Abstract
We use elastic neutron scattering to demonstrate that a sharp increase in the mean-squared atomic displacements, commonly observed in hydrated proteins above 200 K and often referred to as the dynamical transition, is present in the hydrated state of both native and denatured lysozyme. A direct comparison of the native and denatured protein thus confirms that the presence of the transition in the mean-squared atomic displacements is not specific to biologically functional molecules.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The temperature dependence of dynamics of all hydrated biomolecules, such as proteins, DNA, and RNA, exhibits an apparent change at 200–230 K, often referred to as a dynamical transition. In a typical measurement, the transition manifests itself as a sharp increase in the mean-squared atomic displacements, < x 2 >, above 200–230 K [1–12]. The origin of the transition has been extensively debated [13–28], and there have been numerous attempts to relate it to the onset of biological activity. Such a connection might seem intuitive because the onset of biological activity and the transition in the mean-squared displacements are both dependent on temperature and hydration level. This hypothesis has become well known [29–32], even though contra arguments [13, 33–35] have been presented.
In this work, we use elastic neutron scattering to demonstrate that the transition in the atomic mean-squared displacements is observed in the hydrated state of both native and irreversibly denatured protein lysozyme, which clearly confirms the view that this transition is not specific to functional biological molecules. This observation suggests that the onset of bioactivity in functional biological molecules does not depend solely on the transition in the mean-squared displacements. The relationship between the onset of anharmonic dynamics and biological activity must be, at best, indirect, as the anharmonicity is apparent even in the intrinsically inactive, irreversibly denatured protein.
2 Material and methods
The labile hydrogen atoms in chicken egg white lysozyme (Sigma Aldrich L4919; 98% purity) were exchanged for deuterium atoms by dissolving in D2O followed by lyophilization. This process was repeated at least twice for each sample. The denatured lysozyme sample was prepared by dissolving the sample at a concentration of 50 mg/ml in 40 mM NaOD and heating it to 353 K for 30 min followed by lyophilization. Subsequently, the denatured sample underwent one additional D2O exchange step. The samples were hydrated using isopiestic conditions by incubation in a sealed container containing 99.9% D2O. The level of hydration was controlled by varying the incubation time. The final hydration levels of the native and denatured lysozyme were 33.6% and 35.5%, respectively. Circular dichroism spectra were recorded on a Jasco 810 CD spectropolarimeter from 190–240 nm at 298 K. Neutron-scattering measurements were performed on the backscattering spectrometer BASIS (Spallation Neutron Source, Oak Ridge National Laboratory, USA) [36] operated in the regime of elastic intensity scan [37]. The scattering momentum transfer range of 0.5 Å − 1 < Q < 1.7 Å − 1 was used. The Q-averaged energy resolution in the experiment was 3.5 μeV; thus, motions on the time scale of about 0.4 ns and faster were probed. Following cooling down to 20 K, the elastic scattering signal was collected at a heating rate of 1 K/min.
3 Results and discussion
As described in Section 1, the overall aim of this work was to investigate if biological activity in functional molecules is a prerequisite for the dynamical transition that is observed at 200–230 K. Lysozyme was chosen for this study because it has been extensively studied and has well-characterized properties. The protein was denatured under acid and alkali conditions by heating to 353 K, followed by cooling to room temperature. The extent of denaturation of the protein was assessed by circular dichroism spectropolarimetry. After denaturation of lysozyme under acidic conditions, natively folded protein could be detected in the soluble fraction of the sample. Conversely, under alkali conditions, no native lysozyme was detected. A comparison of native and alkali-denatured lysozyme is shown in Fig. 1 and demonstrates clearly that the native conformation of the protein had been disrupted. A deconvolution algorithm [38–40] was used to quantitatively estimate the fraction of each type of secondary structure present in lysozyme in its native and denatured state (Table 1). The α-helical content of the denatured protein was greatly reduced while the β-sheet content and amount of unordered polypeptide had increased, compared to its native counterpart (Table 1). This analysis gives good confidence that the protein is in a non-native conformation after incubation in NaOD at 353 K.
