Abstract
Dynamic Nuclear Polarization solid-state NMR holds the potential to enable a dramatic increase in sensitivity by exploiting the large magnetic moment of the electron. However, applications to biological solids are hampered in uniformly isotopically enriched biomacromolecules due to line broadening which yields a limited spectral resolution at cryogenic temperatures. We show here that high magnetic fields allow to overcome the broadening of resonance lines often experienced at liquid nitrogen temperatures. For a fibril sample of the Alzheimer’s disease β-amyloid peptide, we find similar line widths at low temperature and at room temperature. The presented results open new perspectives for structural investigations in the solid-state.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Within the last decade, MAS (magic angle spinning) solid-state NMR evolved into a powerful technique which allows to determine the structure and dynamics of uniformly isotopically enriched, non-soluble biological macromolecules (Castellani et al. 2002; Wasmer et al. 2008). However, low sensitivity limits many solid-state NMR applications. NMR signal intensities can be enhanced by up to a factor of 660 by making use of dynamic nuclear polarization (DNP) (Maly et al. 2008; Barnes et al. 2009). DNP enhancement is achieved by microwave (MW) irradiation inducing a transfer of polarization from unpaired electrons to the nuclei (Hovav et al. 2011). This signal enhancement allows to investigate systems of biological interest which are not accessible otherwise (Reggie et al. 2011; Renault et al. 2011; Sergeyev et al. 2011; Jacso et al. 2012). Cryogenic temperatures are a pre-requisite for DNP. Low temperatures, on the other hand, yield considerable line broadening in many cases, which is due to suppression of conformational averaging as a result of freezing, or due to non-uniform ordering of water molecules at the glass transition temperature (Ngai et al. 2008; Doster 2010). This line broadening is the major limitation for a wide-spread application of DNP solid-state NMR for the investigation of biological systems. Despite large efforts in improving DNP solid-state NMR, including hardware design (Griffin and Prisner 2010; Rosay et al. 2010) and progress in theory (Hovav et al. 2010; Hovav et al. 2011; Hu et al. 2011), it remains unclear if this technique is suitable to study uniformly isotopically enriched protein samples. We present here field dependent spectra of an uniformly isotopically enriched fibril sample of the Alzheimer’s disease amyloid peptide Aβ. We find that resolution significantly improves at higher external magnetic fields, indicating that conformational heterogeneity is not dominating the achieveable resolution.
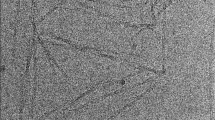
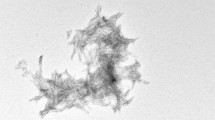
Aggregation of monomeric amyloid–β peptides (Aβ) into soluble oligomers and insoluble fibrils is one of the major pathological hallmarks of Alzheimer’s disease (AD) (Hardy and Selkoe 2002; Walsh et al. 2002). In fact, fibrils formed by the Aβ peptides (36-43 residues) are the primary components of the plaques found in human brains affected by Alzheimer’s disease (Cummings and Cotman 1995). Aβ fibrils are considered a difficult system for solid-state NMR as it is assumed that they can adopt a large variety of different polymorphous forms which in turn might result in line broadening (Fandrich et al. 2009). Fibrils employed in this study are uniformly isotopically enriched and grown using a seeding protocol (Lopez del Amo et al. 2012). An electron microscopic (EM) image of the fibrils obtained by this procedure is shown in Fig. 1a. Figure 1b shows a 1D CPMAS 13C spectra obtained at room temperature, at 96 K and at 96 K with microwave irradiation. A gain in intensity of around fourfold is observed by lowering the temperature only, due to a more favourable Boltzmann distribution. With DNP, a total signal enhancement of a factor of 20 was obtained. In principle, signal enhancements of ε ≈ 660 are possible. Typically, enhancements in biological systems are smaller (Mak-Jurkauskas et al. 2008; Bajaj et al. 2009a), in agreement with the values observed here. Partial sample deuteration can yield significantly larger enhancements on the order of ε = 120–140 (Akbey et al. 2010).
