Abstract
Muscle contraction is powered by myosin interaction with actin-based thin filaments containing Ca2+-regulatory proteins, tropomyosin and troponin. Coiled-coil tropomyosin molecules form a long helical strand that winds around actin filament and either shields actin from myosin binding or opens it. Non-canonical residues G126 and D137 in the central part of tropomyosin destabilize its coiled-coil structure. Their substitutions for canonical ones, G126R and D137L, increase structural stability and the velocity of sliding of reconstructed thin filaments along myosin coated surface. The effect of these stabilizing mutations on force of the actin–myosin interaction is unknown. It also remains unclear whether the stabilization affects single actin–myosin interactions or it modifies the cooperativity of the binding of myosin molecules to actin. We used an optical trap to measure the effects of the stabilization on step size, unitary force and duration of the interactions at low and high load and compared the results with those obtained in an in vitro motility assay. We found that significant prolongation of lifetime of the actin–myosin complex under high load observed at high extent of tropomyosin stabilization, i.e. with double mutant, G126R/D137L, correlates with higher force in the motility assay. Also, the higher the extent of stabilization of tropomyosin, the fewer myosin molecules are needed to propel the thin filaments. The data suggest that the effects of the stabilizing mutations in tropomyosin on the myosin interaction with regulated thin filaments are mainly realized via cooperative mechanisms by increasing the size of cooperative unit.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Muscle contraction is driven by myosin interaction with thin filaments composed of F-actin and regulatory proteins, troponin (Tn) and tropomyosin (Tpm), which control formation of crossbridges, i.e. actin–myosin interactions, through the level of free calcium. Tpm molecules are α-helical coiled-coil dimers which bind each other in a head-to-tail manner and form a strand that winds around the actin filament and binds it. The binding of Ca2+ to Tn causes structural changes in the Tpm–Tn complex and enables interaction of myosin molecules with actin (Lehman and Craig 2008; Gordon et al. 2000). The Ca2+ regulation of the actin–myosin interaction is a cooperative process, which includes crossbridge–crossbridge, Tn–Tn and crossbridge–Tn cooperativities (Gordon et al. 2000). According to the three-state model (McKillop and Geeves 1993), in the absence of Ca2+ Tn binds actin and keeps the Tn–Tpm complex in the blocked or B state where Tpm covers myosin binding sites on actin and blocks actomyosin interaction. Upon Ca2+ binding to Tn, the Tn–Tpm complex moves azimuthally with respect to the axis of the actin filament to the closed or C state where actin sites are partially available for myosin binding. When myosin heads bind actin in a strong stereo-specific manner they move the Tn–Tpm strand further to the open or O state.
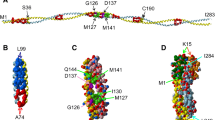
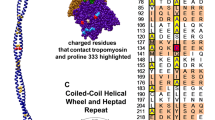
The central part of the Tpm molecule contains conserved non-canonical residues which disrupt the heptad repeat characteristic for a coiled-coil structure, Asp137 (Sumida et al. 2008) and Gly126 (Nevzorov et al. 2011). Replacements of the non-canonical residues with the canonical ones (mutations D137L and G126R) prevent tryptic cleavage of Tpm at the nearby Arg133 (Sumida et al. 2008; Nevzorov et al. 2011; Matyushenko et al. 2014) and result in the increase in thermal stability of the molecule (Nevzorov et al. 2011; Matyushenko et al. 2014, 2015). The D137L and G126R mutations, and especially their combination, G126R/D137L, increase the actin-activated Mg2+-ATPase of myosin (Sumida et al. 2008; Nevzorov et al. 2011; Matyushenko et al. 2014), maximal sliding velocity of regulated thin filaments and calcium sensitivity of the pCa-velocity relationship in an in vitro motility assay (Matyushenko et al. 2014; Shchepkin et al. 2013). The D137L substitution alone or in combination with the G126R one increases the bending stiffness of the thin filament (Nabiev et al. 2015a).
It remains unclear whether changes in the motility caused by the stabilizing substitutions in Tpm result from changes in the interaction of a single myosin molecule with the actin–Tpm–Tn complex as was suggested previously (Shchepkin et al. 2013; Matyushenko et al. 2014) or they are caused by an alteration in the cooperativity of the actin–myosin interaction at an ensemble level. To address this question we, to our knowledge, for the first time applied an optical trap assay (Finer et al. 1994) to study the effect of mutations in the central part of Tpm on the mechanics of interaction of a single myosin molecule with thin filaments and compared the results with force and velocity measurements in the in vitro motility assay.
