Abstract
Fine-scale genetic structure (FSGS) is common in plants, driven by several ecological and evolutionary processes, among which is gene flow. Mangrove trees rely on ocean surface currents to spread their hydrochorous propagules through space. Since pollen dispersal is generally restricted to local scales, high level of short-distance propagule dispersal is expected to result in FSGS in Rhizophora spp. We investigated FSGS, recent bottleneck events, as well as historical and contemporary expansion patterns in Rhizophora racemosa populations from the entire coast of Cameroon, using 11 polymorphic microsatellite markers. Populations of the Cameroon Estuary complex (CEC) showed significant FSGS and significant reduction in effective population sizes (recent bottlenecks), compared to the other areas. Additionally, our results indicate stark differences between historical and contemporary expansion models. These suggest that contemporary processes such as restricted propagule dispersal, bottleneck events from high indirect and direct anthropogenic pressure, and recolonization by founders from ancient local pockets/refugia most plausibly shape the patterns of FSGS in the CEC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dispersal plays an integral role for the persistence of populations and in shaping plant communities (Trakhtenbrot et al., 2005). It influences extinction and colonization events, controls patterns of gene flow, population regeneration and stability, as well as range dynamics (Peakall et al., 2003). Understanding the mechanisms of dispersal can provide insight into how landscapes are being (or have been) colonized by plants and predict range dynamics under future environmental changes (Nilsson et al., 2010). Hydrochory, i.e., dispersal of propagules via water currents, has been reported as an efficient dispersal mechanism over others, evidenced from lower differentiation in riparian meta-populations compared to terrestrial counterparts (Kinlan & Gaines, 2003; Chen et al., 2007). It has been hypothesized that hydrochorous plant populations show reduced genetic structure due to high gene flow (Kudoh & Whigham, 1997; Nilsson et al., 2010).
In mangroves, which occur in intertidal areas along tropical, subtropical, and warm temperate coastlines, dispersal distance and directionality of propagules is determined mainly by the interaction of ambient ocean surface currents and winds (Van der Stocken et al., 2013, 2015a; also, see Pil et al., 2011; Wee et al., 2014; Mori et al., 2015). The spatial extent of dispersal, however, may complicate or prevent direct observation of dispersal patterns, rendering quantification of propagule dispersal between mangrove populations difficult. Dispersal distances generally describe a leptokurtic distribution (Nathan et al., 2008), the minority of propagules actually dispersing over long distances. In mangroves, the short-distance component of the dispersal kernel may be due partly to the densely rooted areas in mangrove forests, which may obstruct the transport of propagules to the open sea (Van der Stocken et al., 2015b). It has been shown that restricted pollen and propagule dispersal may result in fine-scale genetic structure (FSGS) (Nilsson et al., 2010; Cerón-Souza et al., 2012; Jalonen et al., 2014; Mori et al., 2015), i.e. the non-random spatial distribution of genetic variation at fine scale. FSGS is generally explained by ecological drivers such as limited gene flow, adult population density, thinning among cohorts (fragmentation), breeding systems, spatio-temporal patterns of seedling colonization, disturbance history, as well as evolutionary processes such as local adaptation (Loveless & Hamrick, 1984; Vekemans & Hardy, 2004; Chung & Chung, 2013, Millán-Aguilar et al., 2016), of which gene flow remains the most widely studied factor (Wells & Young, 2002; Hardy et al., 2004; also see Cerón-Souza et al., 2012; Mori et al., 2015).
Gene flow is inferred from population genetic differentiation (genetic structure), since direct measurements are not feasible. However, in areas where anthropogenic pressure results in continuous re-expansion/recolonization of species following destruction, inferring gene flow from population structure is not straightforward. Strong anthropogenic pressure such as fragmentation due to wood extraction can negatively affect gene flow, resulting in strong population differentiations both at fine and medium scales. Several estimators of population differentiation exist, with different critics on their applicability and their interpretation. F-statistics and R-statistics are the most widely used, probably due to their estimation of the number of migrants (Nm), although both can be criticized in a number of ways (Balloux & Lugon-Moulin, 2002). Therefore, careful interpretation of both (F- and R-) statistics may provide the most valuable insights on genetic structure of populations (Balloux & Lugon-Moulin, 2002), as well as the levels of gene flow among populations. Also, spatial genetic autocorrelation analyses allow for meaningful estimates of (fine-scale) spatial genetic differentiation (Peakall et al., 2003), even in cases where genetic structure is low (Epperson & Tianquan, 1996).
Species with high dispersal abilities and weak habitat specificity are expected to show little population genetic differentiation (Latch et al., 2014). It is hypothesized that FSGS is unlikely to occur in widely dispersing species and in outcrossing taxa (Peakall et al., 2003; Epperson, 2007). Although Rhizophora propagules have a high probability to be retained, they are potentially capable of long distance dispersal due to their long viability and flotation periods (see De Ryck et al., 2012; Tonné et al., 2016, 2017). Additionally, preliminary findings suggest pollen dispersal to be highly restricted in the (self-compatible and partially protandrous) Rhizophora racemosa G. Mey. populations in the Cameroon Estuary, and these populations were only moderately outcrossing, indicating a mixed mating system of both outcrossing and geitonogamous selfing (unpublished data). Apart from pollen and propagule dispersal limitation, FSGS is expected to be higher in “recent” populations such as those in the periphery of a range, rather than in core populations, due to higher pressure from genetic drift, reduced effective population sizes, founder effects, and restricted gene flow (Pandey & Rajora, 2012). However, the same can occur even in core populations that persist, due to repeated re-expansion/recolonization in certain areas that are highly disturbed, yet contain pockets of original forest or remain conducive for colonization.
Mangrove populations of the Cameroon Estuary complex (CEC) are among the most genetically diverse along the Cameroonian coastline (see Ngeve et al., 2016a, b). Ngeve et al. (2016a) found that (recent) colonization patterns in this estuary might have followed the migrant pool model for most populations, and the propagule pool model for some others. Additionally, gene flow within the CEC was observed to be high such that all individuals make up a single genetic cluster, while propagule dispersal covered several kilometres within the estuary (Ngeve et al., 2016b). However, high levels of gene flow (low differentiation) among populations and seemingly successful establishment of genetically rich founder populations do not prevent the build-up of strong, rather persistent spatial genetic structure within populations (Helsen et al., 2015). Since the seed dispersal curve tends to be leptokurtic, local (re-) expansion into degraded gaps and new patches, as well as the growth of new cohorts, occurs from local more ancient pockets. Therefore, genetic structure at small spatial scales (within population/mangrove forest patch) amidst low differentiation between populations, and high genetic diversity, suggests “hidden founders” (sensu Helsen et al., 2015). “Hidden” refers to the fact that, besides high genetic variability and reduced differentiation among populations, some small patches within the whole are made of founders from colonists of local origin, resulting in the build-up of genetic structure at very small spatial scales (a few tens of metres). Studies assessing FSGS in (Rhizophora) mangroves are rare (but see Cerón-Souza et al., 2012; Mori et al., 2015). While for mangroves along the Cameroonian coast, earlier work assessed genetic differentiation at the inter-population scale (cf., Ngeve et al., 2016a, b), no data are as yet available on the within-population differentiation, i.e., at much smaller geographical scales (e.g. <200 m). Figure 1 provides a conceptual framework of the links and differences of this current study and our two previous studies.
