T94C, C105S, and D142A) got expressed in the soluble fraction and got purified similar to wild type (Fig. 6a). The other three mutants (H114A, E269V, and W255L) were completely expressed in insoluble fractions as inclusion bodies (data not shown). Similarly, attempts to express the deletion construct, MtbMAP A-∆164-176, in soluble fraction too proved unsuccessful (Fig. 6b). The relative enzyme activities of the purified substitution mutants of MtbMAP B in the presence of 100 μM NiCl2 are shown in Fig. 6c. Substitution of metal-binding residue D142 almost completely abrogated the enzyme activity. Substitution of conserved S1′ forming cysteine with serine, C105S, led to nearly 50% reduction of original enzyme activity. Substituting T94 in MtbMAP B, corresponding to the cysteine in other type I bacterial MAPs, i.e., T94C, resulted in slight enhancement in the enzyme activity.
Expression, purification, and activity assay of site-directed mutants of MtbMAPs. a 12% SDS-PAGE showing soluble fractions expressing recombinant MtbMAP B and its site-directed mutants (labeled above each lane) (upper panel) and proteins purified from their soluble fractions (lower panel). b 12% SDS-PAGE showing soluble fractions expressing recombinant MtbMAP A and its deletion mutant (upper panel) and their purification from corresponding fractions (lower panel). c Relative enzyme activities of purified mutant proteins of MtbMAP B with reference to the wild type. Assay was performed using indicated concentrations of apo-enzymes in the presence of 100 μM Ni2+
Discussion
Methionine aminopeptidase, involved in NME process across the living world, plays vital physiological roles in cellular metabolism, which makes it indespensable growth and survival [1, 2]. Although a single map gene was thought to be essential for metabolism and survival of prokaryotic cells, many recently sequenced genomes have multiple map genes. Some of these paralogous MAP proteins were shown to have alternative physiological roles apart from NME [27].
The genome of M. tuberculosis possesses two map genes (Rv2861c and Rv0734), both of which encode functional MAP proteins [17–19]. The genome of rapid-growing, non-pathogenic M. smegmatis encodes four paralogus map genes (http://www.tigr.org). With their physiological relevance and low-sequence similarity with human MAPs, MtbMAPs provide opportunities for developing them as potent antimycobacterial drug targets for screening and developing selective inhibitors. The differential biochemical roles of MAP proteins during different phases of M. tuberculosis infection have not been addressed so well, except for identifying and quantifying the increased expression of MtbMAP A and MtbMAP B in log and stationary phase of its growth, respectively [17, 18].
The higher molecular weight of recombinant MtbMAPs on SDS-PAGE, compared to their calculated molecular weight (32 and 27 kDa), was reported in studies of Kanudia et al. [19]. Native MAP protein purified from M. smegmatis too displayed a molecular mass close to 37 kDa on SDS-PAGE [16]. Using Matrix-assisted laser ionization time-of-flight, the molecular mass of MtbMAP A and MtbMAP B was reported to be 28.8 and 32.5 kDa, respectively [17]. The low solublity of recombinant MtbMAP A compared to MtbMAP B was observed previously by Lu and Ye [28]. Nevertheless, considerable amounts of purified MtbMAP A proteins were available for biochemical characterization. This low solublity of MtbMAP A might be one of the major hinderance for determining its crystal structure.
Among the few studies published on biochemical characterization of MtbMAPs, majority of them were performed using purified holo-enzymes of MtbMAPs, without determining the nature of bound metal ions [17–19]. However, in our studies, we have purified MtbMAPs as apo-enzymes, which allowed us to screen and determine the best activating metal ions and their optimum concentration. All the above studies using holo-enzymes have stated MtbMAP B to be catalytically more efficient compared to MtbMAP A. Using the previously reported unoptimized concentration of Co2+ [17], we too have found MtbMAP B to be more active than MtbMAP A (Fig. 1a). On the other hand, previous studies using apo-enzymes state MtbMAP A to be catlytically more active than MtbMAP B [28, 29]. Interestingly, the present study with apo-enzymes demonstrated Ni2+ as an activator of MtbMAPs, apart from the reported Co2+. Our results also demonstrated the difference in in vitro metal preferences between the two MtbMAPs, which might be true in the cellular environment during their differential expressions at different phases of growth [17]. Mg2+ was reported as the best activator for native MAP enzyme from M. smegmatis [16] and was found true in case of recombinant holo-MtbMAP A in one of the previous studies [17]. However, both the recombinant apo-MtbMAPs were not activated by Mg2+ in the present study.
