Abstract
An absorptive fungal strain for uranium adsorption which is named Q5, was isolated from the drainage of a uranium mine in South China. The morphological, physiological and phylogenetic characterizations of the strain Q5 were investigated. The results showed that, the strain Q5 identified as Penicillium funiculosum (99 % similarity in gene sequence). Furthermore, the adsorption performance of P. funiculosum was greatly improved by the mutational method that combined hydroxylamine hydrochloride and UV light. The results showed that the adsorption capacity of the mutated P. funiculosum for U(VI) was obviously better than the non-mutational one at pH 2.0–9.0.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Microbial behaviour is extraordinarily diverse in environmental bioremediation system, which mainly involves bioreduction, biosorption and bioaccumulation [1]. One way for uranium bioremediation is to reduce soluble U(VI) to insoluble U(IV), another is to adsorb soluble U(VI) by some vectors [1–3]. The microorganisms for reducing U(VI) to U(IV) have been found in several unrelated phylogenetic groups, such as proteobacteria, Firmicutes, Deinococci and Actinobacteria among others [4, 5]. Most studies have focused on the Fe(III)-reducing bacteria (FRB) such as Geobacter, and the sulfate-reducing bacteria such as Desulfovibrio [6–9]. On the other hand, a vast variety of microorganisms have already been conducted for biosorption of U(VI), such as filamentous fungus [10, 11], saccharomycetes [12], actinomycetes and unicellular bacterium [13], alga [14–16] and so on. Aspergillus, Rhizopus and Penicillium [11, 17, 18] have been found to be the important fungal biosorbents. Penicillium citrinum was found to have adsorption capacity of 127.3 mg uranium per g dry weight of the biomass under low concentration uranium of 50 µg mL−1 at 318 K [11].
In order to increase the adsorption capacity of the biosorbents, modification by physical and chemical agents has been extensively investigated. Selatnia et al. [19] used NaOH-treated bacterial dead Streptomyces rimosus to adsorb Cd2+ from aqueous solution. Ultraviolet (UV) exposure, as a common mutation method, has widely been applied to generate genetic mutation for the improvement of microbial strain. For example, UV mutagenesis in Bacillus subtilis greatly improved its biosorption for heavy metals, such as Cd(II), Hg(II), and Pb(II) [20]. Moreover, Joshi et al. [21] employed random UV mutagenesis approach to improve the biodegradation effect of Pseudomonas sp. for sulfonated azo dye, green HE4B.
Penicillium funiculosum is an absorptive fungal strain for uranium in wastewater. It was recently found that P. funiculosum and SiO2-nanoparticles immobilized P. funiculosum had the maximum biosorption capacity for lead of approximately 1,200.0 and 1,266.7 µmol g−1 respectively at pH 5 [22]. In the present study, an absorptive strain for uranium named P. funiculosum Q5 strain was isolated from uranium mine drainage and its characteristics were investigated. In order to improve the biosorption performance, the method that combined with hydroxylamine hydrochloride (NH2OH·HCl) and UV light were used to mutate the P. funiculosum strain. Also, the effect of the mutation on uranium adsorption performance was researched.
Materials and methods
Isolation and morphology observation
Sludge samples were taken from a uranium mine in South China. The samples were diluted and the suspended particles were left precipitated naturally. The supernatants were performed to isolate microorganisms by spreading on agar PDA solid medium, which was prepared as follows: 200 g peeled potato were chopped into pieces, boiled in 1,000 mL distilled water for 15 min, filtrated and then mixed with 20 g sucrose and 20 g agar. The agar plates were incubated at 30 °C for about 6 days and inspected daily for microbial growth. Single colonies were carefully picked and inoculated on PDA solid medium for at least three times. The morphology of the mycelia was observed using a phase-contrast microscope.
Physiological characterization
In order to figure out optimal medium and its pH for the isolated strain, the conidia was inoculated from solid PDA medium to liquid PDA medium, Martin medium and Czapek’s medium with different pH value respectively. Thereafter, it was cultivated in 250 mL shake flasks with 200 r min−1 at 30 °C. The shape and size of the strain mycelia was observed regularly.
28S rDNA amplification, sequencing and phylogenetic analysis
The mycelia were ground into powder with liquid nitrogen and DNA extraction were performed by using cetyltrimethyl ammonium bromide (CTAB) extraction procedure as described by Jeewon et al. [23]. Amplification of 28S rDNA was performed as described previously [23]. The PCR product was separated by gel electrophoresis on a 1 % agarose gel in Tris–acetate–EDTA (TAE) buffer and analyzed by staining with ethidium bromide (EB) under UV light.
