Abstract
A powerful experiment for the investigation of conformational properties of unstructured states of proteins is presented. The method combines a phase sensitive J-resolved experiment with a 1H-15N SOFAST-HMQC to provide a 3D spectrum with an E.COSY pattern originating from splittings due to 3JHNHα and 2JNHα couplings. Thereby an effectively homodecoupled 1H-15N correlation spectrum is obtained with significantly improved resolution and greatly reduced spectral overlap compared to standard HSQC and HMQC experiments. The 3JHNHα is revealed in three independent ways directly from the peak positions, allowing for internal consistency testing. In addition, the natural HN linewidths can easily be extracted from the lineshapes. Thanks to the SOFAST principle, the limited sweep width needed in the J-dimension and the short phase cycle, data accumulation is rapid with excellent sensitivity per time unit. The experiment is demonstrated for the intrinsically unstructured 14 kDa protein α-synuclein.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The roles of partly or completely unstructured proteins in cellular processes and pathological conditions are becoming increasingly recognized (Dyson and Wright 2005; Dunker et al. 2008). Such proteins offer new challenges for structural biology since traditional methods, such as X-ray crystallography, are not applicable. Among the most powerful techniques for characterizing these states is NMR, since it enables to study the proteins with atomic resolution. In unstructured states, the normally encountered size problem in NMR can partially be neglected thanks to fast dynamics. However, the reduced chemical shift dispersion makes spectral overlap an even worse problem than for globular proteins. A variety of NMR-based approaches have so far been used to characterize fully or partially unstructured states of proteins (Meier et al. 2008; Eliezer 2009), including paramagnetic relaxation enhancement (PRE; e.g. Gillespie and Shortle 1997a, b; Dedmon et al. 2005; Bertoncini et al. 2005), residual dipolar couplings (RDCs; e.g. Wells et al. 2008; Shortle and Ackerman 2001; Meier et al. 2007), and J-couplings (e.g. Schwalbe et al. 1997; Massad et al. 2007). The three bond J-couplings of the peptide backbone provide very useful data since they are directly correlated with the distribution of dihedral angles. In particular the 3JHNHα coupling (corresponding to the φ angle) is frequently used in structural characterizations of unfolded proteins (Schwalbe et al. 1997; Meier et al. 2008; Massad et al. 2007).
Several experiments for measuring 3JHNHα have previously been reported (e.g. Kay and Bax 1990; Neri et al. 1990; Billeter et al. 1992; Vuister and Bax 1993; Weisemann et al. 1994; Kelly et al. 1996; Heikkinen et al. 1999; Köver and Batta 2001). Among the most widely employed methods is the HNHA experiment (Vuister and Bax 1993), originally designed for proteins that are too large for other methods. In the HNHA an HMQC signal is split into two signals appearing at δHN and δHα in the third (indirect) proton dimension. The 3JHNHα coupling constants are derived from the intensity ratio of these peaks using a quantitative trigonometric relation. Although this is often an excellent method for globular proteins, spectral overlap makes the measurements unreliable or even impossible for many residues in unstructured polypeptides. Similar problems are as well encountered in many other methods for measuring 3JHNHα, such as COSY (Neuhaus et al. 1985), 2D HMQC-J/HSQC-J (Kay and Bax 1990; Billeter et al. 1992) and HNCA-J (Weisemann et al. 1994).
The J-resolved approach for measuring J-coupling constants is not new (Aue et al. 1976) but has not been widely used for large biological systems because of its inherent phase-twisted lineshape. Recently, it was shown that absorption mode lineshapes can be obtained by applying a selective refocusing pulse on one of the coupled spins and combine this ‘anti J spectrum’ with the data from a standard J-resolved experiment using an echo/anti-echo type of sampling (Pell and Keeler 2007). In this article, a new 3D experiment is presented where a phase sensitive J-resolved experiment, aiming specifically for the amide signals, is combined with a SOFAST-HMQC (Schanda et al. 2005; Schanda and Brutscher 2005). The method resolves the 3JHNHα coupling, a spectroscopic splitting which is always present in 1H-15N-correlation experiments, as a phase sensitive J pattern in a third dimension. As a result, the effective linewidth in the direct proton dimension is significantly reduced compared to regular HSQC or HMQC spectra. In addition, the J-coupling can be estimated in up to three independent ways directly from the peak positions, which provide an opportunity for consistency control. Thanks to the limited sweep width needed in the J-dimension, the fast spin-lattice relaxation properties inherited from the SOFAST-HMQC and the short phase cycle, high resolution 3D data can be acquired within a few hours.