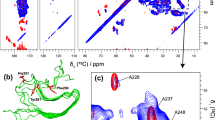
a EM image of the Aβ40 fibrils employed in the MAS solid-state NMR studies. b 1D-13C spectra of Alzheimer’s disease β-amyloid fibrils obtained under DNP conditions, with and without microwave irradiation (MW = on, MW = off), and at room temperature. In total, an enhancement factor of 20 is obtained. c 2D 15N,13C correlation spectrum of β-amyloid fibrils under DNP conditions (100 K). Side chain resonances of lysine, arginine and histidines are well resolved. d DNP 2D 15N,13C correlation spectrum of the histidine side chain region highlighted in b. All spectra were acquired at a magnetic field strength of 9.4 T
In the NCO and NCA experiments recorded for Aβ fibrils (Fig. 1b), most of the backbone resonances are overlapped due to the intrinsic low resolution at 9.4 T. However, well resolved resonances are observed for sidechains. This is especially obvious for the histidine spectral region (Fig. 1c). Here, two spin systems can be identified (labeled as 1 and 2) which are presumably due to His13 and His14 in Aß40. The 13C/15N line widths of the resolved Cδ1-Nε1 correlation peak amounts to 140 Hz in the 15N dimension, and ~150 Hz in the 13C dimension. These values are very similar to the line width which is observed at room temperature (Lopez del Amo et al. 2012). This indicates that structural heterogeneity of the sample is not dominating the achievable resolution at 100 K, but rather the inherent lack of resolution at low magnetic fields (9.4 T).
In order to probe the field dependence of the resonance line widths at low temperatures, MAS solid-state NMR experiments were acquired at 14.1 T and 20.0 T. Figure 2a shows 2D 13C,13C correlation spectra of Aβ40 fibrils that were recorded at low temperatures (~100 K) at the indicated magnetic field strengths. Obviously, the spectral resolution is improving at higher magnetic fields. The significantly overlapped spectrum recorded at 9.4 T is very well resolved at 20.0 T yielding linewidths for individual crosspeaks that are on the order of 140–160 Hz. Figure 2b shows a comparison of the Aβ fibril 2D-13C,13C correlation spectrum recorded at low temperatures (20.0 T) and the spectrum recorded at room temperature (21.1 T). Methyl resonances are typically weak at a temperature of 100 K (Bajaj et al. 2009b). The observed temperature dependent chemical shifts indicate temperature dependent conformational changes that can potentially be probed by NMR. In comparison to the room temperature spectrum, the resolution is only slightly decreased at 100 K. The line width for the resolved 13Cβ serine resonance increases to 160 Hz, in comparison to 130 Hz at room temperature. Structural heterogeneity of the sample is thus not dominating the achievable resolution. We expect similar results for other amyloidogenic peptides and proteins. DNP solid-state NMR was applied recently for the structural characterization of a partially labelled nanocrystallin and a fibrillar sample of the Sup35 derived peptide GNNQQNY (Debelouchina et al. 2010), as well as for the study of acid induced fibrils formed by the PI3 K-SH3 domain (Bayro et al. 2011). These experiments were carried out at 9.4 T and 100 K, and show that line-broadening is in general not an issue for measurements at low temperatures.
a Field dependence of 2D-13C,13C correlation spectra of Aβ40 fibrils at low temperatures (100 K). Spectra were acquired and processed using the same parameters in all cases. In each plot, the lowest contour was drawn at 20 % of the maximum intensity of the isoleucine Cα–Cδ correlation signals. Spectra plotted at different contour levels for each field are shown as part of the Supporting Figure 1, together with 1D traces for selected peaks (Supporting Figure 3). b Superposition of the Aβ fibril 2D-13C,13C correlation spectra recorded at cryogenic (850 MHz, red) and at room temperature (900 MHz, black). The resolution is clearly not compromised at low temperature. A comparison of 2D-13C,13C correlation spectra recorded at cryogenic temperatures (600 and 850 MHz) is shown in Supporting Figure 3
We demonstrate here that homogeneously labelled Alzheimer’s Disease amyloid Aβ40 fibrils retain their spectral resolution at cryogenic temperatures, and are amenable for assignments and a structural characterization in case samples are measured at high enough magnetic field strengths. In the future, signal enhancements of several orders of magnitude combined with high resolution solid-state NMR will be extremely valuable for the characterization of membrane proteins and amyloid structures in complex biological systems. In situ studies of Aβ peptides interacting with native membranes for example might allow to elucidate the mechanism of toxicity involved in Alzheimer’s Disease. Such experiments were impossible in the past due to the low sensitivity as a consequence of the low amount of material that can be packed into an NMR rotor. High-field DNP might be suitable to overcome such problems in the future, with high potential for applications in structural biology research.