The scattering intensity from the samples is dominated by the non-exchangeable H atoms in the protein; the contribution from the D atoms in the protein and hydration water is weaker, though not negligible. The temperature dependence of the mean-squared displacements, averaged over all the atoms in the samples, was estimated using a Gaussian approximation for the elastic intensity [41]:
Here T 0 = 20 K is the lowest temperature point of the elastic scans, at which most of the atomic motions are suppressed. As one can see in Fig. 2, the Gaussian approximation appears to remain satisfactory through the entire Q range of our experiment, even at the highest measured temperature of 300 K. Even though this approximation is typically used for Q values not exceeding 1 Å − 1, it has been suggested to remain applicable to much higher Q values [42].
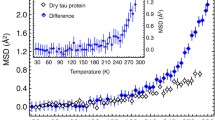
The temperature dependence of the mean-squared displacements for the native lysozyme presented in Fig. 3 is also similar to the previously reported data [7, 10, 12]. Native lysozyme, which possesses a large number of methyl groups, exhibits some anharmonicity (previously attributed to methyl group rotations [7, 10]) above 100 K, followed by the transition and rapidly increasing amplitudes of the atomic displacements above 200 K. Remarkably, denatured lysozyme exhibits a temperature dependence qualitatively similar to that of the native lysozyme, with the transition in the mean-squared atomic displacements clearly present in both states. The difference between the values of the mean-squared displacements for native and denatured lysozyme is rather small, possibly because the large contribution from the methyl groups masks any change of atomic motions that might result from denaturation. The earlier measurement [43] carried out at ambient temperature could not quantify the difference in the mean-squared displacements between native and denatured yeast phosphoglycerate kinase, yielding the same value of (0.42 ± 0.03) Å2 within the experimental error. Nevertheless, in the current temperature-dependent measurement the displacements appear to be systematically larger in the denatured sample. This can be intuitively understood as a result of unfolding of the initially much more compact native protein into a more random-coil-like conformation.
Although we do not suggest that the conformational flexibility that accompanies enhanced atomic dynamics is not important for biological activity [44–46], our data indicate that the onset of the transition in the mean-squared displacement alone is not sufficient for explaining the onset of biological function. It is widely believed that the transition to anharmonicity is driven by the hydration water, which “slaves” the motions of biomolecules. However, the exact mechanism of such “enslavement” has been extensively debated. For example, it has been reported that the transition is coupled to the onset of translational motions in hydration water [47–49]. In addition, it has been debated whether the transition is a real phenomenon, or merely a manifestation of the fact that the measurable atomic dynamics enter the experimentally accessible resolution window above certain temperatures [17, 18, 24, 25, 27]. In such a scenario, the dynamic motions of the biomolecules may increase without any discontinuity. Although the detailed mechanism responsible for the transition remains in question (see Doster [50] for a recent discussion), it should be noted that various hypotheses are not necessarily mutually exclusive. For instance, it has been suggested that hydration water, in general, exhibits a dynamical transition unless the hydration level is too low [51]. Furthermore, the component that gives rise to this transition is physically similar to the translational component in that it requires simultaneous breaking of several hydrogen bonds of a hydration water molecule. However, it has the appearance of a β component because of the localized nature of the motions; on a neutron backscattering spectrometer, this component can be resolved above approximately 200 K [51]. Thus, the first signs of the transition in hydrated biomolecules may coincide with the entrance of this component to the resolution window. At 220–230 K, the hydration water experiences a dynamical transition [20–23] that further increases anharmonicity. Finally, the onset of long-range translational motions of hydration water leads to an even faster increase in anharmonicity at temperatures above 240 K [47–49]. Thus, several different phenomena could contribute to the experimentally observed increase in the mean-squared displacements.
Recent studies of protein dynamics in solutions and solid environments [26, 28] have demonstrated that the internal protein motions are slaved to the β fluctuations of the hydration shell, whereas the large-scale protein motions are slaved to the fluctuations of the bulk solvent. In the framework of this idea, the transition in the mean-squared atomic displacements can be expected for any protein, whether native or denatured, as long as the protein is sufficiently hydrated. Thus, the results of the present study are consistent with the dominant role in the transition of the mean-squared displacements played by the fluctuations of the hydration shell.