At least two other experiments that combine a J-resolved methodology with an HSQC have previously been proposed (Neri et al. 1990; Kelly et al. 1996). The approach presented here is, however, essentially different. The previous experiments do not result in a J-resolved pattern since only the cosine-modulated data of the J evolution is retained. Hence, no resolution enhancement of the 1H-15N correlation map is obtained. The J-resolved SOFAST-HMQC also provides a shorter experimental time (as described above), higher signal intensity (the entire signal is retained) and a method for reducing the effect of radiation damping during the J evolution period (see below). Furthermore, the method presented here is mainly developed for disordered protein states since that will allow for highly selective HN pulses and also experience the highest impact by the enhanced resolution.
Materials and methods
α-synuclein was expressed and purified as described in (Hoyer et al. 2002). 15N-labelled protein was produced using M9 minimal medium supplemented with 15NH4Cl. The experiments were carried out on Bruker Avance 500 and 700 MHz spectrometers equipped with cryoprobes using a sample containing 150 μM α-synuclein in 25 mM phosphate buffer pH 6.5 with 0.1 M NaCl. For the experiments at pH 7.4 the protein was dissolved in 25 mM Tris buffer with 0.1 M NaCl. Data was recorded at 10°C to avoid the enhanced line broadening at higher temperatures (Croke et al. 2008). 3D J-resolved SOFAST-HMQC spectra were typically recorded with 2,048 (or 4,096) × 64 × 14 complex points in ω3 (1H), ω2 (15N) and ω1 (J), respectively. The spectral widths were 12 ppm (ω3), 23 ppm (ω2) and 28 Hz (ω1) and 4, 8 or 16 scans per transient were used. HNHA spectra were recorded with 1,024 × 48 × 48 complex points and 11, 9 and 23 ppm spectral widths in ω3 (1Η), ω2 (1H) and ω1 (15N), respectively. 16 scans per transient were used. HSQC spectra were typically recorded with 8 scans per transient, 1,024 (or 4,096) × 128 complex points and 11 × 23 ppm spectral widths in ω2 (1H) and ω1 (15N), respectively. Data were processed using NMRPIPE (Delaglio et al. 1995) and analyzed using SPARKY (Goddard and Kneller, SPARKY 3, University of California, San Francisco, USA) and CCPNMR (Vranken et al. 2005). Peak positions were determined by default cubic spline interpolation peak picking if nothing else is indicated.
