Abstract
Breast cancer susceptibility gene 1 (BRCA1) is an important tumor suppressor gene in the human which associated with a variety of tumor suppressor genes to inhibit the growth of malignant tumors. Breast cancer is closely related to BRCA1 mutations in humans. Four BRCA1 (856-871) mutants were screened that can interact well with RAD51 (158-180) using computer to simulate mutations at Y 856, R 866 and Q 867. These mutants were synthesized by using Fmoc solid-phase synthesis, and purified by liquid chromatography. According to the results of fluorescence spectroscopy, R866W has the strongest binding ability for the above peptides and RAD51 (158-180). Based on the circular dichroism spectrum results, the sequence of the interaction ability of the above peptides on the secondary structure of RAD51 (158-180) is Y856P, Q867C > Y856P, R866W ≈ R866W > Y856P > BRCA1. All the spectral results show that R866W have the strongest interaction ability with RAD51(158-180).
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
More than 1 to 6 million people are diagnosed with breast cancer every year in our country. Besides, about 1 to 2 million people die from the disease (Fan et al. 2014). At present, the clinical drugs used in the treatment of breast cancer are all peptide drugs, most of which are through inducing DNA damage and inhibiting the growth of tumor cells, thereby achieving the therapeutic effect of tumor cell death (Ma and Jemal 2013).
The incidence of breast cancer is closely related to mutations in breast cancer susceptibility genes: BRCA1 (Miki et al. 1994) and BRCA2 (Wooster et al. 1994). Studies have shown that people or families carrying BRCA1 are more likely to develop breast and ovarian cancer (Tonin et al. 2015). BRCA1 is a tumor suppressor gene, which encodes a protein that can negatively regulate tumor growth (Rohlfs et al. 2018). BRCA1 which is located at 17q21 spans more than 100 000 bp, contains up to 41.5% of Alu repeats and 4.8% of other repeats. It also has 22 coding exons (Nathanson et al. 2001).
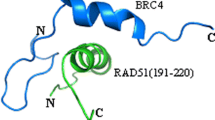
The main function of BRCA1 is to maintain the stability of the genome. As shown in Fig. 1, BRCA1 can directly regulate the cell cycle, DNA and centrosome replication by regulating the effects of RAD51, p53, BRAD1 and other proteins (Savage et al. 2015), thereby inhibiting the generation of malignant tumors (Timms et al. 2014). According to reports, one of the earliest signs that BRCA1 was involved in DNA repair is the observation that BRCA1 binds to and co-localizes with RAD51 in the nuclear foci of mitotic cells (Gudmundsdottir et al. 2006). BRCA1 can not only speed up the conversion and repair of genes with DNA double-strand breaks, but also inhibit spontaneous gene conversion mutations, which also confirms that BRCA1 can respond to and regulate the RAD51 recombination process according to different types of DNA damage (Wang et al. 2010). BRCA1 inhibits SCRS, indicating that the protein can play a role in sister chromatid cohesion (Cousineau et al. 2005).
After BRCA1 is inactivated, once DNA is damaged in response to estrogen-induced damage, loss of this regulation may cause genome instability, which may also be an important initial change in human carcinogenesis (Chu-Xia et al. 2003). However, in current studies, it can only be proved that BRCA1 is involved in such cell regulation, but its specific mode of action and mechanism are being studied (Annals of Oncology 2018).
BRCA1 is composed of 1863 amino acid residues and has high homology. Studies have shown that the peptides with amino acid codes 846-871 in BRCA1 can specifically bind to RAD51 (Clark et al. 2012; Ramirez et al. 2004). In the conserved sequence of BRCA1 (846-871), mutations in five special binding sites are likely to cause breast cancer: Y 856, K 862, R 866, Q 867 and P 871 (Yan et al. 2009).
In this work, BRCA1(856-871) and its mutant peptide were synthesized by solid phase peptide synthesis. Compared with BRCA1(846-871), BRCA1(856-871) not only retains the original 5 active sites that are prone to mutation, but also reduces 10 amino acids. The combination of Y856P and RAD51 (158-180) has been greatly improved, with the more obvious interaction effect. This work provides ideas and theoretical basis for further research and the design of breast cancer suppressive drugs.