The purified PCR product was sequenced by Sangon Biotechnology, Co., Ltd., (Shanghai, China). The 28S rDNA sequence of the isolated strain was submitted to the GenBank. Sequence identification was estimated by using the BLASTN program. All available subsets of 28S rDNA sequences were selected, analyzed and aligned with CLUSTAL X 1.8. The final phylogenetic tree was generated by MEGA 5.0.
Uranium adsorption experiments
Uranium adsorption experiments were conducted with mid-log phase mycelium with different media and different pH. The mycelia at mid-log phase were filtered, washed three times with deionized water, dried at 60 °C and then ground into particles with diameter less than 180 µm. The adsorption experiments were performed using 40 mg mycelia power suspended in 100 mL uranium solution with initial concentration of about 50 µg mL−1 in 250-mL shake flasks with 200 r min−1 at 30 °C. To determine the adsorption capacity, 2 mL of solution samples were centrifuged and the supernatants were analyzed for uranium concentration. This experiment was carried out at 4 and 8 h of adsorption.
Strain mutagenesis experiments
The spore suspension was dealt with hydroxylamine hydrochloride (NH2OH·HCl, 5 mg mL−1) for 20 min. Then it was radiated at the distance of 30 cm by the UV lamp (15 W) for 10 min. The 100 µL conditioning fluid was taken to PDA plating medium, coated well for good distribution. Then it was cultivated in the incubator at 30 °C until the fungal colony grew strong enough.
The mutated strain was inoculated into liquid PDA nutrient medium and cultivated in shake flasks for 72 h at 30 °C. Mid-log phase mycelia were collected by filtration and washed thoroughly with deionized water. They were dried at 60 °C and ground into particles with diameter less than 180 µm. The variation of functional groups before and after mutation was detected by using Fourier transformed infrared spectrometry (FT-IR), and the biosorption experiments were conducted as described previously.
Analytical methods
The concentration of U(VI) in aqueous solution was determined by Br-padap spectrophotometry metrical method. According to Br-padap and U(VI) form purple complex in the water–acetone medium, the sample was screened by 1,2-trans-cyclohexanediaminetetraacetic acid, buffered by triethanolamine, and assayed for the concentration of U(VI) by using spectrophotometry at wavelength of 578 nm.
FT-IR was applied to detect the changes in functional groups after strain mutation. For the FT-IR assay, the dried samples encapsulated with KBr were detected by using a FT-IR spectrometer.
Results and discussion
Morphological characterization
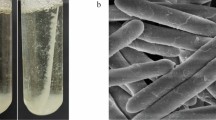
The fungal strain designated Q5 was isolated from the sludge from a uranium mine. After being cultivated on PDA solid medium for 7 days, colonies grew round with their diameter being about 3 cm and flat with concentric rings grain. The mycelia looked like rope and were flocculent. Figure 1 showed that a lot of conidia were generated and they were greyish-green, or blue-green, or black-green.
Figure 2 indicated the microscopic manifestation, from which it could be found that the conidiophore was grown on substrate mycelia or aerial mycelia. Spore stems were superficially smooth, 20–150 µm in length and 2.5–3.2 µm in diameter. Penicillus were lanceolar, mostly double-whorled, and a few single-whorled. There were 4–8 whorled branches on each wheel, and the branches were clung each other, nearly paralleled, 8–11 µm in length and 2.3–3.0 µm in diameter. The conidia were elliptical, superficially smooth, 3.0–3.5 µm in length and 2.0–2.5 µm in diameter.
Physiological characteristics analysis
In order to figure out optimal medium and its pH for the isolated strain, the conidia was inoculated from PDA solid medium to PDA liquid medium, Martin medium and Czapek’s medium with different pH values respectively. As shown in Table 1, the distribution of mycelia was all scattered but not clustered. This indicated that it was not affected by the energy source and pH of the three media. The non-clustering property was probably attributed to the hereditary and physiological characters of the strain Q5. By comparison, the PDA medium and Martin medium were more suitable than the Czapek’s medium for the growth of the strain Q5. The mycelia were plentiful and filled on PDA medium and Martin medium, but they were sparse and low-yielding on the Czapek’s medium. The results showed that the Czapek’s medium was not suitable for the growth of the strain Q5.