In this study, we assess the FSGS hypothesis for mangrove populations along the Cameroonian coastline, which are under varying degrees of anthropogenic pressures. The most disturbed are those on the Cameroon Estuary complex (Corcoran et al., 2007; Nfotabong-Atheull et al., 2011, 2013; Ngeve et al., 2015). Anthropogenic pressures range from encroachment by local people, deforestation via illegal logging, sediment/sand extraction (personal observation; Corcoran et al., 2007; FAO, 2011; Nfotabong-Atheull et al., 2009, 2011, 2013), and pollution (Alemagi et al., 2006, Ngeve et al., 2015). Despite converging ocean currents offshore at the CEC, the overall genetic diversity of this estuary was not significantly higher than others (Ngeve et al., 2016a). Although this could result from the westward jets created by converging currents, transporting propagules seaward rather than landward (Ngeve et al., 2016a); it could also result from natural river-estuary dynamics or high levels of anthropogenic pressure as reported in this area (see Ngeve et al., 2016b). Firstly, we hypothesize that FSGS is high in Rhizophora mangrove areas due to the high propagule retention potential in such forests with a dense stilt root structure (see Sousa et al., 2007; Van der Stocken et al., 2015b; Gillis et al., 2016) as well as restricted pollen dispersal (see Cerón-Souza et al., 2012; Mori et al., 2015). To test this, we evaluate comparative FSGS of the different mangrove areas in Cameroon. Secondly, we hypothesize the mangroves in the CEC, being the most disturbed along the Cameroonian coastline, show stronger patterns of FSGS and reduced effective population sizes (bottlenecks), compared to more pristine areas such as the Rio del Rey Estuary. Therefore, we investigate populations for recent reduction in effective population size to assess whether these have any relationship with FSGS. Additionally, we investigate the expansion pathway (historical migration/dispersal) of mangroves along the entire coastline of Cameroon to test for recent expansion/recolonization of this mangrove area. That is, whether the present mangroves of the CEC were the last to be established by colonization due to the present-day ocean surface current patterns that characterize this coastline, flowing northwards and southwards and converging offshore the CEC (see Ngeve et al., 2016a). This will provide more information as to whether patterns of recent bottleneck events are carry-overs of historical founders or result from recent (anthropogenic) pressure, or both. Thirdly, we hypothesize that due to the differences in the degree of anthropogenic pressures among the 4 different mangrove estuaries, estuarine-scale genetic differentiation would be different. Therefore, we compared the overall genetic structure within the four studied estuaries to elucidate the influence of anthropogenic on genetic differentiation.
Materials and methods
Description of the study area
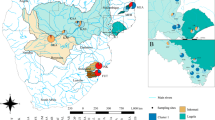
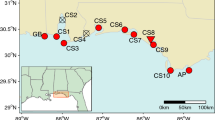
Mangroves of the Cameroonian coastline are primarily distributed along four estuarine areas—the Rio del Rey Estuary in the north (most pristine), the Cameroon Estuary complex (CEC) (most anthropized), and the more (naturally smaller) southerly mangrove areas—the Lokoundje and Ntem estuaries (Fig. 2). The nation’s overall mangrove cover declined from ca. 1119 km2 in 2000 to ca. 1113 km2 in 2014 (Hamilton & Casey, 2016), with the CEC being the most impacted (Corcoran et al., 2007; Nfotabong-Atheull et al., 2013). Despite the decline in its spatial extent, due to coastal development, urbanization, and conversion into farm fields (for planting, e.g. maize and oil palm) (Nfotabong-Atheull et al., 2013, Ndjebet & Ngokoy, 2014), the CEC remains a vast mangrove area ca. 88,000-100,000 ha, second to the Rio del Rey Estuary (ca. 100,000 ha) and larger than the southern estuaries (ca. 2000 ha) (FAO, 2011) (Fig. 2). The mangrove population at Londji (Lokoundje Estuary) was highly fragmented, consisting of sparsely distributed remnant adult stands (see Table S1). Unlike the Lokoundje Estuary, the CEC populations are still relatively dense (see details in Table S1). In the Wouri (CEC), R. racemosa stands could grow as high as 40 m, with about 1 m tree diameter (Saenger & Bellen, 1995). However, for most of the sampled areas, tree height could average about 4—10 m in height, except for a few very tall trees (personal observation). Although tree height must not correlate with age, due to the slow growing nature of Rhizophora spp., the large differences in tree heights, up to about 10-fold magnitude, could indicate the coexistence of trees or cohorts of different age classes within the CEC, such that the taller trees are much older than the relatively shorter trees.
Sampling for plant DNA material
All four (4) mangroves estuaries were studied from along the Cameroonian coastline (see Ngeve et al., 2016a). Eleven (11) study sites in total were sampled for plant material for genetic analysis from the four estuaries. Ekondo-Titi (EKO), Mbongo (MBO), and Bekumu (BEK) make up the 3 sites from the Rio del Rey; Mabeta (MAB), Tiko (TIK), Douala—Bonamoussadi (SADI), Douala—Akwa Nord (AKN), Douala-Bonaberi (BERI), and Douala-Edea Reserve at Mouanko (RSVM) make up the 6 sites from the CEC; Kribi, sampled at Londji and Mpalla, makes up the site from the Lokoundje Estuary. While Campo, sampled at Ipono and at Campo Beach, makes up the site at the Ntem Estuary (Fig. 2). Sampling was done along transects and a minimum distance of 10 m between sampled trees was considered deliberately to avoid sampling the same individual. Distances between trees were measured in the field using a measuring tape and from GPS coordinates of each tree.
DNA extraction and amplification
We extracted genomic DNA from 20 to 30 mg of dry plant leaf tissue, using the E.Z.N.A SP plant DNA Mini Kit (Omega bio-tek). Using the 11 polymorphic microsatellite markers initially isolated from R. racemosa from Cameroon (Tiko) (Ngeve et al., 2016b), a multiplex PCR was performed to amplify DNA samples. The constituent of each of the multiplex PCR was 2.5 µl of H2O, 6.25 µl of the master mix (Qiagen multiplex pcr kit master mix), 1.25 µl of the primer mix, and 3 µl of DNA, in a final volume of 13 µl. The PCR conditions were similar to that employed by Ngeve et al. (2016a, b): initial denaturation of 95°C for 15 min followed by 34 cycles of 30 s denaturation, 90 s of annealing at a temperature of 57°C and elongation of 80 s at 72°C, followed by a final elongation time of 30 min at 60°C, and a cooling to 4°C for 1 min in a Bio-Rad thermal cycler (MJ research PTC-200 and Bio-Rad MyCycler) (Ngeve et al., 2016a, b). Fragment analysis was done by Macrogen Corporation (Seoul, South Korea), and scoring of alleles was done using GeneMarker (SoftGenetics LLC, State College, USA).
Data analyses
Hypothesis 1
Spatial autocorrelation analyses with multi-locus genotypes for assessing FSGS
Spatial autocorrelation allows for the examination of the correlation between pairwise genetic distance of individuals within a given geographic distance class, for multiple user-defined classes. Geographic distance matrices were obtained from the distances between trees measured during sampling in the field, using a measuring tape as well as GPS coordinates. We carried out a spatial autocorrelation analyses for all study sites (11) (transects combined), assuming 6 even distance classes of 30 m, at 9999 permutations and 10,000 bootstraps using GenAlEx v 6.5 (Peakall & Smouse, 2012). Subsequently, patterns were then cross-checked using 6 variable distance classes (10, 30, 60, 120, 300, and 500 m). The x-intercept for each of the spatial autocorrelation analyses (each site) gives a measure of the non-random (positive) genetic structure, i.e. an estimate of the “genetic neighbourhood” (Vergara et al., 2015) or “genetic patch size” (Peakall & Smouse, 2012).
Hypothesis 2
Consequences of expansion pathway (dispersal) and effective population sizes on FSGS
We used the program Migrate-n (Beerli, 2006; Beerli & Palczewski, 2010) to test the following 4 historical migration/expansion models: (1) unidirectional expansion from Rio del Rey (north) to Campo (south) (N → S); (2) unidirectional expansion from Campo (south) to Rio del Rey (north) (S → N); (3) bidirectional expansion from Rio del Rey (north) and from Campo (south) into the CEC (centre), assuming the CEC as the most “recent” mangrove area (N → C ← S); and finally, (4) bidirectional expansion from the CEC (centre) to the north and south, assuming the CEC to be the “oldest” mangrove area (N ← C → S). This was done by the comparison of marginal likelihoods and Bayes factors. Estimates of the historical effective population sizes (Ne) and the historical migration rates per generation (Nem) were determined from the model that best describes the expansion pathway of mangroves along this coastline. To enable comparison, parameter values were deliberately kept the same for the respective analyses: Bayesian inference was the search strategy and the Brownian motion microsatellite mutation model was set as data type with uniform mutation rate at all loci. Uniform prior distribution settings (min, max, delta) were as follows: for Θ = 0.0, 10.0, 0.1 and for M = 0.0, 100, 10.0. The number of recorded steps was 106 at a sampling frequency of 103 after an initial burn-in of 106, and four parallel heated (static) chains (1.0, 1.5, 3.0, and 105). Three replicate runs, each with a different initial seed number, were performed using the same specifications for each model. Several trial runs were performed before these replicate runs to determine the most appropriate settings to allow for convergence. The Bezier approximation to marginal likelihood (mL) was used to assess which of the above-mentioned models best describes the data. Log Bayes factors (LBF) were calculated using the following formula (Beerli & Palczewski, 2010):
The effective number of immigrants per generation (Nem) was calculated as ΘM/4 (see Kennedy et al., 2016). Following the procedure of Kennedy et al. (2016), historical effective population sizes (Ne) were calculated as Θ/4µ, where µ is the plant microsatellite mutation rate of 7.7 × 10−4 (Vigouroux et al., 2002).