The choice of l-Met-p-NA [21] as the substrate for MAP assay in this study might have caused variations in overall kinetic parameters of MtbMAPs from the previous reported range using substrates like tri/tetra-peptides [17, 19] or methionyl aminomethylcoumarin (Met-AMC) [28, 29]. The kinetic paramaters in present study were more similar to those reported with l-Met-Pro-p-NA as substrate [18]. In general, there was a 100-fold reduction in the catalytic efficiency values of MtbMAPs compared to the previous reports with other substrates [17, 19, 28, 29]. Nearly 500-fold decrease in catalytic efficiency was reported in case of E. coli and Pyrococcus furiosus MAPs with l-Met-p-NA, compared to tetra-peptide Met–Gly–Met–Met [21]. l-Met-p-NA substrate was selected because it simplifies MAP assay and we successfully used it in characterizing M. smegmatis MAP [16]. Since the assay does not require enzyme coupling or fluorescent quantification, l-Met-p-NA could be the best choice of substrate for in vitro screening of MtbMAP inhibitors. However, this substrate does not differentiate the preferences of MtbMAPs toward the penultimate amino acids. Glycine, alanine, and proline were reported as the preferred amino acids at the penultimate positions for MtbMAPs [17, 19].
Optimum temperature for activity of MtbMAPs also showed variations depending on the substrates used in previous studies. Studies using tri/tera peptides state 55°C as the optimum temperature for MtbMAP A and around 40°C for MtbMAP B [17, 19]. But using l-Met-Pro-p-NA, optimum temperature for MtbMAP A and MtbMAP B was 42 and 50°C, respectively [18]. Results from present study confirmed this variation in temperature optimum of MtbMAPs depending on the substrate used for enzyme assay. The higher temperature optimum for MtbMAP B activity was very well supported by its higher thermo-stability. This again contradicts the results from circular dichroism (CD) spectroscopy experiments published recently, suggesting MtbMAP A to be more thermo-stable compared to MtbMAP B [19]. The stability determinants in the tertiary structures of MAPs are still not well defined.
Among the different aminopeptidase inhibitors tested, actinonin and bestatin showed considerable inhibition of both MtbMAPs at the concentration tested. Actinonin, a known PDF inhibitor with metal chelating hydroxamate side chains, was previously reported to inhibit M. smegmatis MAP and E. coli MAP [16, 24]. Similarly, the derivatives of bestatin, a known leucine aminopeptidase inhibitor, were reported to inhibit E. coli MAP with high efficiencies [2]. The magnitude of inhibition was more in case of MtbMAP B compared to MtbMAP A, revealing the difference in architecture of metal-binding and/or substrate binding pockets between the two MtbMAPs. Amastatin, not bestatin, was reported to inhibit MtbMAPs in a recent study, with more pronounced inhibition in case of MtbMAP A [19]. Our results completely disagree with this report.
At higher concentrations of l-methionine, in vitro activity of MtbMAP B but not MtbMAP A was inhibited. Among the two MAP proteins in S. cerevisiae, MAP 2 was inhibited at high-methionine concentration and not MAP 1 [30]. Based on experimental evidences, they proposed the role of MAP 1 in methionine salvage for maintaining intracellular methionine pool and suggested this property as the reason for the dominant essentiality of MAP 1 over MAP 2 in S. cerevisiae. The essential roles of multiple map genes in bacteria are not yet completely elucidated. Among the two MtbMAPs, the essential role of MtbMAP A over MtbMAP B in cell viability of M. tuberculosis was confirmed by gene knockdown studies [18]. One possible reason for the dominant essentiality of MtbMAP A in M. tuberculosis could be that MtbMAP B gets inhibited at higher methionine concentrations and MtbMAP A might be playing essential functions in methionine salvage similar to MAP 1 in S. cerevisiae.
Out of the three substitution mutants which we failed to express in soluble fractions, two (H114A and W255L) were successfully expressed and purified in a recent study [19]. These two variants showed negligible activity, proving the importance of these residues in the formation of the S1′ pocket of MtbMAP B. Similar results were reported for substitutions at corresponding sites in E. coli MAP and human MAP 1 [25, 26]. Among the two metal-binding residues substituted, only D142A was successfully expressed. Lack of activity of this mutant clearly supports the structural data demonstrating the role of this residue in binding the first metal ion in the di-nuclear metal center [28]. T94 in MtbMAP B corresponds to the S1′ hydrophobic pocket forming C59 residue in E. coli MAP [25]. This cysteine residue was reported to be characteristic for all bacterial type I MAPs studied and offers discrimination with the type II MAPs lacking it at this position [31]. Both MtbMAPs provide exceptions for this property of type I MAPs. The slight increase in enzyme activity of T94C mutant suggests a favorable rearrangement induced in the substrate binding pocket for better substrate binding. The loss of enzyme activity in case of C105S mutant was highly similar to the previously reported C70S mutation in E. coli MAP and human MAP 1 [25, 26], suggesting a similar role for this residue in formation of S1′ pocket in MtbMAP B.