Experimental details
The Aβ peptide was prepared as described previously (Dasari et al. 2011). For one Aβ fibril sample, approximately 8 mg of uniformly 13C and 15N labelled Aβ40 was fibrilized using a seeding protocol: Seeds were obtained after multiple rounds of sonification and fibril growth in order to select a single amyloid morphology (Lopez del Amo et al. 2012). Aβ fibrils were doped using 30 mM of the biradical TOTAPOL (1-(TEMPO-4-oxy)-3-(TEMPO-4-amino)-propan-2-ol) (Hu et al. 2004). The material was concentrated by centrifugation and packed into a 3.2 mm zirconia NMR rotor. All DNP-NMR experiments were performed using a commercial Bruker DNP spectrometer operating at a 1H frequency of 400 MHz and microwave frequency of 263 GHz. Signal enhancement was achieved in situ, directly within the sample. The microwave power was generated by a Bruker gyrotron oscillator. High field low temperature MAS-NMR experiments were acquired at 600 MHz and 850 MHz spectrometers (Bruker Rheinstetten and Max-Planck-Insitut für Biophysikalische Chemie Göttingen, respectively). The reference room temperature spectrum was acquired at 900 MHz (FMP Berlin). In all cases, spectra were recorded using a triple resonance, low-temperature, HCN probe employing conventional 3.2 mm zirconia rotors. Cryogenic temperatures were achieved and controlled with Bruker low-temperature MAS accessory. All spectra were recorded by setting the MAS frequency to 10 kHz, and employing a 1H/13C CP contact time of 1.5 ms. High-power proton decoupling (ωrf = 85 kHz) was applied during acquisition using SPINAL-64 (Fung et al. 2000). 13C,13C transfers were achieved using a PDSD sequence, setting the mixing time to 15 ms. Magnetization was allowed to evolve for 12 ms in the direct and indirect dimensions, respectively.
References
Akbey Ü, Franks WT, Linden A, Lange S, Griffin RG, van Rossum B-J, Oschkinat H (2010) Dynamic nuclear polarization of deuterated proteins. Angew Chem Int Ed 49:7803–7806
Bajaj VS, Mak-Jurkauskas ML, Belenky M, Herzfeld J, Griffin RG (2009a) Functional and shunt states of bacteriorhodopsin resolved by 250 GHz dynamic nuclear polarization–enhanced solid-state NMR. Proc Natl Acad Sci USA 106:9244–9249
Bajaj VS, van der Wel PCA, Griffin RG (2009b) Observation of a low-temperature, dynamically driven structural transition in a polypeptide by solid-state NMR spectroscopy. J Am Chem Soc 131:118–128
Barnes AB, Mak-Jurkauskas ML, Matsuki Y, Bajaj VS, van der Wel PCA, DeRocher R, Bryant J, Sirigiri JR, Temkin RJ, Lugtenburg J, Herzfeld J, Griffin RG (2009) Cryogenic sample exchange NMR probe for magic angle spinning dynamic nuclear polarization. J Magn Reson 198:261–270
Bayro MJ, Debelouchina GT, Eddy MT, Birkett NR, MacPhee CE, Rosay M, Maas WE, Dobson CM, Griffin RG (2011) Intermolecular structure determination of amyloid fibrils with magic-angle spinning and dynamic nuclear polarization NMR. J Am Chem Soc 133:13967–13974
Castellani F, van Rossum B-J, Diehl A, Schubert M, Rehbein K, Oschkinat H (2002) Structure of a protein determined by solid-state magic-angle spinning NMR. Nature 420:98–102
Cummings BJ, Cotman CW (1995) Image-analysis of beta-amyloid load in Alzheimers-disease and relation to dementia severity. Lancet 346:1524–1528
Dasari M, Espargaro A, Sabate R, Lopez del Amo JM, Fink U, Grelle G, Bieschke J, Ventura S, Reif B (2011) Bacterial inclusion bodies of the Alzheimer disease beta-amyloid peptides can be employed to study native like aggregation intermediate states. ChemBioChem 12:407–423
Debelouchina GT, Bayro MJ, van der Wel PCA, Caporini MA, Barnes AB, Rosay M, Maas WE, Griffin RG (2010) Dynamic nuclear polarization-enhanced solid-state NMR spectroscopy of GNNQQNY nanocrystals and amyloid fibrils. Phys Chem Chem Phys 12:5911–5919
Doster W (2010) The protein-solvent glass transition. Bio Chem Biophys Acta 1804:3–14
Fandrich M, Meinhardt J, Grigorieff N (2009) Structural polymorphism of Alzheimer A-beta and other amyloid fibrils. Prion 3:89–93
Fung BM, Khitrin AK, Ermolaev K (2000) An improved broadband decoupling sequence for liquid crystals and solids. J Magn Reson 142:97–101
