Abstract
The occurrence and development of tumors depend on a complex regulation by not only biochemical cues, but also biomechanical factors in tumor microenvironment. With the development of epigenetic theory, the regulation of biomechanical stimulation on tumor progress genetically is not enough to fully illustrate the mechanism of tumorigenesis. However, biomechanical regulation on tumor progress epigenetically is still in its infancy. Therefore, it is particularly important to integrate the existing relevant researches and develop the potential exploration. This work sorted out the existing researches on the regulation of tumor by biomechanical factors through epigenetic means, which contains summarizing the tumor epigenetic regulatory mode by biomechanical factors, exhibiting the influence of epigenetic regulation under mechanical stimulation, illustrating its existing applications, and prospecting the potential. This review aims to display the relevant knowledge through integrating the existing studies on epigenetic regulation in tumorigenesis under mechanical stimulation so as to provide theoretical basis and new ideas for potential follow-up research and clinical applications.
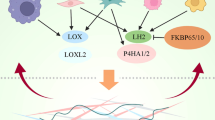
Graphical Abstract
Mechanical factors under physiological conditions stimulate the tumor progress through epigenetic ways, and new strategies are expected to be found with the development of epidrugs and related delivery systems.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The occurrence and development of tumor is a complex process with multi-step. Besides biochemical regulation, the complex regulation of biomechanical factors in tumor microenvironment at the tissue, cell, and even molecular level is one of the factors that determine tumor initiation, malignant development, metastasis, and adhesion, which even show a targeted role in tumor therapy [1]. Because there are many kinds of biomechanical stimuli, such as shear stress, tissue/matrix stiffness, stress/curl effect, and even subcellular nuclear stiffness, the means to realize its regulatory effect are also different. At cell level, the cell/cell or cell/matrix adhesion is modified by the mechanical microenvironment, which makes the cells obtain a more migratory phenotype and ability [2]. At protein level, mechanical stimulation can directly regulate apoptosis related proteins and tumor cell activity [3]. At gene level, biomechanical factors can activate or inactivate proto-oncogenes and regulate tumor initiation [4]. However, these regulatory effects focus on genetic regulation. With the disclosure of epigenetic mechanism, the original regulation mechanism is not enough to fully reveal the mechanism. Studying and summarizing the regulation of biomechanical stimulation on tumor development by epigenetic means will help to enrich the mechanobiological mechanism and provide hope for seeking new therapeutic targets.
Epigenetic regulation is the posttranscriptional regulation of gene expression without interfering with gene transcription. It can be roughly divided into three types: DNA activity regulation, histone modification, and RNA-based regulation [5]. DNA activity can be regulated by methylation/demethylation, acetylation/deacetylation, etc. Histone modification includes histone methylation, acetylation, ubiquitin modification, phosphorylation, isomerization, etc. RNA-based epigenetic regulation can be divided into RNA modification and regulation by special RNA. Methylation usually makes genes lose transcriptional activity, while acetylation activates gene activity or function. As to epigenetic modification based on RNA, the effects on gene activity generally need to be judged according to its type [6].
Although there are many ways of epigenetic regulation, they all likely could be caused by stimulation inside and/or outside cells or tissues. It is reported that some epigenetic regulatory means can be stimulated by mechanical factors, such as DNA methylation and histone modification [7]. Moreover, in the complex mechanical microenvironment, a variety of epigenetic modes also show the potential to regulate cell properties in response to mechanical stimuli. This provides a theoretical basis for the complex tumor mechanical microenvironment to regulate tumor development through epigenetic ways. And based on the existing research, this is partially confirmed. However, the regulation mechanism is not exhaustive and has not been widely concerned because the information or knowledge is too scattered. Therefore, summarizing these regulatory effects can deepen the understanding of this problem.
Having acquired a certain understanding of the mechanism of biomechanical factors regulating tumor development by epigenetic means, it is still an irreplaceable ultimate goal to extend these theories to clinical application, which can improve the outcome of tumor treatment. At present, some epigenetic inhibitors have been developed mainly for different epigenetic modes, such as targeted DNA methylation, acetylation, histone modification, and complex RNA regulation [8]. But this is far from enough, because although a small part of these drugs has been approved by Food and Drug Administration (FDA), most of them are only in clinical trials and the results may not be ideal. What’s more, some inhibitors just stay in the stage of basic research [9]. At the same time, how to accurately and efficiently deliver these drugs to the lesion and how to provide effective intake methods are also major difficulties to be solved. At present, the developed delivery systems have alleviated this problem but they are stretched when it comes to a wide universality or targeted personalized therapy. Therefore, the integration of common epidrugs and delivery systems will help to provide new treatment strategies.
In this review, the existing and potential epigenetic methods, effects, and mechanisms that can stimulate the initiation and development of tumors under biomechanical stimulation are summarized. Then, the applications of epidrugs and their delivery systems are simply reviewed and the scientific research and development directions that may improve the clinical effect are prospected finally. Through this review, the understanding of the mechanism and targeting significance of biomechanical factors regulating tumor development through epigenetic means is intended to be highlighted as well as its clinical potentials.
2 Epigenetic regulation of cancer under biomechanical stimulation
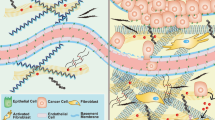
Biomechanical factors play a vital regulatory role in tumorigenesis. There are many ways to achieve it, including regulating the connection between extracellular matrix (ECM) and cells [10], the crosstalk between cells, the transduction of signal molecules between cells [11], the transformation of intracellular signal, and the remodeling of mechanical properties of cells [12]. Mechanistically, besides genetic regulation, epigenetic regulation is indispensable, which refers to model gene expression without disturbing its transcription. In this part, the basic concept and the normal epigenetic regulations are elaborated (Fig. 1) and the process that epigenetic method involved in controlling cell properties by biomechanical factors is summarized based on existing researches.
Normal regulation of epigenetic ways. a Modification of RNA. m6Am methylation, m5C methylation, m1A methylation, amino substitution, hydroxylation, and m6A methylation are listed from left to right. b Modulation of histone. K, lysine; T, threonine; S, serine; R, Arginine. c Epigenetic modification of DNA. DNA methylation, hydroxylation, aldehyde, and carboxylation is shown. d Non-coding RNA discussed in this work including miRNA, lncRNA, and circRNA is shown
2.1 DNA methylation
DNA methylation is a process or an effect that adds a methyl group to the 5-carbon of cytosine residues, which is usually located at the cytosine-phosphate-guanine (CpG) dinucleotide site(s) [13]. The decoration of methylation modifies the conformation and mechanical properties of DNA, which regulates the internal transcription mechanism of cells without altering DNA sequence and usually results to the suppression of gene transcription [14].
There are two modes of methylation regulation under biomechanical stimulation in tumor: One is demethylation within regions of the genome, and another is methylation of special CpG sites [15], during which DNA methyltransferase (DNMT) plays a key role. The methylation changes induced by mechanical factors regulate the activity and proliferation of tumor cells. In an asymmetrical flow field-flow fractionation–based research, the activity changes in DNMT1, but not DNMT3A and DNMT3B, induced by shear stress have been detected recently. Based on the fact that the activity of DNMT1 is further inhibited by combining with a new aptamer, the proliferation of Hela cells and MCF-10A cells were inhibited [16]. DNMT1 is mainly an enzyme for maintenance to ensure the conservation of methylation process during somatic cell division, while DNMT3A and 3B regulate DNA methylation from scratch. In a special mode, DNMT3A, 3B, and nucleosome core particles form a complex with catalytic activity resulting to the interaction with DNA. This function reported previously is involved in mechanical microenvironment stimulated tumor-repopulating cells (TRCs) (cancer stem cell–like cells) selection and further indicates that the complex, especially the core members DNMT3A and DNMT 3B, is sensitive to biomechanical cues [17]. In CpG mode of methylation regulation, although the methylation status of each CpG site under mechanical stimuli is a complex event, the score of cytosine region is an overall evaluation of the compound methylation of multiple cells. As to the regulation, there is usually an inverse relationship between hypermethylation and mRNA expression in the disturbed blood flow region in vivo when CpG is located upstream of the promoter as summarized above, which is conditionally and selectively applied when methylated cytosine is intragenic [18].
To realize the methylated inverse of the transcription, key regulators in biomechanics-driven DNA methylation of cancer cells is necessary. However, only a limited number of key genes/proteins have been reported, which is different with the research on cardiovascular disease [19]. Yes-associated protein (YAP) as a mechanotransducer links the physical and mechanical properties of microenvironment with malignant behavior of tumor cells through the methylation of its promoter. In gastric cancer cells, the interaction between YAP and DNA methylation inhibitors, grainyhead-like transcription factor 2, tet methylcytosine dioxygenase 2 (TET2), and lysine methyltransferase 2A can promote hypomethylation of YAP promoter and rigidity-induced carcinogenic activation of YAP, which could be reversed by softening the matrix stiffness. It indicates that epigenetic reprogramming of biomechanical properties of ECM may be a therapeutic target to preclinical and clinical tumor inhibition [20]. Moreover, Ras association domain family member 1A (RASSF1A) is a tumor suppressor of lung cancer by its promoter methylation and the resulting expression silence [21]. The epigenetic silencing of RASSF1A is due to increased tumor ECM stiffness, and lung cancer cells with RASSF1A promoter methylation showed enhanced expression of prolyl 4-hydroxylase alpha-2 (P4HA2), which increased collagen deposition and ECM rigid. Therefore, a regulatory loop is formed. The higher the methylation degree of RASSF1A, the more ECM deposition, resulting in the higher matrix stiffness of the extracellular microenvironment, which further induces the increase of the methylation degree of CpG, including RASSF1A. However, these can be reversed by re-expressing RASSF1A and inhibiting P4HA2 activity through promoting cancer stem-like cell differentiation [22]. The regulation of RASSF1A inspires that it may be a biomarker related to biomechanical cues in ECM and predicts the risk of cancer.
Considering that only a limited number of studies have reported that mechanical stimulation regulates tumor properties by methylation, it is hypothesized that traditional methylation regulators or genes that can respond to mechanical stimulation may have the potential to explain the mechanobiological mechanism of mechanical stimulation regulating the properties of tumor by methylation. Some of the regulatory factors are fat mass- and obesity-associated protein [23], desmoplakin [24], signal transducer and activator of transcription 3 [25], etc., and more molecules need to be further explored and discovered.
2.2 Histone modification and chromatin remodeling
In the nucleus, chromatin DNA is wrapped around by histone to form a complex unit based on nucleosome, in which about 150 bp of DNA wrapped around the core of each histone octamer. Histone modify could be realized by lysine acetylation, methylation, and serine phosphorylation, because the terminal domain of the histone shows a structure like “tail” [26]. In rare cases, the regulation may be lysine ubiquitination and sulfonation.