Materials and Methods
Reagent
Fmoc-Rink-Amide-MBHA resin, Fmoc-AA-OH, 1-Hydroxybenzotriazole (HOBT), 2-(7-Azabenzotriazol-1-yl)-N,N,N',N'-tetramethyluronium hexafluorophosphate (HATU), were purchased from GL Biochem Ltd. (Shanghai, China). N,N-Dimethylformamide (DMF), Dichloromethane (DCM), N,N-Diisopropylethylamine(DIPEA), Trifluoroacetic acid (TFA) were purchased from Tianjin Kemiou Chemical Reagent Co., Ltd. 2-Hydroxy-1-ethanethiol, Thioanisole, Phenol were obtained from Aladdin reagent Co., Ltd. (Shanghai, China). Piperidine, Acetonitrile, Methanol were provided from Tianjin Siyou Fine Chemicals Company (Tianjin, China). The acetonitrile was chromatographic reagent and other reagents were analytical grade.
Design of BRCA1 (856−871) Mutants
The secondary structure of RAD51 (158-180) protein receptor was obtained from 1N0W provided by PDB bank. The 16 amino acid primary structure of BRCA1 (856-871) is input: YLQNTFKVSKRQSFAP. Other settings are the default parameters. Discovery studio 3.5 software is uesd to scan its active mutation sites for virtual mutations. 2 single mutants and 2 double mutants with good scanning results are selected. In this study, the N-terminus of all peptides was capped with acetic anhydride, and the C-terminus was amidated (Kitamura et al. 2006).
Synthesis of Peptide
Solid phase peptide synthesis (SPPS) is used to synthesize the targeting peptide (Zheng et al. 2013; Zhao and Lu 2015). RAD51 (158-180) (Ac-MYIDTEGTFRPER LLAVAERYG-NH2) is taken as an example. Following the sequence of peptide to link amino acids from the C-terminus to the N-terminus of the peptide chain to construct biomolecules. The synthesis steps are as follows. Taking 1.5 g of Fmoc-Rink-Amide-MBHA (0.34 mmol·g-1) resin and 15 mL of DMF in a solid phase synthesis tube, and blowing with N2 for 40 min to fully swell the resin. After vacuum pumping, adding 15 mL 20% piperidine/DMF solution to fully react for 20 min to remove the Fmoc protecting group. After draining, washing the resin alternately with 10 mL DMF and 10 mL methanol for a total of 3 times, each for 1 min. The Kaiser test is used to detect whether the protecting group is completely removed (it turns black or brown to prove that the protecting group is successfully separated and free amino groups are present). Weighing 0.43 g (4 equiv, 1.44 mmol) of Fmoc-Gly-OH, 0.73 g (6 equiv, 2.88 mmol) of HATU, and 0.19 g (4 equiv, 1.44 mmol) of HOBT with a balance in a small beaker. Adding 15 ml of DMF and stir to dissolve it. Then using a pipette to accurately measure 450 µl of DIPEA (8 equiv, 2.88 mmol) into it which activated for 5 min at room temperature and protected from light. Finally, pouring the activated amino acid into the synthesis tube and bubbling for 4 h with nitrogen protected from light. After the reaction, the resin was drained, and the resin was washed alternately with 10 mL DMF and 10 mL methanol successively, a total of 3 times, 1 min each. The Kaiser test proved that the coupling reaction was completely completed (the color does not change, indicating that the amino acid condensation reaction is complete). Repeating deprotection and coupling steps for the remaining amino acid connections until RAD51 (158-180) is synthesized. Measuring 10 ml of 20% Ac2O/DMF and 450 µl of DIPEA into the synthesis tube. N2 is blown for 4 h to carry out the acetylation reaction to form a stable amide bond. Adding a prepared 15mL cutting reagent with a volume ratio of 82.5% TFA, 5% thioethylene glycol, 5% thioanisole, 5% water and 2.5% phenol. The reaction was stirred for 4h, and the resin was completely cracked from RAD51 (158-180). The RAD51 (158-180) synthesized by the reaction was suction filtered into an Erlenmeyer flask filled with ice ether. It was frozen and allowed to stand overnight in the refrigerator to completely precipitate. The mixed solution after being placed at low temperature is centrifuged and washed 5 times in a centrifuge at a speed of 6500 r/min, each time for 5 min. Finally, the solution was processed in a four-ring freeze dryer to obtain a crude peptide product.