Materials and methods
Protein preparations
All Tpm species used in this work were recombinant proteins with Ala–Ser N-terminal extension to mimic natural N-terminal acetylation of the native Tpm (Monteiro et al. 1994). Recombinant human Tpm1.1 (α-striated Tpm) C190A, D137L/C190A, G126R/C190A, and D137L/G126R/C190A mutants were prepared as described by Matyushenko et al. (2014, 2015). The C190A Tpm mutant was used as a reference which mimics the reduced state of cysteine in the Tpm molecule, which is typical of living muscle (Lehrer et al. 2011). The C190A mutation allows one to avoid the disulfide cross-linking between the two α-helices of Tpm dimer, which was shown to have dramatic effects on the Tpm domain structure and functional properties (Kremneva et al. 2004; Matyushenko et al. 2017). The C190A substitution had no appreciable effect on the tryptic digestion of Tpm (Sumida et al. 2008) as well as on its thermal unfolding and domain structure (Matyushenko et al. 2015); moreover, regulatory properties of the C190A Tpm mutant (Matyushenko et al. 2014) did not significantly differ from those of wild-type Tpm with SH-groups of Cys190 in fully reduced state (Matyushenko et al. 2017). Therefore this non-Cys control C190A Tpm can be successfully used as a ‘wild-type’ protein in its reduced state.
Actin, myosin, and Tn were prepared from skeletal muscles of the rabbit by standard methods (Pardee and Spudich 1982; Margossian and Lowey 1982; Potter 1982). F-actin polymerized by addition of 2 mM ATP, 4 mM MgCl2 and 100 mM KCl was further labeled by a twofold molar excess of TRITC-phalloidin (Sigma-Aldrich Co. LLC). Myosin modified with N-ethylmaleimide (NEM) was prepared as described by Veigel et al. (1998). Regulated thin filaments were reconstituted from F-actin, Tn, and Tpm according to Gordon et al. (1997).
Optical trap
Characteristics of single interactions of myosin molecule with F-actin or reconstructed thin filament were measured using optical trap setup described earlier (Nabiev et al. 2015b). The setup built on the base of an inverted fluorescent microscope (AxioObserver, Carl Zeiss Microscopy GmbH) enabled measurements in both displacement (Molloy et al. 1995; Knight et al. 2001) and force clamp (Takagi et al. 2006) modes using the three-bead assay (Finer et al. 1994). For that, a dumbbell-like probe was assembled from a fragment of F-actin or reconstructed thin filament (F-actin, a Tpm construct and Tn) attached to two polystyrene beads 1.09 µm diameter (Sigma-Aldrich Co. LLC) coated with NEM-modified myosin. The beads were trapped and held by two independent beams of IR laser (Ventus 1064, 5 W, LaserQuantum) focused by high-aperture objective (100×, NA 1.25 oil immersed Carl Zeiss Microscopy GmbH). In the presence of 10 µM ATP at pCa 4 the dumbbell was brought into contact with a myosin molecule bound to the surface of a 2 µm silica bead (Fluka) used as a pedestal.
In the displacement mode, actin–myosin interactions were detected as individual events (Fig. 1a, b) by an abrupt decrease in the amplitude of Brownian noise of one of the beads caused by an increase in the dumbbell stiffness upon myosin binding. Myosin step size during an event and its duration were measured in the displacement mode as described by Knight et al. (2001). The mean step size, d, was taken as deviation of the bead position caused by attachment of myosin molecule averaged over all events (Molloy et al. 1995).