2.2.1 Histone acetylation and deacetylation
When exposed to mechanical stimulation, the most dynamic way of histone regulation to disturb DNA hinder and module tumor progress is lysine acetylation regulated by histone acetylases (HAC) or deacetylases (HDAC). HAC or DHAC in normal human cells responding to mechanical stimulation will have the potential to participate in the regulation of mechanical stimulation on tumorigenesis. Males absent on the first (MOF) is a HAC that plays critical roles in liver fibrosis whose transcription activation is regulated be acetylating histone 3 lysine 16 [27]. As is known, liver fibrosis is a cause to hepatoma, in which the stiffness of liver tissue gradually increases with the development of the disease and cause immune escape [28]. Only if MOF can respond to the stiffness changes mentioned above, it may play a regulatory role in the occurrence and development of hepatoma. The effect of matrix stiffness on tumor cells is significantly different from matrix metalloproteins (MMPs) widely existing in cell microenvironment. The matrix stiffness provides physical stimulation, which can promote cancer stem cells to escape and metastasize, while MMPs can affect both intercellular and cell–matrix communication by regulating the activity of many plasma member–anchored and extracellular proteins, thereby favoring cancer growth and metastasis [29]. Besides, mechanical factors modulate tumorigenesis by acetylating or deacetylating tumor-related genes. In human breast tumor, mechanical stimulation and malignant development are caused by the accumulation of type I collagen, in which the expression of type I collagen gene (COL1A1) is activated by myocardin-related transcription factor A (MRTFA). The interaction of MRTF-A with COL1A1 promoter results to the enhancement of COL1A1 histone acetylation and RNA polymerase II recruitment leading to the transcription of COL1A1, which could be inhibited by the absence of MRTF-A [30]. Similarly, breast tumor cells cultured in stiff 3D matrix (2 kPa) exhibit more accessible chromatin sites, and stiff ECM induces tumorigenic phenotypes through boosting stiffness-mediated tumorigenicity by combining of histone deacetylases 3/8 and transcription factor Sp1 [31]. These regulations indicated that (de)acetylation of some tumor-related genes can be activated by mechanical stimulation and become potential targets for precise treatment.
2.2.2 Histone methylation
Another stable modification of histone tail is lysine and arginine methylation, in which histone-N-methyltransferase involved [32]. In the application of mechanical cues that regulates tumor properties, soft matrix (100 Pa) induces the demethylation of histone 3 lysine 9 (H3K9) in TRCs to maintain their own growth [33]. The expression of focal adhesion kinase (FAK) and H3K9 methylation levels is lower in TRCs in soft fibrin matrix than control melanoma cells, and overexpressing FAK enhances H3K9 methylation, which can be restored by the presence of cell division control protein 42 homolog (CDC42) and Ras homolog family member A (RhoA). Furthermore, silencing FAK, CDC42, or RhoA promotes sex determining region Y-box 2 expression, whose suppression reduces growth of TRCs in soft matrices [34]. In one word, the downregulation of H3K9 methylation mediated by soft fibrin matrix through reduction of CDC42 promotes TRC growth.
2.2.3 Other ways of histone modification
In addition, histone modification that regulates tumor progress under mechanical stimulation may also include phosphorylation, ubiquitination, and ribosylation. Histone phosphorylation involving DNA damage refers to the addition of a phosphate group to the four tails of histones, such as in situ histone H3 phosphorylation (ser10). Histone ubiquitination is usually referred to the ubiquitination nucleosome, the basic structural unit of chromatin, which plays a role in the dynamic gene replication and transcription program [35]. Histone glycosylation was first found in H2B region, which was modified by the addition of O-linked N-acetylglucosamine. However, limited by the current research progress, there is no report about the effect of mechanical factors on the properties and malignant development of tumor explained through ways above. Therefore, this is not only the bottlenecks of the existing research, but also the challenges in the future work.
2.3 RNA-based machinery
The RNA-based epigenetic regulation is mainly realized by non-coding RNAs (ncRNA), which are derived from noncoding sequences in the transcriptome and target to newly synthesized mRNA [36]. According to the size, ncRNA can be divided into two prominent types: (i) long non-coding RNA (lncRNA, molecular size > 200 bp), including long intergenic non-coding RNA, circular RNA (circRNA), natural antisense transcripts and enhancer RNA; (ii) small non-coding RNA (sncRNA, molecular size < 200 bp), including microRNA (miRNA) (consists of 20–25 nucleotides) and P-element-induced wimpy testis-interacting RNA. Here, it is reviewed the role of miRNA, lncRNA, and circRNA in the regulation of tumor properties by biomechanical factors.
2.3.1 MiRNA
MiRNAs function as posttranscriptional regulators by typically inhibiting or silencing their target mRNAs through interacting with their 3′-untranslated regions. They are highly conserved assembled by 20–25 nucleotides [37] and powerful in modeling almost all aspects of cancer biology, which are effective in regulating the cancer development by mechanical factors.
In response to the mechanical stimulation of tumor microenvironment, miRNA regulates the properties of tumor cells by directly inhibiting its target genes. It is illustrated by an in vitro compression model that biomechanical compression (about 7.73 kPa)–induced DNMT3A-dependent methylation decreases the expression of miR-9 in MDA-MB-231 and BT-474 cells and cancer-associated fibroblasts, which enhances the progressed phenotype of breast tumor. Furthermore, target genes of miRNA-9 boosts were accompanied with its flop and result in the enhanced expression of vascular endothelial growth factors (VEGF) and the following malignant phenotype [38]. Likewise, there are other kinds of miRNAs that perform similar functions under mechanical stimulation [39]. This regulatory mechanism can highlight the response of miRNA to mechanical stimulation, reflecting the unique mechanical response properties of some miRNAs. Moreover, the use of this property can better involve targeted therapeutic drugs targeting specific miRNAs, for which miRNA could realize its role in regulating tumor cells under mechanical cues by indirect ways playing as a mediator. The gradually increased matrix stiffness coming along with the tumor progression upregulates miRNA-18a through activating integrin, β-catenin, and MYC proto-oncogene (MYC) leading to the reduction of tumor suppressor phosphatase and tensin homolog (PTEN) and homeobox A9 (HOXA9) in luminal breast cancers, which is also proved by clinical trials in human and mouse [40]. This discovery uncovers the mediator role of miRNA-18a in regulating tumor progression when facing biomechanical stimuli, which suggests the importance of exploring the upstream regulatory factors of miRNA. To do that, it is helpful to enrich the regulatory network and deepen the understanding of mechanical factors in regulating tumor. In addition, miRNA can also participate in the regulation of tumor properties by biomechanical factors in the form of complex. The reason is that miRNAs connect each other as an interactive network and the active/inactive of single miRNA leads to quite limited physiological effects, which indicates a narrower therapeutic potential than that expected [41]. It means that the regulation of tumor properties can be modulated by the combination of certain number of miRNAs or miRNA-mRNA. However, whether it is suitable and applicable in biomechanical microenvironment is worthy of attention in the future work.
2.3.2 LncRNA
Thousands of lncRNAs classified in diverse population in mammalian genome were identified at present, whose potential in epigenetic regulation of tumor under mechanical stimulation has caught the eye of countless people. The regulation of lncRNA on cancer contains epithelial-mesenchymal transition (EMT), cell migration, proliferation [42], resistance to anoikis [43], and angiogenesis [44]. Although lacking the coding potential except for minority ones, lncRNAs are well-characterized in gene silence [45].
There are few reports that confirmed the regulation of lncRNA on tumor cell properties in responding to mechanical stimulation, which neither covers all the regulation modes of lncRNA [46]. LncRNA can directly conduct mechanical stimulation in ECM and regulate mechanotransduction factors resulting to the modulation of tumor properties. In the process of perceiving ECM stiffness and transforming it into biochemical signal, the location of lncRNA nuclear parapecker assembly transcript 1 (NEAT1) defines the mechanical sensitivity of U2OS, 143B, and MDA-MB-231 cells. Compared with stiff hydrogel (40 kPa), the NEAT1 aggregation in the nucleus is increased in cells cultured on soft hydrogel (3 kPa) and leads to the activity/inactivity of YAP and MRTF-A, which is accompanied by enhanced migration of cells on stiff hydrogels [47]. This finding suggests that lncRNA may be an important mechanosensor in cancer mechanobiology. To achieve this goal, relevant target genes are also needed. For example, if cellular-mesenchymal epithelial transition factor and caveolin-1 are blocked, LncRNA HOX transcript antisense intergenic RNA (HOTAIR) will be invalid to help escape from invasion and aggression of various cancer types when exposed to fluid shear stress [48, 49]. Meanwhile, the target genes could also be miRNAs. LncRNA cancer susceptibility candidate 11 promotes bladder cancer proliferation through miRNA-150 [50], lncRNA promoter and pre-rRNA antisense aggravate gastric cancer progression through miRNA-188-5p [51], lncRNA is associated with poor prognosis of hepatic cell carcinoma advances of non-small cell lung cancer progression through miRNA-204 [52], etc. However, there is no mechanism of lncRNA regulating tumor cell properties through miRNA in mechanical microenvironment that has been reported. Associating the abovementioned special mechanosensitive miRNA and lncRNA’s mechanosensor functions, it is easy to speculate that lncRNA and miRNA with mechanoresponse capability may have potential in regulating the tumor progression when exposed to mechanical cues, which would fill the gaps in the above mechanism research [53].
3 The effect of biomechanical cues on tumor through epigenetics
Since the regulation of tumor by biomechanical factors is accompanied with the whole process of tumor initiation and development (Fig. 2), the regulation of tumor by biomechanical factors through epigenetic ways may involve multiple levels. Macroscopically, epigenetic regulation has an impact on the initiation and development of tumor tissue and tissue characteristics [54], while microscopically, epigenetic ways can also regulate cell properties and the characteristics of organelles. This part summarizes the regulation of epigenetic regulation on tumor from the tissue level, cell level, and subcellular level.
3.1 Tissue level
There are countless types of mechanical factors that accompany tumor development. The overall stiffness of solid tumors is higher than that of adjacent tissues, which may be due to the abnormal deposition of ECM in tumor tissues [55]. Meanwhile, there is a stiffness gradient from the outside to the inside of the solid tumor, that is, stiff outside and soft inside [56]. At the same time, the regulation of interstitial fluid pressure existing in specific internal region cannot be ignored, which is one of the main causes of tumor heterogeneity through epigenetic ways [57]. In addition, shear stress and surface tension are also mechanical factors that regulate the malignant development of tumor epigenetically [58].