We also used the program BOTTLENECK v. 1.2.02 (Piry et al., 1999) to test for significant reduction in effective population sizes from recent bottleneck events. The analysis ran 1000 iterations by testing the assumptions that mutation rate of microsatellite markers follow (1) a Stepwise Mutation Model (SMM), or (2) a Two-Phase Model (TPM). TPM assumes 70% SMM rate and a variance of 30%. Significance of this test was considered strong at P < 0.001 or marginal at P < 0.05.
In addition to genetic analyses, a dispersal model was built for simulating propagule dispersal trajectories and estimate dispersal distances (see Ngeve et al., 2016a). The dispersal model was forced using hourly ocean surface currents from a mesoscale and tide-resolving ocean simulation carried out using the Massachusetts Institute of Technology general circulation model (MITgcm; Hill et al., 2007). The MITgcm ocean simulation is initialized from a data-constrained global ocean solution provided by the Estimating the Circulation and Climate of the Ocean, Phase II (ECCO2) project (Menemenlis et al., 2008, also see Rocha et al., 2016). Surface boundary conditions in the MITgcm are from the 0.14° European Centre for Medium-Range Weather Forecasts (ECMWF) operational atmospheric analysis starting in 2011, which is made available on a model grid with ≤15-km horizontal spacing, and bathymetry is a combination of the Smith and Sandwell (1997) v. 14.1 and International Bathymetric Chart of the Arctic Ocean (IBCAO). Horizontal grid spacing is ca. 4 km, while a vertical grid spacing of 1 m near the surface allows to better resolve surface boundary layer currents. For estimating dispersal trajectories, we used MITgcm ocean simulation output for the ocean surface, i.e. the upper vertical level only, which is a reasonable assumption for passively dispersing mangrove propagules that float near the water surface.
Dispersal model output allowed us to investigate whether present-day ocean surface currents are in congruence with historical migration (dispersal) patterns as indicated by genetic data, and to investigate whether propagule dispersal directions and distances have any relationship with FSGS. The dispersal model was used to determine the distances covered by virtual propagules released hourly for the months February, March, April, September, and October 2012 (i.e. 3626 propagules in total), all having a floating period of 3 months (the same model simulation was used by Ngeve et al., 2016a). Dispersal distances were calculated for all 3626 propagules released in the 11 coastal locations, using the spherical law of cosine:
where (x 1, y 1) and (x 2, y 2) are the coordinates of the release and end locations, respectively, and R = 6370 000 m, being the Earth’s radius.
Subsequently, we calculated the Spearman rank correlation between x-intercept values (genetic neighbourhood), as a measure of the FSGS, and the (modelled) dispersal distances (used as a determinant of “open” or “closed” systems). To validate the use of the x-intercept as a measure of FSGS, we estimated the Spearman rank correlation between the spatial autocorrelation coefficients (r) and the x-intercepts.
Hypothesis 3
Comparative estuarine-scale genetic structure
Estuarine-scale statistics of genetic differentiation were done for the 4 different estuaries by assessing AMOVA-Fst and AMOVA-Rst, to compare the genetic differentiation of the 4 different estuaries. Following Hardy et al. (2003), we performed allele size permutation tests for each of the studied estuaries to assess the contribution of allele sizes to genetic structure of the estuaries in SPAGeDi v. 1.4 (Hardy & Vekemans, 2002). For these analyses, we ran a total of 2 × 104 permutations for each estuary. The permutation test calculates a permuted Rst, which is done by randomly assigning allele sizes within each locus and assessing whether stepwise mutation contributes more to the genetic differentiation than migration and drift. For each of all the studied loci, the program then compares estuarine estimates of the permuted Rst vs the normal Rst.
Additionally, to evaluate the impact of population size and disturbance on genetic differentiation, we performed a Spearman rank correlation analysis between the effective population sizes (Ne) vs the genetic differentiation (Fst) of each estuary. We also categorized estuaries as either being in a disturbed state (1) or an undisturbed state (0). Using this categorical variable, we performed another Spearman rank correlation with the Fst values of the estuaries.
Assumptions and limitations
Mindful of the high dispersal capabilities of Rhizophora propagules, and the possibility that colonization and expansion of mangroves along the Cameroonian coastline might have occurred from multiple founders and followed multiple pathways, the lack of samples from other parts of the West African coastline indicates that the expansion model observed herein is the most simplistic of patterns and assumes the Cameroonian coastline (from which all samples were obtained) as an “island” (isolated from all the other mangroves of the entire W. African coastline). Additionally, although it might have been interesting to test more expansion pathways, for example by including some from the recent patterns initially reported by Ngeve et al. (2016a), the additional relevance of these very contemporary patterns in elucidating patterns of historical expansion, on a coastline known to have been altered geomorphologically, is assumed to be of less impact. Additionally, “disturbance” or “disturbed state” as considered in the third hypothesis of this manuscript is based on the subjective assessment of the author (MNN), following insights from extensive literature review (e.g. Corcoran et al., 2007; Nfotabong-Atheull et al., 2009, 2011; Feka & Ajonina, 2011; Nfotabong-Atheull et al., 2013; Ndjebet & Ngokoy, 2014; Feka & Morrison, 2017), as well as insights from comparative field observation of the different mangrove areas.
Results
Hypothesis 1
Spatial autocorrelation analyses (FSGS) in Rhizophora racemosa populations
Details on the genetic variability of the different mangrove populations considered for this work have already been reported by Ngeve et al. (2016a, b). Six (6) out of the 11 sites showed significant FSGS for the first distance class of 30 m, five of which were populations from the CEC (Figs. 3, 4, 5). The five populations from the CEC were MAB, TIK, AKN, BERI, RSVM, while BEK, i.e. the site closest to the CEC, was the only site of the Rio del Rey with a marginally significant (P < 0.05) FSGS. The population SADI (most upstream Wouri) was the only population of the CEC without significant FSGS. The results from spatial autocorrelation analyses using variable distance classes showed patterns that were very comparable to those calculated using even distance classes (data not shown).