Typical experimental records of unitary actin–myosin interactions in the optical trap experiments in the displacement (a, b) and force clamp (c–e) modes. a Displacements of the left (black trace) and right (grey trace) beads during the interaction of myosin molecule with reconstructed thin filament containing the double G126R/D137L/C190A Tpm mutant. b A fragment of the record shown by rectangle in a presented on expanded time scale. c Force applied to the motor bead (black trace) and displacement of the sensor bead (grey trace) during the interactions of a filament containing the double G126R/D137L/C190A Tpm mutant with myosin molecules on a pedestal silica bead. When interaction with myosin caused movement of the sensor bead, the beam of the motor bead was moved in opposite direction with the feedback control system trying to return the sensor bead to its initial position. d, e Fragments of the record shown in C by rectangles 1 and 2 respectively on expanded time scale. Fragment shown by rectangle 3 in c was considered as an event of multiple myosin binding to actin
The force bearing capacity of actin–myosin interaction and the duration of unitary interactions under dynamic load were measured using the isometric force clamp mode (Takagi et al. 2006) which enables real-time tracking of the position of unmovable, ‘sensor’, bead and its deviation from zero position caused by working stroke of attached myosin molecule is compensated by movement of the beam of ‘motor’ bead controlled by acousto-optical deflector (Neos Technologies, Inc.) via feedback. The feedback loop gain was adjusted to set the half-time of the loop response to about 10 ms, which was found to be optimal as the smaller values occasionally led the motor bead to escape from the trap during the interaction. Force on the motor bead rises until returning the sensor bead to zero position (Fig. 1d) or myosin detached from actin before the sensor bead was returned to its initial position (Fig. 1e). The static stiffness of the traps was 0.06–0.07 pN/nm in both modes but in the force clamp mode the feedback increased effective stiffness of the dumbbell. Amplitude of pulling force and duration of the interaction events were measured manually. Force achieved by the time of myosin detachment was averaged over all interaction events in each experiment to obtain the mean value, F. The rate constant of the myosin detachment under load was obtained from the distribution of the event durations (Knight et al. 2001; Nabiev et al. 2015b). Average interaction time, t, under load was taken as the inverse rate constant.
Preparation of an experimental flow cell was described in detail (Nabiev et al. 2015a). Briefly, the flow cell assembled from a microscope slide and a coverslip coated with nitrocellulose containing 2 µm silica beads was filled up as follows: 50 µl of 0.1–1 µg/ml myosin in HIS buffer (KCl 500 mM, imidazole 25 mM, MgCl2 4 mM, EGTA 1 mM. and DTT 10 mM, pH 7.5) for 2 min, 50 µl of 0.5 mg/ml BSA in assay buffer (KCl 25 mM, imidazole 25 mM, MgCl2 4 mM, EGTA 1 mM. and DTT 10 mM, pH 7.5) for 1 min, then 50 µl of assay buffer with the oxygen scavenger system (10 mM DTT, 0.2 mg/ml glucose oxidase, 0.05 mg/ml catalase, and 3 mg/ml glucose), 10 µM ATP, 0.5 µl NEM-myosin coated fluorescent polystyrene beads in assay buffer, 4 nM fluorescently labelled F-actin, and as well 100 nM skeletal Tn and 100 nM a Tpm construct when reconstructed thin filaments. Detail procedure of coating the polystyrene beads with NEM-modified myosin and TRITC-BSA was described previously (Nabiev et al. 2015a). The concentration of myosin added into the flow cell was in the range of 0.1–1 µg/ml. Specific myosin concentration was adjusted for each particular Tpm construct so that we were able to find unitary events on nearly every pedestal scanned. Occasionally pulling force changed abruptly (event 3 in Fig. 1c). We considered such events as suspicious for the simultaneous attachment of more than one myosin molecule to a filament and discarded them from further analysis. All experiments with the optical trap were performed at 30 °C.
In vitro motility assay
Experiments in the in vitro motility assay were performed as described previously (Matyushenko et al. 2014; Shchepkin et al. 2010, 2013; Kopylova et al. 2016). In brief, myosin in AB buffer (25 mM KCl, 25 mM imidazole, 4 mM MgCl2, 1 mM EGTA, and 20 mM DTT, pH 7.5) containing 0.5 M KCl was loaded into the experimental flow cell. After 2 min, 0.5 mg/ml BSA was added for 1 min. Further 50 µg/ml of non-labeled F-actin in AB buffer with 2 mM ATP was added for 5 min to block nonfunctional myosin heads. To form regulated thin filaments, 10 nM TRITC-phalloidin labeled F-actin and 100 nM Tpm/Tn were added for 5 min. Unbound filaments were washed out with AB buffer. Finally, the cell was washed with AB buffer containing 0.5 mg/ml BSA, oxygen scavenger system, 20 mM DTT, 2 mM ATP, 0.5% methylcellulose, 100 nM Tpm, 100 nM Tn, and appropriate Ca2+/EGTA in proportions calculated with the Maxchelator program. The experiments were done at 30 °C and pCa 4.
Fluorescently labeled thin filaments in the experimental flow cell were visualized with an inverted epifluorescence microscope Axiovert 200 (Carl Zeiss) equipped with 100×/1.45 oil-immersed alpha Plan-Fluar objective. Movement of thin filaments over myosin coated surface was recorded with EMCCD iXon-897BV (Andor Technology) camera and their sliding velocity was analyzed with GMimPro software (Mashanov and Molloy 2007).