Strain identification and phylogenetic analysis
The 28S rRNA gene sequence of the strain Q5 (approximately 565 bp) was submitted to GenBank with the accession number KF830260. Phylogenetic relationships based on rRNA gene sequences are depicted in Fig. 3. The closest relatives of the strain Q5 were P. funiculosum strain NRRL 6417 (GQ221866) and P. funiculosum strain bp seven (HQ876766), both with 99 % sequence similarity. Considering that the morphological, physiological characterizations and also the analysis based on 28S rRNA gene sequence of the strain Q5 are similar to P. funiculosum, the strain Q5 was identified as most closely related to P. funiculosum.
Phylogenic tree based on fragments of rRNA gene sequences (The tree rooted with Q5 is constructed by the neighbor-joining method. The numbers at each clustering node indicate the percentage of bootstrap supporting, and in the brackets after each bacterial name are rDNA accession numbers in GenBank. The scale bar 0.005 indicates evolutionary distance.)
Uranium adsorption performance
The adsorption performance of P. funiculosum strain Q5 were detected by using 100 mL uranium solution (50 µg mL−1 initial uranium concentration; pH 6.0.) at 30 °C in shaking flasks. As shown in Table 2, the adsorption capacity of the strain that was cultivated in the same media with different pH was different. By comparison, the adsorption capacity of the strain cultivated in the PDA media with normal pH (5.86) amounted to 96.7 mg g−1, which was higher than that with other pH values. The adsorption capacity of the strain cultivated in the Martin media (pH 4.05) was as high as 92.4 mg g−1 under faintly acidic cultivation conditions, but it was slightly lower than that of the strain cultivated in the PDA media with normal pH. Therefore, the P. funiculosum Q5 showed a good adsorption performance for uranium, and which cultivated by the PDA media at pH (5.86) had the highest adsorption capacity.
Variation of functional groups after mutation
The infrared spectrograms for the strain Q5 before and after mutation were shown in Fig. 4. The wide and superimposed absorption peaks at 3,390–3,450 cm−1 were strengthened and spread after mutation, which indicated that more association –OH or –NH and functional groups in polysaccharide and protein emerged on the cell surface. The strong peak at the frequency of 1,640 cm−1 represented–CHO stretching vibration or N–H in-plane bending vibration [24]. The peak at 1,100 cm−1 enhanced after mutation possibly attributed to the stretching vibration of C–O bond in –COOH, secondary alcohol or C–N [24], which revealed that the numbers of those groups were enlarged. The crest at 2,083 cm−1 was intensified, it probably owing to the high conjugate of cumulative unsaturated bond after mutation. The strong peak at 1,207 cm−1 appeared, possibly because the vibration of C=C or fatty amine was stretched. Mutagenesis of the strain Q5 was induced by NH2OH·HCl and UV rays. Thereafter, the amounts of hydroxyl, amino and unsaturated bonds on the cell wall were increased. These groups were reported to be capable of complexing with U(VI) [25]. Hence, the mutated P. funiculosum showed better adsorption performances than the unmutated one.
Improvement of uranium adsorption capacity after mutation
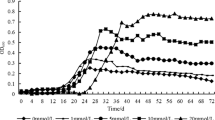
The pH generally influences the activity ionization of functional groups on cell wall and chemical states of heavy metals [25, 26]. Meanwhile, it directly influences the chemical reactions such as hydrolysis, microprecipitation and complexation in organic or inorganic ligands [26, 27]. Considering the impact of pH on the adsorption of P. funiculosum for uranium, a series of adsorption tests were implemented with the pH ranged from 2.0 to 9.0 (adjusted by 0.1–1 mol L−1 HCl and NaOH solutions). As shown in Fig. 5, when pH varied from 2.0 to 6.0, the removal rate of U(VI) by the mutated P. funiculosum increased sharply with pH. The maximum removal was reached at pH 6.0. At this pH, the removal rates were 89.91 and 70.08 % before and after mutagenesis of P. funiculosum, respectively. When pH was less than 4.0, the vast majority of U(VI) existed in the form of UO2 2+ and the adsorption was weak since there was competition between hydrion and uranyl ion. When pH was between 4.0 and 6.0, marginala UO2 2+, (UO2)2(OH) 2+2 , (UO2)3(OH) +5 , (UO2)4(OH) +7 and (UO2)3(OH)7 − coexisted in aqueous solution [24, 28, 29]. When pH was 6.0, (UO2)3(OH) +5 was the main existential state [16]. The observable increase of adsorption efficiency may be defined as the increase of (UO2)3(OH) +5 , because the coulomb power between (UO2)3(OH) +5 and adsorption groups is more than that of simple UO2 2+. Furthermore, the decrease of uranyl ionic charge could result in the decrease of secondary solvation energy [29, 30]. When the pH was above 7.0, a high amount of U(VI) existed in the form of hydrous uranium oxide UO2(OH)2·nH2O and the dissolved solid schoepite 4UO3·9H2O [27], and the precipitates influenced the adsorption process observably. Eliminating the influence of precipitate, the removal rate of U(VI) adsorbed by mutated P. funiculosum was much greater than by P. funiculosum at pH 2.0–9.0, the result proved that mutated P. funiculosum presented a more preferable adsorption for U(VI).