3.1.1 Tissue stiffness regulates tumor progress epigenetically
The difference of stiffness between tumor tissue and surrounding tissue provides a mechanical microenvironment for RNA-based epigenetic regulation. It has been proved that miRNA (as sncRNA) and lncRNA that involved posttranscriptional regulation have emerged in many types of solid tumor. The increase of matrix stiffness in human breast tissue induces high expression of miRNA-18a, which decreases the level of PTEN (a tumor suppressor) and reduces the expression of HOXA9. Based on this, the integrin signal in tumor cells is activated by β-catenin and MYC and then drives tumor progression. Combined with clinical data, the regulation of miRNA-18a is positively correlated with the decreasing of PTEN and HOXA9. Therefore, the boosted tissue mechanics regulates the activation of miRNA dependent PTEN to promote the malignant progression of breast tumor, which also suggests that miRNA-18a, PTEN, and HOXA9 are biomarkers of clinical prognosis for breast tumor [40].
3.1.2 Tissue viscoelasticity regulates tumor progress epigenetically
The mechanical properties of tumor tissue are both elastic and viscous, which determine that its interior is uneven, discontinuous, and anisotropic, so there are often internal and external layers in the structure. In these different layers of structure, the mechanical information is transmitted in different ways. Based on this, tissue viscoelasticity is a key ECM mechanical factor, which regulates tumor gene expression and tumor growth epigenetically by modeling miRNA expression. In breast tumor tissue, the activity of heparanase is regulated by miRNA-1258 directly resulting to the metastasis to brain of tumor cells, in which the key limitation is the impediment provided by the elastic scaffold in breast tumor microenvironment (TME) [59]. The expression of miRNA-1258 is negatively correlated with the expression and activity of heparanase accompanied with the metastasis tendency clinically, which showed a lower expression in nonmalignant tumor or nonmetastatic human mammary tissue. So, the mechanism illustrated above in view of miRNA provides a theoretical basis for the development of miRNA-based therapy for brain metastasis under the influence of TME elasticity and structure [60].
3.1.3 Blood vessel–related biomechanical factors regulate tumor progress epigenetically
The formation of primary tumor and metastasis is inseparable from the supply of nutrients and oxygen by blood vessels. Interstitial fluid pressure with different strengths and fluid shear stresses in blood vessels plays a regulatory role in the angiogenesis and development of blood vessels which are located in the different spatial points of solid tumors [61]. These complex biomechanical factors can regulate tumor development by modeling vascular function, among which epigenetic method stands out. The stiffness of liver tissue in the initial stage of carcinogenesis keeps the high expression of VEGF and protects the angiogenesis effect of VEGF, which leads to the demethylation of the VEGF receptor (VEGFR) resulting to the boost of extracellular VEGF-VEGFR signaling [62, 63]. Consistently, demethylase shows the similar results in hyper-VEGFR-methylation A549 lung cancer cells, which increase the VEGF content and tumor progression [64].
3.1.4 Mechanical properties of tumor-related biomaterials regulate tumor progress epigenetically
Many in vitro tumor models or related biomaterials are designed to research the effects of ECM stiffness on tumor histologically. Patient-derived tumor organoids provide a potentially useful physiological model of tumor tissue [65]. However, traditional organoid cultures that use universal matrix are difficult to adapt to a variety of unique tumor microenvironments [66]. Therefore, modified hydrogel substrates [67], polyacrylamide-gelatin-based microwell with different stiffnesses [68], tumor sphere culture materials with variable stiffness regulated by collagen concentration [69], etc. were developed for further study. From the view of mechanobiological mechanism, the classic mechanotransductive collagen/integrin signal pathway is partially involved in an epigenetic regulation. Collagen in ECM provides mechanical stimulation, with which integrin molecules in the peripheral cells of tumor tissue transmit mechanical signals to cells through their connection. Integrin activation can be eliminated by fibromodulin (FMOD) C-terminal 9-mer wild-type peptide, which is upregulated owning to the lack of methylation of the gene promoter and induces the formation of filamentous actin stress fibers in glioblastoma (GBM) [70]. Upstream of this, mechanosensitive transforming growth factor β (TGF-β) 1 provides the basement for biomechanical regulation [71] and models FMOD expression by epigenetically reconstructing its promoter, which is characterized by the acquisition of active histone markers and promoter demethylation while losing DNMT3A and enhancer of zeste homolog 2. So, the level of FMOD promoter methylation can predict the prognosis of GBM, which is a potential basis for therapeutic intervention [72].
3.1.5 Mechanical property of tumor tissue regulates tumor development–related factors
The epigenetic regulation by biomechanical factors on tumor development also involves other aspects. During the progression of cancer, innate and adaptive immune cells are important components of tumor microenvironment, whose interaction with cancer cells ultimately promotes tumor growth and metastasis [73]. Understanding this crosstalk will help to improve epigenetic treatment methods and propose therapy plans. Moreover, as one of the characteristics of tumor cells is unlimited proliferation, the energy consumption of tumor tissue will be significantly higher than that of adjacent tissues. The metabolism modeled by the stiffness stimulation of tumor tissue and the surrounding tissue through epigenetics intervention will accompany the tumor progression [74]. In addition, cancer-related fibroblast (CAF) is the main component of tumor microenvironment, whose epigenetic regulation directly affects the malignant development of tumor [75]. Besides the secretion regulation of CAF itself, which can provide ECM biofactors for tumor tissue to promote deterioration, the crosstalk of CAF and tumor cells is also mediator affecting tumor progression [76]. The physical force in tumor tissue between tumor cells and CAFs promotes cooperative invasion of both types of cells. Pro-inflammatory cytokines secreted by them, such as leukemia inhibitory factor (LIF) and interleukin-6 (IL-6), mediate the epigenetic modification of CAFs. This enhances the pro-tumorigenic function of CAFs mediated by promoting actomyosin contractility and ECM remodeling to form the tracks used for collective cancer cell migration [77]. A stiff microenvironment can sustain lung fibroblast activation by silencing peroxisome proliferator–activated receptor co-activator-1α (PGC1α) via promoting H3K9 methylation. Fibroblast activation is initiated by a stiff matrix through increasing chromatin accessibility. However, activated myofibroblasts can decrease chromatin accessibility by increasing HDAC activity, which forms a vicious cycle that further contributes to matrix stiffness, while, when faced mechanical stretch, the linker of the nucleoskeleton and cytoskeleton complex controls cell fate by increasing the accessibility of chromatin by transmitting mechanical stretch signals from integrin [78]. When cells in GBM are exposed to mechanical compression, miR-548 family induces a net of epigenetic signaling. The induced signaling was correlated to cell elongation, increased migration, decreased proliferation, and increased tumor aggression characteristics [79]. There may be other ways that biomechanical cues regulating tumor malignant process by epigenetic method; however, the possible regulatory process and mechanobiological mechanism still needs to be explored.
3.2 Cell level
It is a multi-step process from tumor occurrence to multiple organ failures caused by tumor, in which epigenetic regulation is one of the key means for biomechanical factors to regulate the properties of tumor cells involving initiation, growth, morphology deformation, metabolism, migration, drug resistance, reaction to immunity or inflammation, etc. [80]. Understanding the cellular level epigenetic regulatory mechanisms (Fig. 3) can deepen the pathological development of tumor histologically and physiologically, as well as carry out the formulation of precise treatment policies and the development of targeted drugs.
Epigenetic regulatory pathways stimulated by biomechanical factors in tumor cells. Mechanical stimulation is transformed into biochemical signals through intracellular transmission and transduction factors so as to realize the complex regulation of cell properties. Anti-tumor represents the effect of inhibiting tumorigenesis; pro-tumor represents the effect of promoting tumorigenesis. eIF4E, eukaryotic translation initiation factor 4E; EZH2, enhancer of zeste homolog 2; FSS, fluid shear stress; FZD, frizzled; TGFβR, TGF-β receptor; TCF/LEF, T-cell factor/lymphoid enhancer factor; SOX 2, SRY-box 2
3.2.1 Mechanical stimulation regulates tumor initiation
Tumor initiation requires a special subpopulation of cells that experience reprogramming to promote the origination under biomechanical stimulation. When exposed to the stimulation of biomechanics, the conditional dormancy of tumor cells regulated by epigenetic way is an important limiting step of tumor initiation [81]. It is recently illustrated that TRCs could response to mechanical cues and grow immediately when cultured in soft fibrin matrix (90 Pa) but keep dormancy on stiff matrix (1050 Pa) through epigenetic regulation, which is initiated by translocation of CDC42 [82]. In detail, CDC42 is transported into the nucleus and then promotes the transcription of hydroxymethylating enzyme TET2. Mechanically, TET2 motivates the activity of cell cycle–inhibiting genes p27 and p21 and then induces dormancy epigenetically with integrin β3 decreased to maintain this effect. Furthermore, this epigenetic program mediated by matrix stiffness is confirmed in mouse models [83]. Thus, it highlights the potential of epigenetic therapy in tumor treatment by regulating ECM stiffness in tumor initiation stage.
3.2.2 Mechanical stimulation regulates tumor cell proliferate
When tumor cells recover from dormancy, they begin to proliferate and grow. In this process, the ECM is continuously deposited with the increase of the number of cells and the resulting mechanical stimulation regulates the growth of tumor cells through the epigenetic way of methylation. In tumor ECM, histone demethylase jumonji do-main containing 1a (JMJD1a) is downregulated and outflows the nuclear, which results to an epigenetic restriction of breast (MDA-MB-231) and cervical (Hela) carcinoma cell growth. To uncover the molecular mechanism, it is found that JMJD1a positively regulates YAP transcription in a mechanosensitive and stiffness-dependent manner [84]. Besides the mechanical stimulation generated by tumor tissue itself, ECM generated by CAF is often stiffer than that of mammary tissue, of which physical and structural characteristics are also promoters for cancer cell proliferation [85]. Therefore, the growth of tumor cells can be considered as an epigenetic regulatory event regulated by mechanical factors, which also inspires that epigenetic targets are potential biomarkers for inhibiting cell proliferation.
3.2.3 Mechanical stimulation regulates tumor cell morphological changes
Due to the rapid proliferation of tumor cells, the cells gradually undergo morphological changes under the restraint of adjacent tissues (including compression and tension), extravasation, and even hematogenous metastasis, which in most cases lead to cell polarization and malignant phenotype. A significant change of cell morphology is the change of the state and organization of the cytoskeleton composed of actin, which provides awesome opportunities for mechanical factors to regulate cell deformation and skeleton through epigenetic methods and may even produce new cell subtypes [86]. In restraint space, key mechanoresponsive protein non-muscle myosin IIC (MYH14) accumulated under mechanical stress is highly expressed in pancreatic cancer and also makes pancreatic cancer mechanoresponsive, which induces the transformation of globular actin skeleton into cortex and slows down the retrograde flow of actin. Therefore, the correlation of MYH14 to cell polarization indicates that targeting the actin cytoskeleton and other mechanoresponsive proteins is a new strategy to improve the survival rate from inhibiting malignant deformation [87]. In hematogenous metastasis, there is no restraint space, so the morphological changes could be regulated by other ways. It is newly found that the combination of suspension state and shear stress can promote EMT of breast tumor cells through miRNA-29b through TGF-β, which makes the cells obtain tolerance to the adverse environment [88]. It proposes a regulation mechanism of cell deformation under novel stress condition and provides a new idea for tumor treatment.