References
Parak, F., Formanek, H.: Study on vibration and crystal lattice fault of temperature factor in myoglobin by comparative Mossbauer absorption measurements with X-ray structure data. Acta Crystallogr. A 27, 573–578 (1971)
Keller, H., Debrunner, P.G.: Evidence for conformational and diffusional mean-square displacements in frozen aqueous-solution of oxymyoglobin. Phys. Rev. Lett. 45, 68–71 (1980)
Parak, F., Knapp, E.W., Kucheida, D.: Protein dynamics–Mossbauer–spectroscopy on deoxymyoglobin crystals. J. Mol. Biol. 161, 177–194 (1982)
Doster, W., Cusack, S., Petry, W.: Dynamical transition of myoglobin revealed by inelastic neutron-scattering. Nature 337, 754–756 (1989)
Paciaroni, A., Cinelli, S., Cornicchi, E., De Francesco, A., Onori, G.: Fast fluctuations in protein powders: the role of hydration. Chem. Phys. Lett. 410, 400–403 (2005)
Cornicchi, E., Onori, G., Paciaroni, A.: Picosecond-time-scale fluctuations of proteins in glassy matrices: the role of viscosity. Phys. Rev. Lett. 95, 158104 (2005)
Roh, J.H., Novikov, V.N., Gregory, R.B., Curtis, J.E., Chowdhuri, Z., Sokolov, A.P.: Onsets of anharmonicity in protein dynamics. Phys. Rev. Lett. 95, 038101 (2005)
Roh, J.H., Curtis, J.E., Azzam, S., Novikov, V.N., Peral, I., Chowdhuri, Z., Gregory, R.B., Sokolov, A.P.: Influence of hydration on the dynamics of lysozyme. Biophys. J. 91, 2573–2588 (2006)
Cornicchi, E., Marconi, M., Onori, G., Paciaroni, A.: Controlling the protein dynamical transition with sugar-based bioprotectant matrices: a neutron scattering study. Biophys. J. 91, 289–297 (2006)
Caliskan, G., Briber, R.M., Thirumalai, D., Garcia-Sakai, V., Woodson, S.A., Sokolov, A.P.: Dynamic transition in tRNA is solvent induced. J. Am. Chem. Soc. 128, 32–33 (2006)
Cornicchi, E., Capponi, S., Marconi, M., Onori, G., Paciaroni, A.: Temperature dependence of fast fluctuations in single- and double-stranded DNA molecules: a neutron scattering investigation. Philos. Mag. 87, 509–515 (2007)
Roh, J.H., Briber, R.M., Damjanovic, A., Thirumalai, D., Woodson, S.A., Sokolov, A.P.: Dynamics of tRNA at different levels of hydration. Biophys. J. 96, 2755–2762 (2009)
Daniel, R.M., Smith, J.C., Ferrand, M., Hery, S., Dunn, R., Finney, J.L.: Enzyme activity below the dynamical transition at 220 K. Biophys. J. 75, 2504–2507 (1998)
Daniel, R.M., Finney, J.L., Reat, V., Dunn, R., Ferrand, M., Smith, J.C.: Enzyme dynamics and activity: time-scale dependence of dynamical transitions in glutamate dehydrogenase solution. Biophys. J. 77, 2184–2190 (1999)
Sokolov, A.P., Grimm, H., Kisliuk, A., Dianoux, A.J.: Slow relaxation process in DNA. J. Biol. Phys. 27, 313–327 (2001)
Fenimore, P.W., Frauenfelder, H., McMahon, B.H., Parak, F.G.: Slaving: Solvent fluctuations dominate protein dynamics and functions. Proc. Natl. Acad. Sci. USA 99, 16047–16051 (2002)
Daniel, R.M., Finney, J.M., Smith, J.C.: The dynamic transition in proteins may have a simple explanation. Faraday Discuss. 122, 163–169 (2002)
Becker, T., Hayward, J.A., Finney, J.L., Daniel, R.M., Smith, J.C.: Neutron frequency windows and the protein dynamical transition. Biophys. J. 87, 1436–1444 (2004)