Description of the experiment
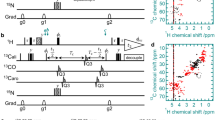
The pulse sequence is displayed graphically in Fig. 1. It is based on the SOFAST-HMQC experiment (Schanda et al. 2005; Schanda and Brutscher 2005) preceded by a phase sensitive J-resolved experiment, aiming at the amide protons. The excellent solvent suppression properties and the time effectiveness of the SOFAST-HMQC are thereby preserved. The excitation should be designed so that the water magnetization ends up parallel to the −z axis. This could be accomplished for example by a classical jump-return element (Plateau and Gueron 1982) with the same phase of both pulses or by a band-selective PC9 pulse (Kupče and Freeman 1993) followed by a hard 180° pulse and a short delay for refocusing the spin evolution that occur during the PC9 pulse. HN excitation is followed by a J-resolved evolution period, t 1, where chemical shift evolution and evolution due to heteronuclear J-couplings are refocused by a hard 180° pulse. This pulse will also turn the water magnetization back to +z, where it stays during the remaining experiment. Radiation damping during t 1 is suppressed by gradient pulses. Beside solvent suppression problems, radiation damping would act like a selective pulse for Hα-resonances close to the water signal and thereby distort the J-resolved pattern. Examples of this were indeed observed when the gradient pulses were omitted (data not shown). To reduce signal attenuation due to diffusion, the gradients are applied with alternating polarity. Two additional gradient pulses of equal polarity are positioned on either side of the hard pulse for selecting the desired coherence transfer pathway. At the end of t 1 the relevant product operators are H N y cos(π J t 1) + 2H N x H α z sin(π J t 1) and the sign of either term can be inverted using a band selective refocusing pulse for the HN-spin. By recording two datasets in the absence and presence of such a band selective pulse the sum and the difference will contain the cosine-modulated in-phase term and the sine-modulated anti-phase term, respectively. Such data handling is obtained by using an echo/anti echo type of sampling in the J dimension (t 1). To ensure balance between the intensities of the echo and anti echo experiments an identical band selective pulse is placed between the excitation element and t 1 in the echo experiment. In the implemented version of the experiment, REBURP shapes were used for these refocusing pulses. Qsim (Helgstrand and Allard 2004) simulations show that the conversion between in-phase and anti-phase operators due to 3JHNHα during the used REBURP pulses is insignificant. This is confirmed by the fact that no phase correction is needed when processing the J dimension. The t1 period, including the band selective refocusing pulses, is followed by the SOFAST-HMQC as described in (Schanda et al. 2005; Schanda and Brutscher 2005). During the HMQC part the spin state of the Hα is preserved while chemical shift evolution for HN is refocused by the band selective pulse. Hence, the J-resolved pattern is preserved.
The 3D J-resolved SOFAST-HMQC experiment. Filled black and open symbols indicate 90° and 180° pulses, respectively. Rectangular symbols are hard pulses while rounded symbols are HN selective pulses. The grey symbols are 180° refocusing pulses at the alternating positions in the echo (E) and anti-echo experiments (A). In our implementation, REBURP shapes (Geen and Freeman 1991) of 3.5 ms (700 MHz) or 4.5 ms (500 MHz) were employed for the band-selective 180° pulses and a jump-return (Plateau and Gueron 1982) element was used for the selective excitation. To suppress radiation damping while avoiding signal attenuation due to diffusion, a train of weak gradient pulses with alternating polarity (±1 Gauss/cm) is used. Gradients G1 and G3 were set to 2.5 and 10 Gauss/cm, respectively. All gradients are 1 ms with half-sine shape. Δ = 1/(2JHN). A four step phase cycle is employed with ϕ2 = x, x, y, y; ϕ3 = x, −x and ϕr = x, −x, −x, x to improve the solvent cancellation (ϕ3) and ensure quadrature balance of the J dimension (ϕ2). All other pulses are applied with x phase. ϕ1 and ϕr are inverted every even increment in t 1 to shift axial peaks in ω1 to the edges of the spectrum. Quadrature detection in t 2 is obtained by phase incrementation of ϕ3 according to States-TPPI. Coherence transfer pathways are shown below the pulse sequence with the broken line indicating the echo variant with selective refocusing previous to the t 1 period and the continuous line the anti-echo with selective refocusing after the t 1 period
After proper processing with Fourier transforms in all three dimensions each amide proton will display a J-resolved doublet structure in ω1−ω3, allowing for the measurement of 3JHNHα both as the peak separation in ω3 and from the absolute positions of each signal in ω1. In ω1, line broadening due to suboptimal shimming is refocused and the signals will thus display their natural linewidths. Since the spin state of the Hα nuclei is preserved throughout the experiment an E.COSY pattern revealing 2JNHα in ω1−ω2 and ω2−ω3 can be observed.
The proposed method results in sharp lines in the directly detected dimension, which motivate long acquisition times. When employing the SOFAST principle, the longitudinal relaxation during the acquisition time is efficient and optimal signal to noise per unit time would result if the excitation followed immediately after the acquisition time (Fig. 2). However, the spectrometer hardware might require longer inter scan delays to avoid overload and minimize heating.