Spatial autocorrelogram of 3 Rhizophora racemosa populations from the Rio del Rey Estuary showing variable fine-scale genetic structure at even distance classes of 30 m. Dashed lines indicate upper and lower 95% confidence intervals (CI) around the spatial autocorrelation coefficient (r) based on 9999 permutations; 95% CI around r from 9999 bootstraps. P values indicate level of significance of fine-scale genetic structure, while significance to recent reduction in effective population sizes (bottlenecks) is indicated by *(P < 0.05) and **(P < 0.001)
Spatial autocorrelogram of 6 Rhizophora racemosa populations from the Cameroon Estuary complex (CEC) showing variable fine-scale genetic structure at even distance classes of 30 m. Dashed lines indicate upper and lower 95% confidence intervals (CI) around the spatial autocorrelation coefficient (r) based on 9999 permutations; 95% CI around r from 9999 bootstraps. P values indicate level of significance of fine-scale genetic structure, while significance to recent reduction in effective population sizes (bottlenecks) is indicated by *(P < 0.05) and **(P < 0.001)
Spatial autocorrelogram of 1 Rhizophora racemosa population from the Lokoundje Estuary (KRIBI), and 1 population from the Ntem Estuary (CAMPO). Dashed lines indicate upper and lower 95% confidence intervals (CI) around the spatial autocorrelation coefficient (r) based on 9999 permutations; 95% CI around r from 9999 bootstraps. P values indicate level of significance of fine-scale genetic structure, while significance to recent reduction in effective population sizes (bottlenecks) is indicated by *(P < 0.05) and **(P < 0.001)
Hypothesis 2
Consequences of expansion pathway (dispersal) and effective population sizes on FSGS
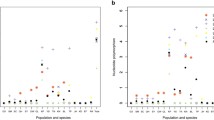
Investigation of recent reduction in effective population sizes showed 6 of the 11 populations (4 sites from the CEC and 2 other sites—BEK and Campo) have signs of recent bottleneck events (Figs. 3, 4, 5). All the populations that showed significant FSGS (except RSVM) also had significantly reduced effective population sizes (bottlenecks). Site 11 (CAMPO) is the only population that had signs of a recent bottleneck event without FSGS. The significance of bottleneck events was observed in these populations following the SMM. Estimates of historical effective population sizes indicate that the Lokoundje Estuary has the lowest historical effective population size (mean Ne = 64, 95% confidence interval = 4–119), while the Ntem Estuary in the South had the highest historical effective population size (mean Ne = 886, 95% confidence interval = 747–1026) (Fig. 6.). Contrary to our expectation, the CEC had the second highest historical effective population size of the entire coastline (mean Ne = 670, 95% confidence interval = 509–709) (Fig. 6). The model of effective historical dispersal pathway that best explains mangrove expansion along this coastline is dispersal from south to north (Table 1). Historical effective migration rates (Nem) are highest from the Ntem Estuary (southernmost) to the Lokoundje Estuary (mean Nem = 60.71, 95% confidence interval = 8.89–167.61) and lowest from Ntem to Rio del Rey (mean Nem = 0.05, 95% confidence interval = 0–0.3) and from CEC to Rio del Rey (mean Nem = 0.08, 95% confidence interval = 0–0.56) (Fig. 6). However, modelled dispersal pathways of virtual propagules indicate a major difference between historical and current migration pattern along the coastline. There is a wide expansion pathway for propagules from the north (Rio del Rey) to the centre (CEC). Although propagules from the Ntem and Lokoundje estuaries are dispersing from south to north, as the historical migration rates, these propagules generally do not reach the CEC and certainly not the Rio del Rey in the north due to contemporary ocean current dynamics (Fig. 7) (also see Ngeve et al., 2016a). The dispersal pathways of virtual propagules from the CEC seem to be highly restricted by the contemporary ocean current patterns, as the propagules from the CEC do disperse out of the CEC. Additionally, calculated propagule dispersal distances indicated the shortest dispersal distances occurring for propagules released at sites within the CEC (Table 2). So, propagules released from populations within the CEC remain in the CEC. A correlation analysis of these distances and the genetic neighbourhood (x-intercept) revealed a negative relationship (scatterplot, Fig. 8).
Mean historical effective population sizes (Ne) and mean effective number of migrants per generation (Nem) between mangrove estuaries of the Cameroonian coastline (brackets = 95% confidence intervals). Estimates were calculated from the best choice model describing mangrove expansion along the Cameroonian coastline based on our data
a Modelled historical mangrove expansion pathway based on comparison of log Bayes factors and log marginal likelihoods (bold lines) and b contemporary simulated dispersal trajectories (indicated as well with dashed-line arrows in (a)). Numbers 1–11 correspond to the different population sites. Historical dispersal pathway a modelled dispersal from south to north seem to best fit the data (bold-line arrows); dashed-line arrows describe contemporary mangrove dispersal pathways as simulated using a propagule dispersal model b, indicating dispersal from both the northern Rio del Rey Estuary, the southerly Ntem, and Lokoundje estuaries towards the centre (CEC), but converging to form westward jets into the open ocean. Propagules from Bekumu (red) and Mbongo (green) in the Rio del Rey Estuary indicate contemporary connectivity with the CEC, while these pathways do not meet with those from the southern Ntem (brown) and Lokoundje estuaries (b). These patterns are in congruence with estimates of contemporary dispersal patterns, using genetic data, as described by Ngeve et al. (2016a)
Hypothesis 3
Comparative estuarine-scale genetic structure
Analysis of molecular variance (AMOVA) F-statistics showed that Fst was low but significant in all estuaries except for the Ntem Estuary (Table S2). Fst was highest for the Lokoundje Estuary (Fst = 0.124) and lowest for the Ntem Estuary (Fst = 0.007) (Table S2). AMOVA R-statistics indicated relatively high and significant Rst values except for the Lokoundje Estuary. Rst was highest for the CEC (Rst = 0.269) and lowest for the Lokoundje Estuary (−0.005) (Table S2). Our results of allele size permutation tests showed that for all 4 estuaries, the permuted Rst was less than the calculated (observed) Rst, although this was significant only in the Rio del Rey Estuary (P < 0.005) (Table S3). Genetic differentiation (Fst) showed an inverse relationship with effective population size (Ne) (r = −0.78), but a strongly positive relationship with disturbed state (r = 0.72), although none of these relationships were significant (at P < 0.05).
Discussion
Hypothesis 1
FSGS as a function of retention in Rhizophora racemosa populations
The difficulty of propagules to enter or leave closed mangrove forest landscapes, due to reduced vector displacement velocity and shorter seed passage times (Van der Stocken et al., 2015b; Nathan et al., 2008), could result in increased FSGS when these propagules eventually strand in the neighbourhood of their parent trees. In Rhizophora spp. however, this can even be exacerbated because of the increased barrier density posed by the prop root network of Rhizophora spp. which may retain large dispersing propagules, such as Rhizophora propagules (see Van der Stocken et al., 2015b). Therefore, it is sensible to assume that forests dominated by Rhizophora species may show FSGS more readily than open release and access forests (e.g. those with less dense root networks), especially in relation to the large and easily trapped propagules of Rhizophora. Our results indicate that 6 out of the 11 studied Rhizophora racemosa populations of the Cameroonian coast have significant within-population FSGS and this appeared to be detectable over a short distance (30 m). This suggests a non-random distribution of genotypes at such fine scale for these populations. Five (5) of these populations were from the CEC and 1 population was from the Rio del Rey Estuary (Bekumu, proximate to the CEC). Populations from the other estuaries did not show significant FSGS.
The absence of FSGS in some Rhizophora mangrove populations suggests that FSGS cannot necessarily be linked to the high retention potential due to the barrier posed by their roots as observed in Rhizophora spp. (see Van der Stocken et al., 2015b). Hydrochory, compared to other dispersal mechanisms, has been reported to be very efficient (Kinlan & Gaines, 2003, Chen et al., 2007). The observation by Van der Stocken et al. (2015b) of heightened retention rates in Rhizophora mangrove forest covered 5 subsequent high tides. Beyond this experimental period however, it is not known whether or not (nor how many) retained propagules have been released with continued dispersal for open water. While their results confirm that generally most propagules disperse over short distances, it does not exclude their potential to exit their neighbourhood and embark on LDD, especially because Rhizophora propagules have potentially long flotation and viability periods (De Ryck et al. 2012; Tonné et al., 2016), ranging from about 90 to 260 days (e.g. see Drexler, 2001; Rabinowitz, 1978; Allen & Krauss, 2006; Steele, 2006). Otherwise, they could disperse over shorter distances, resulting in the connectivity of adjacent populations (e.g. see Ngeve et al., 2016b). Therefore, the occurrence of FSGS in populations of some estuaries and not in others suggests that other local scale processes, rather than the (soley) retention (which is a widespread Rhizophora phenomenon), is responsible for the patterns of FSGS in the affected areas.