Fenimore, P.W., Frauenfelder, H., McMahon, B.H., Young, R.D.: Bulk-solvent and hydration-shell fluctuations, similar to alpha- and beta-fluctuations in glasses, control protein motions and functions. Proc. Natl. Acad. Sci. USA 101, 14408–14413 (2004)
Chen, S.H., Liu, L., Fratini, E., Baglioni, P., Faraone, A., Mamontov, E.: Observation of fragile-to-strong dynamic crossover in protein hydration water. Proc. Natl. Acad. Sci. USA 103, 9012–9016 (2006)
Kumar, P., Yan, Z., Xu, L., Mazza, M.G., Buldyrev, S.V., Chen, S.H., Sastry, S., Stanley, H.E.: Glass transition in biomolecules and the liquid-liquid critical point of water. Phys. Rev. Lett. 97, 177802 (2006)
Chen, S.H., Liu, L., Chu, X., Zhang, Y., Fratini, E., Baglioni, P., Faraone, A., Mamontov, E.: Experimental evidence of fragile-to-strong dynamic crossover in DNA hydration water. J. Chem. Phys. 125, 171103 (2006)
Chu, X.Q., Fratini, E., Baglioni, P., Faraone, A., Chen, S.H.: Observation of a dynamic crossover in RNA hydration water which triggers a dynamic transition in the biopolymer. Phys. Rev. E 77, 011908 (2008)
Khodadadi, S., Pawlus, S., Roh, J.H., Garcia-Sakai, V., Mamontov, E., Sokolov, A.P.: The origin of the dynamic transition in proteins. J. Chem. Phys. 128, 195106 (2008)
Khodadadi, S., Pawlus, S., Sokolov, A.P.: Influence of hydration on protein dynamics: combining dielectric and neutron scattering spectroscopy data. J. Phys. Chem. B 112, 14273–14280 (2008)
Chen, G., Fenimore, P.W., Frauenfelder, H., Mezei, F., Swenson, J., Young, R.D.: Protein fluctuations explored by inelastic neutron scattering and dielectric relaxation spectroscopy. Philos. Mag. 88, 3877–3883 (2008)
Sokolov, A.P., Roh, J.H., Mamontov, E., Garcia-Sakai, V.: Role of hydration water in dynamics of biological macromolecules. Chem. Phys. 345, 212–218 (2008)
Frauenfelder, H., Ghen, G., Berendzen, J., Fenimore, P.W., Jansson, H., McMahon, B.H., Stroe, I.R., Swenson, J., Young, R.D.: A unified model of protein dynamics. Proc. Natl. Acad. Sci. USA 106, 5129–5134 (2009)
Parak, F., Knapp, E.W.: A consistent picture of protein dynamics. Proc. Natl. Acad. Sci. USA 81, 7088–7092 (1984)
Rasmussen, B.F., Stock, A.M., Ringe, D., Petsko, G.A.: Crystalline ribonuclease-A loses function below the dynamic transition at 220K. Nature 357, 423–424 (1992)
Ferrand, M., Dianoux, A.J., Petry, W., Zaccai, G.: Thermal motions and function of bacteriorhodopsin in purple membranes—effects of temperature and hydration studied by neutron-scattering. Proc. Natl. Acad. Sci. USA 90, 9668–9672 (1993)
Ostermann, A., Waschipky, R., Parak, F.G., Nienhaus, G.U.: Ligand binding and conformational motions in myoglobin. Nature 404, 205–208 (2000)
Dunn, R.V., Reat, V., Finney, J., Ferrand, M., Smith, J.C., Daniel, R.M.: Enzyme activity and dynamics: xylanase activity in the absence of fast anharmonic dynamics. Biochem. J. 346, 355–358 (2000)
Bragger, J.M., Dunn, R.V., Daniel, R.M.: Enzyme activity down to −100 degrees C. Biochim. Biophys. Acta 1480, 278–282 (2000)
He, Y.F., Ku, P.I., Knab, J.R., Chen, J.Y., Markelz, A.G.: Protein dynamical transition does not require protein structure. Phys. Rev. Lett. 101, 178103 (2008)
Mamontov, E., Zamponi, M., Hammons, S., Keener, W.S., Hagen, M., Herwig, K.W.: BASIS: A new backscattering spectrometer at the SNS. Neutron News 19, 22–24 (2008)