RP-HPLC Liquid Chromatography Characterization
Instrument model: Agilent 1260 HPLC. The conditions are as follows, Chromatographic column: Agilent Zorbax 300SB-C18 chromatographic column (4.6 mm × 250 mm); Stationary phase: 5 µl silica gel; Eluent A: deionized water containing 0.1% TFA; Eluent B: Acetonitrile; Elution method: gradient elution; Gradient ratio: 0-25-25.1 min correspond to 70:30, 45:55, 0:100 respectively; Scanning wavelength: 220 nm; Flow rate: 1.0 ml·min-1.
Mass Spectrometry Characterization
Instrument model: Thermo ScientificTM LCQ FleetTM. The conditions are as follows, Ion source: ESI; Detection mode: positive ion; Sheath gas flow rate: 20 psi; Scavenging flow rate: 5 psi; Auxiliary gas flow rate: 8 psi; Spray voltage: 4.5 KV; Tube lens voltage: 110 V; Capillary voltage: 35 V; Capillary temperature: 275 ℃; Full scan detection at m/z 150-2000.
Fluorescence Spectrum Characterization
Instrument model: Agilent Cary Eclipse. The conditions are as follows, Excitation wavelength: 275 nm; Excitation gap width: 5 nm; Emission gap width: 5 nm; Test wavelength range: 280–400 nm; Buffer: Tris-HCl solution (pH 7.4, 50 mM); The results are measured twice with samples of each concentration and averaged.
Circular Dichroic Spectrum Characterization
Instrument model: Bio-Logic MOS-500. The conditions are as follows, Scanning wavelength range: 200 nm–260 nm; Gap width: 0.5 nm; Step distance: 1 nm; Scanning speed: 60 nm·min-1; Buffer: Tris-HCl solution (pH 7.4, 50 mM); The results are measured 3 times with samples of each concentration and averaged.
Results and Discussion
Design and Analysis of Peptide
The conserved sequence of BRCA1 (846-871) can interact with RAD51 (158-180). Its truncated peptide BRCA1 (856-871) reduces the workload of coupling amino acids. Besides, it does not affect the active site of the conservative sequence, and it has a strong ability to interact with RAD51 (158-180). The process of docking mutants with RAD51 was simulated by Discovery Studio.
Synthesis and Characterization of Peptide
As shown in Table 1, 856Y in P1 is mutated to P, which enhances the hydrophilicity of the mutant peptide; 866R in P2 is mutated to W; 856Y in P3 is mutated to P and 866R is mutated to W; 856Y in P4 is mutated to P and 867Q is mutated to C. The purity of all peptide synthesized by SPPS is above 95%. As shown in Fig. 1a, b, the purity and mass-to-charge ratio were detected and analyzed by liquid phase and mass spectrometry.
Fluorescence Spectrum Analysis
Fluorescence spectroscopy can be used to analyze the interaction between proteins and other macromolecules, such as hydrogen bonding, hydrophobic interaction, electrostatic interaction, etc., to explore their binding mode and mechanism (Li et al. 2020). Parameters obtained by fluorescence spectroscopy analysis, such as fluorescence intensity, emission spectrum, etc. The characteristics of the polypeptide or protein molecular structure can be calculated from different angles; The Stern-Volmer equation can be used to calculate the quenching method of the fluorescence intensity during the interaction, and obtain the binding constants of the two through the fitting curve. These conclusions support the mechanism of the interaction between proteins in a deeper level (Zhao et al. 2018).