Effect of the Tpm mutants on the force of myosin interaction with regulated thin filaments was assessed using NEM-modified myosin as an external load (Haeberle 1994). The experimental protocol was as that described above except the blocking of nonfunctional myosin heads with non-labeled F-actin was omitted. A mixture of native and NEM-modified myosin in total concentration 100 µg/ml was used in the in vitro motility assay at saturating Ca2+ concentration instead of native myosin. Minimal percentage of NEM-myosin in the mixture at which the filaments stopped moving was taken as a level of isometric force. Criterion of the movement stoppage was the state when only 1 or 2 short (<2 µm long) filaments in the field of view were still moving.
To assess cooperative effects of the Tpm mutants on the actin–myosin interaction, the dependence of the sliding velocity of regulated thin filaments containing the Tpm mutants on the density (c) of myosin on the surface of flow cell was obtained. Surface density of myosin was varied by infusion of different concentrations of myosin in the flow cell. Typically, 30 frames of filament motion in ten different fields in each flow cell were recorded at frame rate 3 s−1. Movement of 5–10 filaments in each field was tracked at least in ten frames, and only movement of filaments moving at a uniform speed (i.e., the standard deviation of the frame-to-frame speed was <0.5 of mean speed) was analyzed. The experiments were repeated three times with each of the Tpm mutants and movement of 50–100 filaments was analyzed in each experiment with each myosin concentration.
The dependence of the sliding velocity V of thin filaments on the concentration of myosin added to the flow cell was fitted with the Hill equation (Brunet et al. 2014) V = V max × c h × (c 50 h + c h)−1 where V max is maximal sliding velocity, c is myosin concentration and c 50 is the concentration required to achieve half-maximal velocity, h is the Hill cooperativity coefficient.
Results
Single molecule measurements
The step size and the average maximal force that single myosin molecule can bear during interaction with reconstructed thin filament containing different Tpm mutants were measured with the optical trap setup at saturating Ca2+ concentration in the presence of 10 µM ATP. Typical records obtained with reconstructed thin filament containing the G126R/D137L/C190A Tpm mutant in the displacement and force modes are shown in Fig. 1.
The results of statistical analysis of all interaction events in two modes of the optical trap operations for F-actin and thin filaments containing Tn and different Tpm mutants are presented in Table 1. None of the Tpm mutants significantly affected the step size d, which was close to that with F-actin alone (Table 1) and was in the range of values observed for the interactions of myosin with F-actin (Tyska and Warshaw 2002). For the thin filaments containing Tpm and Tn, the lifetime of the interactions was longer than that for F-actin alone. The stabilizing substitutions in the Tpm molecule did not induce any significant changes in the step duration (Table 1).
The maximal force that myosin molecule can bear under dynamic load was measured in the isometric force clamp mode. The pulling force achieved by the time of myosin detachment from actin (Fig. 1d, e) was measured for each actomyosin interaction event. Average maximal force of the interactions with regulated thin filaments was somewhat higher than that with F-actin (Table 1) for all Tpm constructs used. As for the lifetime of the actomyosin complex under load, it did not show any certain trend, except that the G126R/D137L/C190A Tpm mutant increased it two-fold longer compared to that with the control C190A Tpm (Table 1, see also Fig. 3a).
In vitro motility experiments
Force-generating capacity of myosin molecules during their interaction with F-actin and with thin filaments containing Tpm mutants was measured in the in vitro motility assay at saturating Ca2+ concentration using NEM-modified myosin as a load (Haeberle 1994). We analyzed the sliding velocity and the fraction of motile filaments (Fig. 2). At increasing concentration of NEM-modified myosin, the sliding velocity and fraction of filaments moving linearly decreased. At any concentration of NEM-myosin no fractioning of the filaments was observed. The correlation coefficients for the linear regression lines in Fig. 2 were in the range from 0.92 to 0.99.
The force, expressed as the fraction of NEM-modified myosin required to stop the filament movement, in case of thin filaments containing C190A Tpm or D137L/C190A Tpm was slightly higher than that with F-actin. With the G126R/D137L/C190A Tpm mutant, force was about twice higher than that observed with either F-actin or thin filaments containing C190A Tpm. With the G126R/C190A Tpm mutant it was lower than that with C190A Tpm or F-actin alone. Force in the in vitro motility assay (Fig. 3b) correlated tightly with the lifetime of single actomyosin complexes under load measured with the same Tpm constructs (Fig. 3a).