Kanudia et al. [19] demonstrated the important functional role of the N-terminal extension of MtbMAP B in enzyme activity, apart from their in vivo role in ribosome binding [10]. Our efforts to identify the functional role of the specific insertion region in MtbMAP A was hindered by its lack of soluble expression (MtbMAP A-∆164-176).
Extensive genome search for MAP proteins with similar insertion region revealed these to be confined to suborder Corynebacterineae. In the absence of any structural information on MtbMAP A, it is highly relevant to characterize the role of this insertion region in the activity and stability of the enzyme. If proved essential, this region could be further explored for selective inhibition of MtbMAP A, either by structural-based inhibitors or by targeting sequence specific anti-sense oligonucleotides.
It was unfortunate that the efforts to solubilize some of these mutant proteins, by changing the expression conditions or by co-expression with molecular chaperons (pGro7 vector), did not produce expected results. Even though this article does not explain the ongoing efforts, we are trying to clone and express them from alternative E. coli expression systems like pMAL vector (Maltose binding), pMV261 expression vector in M. smegmatis, and yeast (Kluyvermyces lactis) vector pKLAC. However, no significant outcome yet. Molecular modeling of MtbMAP A-∆164-176 followed by molecular dynamic simulations and molecular docking studies would be another approach to analyze the importance of this region.
In conclusion, the present study brought out important biochemical features of the two MAP homologues of M. tuberculosis. The variations in the properties of MtbMAPs observed in the present study compared to few available previous reports clearly suggest the necessity for further critical analysis of their properties prior to developing them as drug targets against TB. Furthermore, the study attempted to bring out the importance of unique insertion region in MtbMAP A and proposed ways to further explore this region for designing mycobacterium-specific MAP inhibitors.
Abbreviations
- NME:
-
N-terminal methionine excision
- PDF:
-
Peptide deformylase
- MAP:
-
Methionine aminopeptidase
- MtbMAP:
-
Mycobacterium tuberculosis methionine aminopeptidase
- PCR:
-
Polymerase chain reaction
- l-Met-pNA:
-
l-Methionine-p-nitro anilide
- ORF:
-
Open reading frame
- IPTG:
-
Isopropyl β-d-1-thiogalactopyranoside
- EDTA:
-
Ethylenediaminetetraacetic acid
- BSA:
-
Bovine serum albumin
References
Giglione C, Boularot A, Meinnel T (2004) Protein N-terminal methionine excision. Cell Mol Life Sci 61:1455–1474
Lowther WT, Matthews BW (2000) Structure and function of the methionine aminopeptidases. Biochim Biophys Acta 1477:157–167
Olaleye OA, Bishaie WA, Liu JO (2009) Targeting the role of N-terminal methionine processing enzymes in Mycobacterium tuberculosis. Tuberculosis 89:S55–S59
Chang SY, McGary EC, Chang S (1989) Methionine aminopeptidase gene of Escherichia coli is essential for cell growth. J Bacteriol 171:4071–4072
Miller CG, Kukral AM, Miller JL, Movva NR (1989) pepM is an essential gene in Salmonella typhimurium. J Bacteriol 171:5215–5217
D’Souza VM, Swierczek SI, Cosper NJ et al (2002) Kinetic and structural characterization of manganese (II)-loaded methionyl aminopeptidases. Biochemistry 41:13096–13105
Wingfield P, Graber P, Turcatti G et al (1989) Purification and characterization of a methionine-specific aminopeptidase from Salmonella typhimurium. Eur J Biochem 180:23–32
Arfin SM, Kendall RL, Hall L et al (1995) Eukaryotic methionyl aminopeptidases: two classes of cobalt-dependent enzymes. Proc Natl Acad Sci USA 92:7714–7718
Bradshaw RA, Brickey WW, Walker KW (1998) N-terminal processing: the methionine aminopeptidase and N alpha-acetyl transferase families. Trends Biochem Sci 23:263–267
Addlagatta A, Quillin ML, Omotoso O et al (2005) Identification of an SH3-binding motif in a new class of methionine aminopeptidases from Mycobacterium tuberculosis suggests a mode of interaction with the ribosome. Biochemistry 44:7166–7174