Hypothesis 2
Consequences of expansion pathway (dispersal) and effective population sizes on FSGS
Generally, FSGS is expected to be high when plant species are inbred, when dispersal (gene flow level) is low, at range edges due to genetic drift (Maguire et al., 2000; Arnaud-Haond et al., 2006; also see Islam et al., 2014; Li et al., 2016), founder effects, and anthropogenic disturbance (Pandey & Rajora, 2012), among others. Populations of the most anthropized mangrove estuary (CEC) were primarily those with signs of FSGS. These patterns of FSGS observed here, in spite of the reported high levels of within-estuary genetic connectivity (Ngeve et al., 2016b), could be explained by high anthropogenic disturbance (see Pandey & Rajora, 2012) of the CEC (Alemagi et al., 2006; Nfotabong-Atheull et al., 2009; Feka & Ajonina, 2011; Nfotabong-Atheull et al., 2013; Ngeve et al., 2015; Feka & Morrison, 2017) compared to other mangrove areas in Cameroon. Large mangrove areas are being cleared for coastal development and extensive illegal logging is also in place in many areas, for use as fuel and construction wood by local people (Nfotabong-Atheull et al., 2009, 2011, 2013). It is estimated that about 25% of mangroves of the CEC have been lost due to anthropogenic activities (Ndjebet & Ngokoy, 2014). However, according to Din et al. (2002, 2008), natural regeneration is known to be really high in mangroves of the CEC. Rhizophora racemosa, which is the most exploited mangrove species in this estuary, is characterized by a high colonization capacity in Cameroon, and the colonization of new spaces prevail as a result of accelerated sedimentation rates due to high rainfall (Din et al., 2002, 2008) (sedimentation could be further accentuated by land use/land cover changes and logging). This goes without saying that patches created from exploitation have increased recolonization chances as well. The impact of profound human pressure on this estuary could result in the patterns observed. In an earlier study, Ngeve et al. (2016a) observed that none of the studied Rhizophora populations of the Cameroonian coast were inbred. In another study by the same authors, focusing on within-estuary gene flow, all individuals of the CEC were found to make up a single genetic cluster following Bayesian clustering analysis, suggesting high connectivity within this estuary (Ngeve et al., 2016b). The few studies on FSGS in mangroves, carried out on Avicennia spp. and Rhizophora mangle L., have especially linked FSGS to limited pollen and propagule dispersal (Cerón-Souza et al., 2012; Mori et al., 2015; Millán-Aguilar et al., 2016).
All populations but one with significant FSGS showed signs of recent reduction in effective population sizes (bottleneck events), when assuming the SMM; only 2 populations additionally showed significant bottleneck under the assumption of the TPM. Congruent results when assuming both (SMM and TPM) models may provide more conservative estimates of true bottleneck signatures; however, allele size permutations indicate that the SMM may be the mutation model driving patterns of population differentiation, at least for the more pristine Rio del Rey Estuary—reasons why the bottleneck signatures following the SMM are assumed to be reasonably conclusive for this study. The bottleneck signatures could be the result of extreme fragmentation (e.g. see Millán-Aguilar et al., 2016, for Avicennia germinans L. in Mexico). However, recently, climate change-induced sea level rise has resulted in massive diebacks of seaward mangroves of the CEC (Ellison & Zouh, 2012). Therefore, apart from direct human-induced pressure on this mangrove system, diebacks could as well contribute to the signs of recent bottleneck events observed (e.g. see De Ryck et al., 2016, for South African mangroves). Therefore, it is sensible to assume that the CEC populations of Rhizophora racemosa could be made up of hidden founders due to strong human impacts on them. Otherwise, the reduction in effective population sizes could indicate the expansion and colonization of new sites by local recruitment, making the CEC an old but very “dynamic” meta-population.
Generally, in species with long life spans (like many tree species), the occurrence of many cohorts (of different age classes) within a population is likely, resulting in increased within-population genetic variability (thus reduced FSGS) (Loveless & Hamrick, 1984; Addisalem et al., 2016). However, logging pressure could erase such signatures. The large (comparative) historical effective population size of the CEC (second largest after Ntem), in contrast to signs of recent reduction in effective population sizes in most of its populations, indeed highlights the effects of the current anthropogenic pressure on this estuary. The bottleneck signatures as well as the FSGS support our hypothesis of hidden founders in this estuary, i.e. population (re-)colonization that results in FSGS within populations amidst (relatively) high genetic diversity and low differentiation among populations (high gene flow). These patterns of high within-estuary gene flow in the CEC (Ngeve et al., 2016a, b) together with new colonists from local recruitment are responsible for maintaining an ancient large and thriving mangrove population in the CEC.
Estimates of historical dispersal rates and expansion models indicate that mangroves have expanded from south to north along this coastline, implying a stepping stone model of dispersal. Therefore, the CEC is unlikely to be the “most recent” area to have been colonized. As such, the patterns of FSGS are not as a result of historical founders but rather as a result of recent hidden founders. On the contrary, the modelled virtual propagule dispersal pathways indicate that current dispersal patterns strongly differ from historical patterns (see Ngeve et al., 2016a). The northern Rio del Rey and CEC, found on one side of an ocean current front (converging ocean currents), were highly connected in the recent past, while the Ntem and Lokoundje areas are located on the south of this barrier (Ngeve et al., 2016a). Similar patterns were observed in recent marine sediments, of which the littoral drift acts as a major vector for transporting mangrove pollen northward from the Ntem Estuary, as well as pollen transported southward by the Guinea Current into the Bay of Douala (Van Campo & Bengo, 2004). These findings suggest that before the Last Glacial Maximum (LGM, ca. 21 ka BP), when Bioko Island (Fig. 2) was connected to the mainland, mangrove expansion may have been influenced by the northward flowing longshore current (littoral drift). Therefore, sea level rise-induced changes in the coastal geomorphology could have altered historical ocean current patterns to what they are today. Present-day ocean surface current patterns seem to also make the Rio del Rey Estuary a source area for the CEC, compensating for the pronounced loss due to degradation. This suggests that the CEC’s vast extent persists by continued local colonization of new intertidal areas (see Din et al., 2008), by local recruits (recent local hidden founders), as well as the potential input of propagules from the Rio del Rey Estuary (Ngeve et al., 2016a). Limited propagule founders and subsequent restricted propagule and pollen dispersal were identified as determinants of half-sib family structure in Kandelia candel (L.) Druce (Geng et al., 2008). The estimates of historical expansion of mangroves along the Cameroonian coastline assume that these mangroves are isolated from the mangroves of other areas West African coastline (since only samples from the Cameroonian coastline were used in the simulations). This implies that patterns of multiple colonization/expansion events, with founders from other areas of the W. African coast and beyond, cannot be inferred from this analysis, and the results herein are therefore very simplistic in nature.
As explained earlier, the occurrence of FSGS in populations can result from several factors (see Loveless & Hamrick, 1984; Hamrick & Nason, 1996; Smouse & Peakall, 1999; Pandey & Rajora, 2012). However, all else being equal, unrestricted pollen and seed dispersal among populations would rule out the probability of FSGS occurring. The only site of the CEC that showed no signs of bottleneck or FSGS is that of the upstream Wouri River (site SADI). This upstream area is generally less inhabited and less developed, so the absence of FSGS and bottleneck signatures in this area may reflect reduced anthropogenic pressure. Additionally, following propagule release-recapture experiments by Ngeve et al. (2016b), propagules released at this upstream site were the least recovered (the majority had dispersed far beyond the search area of the Wouri River channel), and even for the minority of propagules retrieved (4%), they were observed to have dispersed over longer distances (see Ngeve et al., 2016b). This indicates that stronger river flow currents in upstream sites could allow more propagules to leave the system and disperse over longer distances, such that there is reduced spatial aggregation of propagules in this site (see Honnay et al., 2010), resulting in the absence of FSGS.
Compared to the CEC, mangroves of the Lokoundje Estuary are also highly disturbed. At Londji (Lokoundje), for instance, only sparsely distributed adult stands remain (and only adult trees were sampled for this study), although recent reforestation efforts (<5 years) resulted in the growth of very young trees in between the adult stands. The absence of FSGS in these mangroves could have resulted from the mass extraction of highly related “near-neighbours” of the remnant stands, creating the large gaps between adult stands that have not (yet) been recolonized. Also, the reduction of tree density would reduce the retention of propagules near their maternal trees, favouring conditions for long(er) distance dispersal events. Unlike the CEC, the Mangroves in Lokoundje are sparse in distribution, while those in the CEC are dense (see Table S1). This variation in forest density could account for the absence of FSGS in the remnant adults of the Lokoundje Estuary.