a Lifetime of the attached state of myosin to actin in the force clamp mode with F-actin or thin filaments containing Tpm mutants (see Table 1). b Relative force developed by myosin during interaction with thin filaments containing the same Tpm mutants assessed in the in vitro motility assay. Force is expressed as percentage of the NEM-modified myosin in the mixture with the native myosin required to stop the filament movement. The columns and error bars are mean ± SEM of three flow cells. Symbols hash and asterisk indicate significant difference with respect to F-actin and to thin filaments with the control C190A Tpm, respectively; p < 0.05
The cooperativity of the interaction of an ensemble of myosin molecules with thin filaments containing the Tpm mutants was evaluated by the dependence of the sliding velocity on the myosin concentration added into the flow cell at pCa 4 (Fig. 4; Table 2). The presence of regulatory proteins, Tn and any of the Tpm mutant used, increased the velocity at saturating myosin concentrations (Fig. 4). In an agreement with previous reports (Shchepkin et al. 2013; Matyushenko et al. 2014), the maximal velocity of the filaments containing Tpm with stabilizing mutations was 20–50% higher than that with the control C190A Tpm.
Dependence of the sliding velocity of thin filaments containing different Tpm mutants on the concentration of myosin added into the flow cell. Each point represents the mean ± SD value obtained in three different flow cells. The data were fitted with the Hill equation (see “Materials and methods”)
Addition of Tpm and Tn to F-actin caused more than tenfold increase in myosin concentration (c 50) required to provide half-maximal filament velocity (Fig. 4; Table 2). For thin filaments containing Tpm with the D137L/C190A or G126R/C190A mutations, the c 50 value was reduced by a factor of 2–2.5 compared to that for the control C190A Tpm (Fig. 4; Table 2), while for the thin filaments with D137L/G126R/C190A Tpm mutant it was as small as that for F-actin alone and at least 10 times less than that for the control C190A Tpm. All these changes were statistically significant.
Discussion
The most interesting and intriguing part of our findings is the effect of the stabilizing Tpm mutations in the in vitro motility experiments where filament velocity was titrated with myosin concentration in the solution added into the flow cell (Fig. 4; Table 2). Both stabilizing mutations, and especially their combination, dramatically reduced the amount of myosin molecules required to propel reconstructed thin filaments.
As was found previously (Homsher et al. 1996; Gordon et al. 1997; Shchepkin et al. 2013; Matyushenko et al. 2014), the maximal sliding velocity at saturating myosin concentration is the smallest for bare F-actin compared to all reconstructed thin filaments (Fig. 4; Table 2). Any of the stabilizing Tpm mutations, G126R and D137L, and, more profoundly, their combination caused further increase in the maximal sliding velocity with respect to the control C190A Tpm as was observed earlier (Shchepkin et al. 2013; Matyushenko et al. 2014). Possible reasons for this are an increase in the bending stiffness of the Tpm strands (Nabiev et al. 2015a) and a change in the myosin–Tpm interface (Matyushenko et al. 2014), both caused by the stabilizing mutations.
The addition of Tpm and Tn to F-actin shifted myosin concentration necessary for achievement of the half-maximal sliding velocity to higher values (Fig. 4; Table 2). Actin–Tpm filaments also require higher myosin concentration to achieve half-maximal sliding velocity compared to bare actin as was shown by VanBuren et al. (1999). This can be explained as follows. Even at saturating Ca2+ concentration myosin binding sites on actin are in the closed state (McKillop and Geeves 1993). Although the binding of myosin heads becomes possible, it is hampered by the Tpm strand. To switch Tpm further on to the open state, myosin heads should bind actin strongly. Such binding pulls Tpm away from the myosin binding sites on actin and opens them for the binding of neighbor myosin heads (Behrmann et al. 2012). The size of the cooperative unit, i.e. the number of neighbor actin sites which become open upon the strong binding of a head was estimated in vitro using biochemical approach (Geeves and Lehrer 1994) and more recently by direct visualization (Desai et al. 2015). At low surface density of myosin heads, the distance between neighbor heads is longer than the length of the cooperative unit. This reduces the probability of actin–myosin binding and decreases the number of heads per unit length of a filament. The sliding velocity is determined by the pulling force of strongly bound myosin heads, their lifetime and drag force arising from non-cycling or disordered myosin heads, nitrocellulose coating, etc. (Gordon et al. 2000). At low myosin surface density, fewer myosin heads are able to bind a thin filament and pull it, especially if the distance between neighbor heads is longer than the size of cooperative unit. An increase in the unit size facilitates myosin binding even at low surface density and should increase the velocity at non-saturating surface density of myosin. The data shown in Fig. 4 and Table 2 indicate that the stabilizing mutations in Tpm cause a significant change in the myosin density needed to achieve the half-maximal filament velocity.