With the continuous increase of the volume of BRCA1 (856-871) added, the fluorescence intensity of the targeting peptide exhibits a regular fluorescence quenching phenomenon at a wavelength of 303 nm. According to the different quenching phenomena of the two interaction modes, it can be divided into two types: static quenching and dynamic quenching. Static quenching refers to the weak binding between the fluorescent molecules in the ground state and the quencher to generate a non-fluorescent complex; Dynamic quenching refers to the phenomenon that the fluorescent molecules in the excited state collide with the quencher to quench their fluorescence. If BRCA1 (856-871) causes dynamic quenching of RAD51 (158-180) fluorescence, the Stern-Volmer equation should be followed (Lakowicz 2006):
In the equation, F0 represents the fluorescence intensity of the targeting peptide RAD51 (158-180) when the parent peptide BRCA1 (856-871) is not added; F represents the fluorescence intensity of the targeting peptide RAD51 (158-180) after adding different volumes of BRCA1 (856-871); τ0 represents the average lifetime of fluorescent molecules when there is no quencher. Generally, the fluorescent lifetime of biological macromolecules is about 10-8 s; [Q] indicates the concentration of the parent peptide BRCA1 (856-871). Plotting F0/F and [Q] fluorescence intensity at a wavelength of 303 nm to obtain the fluorescence quenching curve of BRCA1 (856-871) and targeting peptide RAD51 (158-180) interaction. The quenching constant Kq can be calculated as 1.13 × 1012 L·mol−1·s−1 greater than the maximum controlled diffusion collision constant 2 × 1010 L·mol−1·s−1. It shows that the fluorescence quenching of the targeting peptide RAD51 (158-180) caused by BRCA1 (856-871) is not a dynamic quenching, but the interaction between the two to generate a static quenching of a non-luminescent complex. Similarly, the quenching constants of interaction between P1, P2, P3, P4 and RDA51 (158-180) can be obtained as shown in Fig. 2 and Table 2. All of them are greater than the maximum controlled diffusion collision constant 2 × 1010 L·mol−1·s−1. Therefore, between the peptide and RAD51 (158-180) are formed a non-luminescent complex caused by static quenching of fluorescence. Calculated by the above formula (1), the binding ability between any peptide calculated by the above formula and RAD51 (158-180) is a non-luminescent complex which causes static quenching. Therefore, the binding strength between them can be compared with each other by the following formula (2):
In the equation, F0 represents the fluorescence intensity of the targeting peptide RAD51 (158-180) without any BRCA1 peptide; F represents the fluorescence intensity of the system after adding BRCA1 peptide; [Q] represents the concentration of BRCA1 peptide; K represents the binding constant when the two interact; n represents the number of binding sites. Using lg[(F0-F)/F] to plot lg[Q], the number of binding sites n and the binding constant K are obtained by the slope and intercept of the formula. The binding constant K and the number of binding sites n between all peptide and RAD51 (158-180) are shown in the following Table 2.
According to the data in Table 2, compared with the parent peptide, the K value of P2 and RAD51 (158-180) increases, which indicates that the interaction between RAD51 (158-180) and P2 is stronger. The reason may be that the mutation of 866 R to W reduces the average hydrophilicity, which enhances the hydrophobic interaction between RAD51 (158-180) and W. Moreover, R is mutated to tryptophan W, which has the strongest autofluorescence. The binding ability of P1, P3, P4 and RAD51 (158-180) is slightly weakened. The mutation of 856 Y to P in P1 may be because the amino acid residue of Y itself contains phenolic hydroxyl groups which connected to the benzene ring is an electron-donating group. It is more conducive to interaction, so the binding ability is greatly reduced after the mutation; the decrease in P3 binding constant may be related to the decrease in net charge after mutation, which weakens the electrostatic interaction between molecules. In addition, 866 R in P3 is mutated to W, which may be due to the p-π conjugate of the amino acid residues of R, destroying the electron delocalization and weakening the binding ability; In P4, due to the mutation of 867 Q to C, the amino acid residue of cysteine contains -SH, which may interact with RAD51 (158-180) to form disulfide and hydrogen bonds. In summary, the binding capacity is P2 > BRCA1 > P4 ≈ P1 > P3.
Circular Dichroism Analysis
Circular dichroism can be used to determine the three-dimensional structure of proteins. The ultraviolet region of circular dichroism is 190–240 nm, the peak shape is “W”, and the main chromophore is peptide chain. The circular dichroism of α-helical conformation has a positive peak near 190 nm and negative peaks at 208 nm and 222 nm; The circular dichroism of β-sheet conformation has a strong positive peak at 195–198 nm and a negative peak at 217–278 nm; The circular dichroism of random coiled conformation has a negative band near 198 nm. Besides, there is a small broad positive peak near 220 nm. The change of the ellipticity of RAD51 (158-180) in the circular dichroic spectrum is related to the expansion of its α-helical structure. The circular dichroic spectra of BRCA1 and mutants are shown in Fig. 3a, b.