3.2.4 Mechanical stimulation regulates tumor cell metabolism
In order to provide tumor cells with the energy to realize the changes of above properties, the function changes of mitochondrial function bear the brunt under the specific biomechanical microenvironment. There are potassium channels in the inner membrane of mitochondria, which can regulate mitochondrial membrane potential and intracellular respiration. Membrane tension–derived mechanical stimulation increases the probability of large conductivity calcium activated potassium (mitoBKCa) channel opening in U-87 MG cells. However, its mechanosensitivity is variable. The mitoBKCa gene is epigenetically hypermethylated by DNMT1 and DNMT3b under mechanical stimulation resulting to the accuracy of many splicing subtypes during the splicing process [89]. And only when the subtype derived by stress-regulated exon exists in the controlling unit, the regulatory protein is mechanosensitive [90]. This view preliminarily illustrates that membrane tension can regulate mitochondrial activity through epigenetic splicing and accordingly modulates function of mitochondria and cell metabolism. As mitochondria contains a set of their own DNA, besides the epigenetic regulation of ion channels, the changes of mitochondrial function can also be regulated by other potential epigenetic methods under the stimulation of mechanical factors, such as DNA methylation [91] and ncRNA regulation [92], so these means that there can be the potential regulatory factors for regulating cell metabolism and energy level.
3.2.5 Mechanical stimulation regulates tumor cell migration
After deformation and metabolic adjustment, tumor cells may migrate under the influence of mechanical stimulation. ECM stiffness can not only maintain the integrity of epithelial cells, but it can also trigger tumor cell metastasis in the process of carcinogenesis [93]. In an experimental mouse model, MDA-MB-231 cells expressing inter-alpha-trypsin inhibitor heavy chain 5 (ITIH5) could hardly form lung metastasis. The reason is that ITIH5 regulates integrin mechanical signal in this rare metastatic tumor cells and makes its receptor shift toward integrin β1. This results in the reprograming activation of the promoter regions of tumor suppressor death-associated protein kinase 1 (DAPK1) and the formation of DNA structures with different degrees of methylation, which eventually leads to its high expression [94]. This suggests that the biomechanical stimulation of ECM determines the migration of tumor cells by reprogramming specific gene expression through methylation. Besides, some miRNAs also play a key role in the epigenetic regulation of tumor cell migration when exposed biomechanical stimulation. The microenvironment with compressive solid stress (CSS) endows GBM differential motility and CSS of 23 Pa ensures that LN229 cells reach its migration peak [95]. The following bioinformatics prediction shows that the enhanced migration ability of cancer cells under the modulation of CSS was related to the activation of epigenetic signals. More deeply, miRNA-548 family members are involved in the regulation of CSS induced signal transduction [96].
3.2.6 Mechanical stimulation regulates other properties of tumor cells
In addition, epigenetic regulation of tumor cells by ubiquitous mechanical factors also involves drug resistance and response to inflammation or immunity. Desmosomes confer the resistance to mechanical stimulation of epithelial cells (including epithelial cancer cells), for which desmoplakin (DSP) is down regulated by methylation [97]. Oppositely, overexpression of DSP in H157 cells leads to the decrease of transcription activity of T-cell factor/lymphoid enhancer factor resulting to the declined expressions of axis inhibition protein 2 (AXIN2) and MMP14, thus enhancing the drug resistance to gemcitabine. This indicates that DSP is inactivated by epigenetic mechanism under mechanical stimulation and increases the resistance of cells to anticancer drugs [98]. As for immune regulation, an example is briefly outlined. A microfluidic based “GBM-on-a-chip” microphysiological system verified the heterogeneity of immunosuppression in tumor microenvironment. Different GBM subtypes have different epigenetic and immune characteristics. At the same time, the combination of programmed cell death protein-1 inhibitor nivolumab and colony-stimulating factor 1 receptor inhibitor BLZ945 can promote GBM apoptosis, which provides a way for precise immunotherapy of GBM [99].
3.3 Subcellular
The epigenetic regulation of biomechanical stimulation on cells is not only reflected in the tissue and cell level, but also can be extended to the subcellular level. The nuclear can respond to the stimulation of biomechanics transmitted from cell membrane through the cytoskeleton and modulate its own mechanical properties [100]. This provides a basis for mechanical stimulation to regulate the properties of the nucleus by epigenetic means and also indicates the possibility that the nucleus affects the properties of the cell. The mechanical interaction between nucleus and cell is indispensable in the development of tumor (Fig. 4).
Epigenetic regulatory pathways stimulated by biomechanical factors in nuclear. The mechanical properties of nucleus and cell properties are interrelated. The nucleus not only responds to the mechanical stimulation transmitted through the cytoskeleton, but also affects the overall properties of cells through the changes of its own mechanical properties. KMT, lysine methyltransferase
3.3.1 Epigenetic regulation of nuclear biomechanics
Biomechanical factors regulate the properties of the nucleus epigenetically mainly by the way that transferring mechanical stimulation to the nucleus through the cytoskeleton [101] and resulting in chromatin remodeling or epigenetic modification [102, 103]. Studies show the cytoskeleton effects on chromatin organization and its access to transcription machinery under biomechanical stimulation, whose change of mechanical state is related to the increase of histone acetylation promoting the transcriptional activity of chromatin [104]. By regulating the nuclear shuttling of HDACs, mechanical stress alters gene expression through cytoskeleton, which is impaired by inhibition of actin-myosin contractility [105]. In addition, histone methylation is also involved in the above process. For example, the methylation of histone 3 lysine 4 (H3K4) regulates TGF-β activity in a MRTF-dependent manner, whose silencing reduces disposition of H3K4 and chromatin structure stability [106].
Chromatin is one of the determinants of nuclear stiffness, the role of which could be realized by epigenetic regulation in many ways. For example, exposing to divalent cations can promote the condensation of chromatin that increases the stiffness of nucleus [107], while the use of epigenetic drugs can depolymerize chromatin and then soften nucleus [108]. Similarly, histone tailing and DNA linker induction are also essential for maintaining nuclear rigidity [109]. Specially, in the process of confined migration, chromatin inactivation caused by 3D environment reduces the nuclear stiffness [110].
3.3.2 Effect of nuclear mechanical state on cell properties
The mechanical properties of the nucleus can reversely regulate the properties of the cell epigenetically. Lamin A/C expression in neuroblastoma cells is inhibited which indicates that the nucleus became soft, thus promoting cell proliferation and migration, but exogenous Lmain A/C (encoded by LMNA gene) application inhibits this change [111]. It is found that high-frequency methylation appears in the promoter region of LMNA gene, which contributes to the inactivation of Lamin A/C expression in cells. This shows that the nucleus stiffness can modulate cell properties through methylation [112]. Meanwhile, the drug resistance and cell stiffness could be also regulated by nuclear rigidity. Polyploid giant cancer cells can be found in advanced or post chemotherapy tumor specimens, which show the stemness associated with recurrence [113] and higher nuclear rigidity than primary tumor [114]. Through multi-particle tracking analysis, it is observed that the increase of nuclear stiffness makes the cells more tolerant to mechanical stimulation and more insensitive to paclitaxel (PTX). Mechanistically, the increase of nuclear stiffness improves the actin cytoskeleton contract state, which eventually leads to the increase of cell rigidity and resistance to PTX [115]. The discovery of this biophysical mechanism provides a reliable new idea for the development of epigenetic drugs.
In addition, the regulation of nucleus on cell properties may involve EMT regulation, invasion, growth, etc., but the epigenetic mechanism still needs to be further explored, which provides a theoretical basis for subsequent clinical research and application.
4 Potential for epigenetic treatment of cancer
Having emphasized that biomechanical factors regulate tumorigenesis through epigenetics, these basic research results will hopefully be put into practical application. At present, some tumor drugs targeting epigenetic factors have been developed. The targets include the following: DNA methylation [116], histone acetylation [117], histone methylation [118], special markers [119], RNA modification [120], etc. Although these drugs are not specifically developed for biophysical or biomechanical factors, they can indeed inhibit the epigenetic changes of tumors caused by physical factors by taking alone or in combination. Besides the combination of different epidrugs [121], epidrugs can be combined with radiotherapy, chemotherapy, differentiation agents, and new targeted drugs to treat tumors [122], as well as cytotoxic drugs [123], immunotherapy [124], and multiple combinations [125]. Moreover, some nanoparticle (NP) delivery systems have been developed to meet the needs of clinical treatment [126]. At present, the NPs used mainly include degradable polymer nanoparticles, dendrimers, solid lipid nanoparticles, liposomes, inorganic materials, and natural biomaterials (Fig. 5). It is hoped that the combination of epidrugs and appropriate delivery systems can effectively improve the efficiency of cancer treatment.
Nanoparticles used in epidrug delivery system. a Dendrimers. b Inorganic materials. c Natural biomaterials. d Degradable polymer nanoparticles. e Liposomes. f Solid lipid nanoparticles. NPs are usually used to load epidrugs and target to the lesions by blood injection or in situ injection to inhibit tumor development
5 Conclusions and perspectives
Based on the existing foundation, some outlook should be considered.
-
1.
Map of spatially/timely biomechanical epigenetics. In solid tumors, with the growth of tumors, there are mechanical stimuli such as stiffness gradient acting on tumor cells, which provides a biomechanical microenvironment for epigenetic regulation. Therefore, it reveals an epigenetic regulation change corresponding to the spatial/temporal changes of biomechanical factors, which will help to further explain the regulation of mechanical factors in tumor development through epigenetic pathway.
-
2.
Epidrug loading material with appropriate stiffness. Large or small wounds will be left after tumor surgery. If not treated, the wound will change with the extrusion of surrounding tissues and the biomechanical state of cells here will change, which provides a mechanical basis for the occurrence of epigenetic changes. Therefore, the preparation of epidrug loading materials with suitable stiffness can carry epidrug for wounds and help to alleviate the disorder of epigenetic level.
-
3.
Biomimetic prediction of epigenetically related regulation network. Although the current biomimetic research has been carried out in the field of tumor, it rarely involves epigenetic regulation and let alone mechanical stimulation. Therefore, it will help to reasonably formulate targeted treatment strategies that constructing the epigenetic regulation network of tumor under mechanical stimulation in different stages of tumor development by means of biomimetics. There are many other such research perspectives, which are related to the epigenetic regulation under biomechanical stimulation, and the disclosure of its mechanism will contribute to clinical tumor treatment.
In conclusion, biomechanical factors play an indispensable role in regulating tumorigenesis epigenetically through a variety of modes. Exploring the regulatory mechanism and expanding the application field can develop a practical new approach to cancer treatment. Although further research is needed, the treatment of tumor by biomechanical epigenetic means is expected in the future.