The question remains whether this effect is caused by changes in the interaction characteristics of the single myosin molecule with a thin filament or it results from a change in the cooperativity of the interaction at the ensemble level, i.e. due to changes in the size of the cooperativity unit. Single molecule data (Fig. 1; Table 1) and those obtained in the in vitro motility assay (Figs. 2, 3) allow us to estimate contribution of each of these factors.
Myosin step size d was the same with all filaments and was in the range of previously found values for whole skeletal myosin (Finer et al. 1994; Molloy et al. 1995; Tyska and Warshaw 2002). The duration t on of myosin interaction with thin filament did not depend on the stabilizing mutations in Tpm but it was always longer than that with F-actin (Table 1). Increase in lifetime, t on, in the presence of Tpm and Tn compared with F-actin was previously observed by Kad et al. (2005) although at pCa >8. This can be explained by the fact that Tpm increases myosin affinity for F-actin (Williams and Greene 1983). As it was shown by cryoelectron microscopy (Behrmann et al. 2012) myosin tightly interacts with actin-bound Tpm and forms a strong ternary complex.
The sliding velocity of bare F-actin at saturating myosin concentration was the smallest among all studied filaments (Fig. 4). The filament sliding velocity in the in vitro motility assay is generally thought to be V ≈ d/t on (Tyska and Warshaw 2002). According to this formula and the data obtained with the optical trap, Tpm should inhibit the velocity of the thin filament compared with the velocity of F-actin, not increase it as observed. This discrepancy can be explained as follows. In the motility assay, there is always some drag force arising from non-cycling or disordered myosin heads, nonspecific interaction with nitrocellulose coating, etc (Gordon et al. 2000). Tpm increases t on and myosin affinity for actin thus promotes involvement of a larger number of myosin heads into movement of thin filament compared to F-actin and facilitates overcoming this drag force resulting in an increase in the sliding velocity at constant load.
Under dynamic load, the force of myosin interaction with the reconstructed thin filaments containing any Tpm mutant was higher than that with F-actin alone, although for the G126R/C190A Tpm mutant the difference was not significant (Table 1). No difference was found in the unitary force F between different Tpm mutants (Table 1). The duration of the myosin interaction with thin filaments containing the G126R/D137R/C190A Tpm mutant was about twice longer than that for the control C190A Tpm and those with any of single stabilizing mutations, D137L/C190A or G126R/C190A.
An interesting finding is that under load, the durations of unitary interactions of myosin molecules with F-actin or with thin filaments containing different Tpm mutants (Fig. 3a) tightly correlates with the driving force in the in vitro motility experiments with the same thin filaments (Fig. 3b). The stabilizing mutations in the central part of Tpm did not induce a significant change in the force of myosin interaction with a thin filament (Table 1). Therefore, the correlation indicates that force developed by an ensemble of myosin molecules in vitro is mainly determined by lifetime of the unitary actin–myosin complex and by the number of myosin heads participating in force generation. A significant increase in force of interaction of myosin with reconstructed thin filaments containing both stabilizing mutations compared to F-actin alone and single Tpm mutants (Fig. 3b) can be explained by an increase in lifetime of the actin-myosin bond under load caused by this double mutation (Fig. 1; Table 1). Earlier we have found that the double G126R/D137L/C190A Tpm mutation leads to a significant increase in the sliding velocity of reconstructed thin filaments in vitro (Matyushenko et al. 2014) and in the bending stiffness of the reconstructed filaments (Nabiev et al. 2015a). The data shown in Fig. 3 demonstrate that the stabilizing mutations in the Tpm molecule similarly affect the load-bearing capacity of actomyosin interaction at the single molecule and ensemble levels.
Detailed mechanistic theory (Metalnikova and Tsaturyan 2013) based on the approach developed by Smith and colleagues (Smith et al. 2003; Smith and Geeves 2003) considers a Tpm strand as an bendable elastic bar. It suggests that an increase in the bending rigidity of the Tpm strand increases the size of cooperative unit. Recently, we have measured the bending stiffness of thin filaments with the same Tpm mutants as used here and found that the D137L/C190A and G126R/D137L/C190A Tpm mutants significantly increased the filament bending stiffness most probably due to an increase in the bending stiffness of Tpm (Nabiev et al. 2015a). Those observations readily explain the data shown in Fig. 4 and Table 2. Indeed, an increase in the Tpm stiffness upon the stabilizing of Tpm, especially with the G126R/D137L/C190A mutant, elongates the cooperative unit on thin filament. Within the unit, the strong binding of a myosin head to actin promotes binding of other heads to neighbor actin monomers. An increase in the length of the cooperative unit facilitates myosin binding, so that the number of myosin heads sufficient for propelling a filament at maximal velocity can be achieved at a lower myosin surface density (Fig. 4; Table 2).