Intensity (integral) per unit time plotted as a function of the scan time (T scan) for the first 1D trace of J-resolved SOFAST-HMQC recorded for α-synuclein at pH 6.5. The broken line indicates the employed acquisition time (0.34 s). Data was recorded without 15N decoupling to avoid spectrometer hardware damage
Results and discussion
As a proof of principle, the J-resolved SOFAST-HMQC was employed to measure J-coupling constants and HN linewidths in the 14 kDa intrinsically unstructured protein α-synuclein at pH 6.5. The enhanced resolution in this method allowed for accurate identification of several additional resonances in the 1H-15N correlation map compared to HSQC data (Fig. 3). In total, the experimental data allowed for measurements of 3JHNHα for 120 out of 135 non-proline residues in α-synuclein. In Fig. 4a, the J-coupling constants derived from the indirect and direct dimensions are compared. Average 3JHNHα coupling constants for glycine residue were estimated from the two outer components of the pseudo triplet. The correlation is very good and suspicious outliers, which may deserve manual inspection, may be readily identified. The RMSD between the direct and indirect dimensions is 0.38 Hz, indicating a standard error of 0.19 Hz in the average. 3JHNHα data were also derived from an HNHA experiment (Vuister and Bax 1993) recorded for the same α-synuclein sample to investigate potential systematic differences between the two methods. To avoid errors due to spectral overlap, the J-coupling constants from 49 residues displaying well-resolved signals both in the HNHA and the J-resolved SOFAST-HMQC were selected. The data from the two methods are in close agreement (Fig. 4b), with an RMSD of 0.31 Hz and a Pearson correlation coefficient of 0.92. Hence, large systematic differences can be excluded and the average of the observed splitting in the direct and indirect dimensions can be used in the same way as HNHA data are used.
Illustration of the improved spectral resolution obtained using the J-resolved SOFAST-HMQC (blue contours) compared to standard HSQC spectrum (red contours). The upper panels show HN-N correlation planes and the lower panels HN-J planes. Circles and triangles indicate different spin systems. Open and solid symbols indicate the peaks belonging to the same J doublet. a Non-overlapped Glu137. The doublet originating from the 3JHNHα coupling is resolved in J-dimension resulting in a significantly narrower apparent linewidth than in the HSQC spectrum. The 3JHNHα coupling constant can be measured independently in the HN and J dimensions. b, c Overlapping resonances are resolved thanks to the improved spectral resolution and the 3JHNHα coupling constants can be measured. b Val52 (circles) and Val74 (triangles). c Ile88 (circles) and Gln134 (triangles). The J-resolved data was recorded on the 500 MHz spectrometer with spectral resolutions of 1.5 and 1 Hz for the HN (ω3) and J (ω1) dimensions, respectively. The HSQC was recorded on the 700 MHz spectrometer with a spectral resolution of 1.0 Hz in the HN dimension. Data were processed using exponential window functions with line broadening factors of 1 Hz and zero-filled to 16 k points in the direct detected dimension and 64 points in the J dimension. The lowest contour levels correspond to half the maximum peak height in the displayed spectral region
a Correlation between the measured 3JHΝHα coupling constants in the indirect J dimension (ω1) and the direct detected 1H dimension (ω3) of the J-resolved SOFAST-HMQC experiment for 120 assigned residues (pair wise RMSD = 0.38 Hz; correlation coefficient, R = 0.92). b Correlation between the 3JHΝHα coupling constants measured using the J-resolved experiment (averages of the data derived from ω1 and ω3) and the HNHA experiment for 49 well resolved residues (pair wise RMSD = 0.31 Hz; correlation coefficient, R = 0.92)
Increased line broadening, for example due to hydrogen exchange at higher pH or temperature, could make it more difficult to determine the correct peak positions using default cubic spline interpolation peak picking, especially for small coupling constants. However, applying a lineshape fitting algorithm significantly improves the accuracy of the analysis in such cases. For data recorded on α-synuclein at pH 7.4 the RMSD compared to HNHA data for 36 well resolved residues were reduced from 1.1 to 0.32 Hz when using the Lorentzian fitting procedure in SPARKY (Goddard and Kneller, SPARKY 3, University of California, San Francisco, USA) instead of the automatic peak picking (data not shown). For pH 6.5 data, however, the apparent peak maxima and Lorentzian lineshape fitting give equally good results. Furthermore, the intrinsic symmetry of the J-resolved data offers a possibility to design even more sophisticated fitting algorithms.