As shown in Table 3, after adding different concentrations of mutants to RAD51 (158-180), the content of α-helix decreased. It may be due to the weak non-covalent interaction with RAD51 (158-180) and the hydrophobic influence of the peptide, which caused the protein structure of RAD51 (158-180) to shrinking. With the addition of P4, the content of α-helix in the secondary structure of RAD51(158-180) increased slightly, which made the secondary structure of the polypeptide more orderly. The content of β-sheets ranges from 35.3 to 19.5%, 35.3 to 18.2%, 35.3 to 9.5%, 35.3 to 5.3%, respectively. In β-sheets, peptide bonds are involved in the formation of interchain hydrogen bonds and hydrophobic interaction. The substantial decrease in β-sheet content may be the result of the interaction with RAD51 (158-180), which changes the secondary structure. The content of β-turns ranges from 5.1 to 26.3%, 5.1 to 35.1%, 5.1 to 37.2%, 5.1 to 37.8%. The β-turn is a simple non-repetitive structure. The sites that have been found for protein antibody recognition, phosphorylation or glycosylation are often at or near the corner. The increase of β-turn content is more conducive to further optimization of the polypeptide structure after screening. All above shows that the targeting peptide RAD51 (158-180) has formed a secondary structure dominated by α-helix and β-turn after interacting with the mutants. In addition, the circular dichroism spectrum of the peptide and RAD51 (158-180) shows that the peak height and peak position have changed. The red shift occurs at the peak position. The height of the peak and the binding force of the hydrogen bond decreases continuously, resulting in the tightening of the polypeptide structure and the rearrangement of the peptide chain. It can be observed from the Fig. 3 that the secondary structure of the targeting peptide RAD51 (158-180) has changed to a certain extent. RAD51 (158-180) is a peptide with α-helix structure. According to the addition of the same gradient concentration and different mutants, the content of the α-helix structure of the targeting peptide can be changed. It is concluded that the order of the interaction effect between the above peptide and RAD51 (158-180) is P4 > P3 ≈ P2 > P1 > BRCA1 (Fig. 4).
Conclusion
BRCA1 (846-871) in the binding region of RAD51 (158-180) was selected as the template, combined with the reported active site of BRCA1 (846-871), and truncated to BRCA1 (856-871). Four mutant peptide computer simulation were selected according computer simulation design results. The parent peptide BRCA1 (856-871), the mutants and targeting peptide RAD51 (158-180) were synthesized by solid phase peptide synthesis. The crude peptide was separated and purified with a reversed-phase high performance solution, and then the separated fractions were characterized by ESI-MS mass spectrometry. The results show that the main peaks are all required targeting peptide, and the purity is above 95%.
The results of fluorescence spectroscopy and circular dichroic spectroscopy show that under the condition that the concentration of the fixed targeting peptide remains unchanged, as the concentration of any parent peptide or mutant is added, the targeting peptide is statically quenched to varying degrees. Besides, after adding the mutants, the secondary structure of RAD51 (158-180) has undergone significant changes, with a slight decrease in α-helix content and a significant increase in β-turn content. Among them, R866W and RAD51 (158-180) have the strongest affinity and the best binding ability.
Compared with previous work in the laboratory, this work uses truncated peptides. It not only retains the mutation site of the peptide, but also does not affect its activity. The truncated peptides can reduce the experimental workload to the greatest extent. Studies of the interaction between BRCA1 and different RAD51 peptides provide molecular design directions for exploring RAD51 inhibitors and drugs for breast cancer.