In conclusion, the results of the present work demonstrate that the stabilizing mutations in the central part of Tpm molecule affect the force generation and sliding velocity of thin filament mainly via cooperative activation of the actin–myosin interaction. Although the effect of these mutations appears at the single molecule level, when both stabilizing mutations, G126R and D137L, are introduced into Tpm, it is more significantly expressed at the ensemble level. The data obviously indicate an importance of the cooperativity mechanisms in force generation by muscle and the role of non-canonical amino-acid residues in the Tpm structure in these mechanisms.
Abbreviations
- Tpm:
-
Tropomyosin
- Tn:
-
Troponin
- NEM:
-
N-ethylmaleimide
References
Behrmann E, Müller M, Penczek PA, Mannherz HG, Manstein DJ, Raunser S (2012) Structure of the rigor actin–tropomyosin–myosin complex. Cell 150:327–338
Brunet NM, Chase PB, Mihajlović G, Schoffstall B (2014) Ca2+-regulatory function of the inhibitory peptide region of cardiac troponin I is aided by the C-terminus of cardiac troponin T: effects of familial hypertrophic cardiomyopathy mutations cTnI R145G and cTnT R278C, alone and in combination, on filament sliding. Arch Biochem Biophys 552–553:11–20
Desai R, Geeves MA, Kad NM (2015) Using fluorescent myosin to directly visualize cooperative activation of thin filaments. J Biol Chem 290:1915–1925
Finer JT, Simmons RM, Spudich JA (1994) Single myosin molecule mechanics: piconewton forces and nanometre steps. Nature 368:113–119
Geeves MA, Lehrer SS (1994) Dynamics of the muscle thin filament regulatory switch: the size of the cooperative unit. Biophys J 67:273–282
Gordon AM, LaMadrid MA, Chen Y et al (1997) Calcium regulation of skeletal muscle thin filament motility in vitro. Biophys J 72:1295–1307
Gordon AM, Homsher E, Regnier M (2000) Regulation of contraction in striated muscle. Physiol Rev 80:853–924
Haeberle JR (1994) Calponin decreases the rate of cross-bridge cycling and increases maximum force production by smooth muscle myosin in an in vitro motility assay. J Biol Chem 269:12424–12431
Homsher E, Kim B, Bobkova A, Tobacman LS (1996) Calcium regulation of thin filament movement in an in vitro motility assay. Biophys J 70:1881–1892
Kad NM, Kim S, Warshaw DM et al (2005) Single-myosin crossbridge interactions with actin filaments regulated by troponin–tropomyosin. Proc Natl Acad Sci USA 102:16990–16995
Knight AE, Veigel C, Chambers C, Molloy JE (2001) Analysis of single-molecule mechanical recordings: application to acto-myosin interactions. Prog Biophys Mol Biol 77:45–72
Kopylova G, Nabiev S, Nikitina L et al (2016) The properties of the actin–myosin interaction in the heart muscle depend on the isoforms of myosin but not of α-actin. Biochem Biophys Res Commun 476:648–653
Kremneva E, Boussouf S, Nikolaeva O, Maytum R, Geeves MA, Levitsky DI (2004) Effects of two familial hypertrophic cardiomyopathy mutations in alpha-tropomyosin, Asp175Asn and Glu180Gly, on the thermal unfolding of actin-bound tropomyosin. Biophys J 87:3922–3933
Lehman W, Craig R (2008) Tropomyosin and the steric mechanism of muscle regulation. Adv Exp Med Biol 644:95–109
Lehrer SS, Ly S, Fuchs F (2011) Tropomyosin is in a reduced state in rabbit psoas muscle. J Muscle Res Cell Motil 32:19–21
Margossian SS, Lowey S (1982) Preparation of myosin and its subfragments from rabbit skeletal muscle. Methods Enzymol 85(B):55–71
Mashanov GI, Molloy JE (2007) Automatic detection of single fluorophores in live cells. Biophys J 92:2199–2211
Matyushenko AM, Artemova NV, Shchepkin DV, Kopylova GV, Bershitsky SY, Tsaturyan AK, Sluchanko NN, Levitsky DI (2014) Structural and functional effects of two stabilizing substitutions, D137L and G126R, in the middle part of α-tropomyosin molecule. FEBS J 281:2004–2016