Lineshape fitting also provides measurements of the natural HN linewidths, which can be related to exchange, flexibility or local proton density. The linewidths may also report on the PRE in designed systems, which is a powerful approach for investigating structural properties of unstructured proteins (Gillespie and Shortle 1997a, b). A variation between 2 and 8 Hz is observed for the HN linewidths along the sequence of α-synuclein (Fig. 5c) which can be compared with the apparent linewidths (including the doublet structure) of approximately 15 Hz in the HSQC spectrum (Fig. 3a). Hence the proposed J-resolved SOFAST-HMQC has at least twice the resolving power of a regular HSQC for α-synuclein.
Structural information derived from a single J-resolved experiment. a Helix propensity prediction of α-synuclein at pH 6.5 using AGADIR (Lacroix et al. 1998). b 3JHNHα coupling constants with error bars calculated as |Jω1-Jω3|/2, i.e. the standard error of the mean. c HN linewidths in the indirect J dimension. The numbers are the average of the upfield and downfield linewidths and were obtained by fitting the peaks to Lorentzian lineshapes in SPARKY (Goddard and Kneller, SPARKY 3, University of California, San Francisco, USA) and the error bars represent the standard error of the mean, i.e. |Δνupfield-Δνdownfield|/2. Only successfully fitted data (RMS < 10%) are included. d 2JNHα coupling constants derived from the E.COSY pattern in the 1H-15N plane. Resonances with substantial overlap in the 15N dimension are not included
Finally, 2JNHα couplings can be measured from the E.COSY pattern observed in the 1H-15N plane of the J-resolved experiment. This coupling constant is between 0 and 2 Hz in α-synuclein (Fig. 5d). Due to the wider peaks in the 15N-dimension, and because these couplings are only revealed once, the uncertainty is approximately twice as large as the uncertainty of 3JHNHα in either dimension. The uncertainty in either 1H-dimension is approximately 0.27 Hz, i.e. the RMSD/20.5. Hence the uncertainty in 2JNHα is approximately 0.5 Hz. However, the structural information that can be derived from 2JNHα data remains to be investigated.
The results obtained for α-synuclein are summarized in Fig. 5 and there are some obvious observations: The regions with the smallest 3J-coupling constants (residues 18, 56 and 90 with neighbours) coincide very well with the regions of highest predicted α-helical propensity (Fig. 5a) and highest helix content indicated by Cα secondary chemical shifts (Eliezer et al. 2001). The overall smallest 3JHNHα are observed for the region around residue 20. Interestingly, this region also corresponds to the largest HN linewidths in the protein indicating dynamic properties that are different from the surrounding regions.
Conclusions
A novel method for measuring 3JHNHα coupling constants that primarily aims at unstructured states of proteins is presented. The method is significantly faster than HNHA and more important; it provides an outstanding spectral resolution in the HN dimension compared to HSQC-based techniques. This increases the completeness of the measured data which is indeed essential for highly underdetermined systems such as unstructured proteins. Furthermore, the demonstration of the method on the intrinsically unstructured protein α-synuclein clearly illustrates that the data from a single J-resolved SOFAST-HMQC experiment provide substantial information about the conformational behaviour of the protein. 2JNHα and 3JHNHα coupling constants, as well as the natural HN linewidths can easily be extracted from the spectrum. The method therefore provides a very useful tool for investigations of the conformational properties of disordered states of proteins.