Availability of data and material
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study.
Code availability
Not applicable.
References
Lei, K., Kurum, A., Tang, L.: Mechanical immunoengineering of T cells for therapeutic applications. Acc. Chem. Res. 53, 2777–2790 (2020)
Dong, Y., Zheng, Q., Wang, Z., Lin, X., You, Y., Wu, S., Wang, Y., Hu, C., Xie, X., Chen, J., Gao, D., Zhao, Y., Wu, W., Liu, Y., Ren, Z., Chen, R., Cui, J.: Higher matrix stiffness as an independent initiator triggers epithelial-mesenchymal transition and facilitates HCC metastasis. J. Hematol. Oncol. 12, 112 (2019)
Yao, B., Niu, Y., Li, Y., Chen, T., Wei, X., Liu, Q.: High-matrix-stiffness induces promotion of hepatocellular carcinoma proliferation and suppression of apoptosis via miR-3682-3p-PHLDA1-FAS pathway. J. Cancer 11, 6188–6203 (2020)
Chang, L., Azzolin, L., Di Biagio, D., Zanconato, F., Battilana, G., Lucon Xiccato, R., Aragona, M., Giulitti, S., Panciera, T., Gandin, A., Sigismondo, G., Krijgsveld, J., Fassan, M., Brusatin, G., Cordenonsi, M., Piccolo, S.: The SWI/SNF complex is a mechanoregulated inhibitor of YAP and TAZ. Nature 563, 265–269 (2018)
Dawson, M.A., Kouzarides, T.: Cancer epigenetics: from mechanism to therapy. Cell 150, 12–27 (2012)
Liu, Z., Zhou, Y., Liang, G., Ling, Y., Tan, W., Tan, L., Andrews, R., Zhong, W., Zhang, X., Song, E., Gong, C.: Circular RNA hsa_circ_001783 regulates breast cancer progression via sponging miR-200c-3p. Cell Death. Dis. 10, 55 (2019)
Song, Y., Soto, J., Li, S.: Mechanical regulation of histone modifications and cell plasticity. Curr. Opin. Solid. State. Mater. Sci. 24, 100872 (2020)
Cartron, P.F., Cheray, M., Bretaudeau, L.: Epigenetic protein complexes: the adequate candidates for the use of a new generation of epidrugs in personalized and precision medicine in cancer. Epigenomics 12, 171–177 (2020)
Ghasemi, S.: Cancer’s epigenetic drugs: where are they in the cancer medicines? Pharmacogenomics J. 20, 367–379 (2020)
Wolf, K.J., Shukla, P., Springer, K., Lee, S., Coombes, J.D., Choy, C.J., Kenny, S.J., Xu, K., Kumar, S.: A mode of cell adhesion and migration facilitated by CD44-dependent microtentacles. Proc. Natl. Acad. Sci. USA 117, 11432–11443 (2020)
Patkunarajah, A., Stear, J.H., Moroni, M., Schroeter, L., Blaszkiewicz, J., Tearle, J.L., Cox, C.D., Fürst, C., Sánchez-Carranza, O., Ocaña Fernández, M.D.Á., Fleischer, R., Eravci, M., Weise, C., Martinac, B., Biro, M., Lewin, G.R., Poole, K.: TMEM87a/Elkin1, a component of a novel mechanoelectrical transduction pathway, modulates melanoma adhesion and migration. Elife 9, e53308 (2020)
Park, J.S., Burckhardt, C.J., Lazcano, R., Solis, L.M., Isogai, T., Li, L., Chen, C.S., Gao, B., Minna, J.D., Bachoo, R., DeBerardinis, R.J., Danuser, G.: Mechanical regulation of glycolysis via cytoskeleton architecture. Nature 578, 621–626 (2020)
Fuemmeler, B.F., Dozmorov, M.G., Do, E.K., Zhang, J.J., Grenier, C., Huang, Z., Maguire, R.L., Kollins, S.H., Hoyo, C., Murphy, S.K.: DNA methylation in babies born to nonsmoking mothers exposed to secondhand smoke during pregnancy: an epigenome-wide association study. Environ. Health Perspect. 129, 57010 (2021)
Klemm, S.L., Shipony, Z., Greenleaf, W.J.: Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet. 20, 207–220 (2019)
Traube, F.R., Carell, T.: The chemistries and consequences of DNA and RNA methylation and demethylation. RNA Biol. 14, 1099–1107 (2017)
Wang, L., Lee, J.Y., Gao, L., Yin, J., Duan, Y., Jimenez, L.A., Adkins, G.B., Ren, W., Li, L., Fang, J., Wang, Y., Song, J., Zhong, W.: A DNA aptamer for binding and inhibition of DNA methyltransferase 1. Nucleic Acids Res. 47, 11527–11537 (2019)
Huang, W., Hu, H., Zhang, Q., Wang, N., Yang, X., Guo, A.Y.: Genome-wide DNA methylation enhances stemness in the mechanical selection of tumor-repopulating cells. Front. Bioeng. Biotechnol. 8, 88 (2020)
Salgado, C., Oosting, J., Janssen, B., Kumar, R., Gruis, N., van Doorn, R.: Genome-wide characterization of 5-hydoxymethylcytosine in melanoma reveals major differences with nevus. Genes Chromosom. Cancer 59, 366–374 (2020)
Dunn, J., Qiu, H., Kim, S., Jjingo, D., Hoffman, R., Kim, C.W., Jang, I., Son, D.J., Kim, D., Pan, C., Fan, Y., Jordan, I.K., Jo, H.: Flow-dependent epigenetic DNA methylation regulates gene expression and atherosclerosis. J. Clin. Invest. 124, 3187–3199 (2014)
Jang, M., An, J., Oh, S.W., Lim, J.Y., Kim, J., Choi, J.K., Cheong, J.H., Kim, P.: Matrix stiffness epigenetically regulates the oncogenic activation of the Yes-associated protein in gastric cancer. Nat. Biomed. Eng. 5, 114–123 (2021)
Kawamoto, K., Okino, S.T., Place, R.F., Urakami, S., Hirata, H., Kikuno, N., Kawakami, T., Tanaka, Y., Pookot, D., Chen, Z., Majid, S., Enokida, H., Nakagawa, M., Dahiya, R.: Epigenetic modifications of RASSF1A gene through chromatin remodeling in prostate cancer. Clin. Cancer Res. 27, 2665 (2021)
Pankova, D., Jiang, Y., Chatzifrangkeskou, M., Vendrell, I., Buzzelli, J., Ryan, A., Brown, C., O’Neill, E.: RASSF1A controls tissue stiffness and cancer stem-like cells in lung adenocarcinoma. EMBO J. 38, e100532 (2019)
Shan, K., Zhou, R.M., Xiang, J., Sun, Y.N., Liu, C., Lv, M.W., Xu, J.J.: FTO regulates ocular angiogenesis via m6A-YTHDF2-dependent mechanism. Exp. Eye. Res. 197, 108107 (2020)
Qu, J., Zhu, L., Zhou, Z., Chen, P., Liu, S., Locy, M.L., Thannickal, V.J., Zhou, Y.: Reversing mechanoinductive DSP expression by CRISPR/dCas9-mediated epigenome editing. Am. J. Respir. Crit. Care Med. 198, 599–609 (2018)
Cai, C., Gu, S., Yu, Y., Zhu, Y., Zhang, H., Yuan, B., Shen, L., Yang, B., Feng, X.H.: PRMT5 enables robust STAT3 activation via arginine symmetric dimethylation of SMAD7. Adv. Sci. (Weinh.) 8, 2003047 (2021)
Kragesteen, B.K., Amit, I.: Heads or tails: histone tail clipping regulates macrophage activity. Nat. Immunol. 22, 678–680 (2021)
Zhang, J., Liu, H., Pan, H., Yang, Y., Huang, G., Yang, Y., Zhou, W.P., Pan, Z.Y.: The histone acetyltransferase hMOF suppresses hepatocellular carcinoma growth. Biochem. Biophys. Res. Commun. 452, 575–580 (2014)
Ke, M.Y., Xu, T., Fang, Y., Ye, Y.P., Li, Z.J., Ren, F.G., Lu, S.Y., Zhang, X.F., Wu, R.Q., Lv, Y., Dong, J.: Liver fibrosis promotes immune escape in hepatocellular carcinoma via GOLM1-mediated PD-L1 upregulation. Cancer Lett. 513, 14–25 (2021)
Niland, S., Riscanevo, A.X., Eble, J.A.: Matrix metalloproteinases shape the tumor microenvironment in cancer progression. Int. J. Mol. Sci. 23, 146 (2021)
Meng, C., He, Y., Wei, Z., Lu, Y., Du, F., Ou, G., Wang, N., Luo, X.G., Ma, W., Zhang, T.C., He, H.: MRTF-A mediates the activation of COL1A1 expression stimulated by multiple signaling pathways in human breast cancer cells. Biomed. Pharmacother. 104, 718–728 (2018)
Stowers, R.S., Shcherbina, A., Israeli, J., Gruber, J.J., Chang, J., Nam, S., Rabiee, A., Teruel, M.N., Snyder, M.P., Kundaje, A., Chaudhuri, O.: Matrix stiffness induces a tumorigenic phenotype in mammary epithelium through changes in chromatin accessibility. Nat. Biomed. Eng. 3, 1009–1019 (2019)
Ren, W., Fan, H., Grimm, S.A., Kim, J.J., Li, L., Guo, Y., Petell, C.J., Tan, X.F., Zhang, Z.M., Coan, J.P., Yin, J., Kim, D.I., Gao, L., Cai, L., Khudaverdyan, N., Çetin, B., Patel, D.J., Wang, Y., Cui, Q., Strahl, B.D., Gozani, O., Miller, K.M., O’Leary, S.E., Wade, P.A., Wang, G.G., Song, J.: DNMT1 reads heterochromatic H4K20me3 to reinforce LINE-1 DNA methylation. Nat. Commun. 12, 2490 (2021)
Liu, J., Tan, Y., Zhang, H., Zhang, Y., Xu, P., Chen, J., Poh, Y.C., Tang, K., Wang, N., Huang, B.: Soft fibrin gels promote selection and growth of tumorigenic cells. Nat. Mater. 11, 734–741 (2012)
Tan, Y., Tajik, A., Chen, J., Jia, Q., Chowdhury, F., Wang, L., Chen, J., Zhang, S., Hong, Y., Yi, H., Wu, D.C., Zhang, Y., Wei, F., Poh, Y.C., Seong, J., Singh, R., Lin, L.J., Doğanay, S., Li, Y., Jia, H., Ha, T., Wang, Y., Huang, B., Wang, N.: Matrix softness regulates plasticity of tumour-repopulating cells via H3K9 demethylation and Sox2 expression. Nat. Commun. 5, 4619 (2014)
Zhu, F., Zykova, T.A., Peng, C., Zhang, J., Cho, Y.Y., Zheng, D., Yao, K., Ma, W.Y., Lau, A.T., Bode, A.M., Dong, Z.: Phosphorylation of H2AX at Ser139 and a new phosphorylation site Ser16 by RSK2 decreases H2AX ubiquitination and inhibits cell transformation. Cancer Res. 71, 393–403 (2011)