Matyushenko AM, Artemova NV, Sluchanko NN, Levitsky DI (2015) Effects of two stabilizing substitutions, D137L and G126R, in the middle part of α-tropomyosin on the domain structure of its molecule. Biophys Chem 196:77–85
Matyushenko AM, Artemova NV, Shchepkin DV, Kopylova GV, Nabiev SR, Nikitina LV, Levitsky DI, Bershitsky SY (2017) The interchain disulfide cross-linking of tropomyosin alters its regulatory properties and interaction with actin filament. Biochem Biophys Res Commun 482:305–309
McKillop DF, Geeves MA (1993) Regulation of the interaction between actin and myosin subfragment 1: evidence for three states of the thin filament. Biophys J 65:693–701
Metalnikova NA, Tsaturyan AK (2013) A mechanistic model of Ca regulation of thin filaments in cardiac muscle. Biophys J 105:941–950
Molloy JE, Burns JE, Kendrick-Jones J et al (1995) Movement and force produced by a single myosin head. Nature 378:209–212
Monteiro PB, Lataro RC, Ferro JA, Reinach FdeC (1994) Functional α-tropomyosin produced in Escherichia coli. A dipeptide extension can substitute the amino-terminal acetyl group. J Biol Chem 269:10461–10466
Nabiev SR, Ovsyannikov DA, Kopylova GV et al (2015a) Stabilizing of the central part of tropomyosin increases bending stiffness of thin filament. Biophys J 109:373–379
Nabiev SR, Ovsyannikov DA, Tsaturyan AK, Bershitsky SY (2015b) The lifetime of the actomyosin complex in vitro under load corresponding to stretch of contracting muscle. Eur Biophys J 44:457–463
Nevzorov IA, Nikolaeva OP, Kainov YA, Redwood CS, Levitsky DI (2011) Conserved noncanonical residue Gly-126 confers instability to the middle part of the tropomyosin molecule. J Biol Chem 286:15766–15772
Pardee JD, Spudich JA (1982) Purification of muscle actin. Methods Cell Biol 24:271–289
Potter JD (1982) Preparation of troponin and its subunits. Methods Enzymol 85(B):241–263
Shchepkin DV, Kopylova GV, Nikitina LV (2010) Effects of cardiac myosin binding protein-C on the regulation of interaction of cardiac myosin with thin filament in an in vitro motility assay. Biochem Biophys Res Commun 401:159–163
Shchepkin DV, Matyushenko AM, Kopylova GV et al (2013) Stabilization of the central part of tropomyosin molecule alters the Ca2+-sensitivity of actin–myosin interaction. Acta Nat 5:126–129
Smith DA, Geeves MA (2003) Cooperative regulation of myosin–actin interactions by a continuous flexible chain II: actin–tropomyosin–troponin and regulation by calcium. Biophys J 84:3168–3180
Smith DA, Maytum R, Geeves MA (2003) Cooperative regulation of myosin–actin interactions by a continuous flexible chain I: actin–tropomyosin systems. Biophys J 84:3155–3167
Sumida JP, Wu E, Lehrer SS (2008) Conserved Asp-137 imparts flexibility to tropomyosin and affects function. J Biol Chem 283:6728–6734
Takagi Y, Homsher EE, Goldman YE, Shuman H (2006) Force generation in single conventional actomyosin complexes under high dynamic load. Biophys J 90:1295–1307
Tyska MJ, Warshaw DM (2002) The myosin power stroke. Cell Motil Cytoskeleton 51:1–15
VanBuren P, Palmiter KA, Warshaw DM (1999) Tropomyosin directly modulates actomyosin mechanical performance at the level of a single actin filament. Proc Natl Acad Sci USA 96:12488–12493
Veigel C, Bartoo ML, White DC, Sparrow JC, Molloy JE (1998) The stiffness of rabbit skeletal actomyosin cross-bridges determined with an optical tweezers transducer. Biophys J 75:1424–1438
Williams DL Jr, Greene LE (1983) Comparison of the effects of tropomyosin and troponin–tropomyosin on the binding of myosin subfragment 1 to actin. BioChemistry 22:2770–2774
Acknowledgements
The work was supported by Grant 16-14-10199 of Russian Science Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shchepkin, D.V., Nabiev, S.R., Kopylova, G.V. et al. Cooperativity of myosin interaction with thin filaments is enhanced by stabilizing substitutions in tropomyosin. J Muscle Res Cell Motil 38, 183–191 (2017). https://doi.org/10.1007/s10974-017-9472-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10974-017-9472-x