References
Aue WP, Karhan L, Ernst RR (1976) Homonuclear broad band decoupling and two-dimensional J-resolved NMR spectroscopy. J Chem Phys 64:4226–4227
Bertoncini CW, Jung Y, Fernandez CO, Hoyer W, Griesinger C, Jovin TM, Zweckstetter M (2005) Release of long-range tertiary interactions potentiates aggregation of natively unstructured α-synuclein. Proc Natl Acad Sci USA 102:1430–1435
Billeter M, Neri D, Otting G, Qian YQ, Wüthrich K (1992) Precise vicinal coupling constants 3JHNα in proteins from nonlinear fits of J-modulated [15N,1H]-COSY experiments. J Biomol NMR 2:257–274
Croke RL, Sallum CO, Watson E, Watt ED, Alexandrescu AT (2008) Hydrogen exchange of monomeric α-synuclein shows unfolded structure persists at physiological temperature and is independent of molecular crowding in Escherichia coli. Protein Sci 17:1434–1445
Dedmon MM, Lindorff-Larsen K, Christodoulou J, Vendruscolo M, Dobson CM (2005) Mapping long-range interactions in α-synuclein using spin-label NMR and ensemble molecular dynamics simulations. J Am Chem Soc 127:476–477
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Dunker AK, Oldfield CJ, Meng J, Romero P, Yang JY, Chen JW, Vacic V, Obradovic Z, Uversky VN (2008) The unfoldomics decade: an update on intrinsically disordered proteins. BMC Genomics 9(Suppl 2):S1
Dyson HJ, Wright PE (2005) Intrinsically unstructured proteins and their functions. Nat Rev Mol Cell Biol 6:197–208
Eliezer D (2009) Biophysical characterization of intrinsically disordered proteins. Curr Opin Struct Biol 19:23–30. doi:10.1016/j.sbi.2008.12.004
Eliezer D, Kutluay E, Bussell RJ, Browne G (2001) Conformational properties of α-synuclein in its free and lipid-associated states. J Mol Biol 307:1061–1073
Geen H, Freeman R (1991) Band-selective radiofrequency pulses. J Magn Reson 93:93–141
Gillespie JR, Shortle D (1997a) Characterization of long-range structure in the denatured state of staphylococcal nuclease. I. Paramagnetic relaxation enhancement by nitroxide spin labels. J Mol Biol 268:158–169
Gillespie JR, Shortle D (1997b) Characterization of long-range structure in the denatured state of staphylococcal nuclease. II. Distance restraints from paramagnetic relaxation and calculation of an ensemble of structures. J Mol Biol 268:170–184
Heikkinen S, Aitio H, Permi P, Folmer R, Lappalainen K, Kilpeläinen I (1999) J-Multiplied HSQC (MJ-HSQC): a new method for measuring 3 J(HNHα) couplings in 15N-labeled proteins. J Magn Reson 137:243–246
Helgstrand M, Allard P (2004) QSim, a program for NMR simulations. J Biomol NMR 30:71–80
Hoyer W, Antony T, Cherny D, Heim G, Jovin TM, Subramaniam V (2002) Dependence of α-synuclein aggregate morphology on solution conditions. J Mol Biol 322:383–393
Kay LE, Bax A (1990) New methods for the measurement of NH-CαH coupling constants in 15N-labeled proteins. J Magn Reson 86:110–126
Kelly GP, Muskett FW, Whitford D (1996) 3D J-Resolved HSQC, a novel approach to measuring 3JHNα. Application to paramagnetic proteins. J Magn Reson B 113:88–90
Köver KE, Batta G (2001) J-modulated TROSY experiment extends the limits of homonuclear coupling measurements for larger proteins. J Magn Reson 151:60–64
Kupče Ē, Freeman R (1993) Polychromatic selective pulses. J Magn Reson A 102:122–126
Lacroix E, Viguera AR, Serrano L (1998) Elucidating the folding problem of α-helices: local motifs, long-range electrostatics, ionic-strength dependence and prediction of NMR parameters. J Mol Biol 284:173–191