Yu, Y., Zhang, Y., Chen, X., Chen, Y.: Plant noncoding RNAs: hidden players in development and stress responses. Annu. Rev. Cell Dev. Biol. 35, 407–431 (2019)
Rupaimoole, R., Slack, F.J.: MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat. Rev. Drug Discov. 16, 203–222 (2017)
Kim, B.G., Gao, M.Q., Kang, S., Choi, Y.P., Lee, J.H., Kim, J.E., Han, H.H., Mun, S.G., Cho, N.H.: Mechanical compression induces VEGFA overexpression in breast cancer via DNMT3A-dependent miR-9 downregulation. Cell Death Dis. 8, e2646 (2017)
Northey, J.J., Barrett, A.S., Acerbi, I., Hayward, M.K., Talamantes, S., Dean, I.S., Mouw, J.K., Ponik, S.M., Lakins, J.N., Huang, P.J., Wu, J., Shi, Q., Samson, S., Keely, P.J., Mukhtar, R.A., Liphardt, J.T., Shepherd, J.A., Hwang, E.S., Chen, Y.Y., Hansen, K.C., Littlepage, L.E., Weaver, V.M.: Stiff stroma increases breast cancer risk by inducing the oncogene ZNF217. J. Clin. Invest. 130, 5721–5737 (2020)
Mouw, J.K., Yui, Y., Damiano, L., Bainer, R.O., Lakins, J.N., Acerbi, I., Ou, G., Wijekoon, A.C., Levental, K.R., Gilbert, P.M., Hwang, E.S., Chen, Y.Y., Weaver, V.M.: Tissue mechanics modulate microRNA-dependent PTEN expression to regulate malignant progression. Nat. Med. 20, 360–367 (2014)
Daniel, J.M., Penzkofer, D., Teske, R., Dutzmann, J., Koch, A., Bielenberg, W., Bonauer, A., Boon, R.A., Fischer, A., Bauersachs, J., van Rooij, E., Dimmeler, S., Sedding, D.G.: Inhibition of miR-92a improves re-endothelialization and prevents neointima formation following vascular injury. Cardiovasc. Res. 103, 564–572 (2014)
Lu, W., Zhang, H., Niu, Y., Wu, Y., Sun, W., Li, H., Kong, J., Ding, K., Shen, H.M., Wu, H., Xia, D., Wu, Y.: Long non-coding RNA linc00673 regulated non-small cell lung cancer proliferation, migration, invasion and epithelial mesenchymal transition by sponging miR-150-5p. Mol. Cancer 16, 118 (2017)
Dai, W., Tian, C., Jin, S.: Effect of lncRNA ANRIL silencing on anoikis and cell cycle in human glioma via microRNA-203a. Onco. Targets Ther. 11, 5103–5109 (2018)
Zhang, Q., Li, T., Wang, Z., Kuang, X., Shao, N., Lin, Y.: lncRNA NR2F1-AS1 promotes breast cancer angiogenesis through activating IGF-1/IGF-1R/ERK pathway. J. Cell Mol. Med. 24, 8236–8247 (2020)
Li, X.L., Pongor, L., Tang, W., Das, S., Muys, B.R., Jones, M.F., Lazar, S.B., Dangelmaier, E.A., Hartford, C.C., Grammatikakis, I., Hao, Q., Sun, Q., Schetter, A., Martindale, J.L., Tang, B., Jenkins, L.M., Robles, A.I., Walker, R.L., Ambs, S., Chari, R., Shabalina, S.A., Gorospe, M., Hussain, S.P., Harris, C.C., Meltzer, P.S., Prasanth, K.V., Aladjem, M.I., Andresson, T., Lal, A.: A small protein encoded by a putative lncRNA regulates apoptosis and tumorigenicity in human colorectal cancer cells. Elife 9, e53734 (2020)
Leisegang, M.S., Bibli, S.I., Günther, S., Pflüger-Müller, B., Oo, J.A., Höper, C., Seredinski, S., Yekelchyk, M., Schmitz-Rixen, T., Schürmann, C., Hu, J., Looso, M., Sigala, F., Boon, R.A., Fleming, I., Brandes, R.P.: Pleiotropic effects of laminar flow and statins depend on the Krüppel-like factor-induced lncRNA MANTIS. Eur. Heart J. 40, 2523–2533 (2019)
Todorovski, V., Fox, A.H., Choi, Y.S.: Matrix stiffness-sensitive long noncoding RNA NEAT1 seeded paraspeckles in cancer cells. Mol. Biol. Cell 31, 1654–1662 (2020)
Qu, X., Alsager, S., Zhuo, Y., Shan, B.: HOX transcript antisense RNA (HOTAIR) in cancer. Cancer Lett. 454, 90–97 (2019)
Topel, H., Bagirsakci, E., Comez, D., Bagci, G., Cakan-Akdogan, G., Atabey, N.: lncRNA HOTAIR overexpression induced downregulation of c-Met signaling promotes hybrid epithelial/mesenchymal phenotype in hepatocellular carcinoma cells. Cell Commun. Signal 18, 110 (2020)
Luo, H., Xu, C., Le, W., Ge, B., Wang, T.: lncRNA CASC11 promotes cancer cell proliferation in bladder cancer through miRNA-150. J. Cell Biochem. 120, 13487–13493 (2019)
Shi, X., You, X., Zeng, W.C., Deng, Y.J., Hong, H.L., Huang, O.X., Wang, M.F.: LncRNA PAPAS aggravates the progression of gastric cancer through regulating miRNA-188-5p. Eur. Rev. Med. Pharmacol. Sci. 23, 10761–10768 (2019)
Wu, D., Qin, B.Y., Qi, X.G., Hong, L.L., Zhong, H.B., Huang, J.Y.: LncRNA AWPPH accelerates the progression of non-small cell lung cancer by sponging miRNA-204 to upregulate CDK6. Eur. Rev. Med. Pharmacol. Sci. 24, 4281–4287 (2020)
Olivero, C.E., Martínez-Terroba, E., Zimmer, J., Liao, C., Tesfaye, E., Hooshdaran, N., Schofield, J.A., Bendor, J., Fang, D., Simon, M.D., Zamudio, J.R., Dimitrova, N.: p53 activates the long noncoding RNA Pvt1b to inhibit Myc and suppress tumorigenesis. Mol. Cell 77, 761-774.e8 (2020)
Leal, A., Sidransky, D., Brait, M.: Tissue and cell-free DNA-based epigenomic approaches for cancer detection. Clin. Chem. 66, 105–116 (2020)
Saini, H., Rahmani Eliato, K., Veldhuizen, J., Zare, A., Allam, M., Silva, C., Kratz, A., Truong, D., Mouneimne, G., LaBaer, J., Ros, R., Nikkhah, M.: The role of tumor-stroma interactions on desmoplasia and tumorigenicity within a microengineered 3D platform. Biomaterials 247, 119975 (2020)
Plodinec, M., Loparic, M., Monnier, C.A., Obermann, E.C., Zanetti-Dallenbach, R., Oertle, P., Hyotyla, J.T., Aebi, U., Bentires-Alj, M., Lim, R.Y., Schoenenberger, C.A.: The nanomechanical signature of breast cancer. Nat. Nanotechnol. 7, 757–765 (2012)
Hansem, L.M.K., Huang, R., Wegner, C.S., Simonsen, T.G., Gaustad, J.V., Hauge, A., Rofstad, E.K.: Intratumor heterogeneity in interstitial fluid pressure in cervical and pancreatic carcinoma xenografts. Transl. Oncol. 12, 1079–1085 (2019)
Tomasini, M.D., Rinaldi, C., Tomassone, M.S.: Molecular dynamics simulations of rupture in lipid bilayers. Exp. Biol. Med. (Maywood) 235, 181–188 (2010)
Lemma, E.D., Spagnolo, B., Rizzi, F., Corvaglia, S., Pisanello, M., De Vittorio, M., Pisanello, F.: Microenvironmental stiffness of 3D polymeric structures to study invasive rates of cancer cells. Adv. Health. Mater. 6, 1700888 (2017)
Zhang, L., Sullivan, P.S., Goodman, J.C., Gunaratne, P.H., Marchetti, D.: MicroRNA-1258 suppresses breast cancer brain metastasis by targeting heparanase. Cancer Res. 71, 645–654 (2011)
Bordeleau, F., Mason, B.N., Lollis, E.M., Mazzola, M., Zanotelli, M.R., Somasegar, S., Califano, J.P., Montague, C., LaValley, D.J., Huynh, J., Mencia-Trinchant, N., Negrón Abril, Y.L., Hassane, D.C., Bonassar, L.J., Butcher, J.T., Weiss, R.S., Reinhart-King, C.A.: Matrix stiffening promotes a tumor vasculature phenotype. Proc. Natl. Acad. Sci. USA 114, 492–497 (2017)
Dong, Y., Xie, X., Wang, Z., Hu, C., Zheng, Q., Wang, Y., Chen, R., Xue, T., Chen, J., Gao, D., Wu, W., Ren, Z., Cui, J.: Increasing matrix stiffness upregulates vascular endothelial growth factor expression in hepatocellular carcinoma cells mediated by integrin β1. Biochem. Biophys. Res. Commun. 444, 427–432 (2014)
Barr, M.P., O’Byrne, K.J., Al-Sarraf, N., Gray, S.G.: VEGF-mediated cell survival in non-small-cell lung cancer: implications for epigenetic targeting of VEGF receptors as a therapeutic approach. Epigenomics 7, 897–910 (2015)
Kim, J., Hwang, J., Jeong, H., Song, H.J., Shin, J., Hur, G., Park, Y.W., Lee, S.H., Kim, J.: Promoter methylation status of VEGF receptor genes: a possible epigenetic biomarker to anticipate the efficacy of intracellular-acting VEGF-targeted drugs in cancer cells. Epigenetics 7, 191–200 (2012)
Lai Benjamin, F.L., Lu Rick, X., Hu, Y., Davenport, H.L., Dou, W., Wang, E.Y., Radulovich, N., Tsao, M.S., Sun, Y., Radisic, M.: Recapitulating pancreatic tumor microenvironment through synergistic use of patient organoids and organ-on-a-chip vasculature. Adv. Funct. Mater. 30, 2000545 (2020)
Ng, S., Tan, W.J., Pek, M.M.X., Tan, M.H., Kurisawa, M.: Mechanically and chemically defined hydrogel matrices for patient-derived colorectal tumor organoid culture. Biomaterials 219, 119400 (2019)
Jiang, T., Zhao, J., Yu, S., Mao, Z., Gao, C., Zhu, Y., Mao, C., Zheng, L.: Untangling the response of bone tumor cells and bone forming cells to matrix stiffness and adhesion ligand density by means of hydrogels. Biomaterials 188, 130–143 (2019)
Casey, J., Yue, X., Nguyen, T.D., Acun, A., Zellmer, V.R., Zhang, S., Zorlutuna, P.: 3D hydrogel-based microwell arrays as a tumor microenvironment model to study breast cancer growth. Biomed. Mater. 12, 025009 (2017)