Additionally, FSGS can also be explained by landscape features and estuarine geomorphology (Cerón-Souza et al., 2012). The geomorphology of a coastline could be implicated in prohibiting the effective dispersal of mangrove propagules, such as observed for Avicennia marina in some South African populations (De Ryck et al., 2016). Compared to other areas of the coastline, populations of the CEC generally showed the shortest propagule dispersal distances, as virtual propagules released at these sites dispersed rather locally. While the dispersal model used herein does not consider pre- or post-dispersal retention by dense root systems, estuarine and local landscape features (as well as the contemporary ocean current patterns) may contribute to the reduced dispersal distances from populations of the CEC with severe implications on the FSGS, as observed. Hence, the mangroves of the CEC may act more as a sink, rather than as a source; although within the estuary, “microrefugia” may be responsible for rejuvenation and expansion from internal input (hidden founders). We also found a pronounced negative relationship (R = −0.71) between the calculated dispersal distances and the genetic neighbourhood (a measure of FSGS), suggesting that limited propagule dispersal may accentuate the development of FSGS in populations of the CEC.
Hypothesis 3
Comparative estuarine-scale genetic structure
Comparative genetic differentiation among estuaries shows low to moderate levels of differentiation within all estuaries (Fst < 0.15, P = < 0.001). This could be explained by the high dispersal potential of R. racemosa (once released from the system). Results of allele size permutation test do not support longer divergence times (accumulation of stepwise mutation) of most (CEC, Lokoundje, and Ntem) mangrove areas. This is likely due to the high anthropogenic pressure in these estuaries resulting in either more frequent recolonization events and/or in less genetic variation, since the most pristine area (Rio del Rey) seems to have significantly accumulated more mutations than the disturbed ones. According to Hardy et al. (2003), when the normal Rst value is significantly greater than the permuted Rst value, populations have undergone longer divergence times such that there has been sufficient accumulation of mutations to contribute more to the observed patterns of differentiation. The normal Rst value was only significantly greater than the permuted Rst value in the most pristine estuary, i.e. Rio del Rey. The variation in anthropogenic pressure in different estuaries could also restrict the accumulation of mutations, i.e. genetic variation, in some areas as compared to others.
Analyses of molecular variance (AMOVA) showed great variation among estuaries. Although the correlation between “disturbance” and genetic differentiation was not significant, the patterns revealed from AMOVA could be interpreted by the varying degrees of human disturbance on these different estuaries. AMOVA-Fst indicated that the Lokoundje Estuary had the highest variation among individuals (27%) and among populations (12%), compared to other estuaries. This is likely the result from the mass extraction of trees that have left remnants of distantly located unrelated individuals, as we have discussed above. The Rio del Rey Estuary being the most pristine mangrove area (Corcoran et al., 2007; FAO, 2011), as well as the Ntem Estuary which is protected within the confines of the Campo Ma’an National Park, had the lowest genetic variation among populations. High levels of gene flow among populations of the Rio del Rey (Nm = 7) and Ntem (Nm = 15) estuaries, and reduced deforestation in these areas as compared to the CEC and the Lokoundje Estuary, are concordant with reduced genetic structuring of these estuaries. The impact of mangrove forest fragmentation on genetic structuring is more pronounced for naturally small populations, due to the higher effects of genetic drift on smaller populations. Therefore, the Lokoundje Estuary, which is small, marginal, and an estuary in a disturbed state (with some highly fragmented areas e.g. Londji), appeared to be more genetically structured than the CEC which has been reported as the most anthropized. Indeed, genetic drift severely affects smaller and more fragmented populations, resulting in strong differentiation, while gene flow in turn balances out genetic drift in large and less fragmented populations (see Leonardi et al., 2012) like the CEC. The availability of suitable sites for (re-)colonization (see Din et al., 2002, 2008) as well as the combination of high levels of fragmentation, multiple founder events (locally derived and potential input from the Rio del Rey), and high within-estuary dispersal (Ngeve et al., 2016b) interplay to maintain the mangrove genetic differentiation of the CEC. On the contrary, strong genetic differentiation of the Lokoundje Estuary could have arisen as a result of extreme fragmentation and subsequent reduction in gene flow, on a naturally small mangrove area. Logging disturbance has been shown to increase genetic differentiation among populations, especially in species with limited (pollen and propagule) dispersal (Jalonen et al., 2014).
Conclusion
Despite the efficiency of hydrochory as a dispersal mechanism in Rhizophora populations, a high retention potential (mediated by their roots) may not be a driving force for the induction of FSGS. Other local features, such as anthropogenic pressure, could be more significant drivers at that scale. Compared to pristine areas, higher FSGS was observed in areas with strong (direct and indirect) anthropogenic pressure, for areas that still sustain a dense mangrove forest. Disturbed areas that were very sparse (like Lokoundje) on the contrary did not show signs of FSGS. This indicates that in the presence of high gene flow and when local conditions allow for high (re-)colonization rates, fragmentation-induced genetic bottlenecks and founder events can create spatial structure at fine scale (e.g. CEC). In areas with reduced chances of (re-)colonization, fragmentation is likely to result in the removal of highly related near-neighbouring individuals and evade signatures of FSGS in areas with sparsely distributed remnant stands (e.g. Lokoundje). In addition to anthropogenic pressure, landscape elements such as forest density, continuous re-expansion/recolonization of mangroves, and coastal/estuarine geomorphology, can result in the observed patterns of FSGS even in core populations. These insights have wide applications in strategizing for the conservation of these populations.
References
Addisalem, A. B., J. Duminil, D. Wouters, D. F. Bongers & M. J. M. Smulders, 2016. Fine-scale spatial genetic structure in the frankincense tree Boswellia papyrifera (Del.) Hochst. and implications for conservation. Tree Genetics & Genomes 12: 86.
Alemagi, D., P. M. Oben & J. Ertel, 2006. Mitigating industrial pollution along the atlantic coast of cameroon: an overview of government efforts. Environmentalist 26: 41–50.
Allen, J. A. & K. W. Krauss, 2006. Influence of propagule flotation longevity and light availability on establishment of introduced mangrove species in Hawaii. Pacific Science 60: 367–376.
Arnaud-Haond, S., S. Teixeira, S. I. Massa, C. Billot, P. Saenger, G. Coupland, C. M. Duarte & E. A. Serrão, 2006. Genetic structure at range edge: low diversity and high inbreeding in Southeast Asian mangrove (Avicennia marina) populations. Molecular Ecology 15: 3515–3525.
Balloux, F. & N. Lugon-Moulin, 2002. The estimation of population differentiation with microsatellite markers. Molecular Ecology 11: 155–165.
Beerli, P., 2006. Comparison of Bayesian and maximum likelihood inference of population genetic parameters. Bioinformatics 22: 341–345.
Beerli, P. & M. Palczewski, 2010. Unified framework to evaluate panmixia and migration direction among multiple sampling locations. Genetics 185: 313–326.
Cerón-Souza, I., E. Bermingham, W. O. McMillan & F. A. Jones, 2012. Comparative genetic structure of two mangrove species in Caribbean and Pacific estuaries of Panama. BMC Evolutionary Biology 12: 205.
Chen, L., L. Xu & H. Huanga, 2007. Genetic diversity and population structure in Vallisneria spinulosa (Hydrocharitaceae). Aquatic Botany 86: 46–52.
Chung, M. Y. & M. G. Chung, 2013. Significant spatial aggregation and fine-scale genetic structure in the homosporous fern Cyrtomium falcatum (Dryopteridaceae). New Phytologist 199: 663–672.
Corcoran, E., C. Ravilious, & M. Skuja, 2007. Mangroves of Western and Central Africa. By ISBN: 978-92-807-2792-0. UNEP-WCMC Biodiversity Series 26 (2007).
De Ryck, D. J. R., E. M. R. Robert, N. Schmitz, T. Van der Stocken, D. Di Nitto, F. Dahdouh-Guebas & N. Koedam, 2012. Size does matter, but not only size: two alternative dispersal strategies for viviparous mangrove propagules. Aquatic Botany 103: 66–73.
De Ryck, D. J. R., N. Koedam, T. Van der Stocken, R. M. van der Ven, J. Adams & L. Triest, 2016. Dispersal limitation of the mangrove Avicennia marina at its South African range limit in strong contrast to connectivity in its core East African region. Marine Ecology Progress Series 545: 123–134.
Din, N., R. J. Priso, M. Kenne, D. E. Ngollo & F. Blasco, 2002. Early growth stages and natural regeneration of Avicennia germinans (L.) stearn in the Wouri estuarine mangroves (Douala-Cameroon). Wetlands Ecology and Management 10: 461–472.
Din, N., P. Saenger, R. J. Priso, D. S. Dibong & F. Blasco, 2008. Logging activities in mangrove forests: a case study of Douala Cameroon. African Journal of Environmental Science and Technology 2: 22–30.