Massad T, Jarvet J, Tanner R, Tomson K, Smirnova J, Palumaa P, Sugai M, Kohno T, Vanatalu K, Damberg P (2007) Maximum entropy reconstruction of joint φ, ψ-distribution with a coil-library prior: the backbone conformation of the peptide hormone motilin in aqueous solution from φ and ψ-dependent J-couplings. J Biomol NMR 38:107–123
Meier S, Grzesiek S, Blackledge M (2007) Mapping the conformational landscape of urea-denatured ubiquitin using residual dipolar couplings. J Am Chem Soc 129:9799–9807
Meier S, Blackledge M, Grzesiek S (2008) Conformational distributions of unfolded polypeptides from novel NMR techniques. J Chem Phys 128:052204
Neri D, Otting G, Wüthrich K (1990) New nuclear magnetic resonance experiment for measurements of the vicinal coupling constants 3JHNα in proteins. J Am Chem Soc 112:3663–3665
Neuhaus D, Wagner G, Vasák M, Kägi JH, Wüthrich K (1985) Systematic application of high-resolution, phase-sensitive two-dimensional 1H-NMR techniques for the identification of the amino-acid-proton spin systems in proteins. Rabbit metallothionein-2. Eur J Biochem 151:257–273
Pell AP, Keeler J (2007) Two-dimensional J-spectra with absorption-mode lineshapes. J Magn Reson 189:293–299
Plateau P, Gueron M (1982) Exchangeable proton NMR without base-line distortion, using new strong-pulse sequences. J Am Chem Soc 104:7310–7311
Schanda P, Brutscher B (2005) Very fast two-dimensional NMR spectroscopy for real-time investigation of dynamic events in proteins on the time scale of seconds. J Am Chem Soc 127:8014–8015
Schanda P, Kupče Ē, Brutscher B (2005) SOFAST-HMQC experiments for recording two-dimensional heteronuclear correlation spectra of proteins within a few seconds. J Biomol NMR 33:199–211
Schwalbe H, Fiebig KM, Buck M, Jones JA, Grimshaw SB, Spencer A, Glaser SJ, Smith LJ, Dobson CM (1997) Structural and dynamical properties of a denatured protein. Heteronuclear 3D NMR experiments and theoretical simulations of lysozyme in 8 M urea. Biochemistry 36:8977–8991
Shortle D, Ackerman MS (2001) Persistence of native-like topology in a denatured protein in 8 M urea. Science 293:487–489
Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas M, Ulrich EL, Markley JL, Ionides J, Laue ED (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696
Vuister GW, Bax A (1993) Quantitative J correlation: a new approach for measuring homonuclear three-bond J(HNHα) coupling constants in 15N-enriched proteins. J Am Chem Soc 115:7772–7777
Weisemann R, Rüterjans H, Schwalbe H, Schleucher J, Bermel W, Griesinger C (1994) Determination of HN,Hα and HN,C′ coupling constants in 13C,15N-labeled proteins. J Biomol NMR 4:231–240
Wells M, Tidow H, Rutherford TJ, Markwick P, Jensen MR, Mylonas E, Svergun DI, Blackledge M, Fersht AR (2008) Structure of tumor suppressor p53 and its intrinsically disordered N-terminal transactivation domain. Proc Natl Acad Sci USA 105:5762–5767
Acknowledgments
We thank Dr. Carlos W. Bertoncini (University of Cambridge) for assistance with the protein production, Torbjörn Astlind (Stockholm University) for NMR spectrometer service and Dr. Jens Danielsson (Stockholm University) for critically reading the manuscript. The Carl Trygger foundation, the Magn. Bergvall foundation and the European Molecular Biology Organization are acknowledged for financial support.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lendel, C., Damberg, P. 3D J-resolved NMR spectroscopy for unstructured polypeptides: fast measurement of 3JHNHα coupling constants with outstanding spectral resolution. J Biomol NMR 44, 35–42 (2009). https://doi.org/10.1007/s10858-009-9313-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-009-9313-3