Akkari, L., Haouzi, D., Binamé, F., Floc’h, N., Lassus, P., Baghdiguian, S., Hibner, U.: Cell shape and TGF-beta signaling define the choice of lineage during in vitro differentiation of mouse primary hepatic precursors. J. Cell Physiol. 225, 186–195 (2010)
Tang, X., Xu, P., Wang, B., Luo, J., Fu, R., Huang, K., Dai, L., Lu, J., Cao, G., Peng, H., Zhang, L., Zhang, Z., Chen, Q.: Identification of a specific gene module for predicting prognosis in glioblastoma patients. Front. Oncol. 9, 812 (2019)
Subramanian, A., Kanzaki, L.F., Galloway, J.L., Schilling, T.F.: Mechanical force regulates tendon extracellular matrix organization and tenocyte morphogenesis through TGFbeta signaling. Elife 7, e38069 (2018)
Mondal, B., Patil, V., Shwetha, S.D., Sravani, K., Hegde, A.S., Arivazhagan, A., Santosh, V., Kanduri, M., Somasundaram, K.: Integrative functional genomic analysis identifies epigenetically regulated fibromodulin as an essential gene for glioma cell migration. Oncogene 36, 71–83 (2017)
Medina-Echeverz, J., Hinterberger, M., Testori, M., Geiger, M., Giessel, R., Bathke, B., Kassub, R., Gräbnitz, F., Fiore, G., Wennier, S.T., Chaplin, P., Suter, M., Hochrein, H., Lauterbach, H.: Synergistic cancer immunotherapy combines MVA-CD40L induced innate and adaptive immunity with tumor targeting antibodies. Nat. Commun. 10, 5041 (2019)
Nia, H.T., Liu, H., Seano, G., Datta, M., Jones, D., Rahbari, N., Incio, J., Chauhan, V.P., Jung, K., Martin, J.D., Askoxylakis, V., Padera, T.P., Fukumura, D., Boucher, Y., Hornicek, F.J., Grodzinsky, A.J., Baish, J.W., Munn, L.L., Jain, R.K.: Solid stress and elastic energy as measures of tumour mechanopathology. Nat. Biomed. Eng. 1, 0004 (2016)
Dou, C., Liu, Z., Tu, K., Zhang, H., Chen, C., Yaqoob, U., Wang, Y., Wen, J., van Deursen, J., Sicard, D., Tschumperlin, D., Zou, H., Huang, W.C., Urrutia, R., Shah, V.H., Kang, N.: P300 acetyltransferase mediates stiffness-induced activation of hepatic stellate cells into tumor-promoting myofibroblasts. Gastroenterology 154, 2209-2221.e14 (2018)
Bayer, S.V., Grither, W.R., Brenot, A., Hwang, P.Y., Barcus, C.E., Ernst, M., Pence, P., Walter, C., Pathak, A., Longmore, G.D.: DDR2 controls breast tumor stiffness and metastasis by regulating integrin mediated mechanotransduction in CAFs. Elife 8, e45508 (2019)
Yoshida, G.J.: Regulation of heterogeneous cancer-associated fibroblasts: the molecular pathology of activated signaling pathways. J. Exp. Clin. Cancer Res. 39, 112 (2020)
Wang, C., Yang, J.: Mechanical forces: the missing link between idiopathic pulmonary fibrosis and lung cancer. Eur. J. Cell Biol. 101, 151234 (2022)
Grossen, A., Smith, K., Coulibaly, N., Arbuckle, B., Evans, A., Wilhelm, S., Jones, K., Dunn, I., Towner, R., Wu, D., Kim, Y.T., Battiste, J.: Physical forces in glioblastoma migration: a systematic review. Int. J. Mol. Sci. 23, 4055 (2022)
Veerasubramanian, P.K., Trinh, A., Akhtar, N., Liu, W.F., Downing, T.L.: Biophysical and epigenetic regulation of cancer stemness, invasiveness and immune action. Curr. Tissue Microenviron. Rep. 1, 277–300 (2020)
Cha, S.T., Tan, C.T., Chang, C.C., Chu, C.Y., Lee, W.J., Lin, B.Z., Lin, M.T., Kuo, M.L.: G9a/RelB regulates self-renewal and function of colon-cancer-initiating cells by silencing Let-7b and activating the K-RAS/β-catenin pathway. Nat. Cell Biol. 18, 993–1005 (2016)
Jia, Q., Zhou, W., Yao, W., Yang, F., Zhang, S., Singh, R., Chen, J., Chen, J.J., Zhang, Y., Wei, F., Zhang, Y., Jia, H., Wang, N.: Downregulation of YAP-dependent Nupr1 promotes tumor-repopulating cell growth in soft matrices. Oncogenesis 5, e220 (2016)
Liu, Y., Lv, J., Liang, X., Yin, X., Zhang, L., Chen, D., Jin, X., Fiskesund, R., Tang, K., Ma, J., Zhang, H., Dong, W., Mo, S., Zhang, T., Cheng, F., Zhou, Y., Xie, J., Wang, N., Huang, B.: Fibrin stiffness mediates dormancy of tumor-repopulating cells via a Cdc42-driven Tet2 epigenetic program. Cancer Res. 78, 3926–3937 (2018)
Bertero, T., Oldham, W.M., Grasset, E.M., Bourget, I., Boulter, E., Pisano, S., Hofman, P., Bellvert, F., Meneguzzi, G., Bulavin, D.V., Estrach, S., Feral, C.C., Chan, S.Y., Bozec, A., Gaggioli, C.: Tumor-stroma mechanics coordinate amino acid availability to sustain tumor growth and malignancy. Cell Metab. 29, 124-140.e10 (2019)
Kaukonen, R., Mai, A., Georgiadou, M., Saari, M., De Franceschi, N., Betz, T., Sihto, H., Ventelä, S., Elo, L., Jokitalo, E., Westermarck, J., Kellokumpu-Lehtinen, P.L., Joensuu, H., Grenman, R., Ivaska, J.: Normal stroma suppresses cancer cell proliferation via mechanosensitive regulation of JMJD1a-mediated transcription. Nat. Commun. 7, 12237 (2016)
Sroka, J., von Gunten, M., Dunn, G.A., Keller, H.U.: Phenotype modulation in non-adherent and adherent sublines of Walker carcinosarcoma cells: the role of cell-substratum contacts and microtubules in controlling cell shape, locomotion and cytoskeletal structure. Int. J. Biochem. Cell Biol. 34, 882–899 (2002)
Surcel, A., Schiffhauer, E.S., Thomas, D.G., Zhu, Q., DiNapoli, K.T., Herbig, M., Otto, O., West-Foyle, H., Jacobi, A., Kräter, M., Plak, K., Guck, J., Jaffee, E.M., Iglesias, P.A., Anders, R.A., Robinson, D.N.: Targeting mechanoresponsive proteins in pancreatic cancer: 4-hydroxyacetophenone blocks dissemination and invasion by activating MYH14. Cancer Res. 79, 4665–4678 (2019)
Zhao, B., Lv, Y.: Suspension state and shear stress enhance breast tumor cells EMT through YAP by microRNA-29b. Cell Biol. Toxicol. (2021). https://doi.org/10.1007/s10565-021-09661-6
Zhang, Y., Chen, Y., Xu, Z., Wu, Y., Zhang, Y., Shi, L.: Chronic exercise mediates epigenetic suppression of L-type Ca2+ channel and BKCa channel in mesenteric arteries of hypertensive rats. J. Hypertens. 38, 1763–1776 (2020)
Walewska, A., Kulawiak, B., Szewczyk, A., Koprowski, P.: Mechanosensitivity of mitochondrial large-conductance calcium-activated potassium channels. Biochim. Biophys. Acta Bioenerg. 1859, 797–805 (2018)
Hao, Z., Wu, T., Cui, X., Zhu, P., Tan, C., Dou, X., Hsu, K.W., Lin, Y.T., Peng, P.H., Zhang, L.S., Gao, Y., Hu, L., Sun, H.L., Zhu, A., Liu, J., Wu, K.J., He, C.: N6-deoxyadenosine methylation in mammalian mitochondrial DNA. Mol. Cell 78, 382-395.e8 (2020)
Noh, J.H., Kim, K.M., Abdelmohsen, K., Yoon, J.H., Panda, A.C., Munk, R., Kim, J., Curtis, J., Moad, C.A., Wohler, C.M., Indig, F.E., de Paula, W., Dudekula, D.B., De, S., Piao, Y., Yang, X., Martindale, J.L., de Cabo, R., Gorospe, M.: HuR and GRSF1 modulate the nuclear export and mitochondrial localization of the lncRNA RMRP. Genes Dev. 30, 1224–1239 (2016)
Tian, C., Öhlund, D., Rickelt, S., Lidström, T., Huang, Y., Hao, L., Zhao, R.T., Franklin, O., Bhatia, S.N., Tuveson, D.A., Hynes, R.O.: Cancer cell-derived matrisome proteins promote metastasis in pancreatic ductal adenocarcinoma. Cancer Res. 80, 1461–1474 (2020)
Rose, M., Kloten, V., Noetzel, E., Gola, L., Ehling, J., Heide, T., Meurer, S.K., Gaiko-Shcherbak, A., Sechi, A.S., Huth, S., Weiskirchen, R., Klaas, O., Antonopoulos, W., Lin, Q., Wagner, W., Veeck, J., Gremse, F., Steitz, J., Knüchel, R., Dahl, E.: ITIH5 mediates epigenetic reprogramming of breast cancer cells. Mol. Cancer 16, 44 (2017)
Nia, H.T., Datta, M., Seano, G., Zhang, S., Ho, W.W., Roberge, S., Huang, P., Munn, L.L., Jain, R.K.: In vivo compression and imaging in mouse brain to measure the effects of solid stress. Nat. Protoc. 15, 2321–2340 (2020)
Calhoun, M.A., Cui, Y., Elliott, E.E., Mo, X., Otero, J.J., Winter, J.O.: MicroRNA-mRNA interactions at low levels of compressive solid stress implicate mir-548 in increased glioblastoma cell motility. Sci. Rep. 10, 311 (2020)
Han, X., Jiang, H., Qi, J., Li, J., Yang, J., Tian, Y., Li, W., Jing, Q., Wang, C.: Novel lncRNA UPLA1 mediates tumorigenesis and prognosis in lung adenocarcinoma. Cell Death Dis. 11, 999 (2020)
Yang, L., Chen, Y., Cui, T., Knösel, T., Zhang, Q., Albring, K.F., Huber, O., Petersen, I.: Desmoplakin acts as a tumor suppressor by inhibition of the Wnt/β-catenin signaling pathway in human lung cancer. Carcinogenesis 33, 1863–1870 (2012)