Mamontov, E., Luo, H., Dai, S.: Proton dynamics in N,N,Nʹ,Nʹ-Tetramethylguanidinium Bis(perfluoroethylsulfonyl)imide protic ionic liquid probed by quasielastic neutron scattering. J. Phys. Chem. B 113, 159–169 (2009)
Provencher, S.W., Glockner, J.: Estimation of globular protein secondary structure from circular dichroism. Biochem. 20, 33–37 (1981)
Van Stokkum, I.H.M., Spoelder, H.J.W., Bloemendal, M., Van Grondelle, R., Groen, F.C.A.: Estimation of protein secondary structure and error analysis from CD spectra. Anal. Biochem. 191, 110–118 (1990)
Whitmore, L., Wallace, B.A.: Protein secondary structure analysis from circular dichroism spectroscopy: methods and reference databases. Biopolymers 89, 392–400 (2008)
Bée, M.: Quasielastic Neutron Scattering. Adam Hilger, Philadelphia (1988)
Nakagawa, H., Joti, Y., Kitao, A., Kataoka, M.: Hydration affects both harmonic and anharmonic nature of protein dynamics. Biophys. J. 95, 2916–2923 (2008)
Receveur, V., Calmettes, P., Smith, J.C., Desmadril, M., Coddens, G., Durand, D.: Picosecond dynamical changes on denaturation of yeast phosphoglycerate kinase revealed by quasielastic neutron scattering. Proteins: Struct. Funct. Bioinformatics 28, 380–387 (1997)
Zaccai, G.: How soft is a protein? A protein dynamics force constant measured by neutron scattering. Science 288, 1604–1607 (2000)
Parak, F.G.: Physical aspects of protein dynamics. Rep. Prog. Phys. 66, 103–129 (2003)
Frauenfelder, H., Fenimore, P.W., Chen, G., McMahon, B.H.: Protein folding is slaved to solvent motions. Proc. Natl. Acad. Sci. USA 103, 15469–15472 (2006)
Tsai, A.M., Neumann, D.A., Bell, L.N.: Molecular dynamics of solid-state lysozyme as affected by glycerol and water: a neutron scattering study. Biophys. J. 79, 2728–2732 (2000)
Tarek, M., Tobias, D.J.: Role of protein-water hydrogen bond dynamics in the protein dynamical transition. Phys. Rev. Lett. 88, 138101 (2002)
Tournier, A.L., Xu, J., Smith, J.C.: Translational hydration water dynamics drives the protein glass transition. Biophys. J. 85, 1871–1875 (2003)
Doster, W.: The dynamical transition of proteins, concepts and misconceptions. Eur. Biophys. J. 37, 591–602 (2008)
Mamontov, E., Vlcek, L., Wesolowski, D.J., Cummings, P.T., Rosenqvist, J., Wang, W., Cole, D.R., Anovitz, L.M., Gasparovic, G.: Suppression of the dynamic transition in surface water at low hydration levels: a study of water on rutile. Phys. Rev. E 79, 051504 (2009)
Acknowledgements
The authors are thankful to K. W. Herwig and C. Hoffmann for critical reading of the manuscript, and K. L. Weiss for useful technical discussions. This work was supported by the US Department of Energy Basic Energy Sciences and the Office of Biological and Environmental Research, using facilities supported by Oak Ridge National Laboratory, and managed by UT-Battelle, LLC, for the US DOE under Contract No. DE-AC05-00OR22725.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mamontov, E., O’Neill, H. & Zhang, Q. Mean-squared atomic displacements in hydrated lysozyme, native and denatured. J Biol Phys 36, 291–297 (2010). https://doi.org/10.1007/s10867-009-9184-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10867-009-9184-6