Griffith EC, Su Z, Turk BE et al (1997) Methionine aminopeptidase (type 2) is the common target for angiogenesis inhibitors AGM-1470 and ovalicin. Chem Biol 4:461–471
Bradshaw RA, Yi E (2002) Methionine aminopeptidases and angiogenesis. Essays Biochem 38:65–78
Chen X, Chong CR, Shi L et al (2006) Inhibitors of Plasmodium falciparum methionine aminopeptidase 1b possess antimalarial activity. Proc Natl Acad Sci USA 103:14548–14553
Yuan H, Chai SC, Lam CK et al (2011) Two methionine aminopeptidases from Acinetobacter baumannii are functional enzymes. Bioorg Med Chem Lett 21:3395–3398
Chai SC, Wang WL, Ding DR, Ye QZ (2011) Growth inhibition of Escherichia coli and methicillin-resistant Staphylococcus aureus by targeting cellular methionine aminopeptidase. Eur J Med Chem 46:3537–3540
Narayanan SS, Ramanujam A, Krishna S, Nampoothiri KM (2008) Purification and biochemical studies on Methionine aminopeptidaes from Mycobacterium smegmatis mc2 155. Appl Biochem Biotechnol 151:512–521
Zhang X, Chen S, Hu Z et al (2009) Expression and characterization of two functional methionine aminopeptidases from Mycobacterium tuberculosis H37Rv. Curr Microbiol 59:520–525
Olaleye O, Raghunand TR, Bhat S et al (2010) Methionine aminopeptidases from Mycobacterium tuberculosis as novel antimycobacterial targets. Chem Biol 29:86–97
Kanudia P, Mittal M, Kumaran S, Charaborti PK (2011) Amino-terminal extension present in the Methionine aminopeptidase type 1c of Mycobacterium tuberculosis is indispensible for its activity. BMC Biochem 12:35
Sambrook J, Fritsch FE, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, New York
Mitra S, Dygas-Holz AM, Jiracek J et al (2006) New colorimetric assay for methionyl aminopeptidases: examination of the binding of a new class of pseudopeptide analog inhibitors. Anal Biochem 357:43–49
Rich DH, Moon BJ, Harbeson S (1984) Inhibition of aminopeptidases by amastatin and bestatin derivatives. Effect of inhibitor structure on slow-binding processes. J Med Chem 27:417–422
Chen DZ, Patel DV, Hackbarth CJ et al (2000) Actinonin, a naturally occurring antibacterial agent, is potent deformylase inhibitor. Biochemistry 39:1256–1262
Lowther WT, Mcmillen DA, Orville AM, Matthews BW (1998) The anti-angiogenic agent fumagillin covalently modifies a conserved active-site histidine in the Escherichia coli methionine aminopeptidase. Proc Natl Acad Sci USA 95:12153–12157
Chiu CH, Lee CZ, Lin KS et al (1999) Amino acid residues involved in the functional integrity of Escherichia coli methionine aminopeptidase. J Bacteriol 181:4686–4689
Li JY, Cui YM, Chen LL et al (2004) Mutations at the S1 sites of methionine aminopeptidases from Escherichia coli and Homo sapiens reveal the residues critical for substrate specificity. J Biol Chem 279:21128–21134
Drath M, Baier K, Forchhammer K (2009) An alternative methionine aminopeptidase, MAP-A, is required for nitrogen starvation and high-light acclimation in the cyanobacterium Synechocystis sp. PCC 6803. Microbiology 155:1427–1439
Lu JP, Chai SC, Ye QZ (2010) Catalysis and inhibition of Mycobacterium tuberculosis methionine aminopeptidase. J Med Chem 53:1329–1337
Lu JP, Ye QZ (2010) Expression and characterization of Mycobacterium tuberculosis methionine aminopeptidase type 1a. Bioorg Med Chem Lett 20:2776–2779
Dummitt B, Micka WS, Chang YH (2003) N-terminal methionine removal and methionine metabolism in Saccharomyces cerevisiae. J Cell Biochem 89:964–974
Swierczek K, Copik AJ, Swierczek SI, Holz RC (2005) Molecular discrimination of type-I over type-II methionyl aminopeptidases. Biochemistry 44:12049–12056
Acknowledgments
KMN acknowledges the Department of Biotechnology (DBT), New Delhi, India, for the research Grant. SSN expresses his gratitude to CSIR, India, for the senior research fellowship (SRF).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Narayanan, S.S., Nampoothiri, K.M. Biochemical characterization of recombinant methionine aminopeptidases (MAPs) from Mycobacterium tuberculosis H37Rv. Mol Cell Biochem 365, 191–202 (2012). https://doi.org/10.1007/s11010-012-1260-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-012-1260-8