Cui, X., Ma, C., Vasudevaraja, V., Serrano, J., Tong, J., Peng, Y., Delorenzo, M., Shen, G., Frenster, J., Morales, R.T., Qian, W., Tsirigos, A., Chi, A.S., Jain, R., Kurz, S.C., Sulman, E.P., Placantonakis, D.G., Snuderl, M., Chen, W.: Dissecting the immunosuppressive tumor microenvironments in glioblastoma-on-a-chip for optimized PD-1 immunotherapy. eLife 9, e52253 (2020)
Nava, M.M., Miroshnikova, Y.A., Biggs, L.C., Whitefield, D.B., Metge, F., Boucas, J., Vihinen, H., Jokitalo, E., Li, X., García Arcos, J.M., Hoffmann, B., Merkel, R., Niessen, C.M., Dahl, K.N., Wickström, S.A.: Heterochromatin-driven nuclear softening protects the genome against mechanical stress-induced damage. Cell 181, 800-817.e22 (2020)
Wan, Q., Cho, E., Yokota, H., Na, S.: Rac1 and Cdc42 GTPases regulate shear stress-driven β-catenin signaling in osteoblasts. Biochem. Biophys. Res. Commun. 433, 502–507 (2013)
Rabineau, M., Flick, F., Ehlinger, C., Mathieu, E., Duluc, I., Jung, M., Senger, B., Kocgozlu, L., Schaaf, P., Lavalle, P., Freund, J.N., Haikel, Y., Vautier, D.: Chromatin de-condensation by switching substrate elasticity. Sci. Rep. 8, 12655 (2018)
Yang, G., Feng, J., Liu, Y., Zhao, M., Yuan, Y., Yuan, H., Yun, H., Sun, M., Bu, Y., Liu, L., Liu, Z., Niu, J.Q., Yin, M., Song, X., Miao, Z., Lin, Z., Zhang, X.: HAT1 signaling confers to assembly and epigenetic regulation of HBV cccDNA minichromosome. Theranostics 9, 7345–7358 (2019)
Graham, D.M., Burridge, K.: Mechanotransduction and nuclear function. Curr. Opin. Cell Biol. 40, 98–105 (2016)
Rolando, M., Stefani, C., Doye, A., Acosta, M.I., Visvikis, O., Yevick, H.G., Buchrieser, C., Mettouchi, A., Bassereau, P., Lemichez, E.: Contractile actin cables induced by Bacillus anthracis lethal toxin depend on the histone acetylation machinery. Cytoskeleton (Hoboken) 72, 542–556 (2015)
Tian, W., Fan, Z., Li, J., Hao, C., Li, M., Xu, H., Wu, X., Zhou, B., Zhang, L., Fang, M., Xu, Y.: Myocardin-related transcription factor A (MRTF-A) plays an essential role in hepatic stellate cell activation by epigenetically modulating TGF-β signaling. Int. J. Biochem. Cell Biol. 71, 35–43 (2016)
Pajerowski, J.D., Dahl, K.N., Zhong, F.L., Sammak, P.J., Discher, D.E.: Physical plasticity of the nucleus in stem cell differentiation. Proc. Natl. Acad. Sci. U. S. A. 104, 15619–15624 (2007)
Furusawa, T., Rochman, M., Taher, L., Dimitriadis, E.K., Nagashima, K., Anderson, S., Bustin, M.: Chromatin decompaction by the nucleosomal binding protein HMGN5 impairs nuclear sturdiness. Nat. Commun. 6, 6138 (2015)
Shimamoto, Y., Tamura, S., Masumoto, H., Maeshima, K.: Nucleosome-nucleosome interactions via histone tails and linker DNA regulate nuclear rigidity. Mol. Biol. Cell 28, 1580–1589 (2017)
Krause, M., Yang, F.W., Te Lindert, M., Isermann, P., Schepens, J., Maas, R.J.A., Venkataraman, C., Lammerding, J., Madzvamuse, A., Hendriks, W., Te Riet, J., Wolf, K.: Cell migration through three-dimensional confining pores: speed accelerations by deformation and recoil of the nucleus. Philos. Trans. R. Soc. Lond. B Biol. Sci. 374, 20180225 (2019)
Maresca, G., Natoli, M., Nardella, M., Arisi, I., Trisciuoglio, D., Desideri, M., Brandi, R., D’Aguanno, S., Nicotra, M.R., D’Onofrio, M., Urbani, A., Natali, P.G., Del Bufalo, D., Felsani, A., D’Agnano, I.: LMNA knock-down affects differentiation and progression of human neuroblastoma cells. PLoS One 7, e45513 (2012)
Rauschert, I., Aldunate, F., Preussner, J., Arocena-Sutz, M., Peraza, V., Looso, M., Benech, J.C., Agrelo, R.: Promoter hypermethylation as a mechanism for Lamin A/C silencing in a subset of neuroblastoma cells. PLoS One 12, e0175953 (2017)
Niu, N., Mercado-Uribe, I., Liu, J.: Dedifferentiation into blastomere-like cancer stem cells via formation of polyploid giant cancer cells. Oncogene 36, 4887–4900 (2017)
Xuan, B., Ghosh, D., Dawson, M.R.: Contributions of the distinct biophysical phenotype of polyploidal giant cancer cells to cancer progression. Semin. Cancer Biol. 81,64–72 (2022)
Xuan, B., Ghosh, D., Cheney, E.M., Clifton, E.M., Dawson, M.R.: Dysregulation in actin cytoskeletal organization drives increased stiffness and migratory persistence in polyploidal giant cancer cells. Sci. Rep. 8, 11935 (2018)
Crabb, S., Danson, S.J., Catto, J.W.F., McDowell, C., Lowder, J.N., Caddy, J., Dunkley, D., Rajaram, J., Ellis, D., Hill, S., Hathorn, D., Whitehead, A., Kalevras, M., Huddart, R., Griffiths, G.: SPIRE - combining SGI-110 with cisplatin and gemcitabine chemotherapy for solid malignancies including bladder cancer: study protocol for a phase Ib/randomised IIa open label clinical trial. Trials 19, 216 (2018)
Li, X., Jiang, Y., Peterson, Y.K., Xu, T., Himes, R.A., Luo, X., Yin, G., Inks, E.S., Dolloff, N., Halene, S., Chan, S.S.L., Chou, C.J.: Design of hydrazide-bearing HDACIs based on panobinostat and their p53 and FLT3-ITD dependency in antileukemia activity. J. Med. Chem. 63, 5501–5525 (2020)
Fang, Y., Liao, G., Yu, B.: LSD1/KDM1A inhibitors in clinical trials: advances and prospects. J. Hematol. Oncol. 12, 129 (2019)
Pearson, S., Guo, B., Pierce, A., Azadbakht, N., Brazzatti, J.A., Patassini, S., Mulero-Navarro, S., Meyer, S., Flotho, C., Gelb, B.D., Whetton, A.D.: Proteomic analysis of an induced pluripotent stem cell model reveals strategies to treat juvenile myelomonocytic leukemia. J. Proteome. Res. 19, 194–203 (2020)
Adams, D., Polydefkis, M., González-Duarte, A., Wixner, J., Kristen, A.V., Schmidt, H.H., Berk, J.L., Losada López, I.A., Dispenzieri, A., Quan, D., Conceição, I.M., Slama, M.S., Gillmore, J.D., Kyriakides, T., Ajroud-Driss, S., Waddington-Cruz, M., Mezei, M.M., Planté-Bordeneuve, V., Attarian, S., Mauricio, E., Brannagan, T.H. 3rd, Ueda, M., Aldinc, E., Wang, J.J., White, M.T., Vest, J., Berber, E., Sweetser, M.T., Coelho, T., patisiran Global OLE study group.: Long-term safety and efficacy of patisiran for hereditary transthyretin-mediated amyloidosis with polyneuropathy: 12-month results of an open-label extension study. Lancet Neurol. 20, 49–59 (2021)
Ahuja, N., Sharma, A.R., Baylin, S.B.: Epigenetic therapeutics: a new weapon in the war against cancer. Annu. Rev. Med. 67, 73–89 (2016)
Morel, D., Jeffery, D., Aspeslagh, S., Almouzni, G., Postel-Vinay, S.: Combining epigenetic drugs with other therapies for solid tumours - past lessons and future promise. Nat. Rev. Clin. Oncol. 17, 91–107 (2020)
Ferrari, A.C., Alumkal, J.J., Stein, M.N., Taplin, M.E., Babb, J., Barnett, E.S., Gomez-Pinillos, A., Liu, X., Moore, D., DiPaola, R., Beer, T.M.: Epigenetic therapy with panobinostat combined with bicalutamide rechallenge in castration-resistant prostate cancer. Clin. Cancer Res. 25, 52–63 (2019)
Gomez, S., Tabernacki, T., Kobyra, J., Roberts, P., Chiappinelli, K.B.: Combining epigenetic and immune therapy to overcome cancer resistance. Semin. Cancer Biol. 65, 99–113 (2020)
Llopiz, D., Ruiz, M., Villanueva, L., Iglesias, T., Silva, L., Egea, J., Lasarte, J.J., Pivette, P., Trochon-Joseph, V., Vasseur, B., Dixon, G., Sangro, B., Sarobe, P.: Enhanced anti-tumor efficacy of checkpoint inhibitors in combination with the histone deacetylase inhibitor Belinostat in a murine hepatocellular carcinoma model. Cancer Immunol. Immunother. 68, 379–393 (2019)
Jain, V., Kumar, H., Anod, H.V., Chand, P., Gupta, N.V., Dey, S., Kesharwani, S.S.: A review of nanotechnology-based approaches for breast cancer and triple-negative breast cancer. J. Control. Release 326, 628–647 (2020)
Funding
This work was supported by National Natural Science Foundation of China (Grant reference number: 11872134), Natural Science Foundation of Hubei, China (Grant reference number: 2022CFA023), and Natural Science Foundation of Chongqing, China (Grant reference number: cstc2020jcyj-msxmX0035).
Author information
Authors and Affiliations
Contributions
Both authors had the idea for the article, performed the literature search and data analysis and drafted and critically revised the work.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Both authors agree to publish this work.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, B., Lv, Y. A biomechanical view of epigenetic tumor regulation. J Biol Phys 49, 283–307 (2023). https://doi.org/10.1007/s10867-023-09633-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10867-023-09633-3