Drexler, J. Z., 2001. Maximum longevities of Rhizophora apiculata and R. mucronata propagules. Pacific Science 55: 17–22.
Ellison, J. C., & I. Zouh, 2012. Vulnerability to climate change of mangroves: Assessment from Cameroon, Central Africa. Biology 1: 617–638. doi:10.3390/biology1030617.
Epperson, B. K., 2007. Plant dispersal, neighbourhood size and isolation by distance. Molecular Ecology 16: 3854–3865.
Epperson, B. K. & L. Tianquan, 1996. Measurement of genetic structure within populations using Moran’s spatial autocorrelation statistics. Proceedings of the National Academy of Science USA 93: 10528–10532.
FAO, 2011. FAO/Global environment facility project document. CBSP—sustainable community-based management and conservation of mangrove ecosystems in Cameroon. GEF project ID: 3821, FAO project symbol: GCP/CMR/030/GFF.
Feka, N. Z. & A. G. Ajonina, 2011. Drivers causing decline of mangrove in West-Central Africa: a review. International Journal of Biodiversity Science, Ecosystem Services & Management 7: 217–230.
Feka, Z. N. & I. Morrison, 2017. Managing mangroves for coastal ecosystems change: a decade and beyond of conservation experiences and lessons for and from west-central Africa. Journal of Ecology and The Natural Environment 9: 99–123.
Geng, Q., C. Lian, S. Goto, J. Tao, M. Kimura, M. S. Islam & T. Hogetsu, 2008. Mating system, pollen and propagule dispersal, and spatial genetic structure in a high-density population of the mangrove tree Kandelia candel. Molecular Ecology 17: 4724–4739.
Gillis, L. G., M. Zimmer & T. J. Bouma, 2016. Mangrove leaf transportation: do mimic Avicennia and Rhizophora roots retain or donate leaves? Marine Ecology Progress Series 551: 107–115.
Hamilton, S. E. & D. Casey, 2016. Creation of a high spatio-temporal resolution global database of continuous mangrove forest cover for the 21st century. Global Ecology and Biogeography 25: 729–738.
Hamrick, J. L. & J. D. Nason, 1996. Consequence of dispersal in plants. In Rhodes, O. E., K. C. Ronald & M. H. Smith (eds), Population Dynamics in Ecological Space and Time. The University of Chicago Press, Chicago: 203–235.
Hardy, O. J. & X. Vekemans, 2002. SPAGeDi: a versatile compute program to analyse spatial genetic structure at the individual or population levels. Molecular Ecology Notes 2: 618–620.
Hardy, O. J., N. Charbonnel, H. Fréville & M. Heuertz, 2003. Microsatellite allele sizes: a simple test to assess their significance on genetic differentiation. Genetics 163: 1467–1482.
Hardy, O. J., S. Gonzalphalez-Martnunez, B. Colas, H. Friotaville, A. Mignot & I. Olivieri, 2004. Fine-scale genetic structure and gene dispersal in Centaurea corymbosa (Asteraceae). II. Correlated paternity within and among sibships. Genetics 168: 1601–1614.
Helsen, K., H. Jacquemyn & O. Honnay, 2015. Hidden founder effects: small-scale spatial genetic structure in recently established populations of the grassland specialist plant Anthyllis vulneraria. Molecular Ecology 24: 2715–2728.
Hill, C., D. Menemenlis, B. Ciotti & C. Henze, 2007. Investigating solution convergence in a global ocean model using a 2048-processor cluster of distributed shared memory machines. Scientific Programming 12: 107–115.
Honnay, O., H. Jacquemy, K. Nackaerts, P. Breyne & K. Van Looy, 2010. Patterns of population genetic diversity in riparian and aquatic plant species along rivers. Journal of Biogeography 37: 1730–1739.
Islam, M. S., C. Lian, N. Kameyama & T. Hogetsu, 2014. Low genetic diversity and limited gene flow in a dominant mangrove tree species (Rhizophora stylosa) at its northern biogeographical limit across the chain of three Sakishima islands of the Japanese archipelago as revealed by chloroplast and nuclear SSR analysis. Plant Systematics and Evolution 300: 1123–1136.
Jalonen, R., L. T. Hong, S. L. Lee, J. Loo & L. Snook, 2014. Integrating genetic factors into management of tropical Asian production forests: a review of current knowledge. Forest Ecology and Management 315: 191–201.
Kennedy, J. P., M. W. Pil, C. E. Proffitt, W. A. Boeger, A. M. Stanford & D. J. Devlin, 2016. Postglacial expansion pathways of red mangrove, Rhizophora mangle, in the Caribbean Basin and Florida. American Journal of Botany 103: 260–276.
Kinlan, B. P. & S. D. Gaines, 2003. Propagule dispersal in marine and terrestrial environments: a community perspective. Ecology 84: 2007–2020.
Kudoh, H. & D. F. Whigham, 1997. Microgeographic genetic structure and gene flow in Hibiscus moscheutos (Malvaceae) populations. American Journal of Botany. 84: 1285–1293.
Latch, E. K., D. M. Reding, J. R. Heffelfinger, C. H. Alcalá-Galván & O. E. Rhodes, 2014. Range-wide analysis of genetic structure in a widespread, highly mobile species (Odocoileus hemionus) reveals the importance of historical biogeography. Molecular Ecology 23: 3171–3190. PMID: 24863151.
Leonardi, S., P. Piovani, M. Scalfi, A. Piotti, R. Giannini & P. Menozzi, 2012. Effect of habitat fragmentation on the genetic diversity and structure of peripheral populations of beech in Central Italy. Journal of Heredity 103: 408–417.
Li, J., Y. Yang, Q. Chen, L. Fang, Z. He, W. Guo, S. Qiao, Z. Wang, M. Guo, C. Zhong, R. Zhou & S. Shi, 2016. Pronounced genetic differentiation and recent secondary contact in the mangrove tree Lumnitzera racemosa revealed by population genomic analyses. Scientific Reports 6: 29486.
Loveless, M. D. & J. L. Hamrick, 1984. Ecological determinants of genetic structure in plant populations. Annual Review of Ecology and Systematics 15: 65–95.
Maguire, T. L., P. Saenger, P. Baverstock & R. Henry, 2000. Microsatellite analysis of genetic structure in the mangrove species Avicennia marina (Forsk.) Vierh. (Avicenniaceae). Molecular Ecology 9: 1853–1862.
Menemenlis, D., J. M. Campin, P. Heimbach, C. Hill, T. Lee, A. Nguyen, M. Schodlok & H. Zhang, 2008. ECCO2: high resolution global ocean and sea ice data synthesis. Mercator Ocean Quarterly Newsletter 31: 13–21.
Millán-Aguilar, O., M. Manzano-Sarabia, A. Nettel-Hernanz, R. S. Dodd, M. Á. Hurtado-Oliva & E. Velázquez-Velázquez, 2016. Genetic Diversity of the Black Mangrove Avicennia germinans (L.) Stearn in Northwestern Mexico. Forests 7: 197.
Mori, G. M., M. I. Zucchi & A. P. Souza, 2015. Multiple-geographic-scale genetic structure of two mangrove tree species: the roles of mating system, hybridization, limited dispersal and extrinsic factors. PLoS ONE 10(2): e0118710.
Nathan, R., F. M. Schurr, O. Spiegel, O. Steinitz, A. Trakhtenbrot & A. Tsoar, 2008. Mechanisms of long-distance seed dispersal. Trends in Ecology & Evolution 23: 638–647.
Ndjebet., C. & P. Ngokoy, 2014. Master plan for Cameroon Estuary mangrove forests. ITTO Tropical Forest Update 23:13–15. http://www.itto.int/partner/id=4110.
Nfotabong-Atheull, A., N. Din, S. N. Longonje, N. Koedam & F. Dahdouh-Guebas, 2009. Commercial activities and subsistence utilization of mangrove forests around the Wouri estuary and the Douala-Edea reserve (Cameroon). Journal of Ethnobiology and Ethnomedicine. doi:10.1186/1746-4269-5-35.