Conclusions
An absorptive fungal strain for uranium adsorption which is named Q5, was isolated and characterized. According to the morphological and physiological characteristics and also the phylogenetic analysis based on 28S rDNA sequence (99 % similarity to P. funiculosum), the strain Q5 was identified as P. funiculosum. The results showed that the strain Q5 cultivated by the PDA media at pH (5.86) had the highest adsorption capacity for U(VI).
Furthermore, the adsorption performance of P. funiculosum for U(VI) was greatly improved by the mutational method that combined hydroxylamine hydrochloride and UV light. The results showed that the adsorption capacity of the mutated P. funiculosum for U(VI) was obviously better than non-mutational one at pH 2.0–9.0.
References
Gadd GM (2010) Metals, minerals and microbes: geomicrobiology and bioremediation. Microbiology 156:609–643
Barlett M, Zhuang K, Mahadevan R, Lovley D (2012) Integrative analysis of Geobacter spp. and sulfate-reducing bacteria during uranium bioremediation. Biogeosciences 9:1033–1040
Williams KH, Bargar JR, Lloyd JR, Lovley DR (2013) Bioremediation of uranium-contaminated groundwater: a systems approach to subsurface biogeochemistry. Curr Opin Biotechnol 24:489–497
Wall JD, Krumholz LR (2006) Uranium reduction. Annu Rev Microbiol 60:149–166
Cardenas E, Wu WM, Leigh MB, Carley J, Carroll S, Gentry T, Luo J, Watson D, Gu B, Ginder-Vogel M, Kitanidis PK, Jardine PM, Zhou J, Criddle CS, Marsh TL, Tiedje JM (2010) Significant association between sulfate-reducing bacteria and uranium-reducing microbial communities as revealed by a combined massively parallel sequencing-indicator species approach. Appl Environ Microbiol 76:6778–6786
Barlett M, Zhuang K, Mahadevan R, Lovley D (2012) Integrative analysis of Geobacter spp. and sulfate-reducing bacteria during uranium bioremediation. Biogeosciences 9:1033–1040
Mtimunye PJ, Chirwa EMN (2014) Characterization of the biochemical-pathway of uranium(VI) reduction in facultative anaerobic bacteria. Chemosphere 113:22–29
Cardenas E, Wu WM, Leigh MB, Carley J, Carroll S, Gentry T, Luo J, Watson D, Gu B, Ginder-Vogel M, Kitanidis PK, Jardine PM, Zhou J, Criddle CS, Marsh TL, Tiedje JM (2008) Microbial communities in contaminated sediments, associated with bioremediation of uranium to submicromolar levels. Appl Environ Microbiol 74:3718–3729
Xu M, Wu WM, Wu L, He Z, Van Nostrand JD, Deng Y, Luo J, Carley J, Ginder-Vogel M, Gentry TJ, Gu B, Watson D, Jardine PM, Marsh TL, Tiedje JM, Hazen T, Criddle CS, Zhou J (2010) Responses of microbial community functional structures to pilot-scale uranium in situ bioremediation. ISME J 4:1060–1070
Akhtar K, Akhtar MW, Khalidc AM (2007) Removal and recovery of uranium from aqueous solutions by Trichoderma harzianum. Water Res 41:1366–1378
Pang C, Liu YH, Cao XH, Li M, Huang GL, Hua R, Wang CX, Liu YT, An XF (2011) Biosorption of uranium(VI) from aqueous solution by dead fungal biomass of Penicillium citrinum. Chem Eng J 170:1–6
Liu MX, Dong FQ, Yan XY, Zeng WM, Hou LY, Pang XF (2010) Biosorption of uranium by Saccharomyces cerevisiae and surface interactions under culture conditions. Bioresour Technol 101:8573–8580
Sar P, Kazy SK, Souza SFD (2004) Radionuclide remediation using a bacterial biosorbent. Int Biodeterior Biodegrad 54:193–202
Bhat SV, Melo JS, Chaugule BB, Souza SFD (2008) Biosorption characteristics of uranium(VI) from aqueous medium onto Catenella repens, a red alga. J Hazard Mater 158:628–635