Reference
Chu-Xia D, Rui-Hong W (2003) Roles of brca1 in dna damage repair: a link between development and cancer. Human Mol Genet. https://doi.org/10.1093/hmg/ddg08
Clark SL, Rodriguez AM, Snyder, et al (2012) Structure-function of the tumor suppressor BRCA1. Comput Struct Biotechnol J. https://doi.org/10.5936/csbj.201204005
Cousineau I (2005) Brca1 regulates rad51 function in response to dna damage and suppresses spontaneous sister chromatid replication slippage: implications for sister chromatid cohesion, genome stability, and carcinogenesis. Cancer Res 65(24):11384–11391. https://doi.org/10.1158/0008-5472.CAN-05-2156
Fan L, Strasser-Weippl et al (2014) Breast cancer in China. Lancet Oncol 15(7):e279–e289. https://doi.org/10.1016/S1470-2045(13)70567-9
Gudmundsdottir K, Ashworth A (2006) The roles of BRCA1 and BRCA2 and associated proteins in the maintenance of genomic stability. Oncogene 25(43):5864–5874. https://doi.org/10.1038/sj.onc.1209874
Kitamura A, Kiyota T, Lee S, Sugihara G (2006) N- and c-terminal effect of amphiphilic. Alpha.helical peptides on the interaction with model and bio-membranes. Bull Chem Soc Jap 71(5):1151–1158. https://doi.org/10.1246/bcsj.71.1151
Lakowicz JR (2006) Principles of fluorescence spectroscopy. Springer, New York. https://doi.org/10.1007/978-0-387-46312-4
Li J, Lu K, Lv M, Liu G, Qi J (2020) Design, synthesis and interaction of brc4 analogous peptides with rad51(241–260). Amino Acids. https://doi.org/10.1007/s00726-019-02813-3
Ma J, Jemal A (2013) Breast cancer statistics. https://doi.org/https://doi.org/10.1007/978-1-4614-5647-6_1
Miki Y et al (1994) A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 266:66–71. https://doi.org/10.1126/science.7545954
Nathanson KL, Wooster R, Weber BL, Nathanson KN (2001) Breast cancer genetics: what we know and what we need. Nat Med 7(5):552–556. https://doi.org/10.1038/87876
Rad51 foci as a functional biomarker of homologous recombination repair and parp inhibitor resistance in germline brca mutated breast cancer. Ann Oncol (2018). https://doi.org/10.1093/annonc/mdy099
Ramirez CJ, Fleming MA, Potter et al (2004) Marsupial brca1: conserved regions in mammals and the potential effect of missense changes. Oncogene 23(9):1780–1788. https://doi.org/10.1038/sj.onc.1207292
Rohlfs EM, Learning WG et al (2018) Direct detection of mutations in the breast and ovarian cancer susceptibility gene brca1 by pcr-mediated site-directed mutagenesis. Clin Chem. https://doi.org/10.1016/S0009-8981(96)06438-8
Savage KI, Harkin DP (2015) Brca1, a ‘complex’ protein involved in the maintenance of genomic stability. FEBS J. https://doi.org/10.1111/febs.13150
Timms KM, Abkevich V, Hughes E, Neff C, Reid J, Morris B et al (2014) Association of brca1/2 defects with genomic scores predictive of dna damage repair deficiency among breast cancer subtypes. Breast Cancer Res 16(6):475. https://doi.org/10.1186/s13058-014-0475-x
Tonin PN, Perret et al (2015) Founder brca1 and brca2 mutations in early-onset french canadian breast cancer cases unselected for family history. Int J Cancer J Int Du Cancer 95(3):189–193. https://doi.org/10.1002/1097-0215(20010520)95:3%3c189::AID-IJC1032%3e3.0.CO;2-N
Wang H, Yang ES, Jiang J et al (2010) Dna damage-induced cytotoxicity is dissociated from brca1’s dna repair function but is dependent on its cytosolic accumulation. Cancer Res 70(15):6258. https://doi.org/10.1158/0008-5472.CAN-09-4713
Wooster R, Neuhausen SL, Mangion J et al (1994) (1994) Localization of a breast cancer susceptibility gene, BRCA2, to chromosome 13q12-13. Science 265(5181):2088–2090. https://doi.org/10.1126/science.8091231
Yan Z, Bao-Long H (2009) Brca1 gene exon11 mutation in 50 cases with breast cancer. China Cancer
Zhao DX, Lu K (2015) Design, synthesis, and characterization of BRC4 mutants based on the crystal structure of BRC4-RAD51(191–220). J Mol Model. https://doi.org/10.1007/s00894-015-2831-x
Zhao D, Lu K, Liu G et al (2018) Design and synthesis of brc analogous peptides and their interactions with a key p53 peptide. FEBS Lett. https://doi.org/10.1002/1873-3468.13256
Zheng JS, Tang S, Qi YK, Wang ZP, Liu L (2013) Chemical synthesis of proteins using peptide hydrazides as thioester surrogates. Nat Protoc 8:2483–2495. https://doi.org/10.1038/nprot.2013.152
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 21572046, 21708005).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no confict of interest.
Research Involving Human Participants and/or Animal
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, Y., Sun, G., Yuan, L. et al. Design and Synthesis of BRCA1 (856-871) Analogous and their Interactions with RAD51 (158-180). Int J Pept Res Ther 27, 1343–1350 (2021). https://doi.org/10.1007/s10989-021-10172-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-021-10172-5