Griffin RG, Prisner TF (2010) High field dynamic nuclear polarization-the renaissance. Phys Chem Chem Phys 12:5737–5740
Hardy JA, Selkoe DJ (2002) The amyloid hypothesis of Alzheimer’s disease: progress and problems on the road to therapeutics. Science 298:962–964
Hovav Y, Feintuch A, Vega S (2010) Theoretical aspects of dynamic nuclear polarization in the solid state—the solid effect. J Magn Reson 207:176–189
Hovav Y, Feintuch A, Vega S (2011) Dynamic nuclear polarization assisted spin diffusion for the solid effect case. J Chem Phys 134:074509
Hu KN, Yu HH, Swager TM, Griffin RG (2004) Dynamic nuclear polarization with biradicals. J Am Chem Soc 126:10844–10845
Hu K-N, Debelouchina GT, Smith AA, Griffin RG (2011) Quantum mechanical theory of dynamic nuclear polarization in solid dielectrics. J. Chem. Phys. 134:125105
Jacso T, Franks WT, Rose H, Fink U, Broecker J, Keller S, Oschkinat H, Reif B (2012) Characterization of membrane proteins in isolated native cellular membranes by dynamic nuclear polarization solid-state NMR spectroscopy without purification and reconstitution. Angew Chem Int Ed 51:432–435
Lopez del Amo JM, Schmidt M, Fink U, Dasari M, Fändrich M, Reif B (2012) The basic subunit in Alzheimer’s disease beta-amyloid fibrils can be an asymmetric dimer. Angew Chem Int Ed 51:6136–6139
Mak-Jurkauskas ML, Bajaj VS, Hornstein MK, Belenky M, Griffin RG, Herzfeld J (2008) Energy transformations early in the bacteriorhodopsin photocycle revealed by DNP-enhanced solid-state NMR. Proc Natl Acad Sci USA 105:883–888
Maly T, Debelouchina GT, Bajaj VS, Hu K-N, Joo C-G, Mak-Jurkauskas ML, Sirigiri JR, van der Wel PCA, Herzfeld J, Temkin RJ, Griffin RG (2008) Dynamic nuclear polarization at high magnetic fields. J Chem Phys 128:052211
Ngai KL, Capaccioli S, Shinyashiki N (2008) The protein “glass” transition and the role of the solvent. J Phys Chem B 112:3826–3832
Reggie L, Lopez JJ, Collinson I, Glaubitz C, Lorch M (2011) Dynamic nuclear polarization-enhanced solid-state NMR of a C-13-labeled signal peptide bound to lipid-reconstituted Sec Translocon. J Am Chem Soc 133:19084–19086
Renault M, Pawsey S, Bos MP, Koers EJ, Nand D, Tommassen-van Boxtel R, Rosay M, Tommassen J, Maas WE, Baldus M (2011) Solid-state NMR spectroscopy on cellular preparations enhanced by dynamic nuclear polarization. Angew Chem Int Ed Engl 51:2998–3001
Rosay M, Tometich L, Pawsey S, Bader R, Schauwecker R, Blank M, Borchard PM, Cauffman SR, Felch KL, Weber RT, Temkin RJ, Griffin RG, Maas WE (2010) Solid-state dynamic nuclear polarization at 263 GHz: spectrometer design and experimental results. Phys Chem Chem Phys 12:5850–5860
Sergeyev IV, Day LA, Goldbourt A, McDermott AE (2011) Abstract: chemical Shifts for the Unusual DNA Structure in Pf1 bacteriophage from dynamic-nuclear-polarization-enhanced solid-state NMR spectroscopy. J Am Chem Soc 133:20208–20217
Walsh DM, Klyubin I, Fadeeva JV, Cullen WK, Anwyl R, Wolfe MS, Rowan MJ, Selkoe DJ (2002) Naturally secreted oligomers of amyloid beta protein potently inhibit hippocampal long-term potentiation in vivo. Nature 416:535–539
Wasmer C, Lange A, Van Melckebeke H, Siemer AB, Riek R, Meier BH (2008) Amyloid fibrils of the HET-s(218–289) prion form a beta solenoid with a triangular hydrophobic core. Science 319:1523–1526
Acknowledgments
This research was supported by the Helmholtz-Gemeinschaft, the Leibniz-Gemeinschaft and the DFG (Re1435, SFB449, SFB740). We are grateful to the Center for Integrated Protein Science Munich (CIPS-M) for financial support. J.M.L.d.A. acknowledges a postdoctoral stipend of the DFG (Lo1546). We thank Arne Linden for assistance in performing the DNP experiments at the Leibniz-Institut für Molekulare Pharmakologie (FMP), and Antoine Loquet, Max-Planck-Institut für Biophysikalische Chemie in Göttingen, for assitance in the low temperature measurements at 850 MHz.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lopez del Amo, JM., Schneider, D., Loquet, A. et al. Cryogenic solid state NMR studies of fibrils of the Alzheimer’s disease amyloid-β peptide: perspectives for DNP. J Biomol NMR 56, 359–363 (2013). https://doi.org/10.1007/s10858-013-9755-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-013-9755-5