Khani MH, Keshtkar AR, Ghannadi M, Pahlavanzadeh H (2008) Equilibrium, kinetic and thermodynamic study of the biosorption of uranium onto Cystoseria indica algae. J Hazard Mater 150:612–618
Vogel M, Günther A, Rossberg A, Li B, Bernhard G, Raff J (2010) Biosorption of U(VI) by the green algae Chlorella vulgaris in dependence of pH value and cell activity. Sci Total Environ 409:384–395
Wang J, Hu X, Liu Y, Xie S, Bao Z (2010) Biosorption of uranium(VI) by immobilized Aspergillus fumigatus beads. J Environ Radioact 101:504–508
Wang J, Hu X, Wang J, Bao Z, Xie S, Yang J (2010) The tolerance of Rhizopus arrihizus to U(VI) and biosorption behavior of U(VI) onto R. arrihizus. Biochem Eng J 51:19–23
Selatnia A, Bakhti MZ, Madani A, Kertous L, Mansouri Y (2004) Biosorption of Cd2+ from aqueous solution by a NaOH-treated bacterial dead Streptomyces rimosus biomass. Hydrometallurgy 75:11–24
Wang T, Sun H (2013) Biosorption of heavy metals from aqueous solution by UV-mutant Bacillus subtilis. Environ Sci Pollut Res 20:7450–7463
Joshi SM, Inamdar SA, Jadhav JP, Govindwar SP (2013) Random UV mutagenesis approach for enhanced biodegradation of sulfonated azo dye, green HE4B. Appl Biochem Biotechnol 169:1467–1481
Mahmoud ME, Yakout AA, Abdel-Aal H, Osman MM (2012) High performance SiO2-nanoparticles-immobilized-Penicillium funiculosum for bioaccumulation and solid phase extraction of lead. Bioresour Technol 106:125–132
Jeewon R, Liew ECY, Hyde KD (2002) Phylogenetic relationships of Pestalotiopsis and allied genera inferred from ribosomal DNA sequences and morphological characters. Mol Phylogenet Evol 25:378–392
Gondhalekar SC, Shukla SR (2014) Equilibrium and kinetics study of uranium(VI) from aqueous solution by Citrus limetta peels. J Radioanal Nucl Chem. doi:10.1007/s10967-014-3165-3
Wang J, Chen C (2009) Biosorbents for heavy metals removal and their future. Biotechnol Adv 27:195–226
Rzymski P, Niedzielski P, Karczewski J, Poniedziałek B (2014) Biosorption of toxic metals using freely suspended Microcystis aeruginosa biomass. Cent Eur J Chem 12:1232–1238
Mezaguer M, Kamel N, Lounici H, Kamel Z (2013) Characterization and properties of Pleurotus mutilus fungal biomass as adsorbent of the removal of uranium(VI) from uranium leachate. J Radioanal Nucl Chem 295(1):393–403
Peyvandi S, Faghihian H (2014) Biosorption of uranyl ions from aqueous solution by Saccharomyces cerevisiae cells immobilized on clinoptilolite. J Radioanal Nucl Chem. doi:10.1007/s10967-014-3184-0
Li X, Ding C, Liao J, Lan T, Li F, Zhang D, Yang J, Yang Y, Luo S, Tang J, Liu N (2014) Biosorption of uranium on Bacillus sp. dwc-2: preliminary investigation on mechanism. J Env Radioact 135:6–12
Moghaddam MR, Fatemi S, Keshtkar A (2013) Adsorption of lead (Pb2+) and uranium (UO2 2+) cations by brown algae; experimental and thermodynamic modeling. Chem Eng J 231:294–303
Acknowledgments
This work was supported by the National Natural Science Foundation of China (51274124), Key Project for Fundamental Science of National Defense (B3720132001), the Program of Science and Technology Department of Hunan Province (2010GK2025) and the Program of Scientific Research Foundation of Education Department of Hunan Province (10A103 and 10C1134).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sun, J., Li, Q., Wang, Y. et al. Isolation of a strain of Penicillium funiculosum and mutational improvement for UO2 2+ adsorption. J Radioanal Nucl Chem 303, 427–432 (2015). https://doi.org/10.1007/s10967-014-3389-2
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10967-014-3389-2