Nfotabong-Atheull, A., N. Din, L. G. E. Koum, B. Satyanarayana, N. Koedam & F. Dahdouh-Guebas, 2011. Assessing forest products usage and local residents’ perception of environmental changes in peri-urban and rural mangroves of Cameroon, Central Africa. Journal of Ethnobiology and Ethnomedicine. doi:10.1186/1746-4269-7-41.
Nfotabong-Atheull, A., N. Din & F. Dahdouh-Guebas, 2013. Qualitative and quantitative characterization of mangrove vegetation structure and dynamics in a peri-urban settings of Douala (Cameroon): an approach using airborn imagery. Estuaries and Coasts 36: 1181–1192.
Ngeve, M. N., M. Leermakers, M. Elskens & M. Kochzius, 2015. Assessment of trace metal pollution in sediments and intertidal fauna at the coast of Cameroon. Environmental Monitoring and Assessment 187: 337.
Ngeve, M. N., T. Van der Stocken, D. Menemenlis, N. Koedam & L. Triest, 2016a. Contrasting effects of historical sea level rise and contemporary ocean currents on regional gene FLOW of Rhizophora racemosa in Eastern Atlantic Mangroves. PLoS ONE 11(3): e0150950.
Ngeve, M. N., T. Van der Stocken, T. Sierens, N. Koedam & L. Triest, 2016b. Bidirectional gene flow on a mangrove river landscape and between-catchment dispersal of Rhizophora racemosa (Rhizophoraceae). Hydrobiologia. doi:10.1007/s10750-016-3021-2.
Nilsson, C., R. L. Brown, R. Jansson & D. M. Merritt, 2010. The role of hydrochory in structuring riparian and wetland vegetation. Biological reviews of the Cambridge Philosophical Society 85: 837–858.
Pandey, M. & O. P. Rajora, 2012. Higher fine-scale genetic structure in peripheral than in core populations of a long-lived and mixed-mating conifer—eastern white cedar (Thuja occidentalis L.). BMC Evolutionary Biology 12: 48.
Peakall, R. & P. E. Smouse, 2012. GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28: 2537–2539.
Peakall, R., M. Ruibal & D. B. Lindenmayer, 2003. Spatial autocorrelation analysis offers new insights into gene flow in the Australian bush rat, Rattus fuscipes. Evolution 57: 1182–1195.
Pil, M. W., M. R. T. Boeger, V. C. Muschner, M. R. Pie, A. Ostrensky & W. A. Boeger, 2011. Postglacial north–south expansion of populations of Rhizophora mangle (Rhizophoraceae) along the Brazilian coast revealed by microsatellite analysis. American Journal of Botany 98: 1031–1039.
Piry, S., G. Luikart, & J.-M. Cornuet, 1999. BOTTLENECK: A program for detecting recent effective population size reductions from allele data frequencies. http://www1.montpellier.inra.fr/CBGP/software/Bottleneck/bottleneck.html.
Rabinowitz, D., 1978. Dispersal properties of mangrove propagules. Biotropica 10: 47–57.
Rocha, C. B., T. K. Chereskin, S. T. Gille & D. Menemenlis, 2016. Mesoscale to submesoscale wavenumber spectra in Drake Passage. Journal of Physical Oceanography 46: 601–620.
Saenger, P., & M. F. Bellan, 1995. The mangrove vegetation of the Atlantic Coast of Africa: a review. Universite’ de Toulouse, Toulouse, France. Copyright P. Saenger 1995.
Smith, W. H. F., & D. T. Sandwell, 1977. Global seafloor topography from satellite altimetry and ship depth soundings. Science 277: 1957–1962.
Smouse, P. E. & R. Peakall, 1999. Spatial autocorrelation analysis of individual multiallele and multilocus genetic structure. Heredity 82: 561–573.
Sousa, W. P., P. G. Kennedy, B. J. Mitchell & L. Ordóñez, 2007. Supply-side ecology in mangroves: do propagule dispersal and seedling establishment explain forest structure? Ecological Monographs 77: 53–76.
Steele, O., 2006. Natural and anthropogenic biogeography of mangroves in the Southwest Pacific. PhD Dissertation. University of Hawaii, Manoa.
Tonné, N., N. Koedam, N. Buls, J. De Mey, H. Beeckman & E. M. R. Robert, 2016. Computed tomography and light microscopy: combining visualisation techniques in the study of mangrove seedling development. The International Association of Wood Anatomists 37: 28–47.
Tonné, N., H. Beeckman, E. M. R. Robert & N. Koedam, 2017. Towards an unknown fate: the floating behaviour of recently abscised propagules from wide ranging Rhizophoraceae mangrove species. Aquatic Botany. doi:10.1016/j.aquabot.2017.01.008.
Trakhtenbrot, A., R. Nathan, G. Perry & D. M. Richardson, 2005. The importance of long-distance dispersal in biodiversity conservation. Diversity and Distributions 11: 173–181.
Van Campo, E. & M. D. Bengo, 2004. Mangrove palynology in recent marine sediments off Cameroon. Marine Geology 208: 315–330.
Van der Stocken, T., D. J. R. De Ryck, T. Balke, T. J. Bouma, F. Dahdouh-Guebas & N. Koedam, 2013. The role of wind in hydrochorous mangrove propagule dispersal. Biogeosciences 10: 3635–3647.
Van der Stocken, T., B. Vanschoenwinkel, D. J. R. De Ryck, T. J. Bouma, F. Dahdouh-Guebas & N. Koedam, 2015a. Interaction between water and wind as a driver of passive dispersal in mangroves. PLoS One 10(3): e0121593.
Van der Stocken, T., D. J. R. De Ryck, B. Vanschoenwinkel, E. Deboelpaep, T. J. Bouma, F. Dahdouh-Guebas & N. Koedam, 2015b. Impact of landscape structure on propagule dispersal in mangrove forests. Marine Ecology Progress Series 524: 95–106.
Vekemans, X. & O. J. Hardy, 2004. New insights from fine-scale spatial genetic structure analyses in plant populations. Molecular Ecology 13: 921–935.
Vergara, M., M. P. Basto, M. J. Madeira, B. J. Gómez-Moliner, M. Santos-Reis, C. Fernandes & A. Ruiz-González, 2015. Inferring population genetic structure in widely and continuously distributed carnivores: the stone marten (Martes foina) as a case study. PLoS ONE 10: e0134257.
Vigouroux, Y., J. S. Jaqueth, Y. Matsuoka, O. S. Smith, W. D. Beavis, J. S. C. Smith & J. Doebley, 2002. Rate and pattern of mutation at microsatellite loci in maize. Molecular Biology and Evolution 19: 1251–1260.
Wee, A. K. S., K. Takayama, T. Asakawa, B. Thompson, S. Sungkaew, N. X. Tung, M. Nazre, K. K. Soe, H. T. W. Tan, Y. Watano, S. Baba, T. Kajita & E. L. Webb, 2014. Oceanic currents, not land masses, maintain the genetic structure of the mangrove Rhizophora mucronata Lam. (Rhizophoraceae) in Southeast Asia. Journal of Biogeography 41: 954–964.
Wells, G. P. & A. G. Young, 2002. Effect of seed dispersal on genetic structure in populations of Rutidosis leptorrhychoides with different levels of correlated paternity. Genetics Research 79: 219–226.
Acknowledgements
This study was funded by the Vrije Universiteit Brussel—International Relations and Mobility Office (VUB—IRMO) Doctoral Scholarship awarded to Magdalene N. Ngeve. The modelling component of this study was done at the Jet Propulsion Laboratory (JPL), California Institute of Technology (during a JPL Visiting Student Researchers Program (JVSRP)), under a contract with the National Aeronautics and Space Administration (NASA). High-end computing resources were provided by the NASA Advanced Supercomputing (NAS) Division of the Ames Research Center.
Author information
Authors and Affiliations
Corresponding author
Additional information
Guest editors: K. W. Krauss, I. C. Feller, D. A. Friess, & R. R. Lewis III / Causes and Consequences of Mangrove Ecosystem Responses to an Ever-Changing Climate
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ngeve, M.N., Van der Stocken, T., Menemenlis, D. et al. Hidden founders? Strong bottlenecks and fine-scale genetic structure in mangrove populations of the Cameroon Estuary complex. Hydrobiologia 803, 189–207 (2017). https://doi.org/10.1007/s10750-017-3369-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10750-017-3369-y