Abstract
The size and shape of tomato (Solanum lycopersicum L.) fruit are determined by locule number. Gibberellin (GA) can increase locule number in tomato, but the underlying molecular mechanism is unclear. Therefore, in this study, multi-locule ‘MLK1’ tomato seedlings with two to three true leaves (pre-flower bud differentiation) were sprayed with GA1, GA3, GA4, GA7, PAC (paclobutrazol; an inhibitor of GA biosynthesis) and H2O, as a control. We found that GA4 resulted in was the most significant increase in tomato locule number among all bioactive GAs, while PAC decreased the locule number. We analyzed the change in locule number by RNA-seq, quantitative real-time PCR and ultra-performance liquid chromatography–tandem mass spectrometry. The categories ‘Phenylpropanoid biosynthesis’, ‘Plant hormone signal transduction’, and ‘Diterpenoid biosynthesis’ were considerably activated after spraying with GA4. Additionally, indole-3-acetic acid (IAA) content significantly increased and trans-Zeatin-riboside (tZ) content significantly reduced after exogenous GA4 application. We conclude that exogenous GA4 application changed the dynamic balance of hormones in the tomato shoot apex. Furthermore, 53 differentially expressed transcription factors were identified in tomato upon exogenous GA4 treatment during floral bud differentiation, including, YABBYs, TCP, NAC, and ARR (some directly regulate lateral organ development). Our results provide novel insights into how exogenous GA4 affects plant hormone homeostasis in the tomato shoot apex and the underlying mechanism of locule number regulation by GA4 in tomato.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The number of locules in tomato (Solanum lycopersicum L.) is an important factor affecting the size and shape of tomato fruit and the incidence of fruit malformation (Liu and Li 2012; Tanksley 2004). In the dicot plant tomato, the shoot apical meristem (SAM) includes the following three functional regions: the subjacent organizing center (OC) of the central zone, peripheral zone (PZ) cells on both sides, and rib zone (RZ) cells at the base (Galli and Gallavotti 2016). The daughters of stem cells that are distributed in the PZ begin to differentiation and eventually form leaves, lateral branches, and flowers, and cells in the RZ form pith tissue of the stem (Bowman and Eshed 2000; Holt et al. 2014). The tomato locules develop from the inner whorl of flower organs, carpel (Lozano et al. 2009). The number of carpels in a tomato flower determines the final number of locules in a mature fruit (Cong et al. 2008a).
The normal SAM function requires the maintenance of a delicate balance between the lateral organs produced from its flanks and uncertain growth at its center. The classic phytohormones gibberellin (GA), cytokinin (CK) and auxin play essential roles in maintaining stem-cell systems in shoot meristems, and exhibit complex functional interactions (Shani et al. 2006). In the SAM, CK accumulates in the OC, the apex of the SAM, while GA and auxin accumulates in the PZ. The resulting high CK: auxin ratio and low GA concentration promotes indeterminate growth in the OC, whereas low CK: auxin ratio, and high GA level, promote initiation of lateral organ (Shani et al. 2006). However, very little is known about the role of GA in locule formation.
Gibberellins are phytohormones that regulate multiple developmental processes, such as, seed germination, stem elongation, flowering, and fruit development (Eriksson et al. 2006; Fleet and Sun 2005; Gou et al. 2010; Yu et al. 2004). Due to their important roles in plant development, The metabolism and signaling pathways of GA have been dissected in depth (Olszewski et al. 2002). In the early steps of GA biosynthesis, GGDP is converted into GA12 by four enzymes, ent -copalyl diphosphate synthase (CPS), ent -kaurene synthase (KS), ent -kaurene oxidase (KO) and ent -kaurenoic acid oxidase (KAO) (Helliwell et al. 2001; Sun and Kamiya 1997). Subsequently, GA 20-oxidases (GA20ox) and GA 3-oxidases (GA3ox) catalyze the final steps in the synthesis of bioactive GA, whereas GA 2-oxidase (GA2ox) is the major GA deactivation enzyme (Hedden and Thomas 2012; Yamaguchi 2008). Once bioactive GAs are synthesized, their signals are perceived by the soluble GA receptor, GID1 (Shimada et al. 2008). By perceiving GA signals, GID1 can interact with the DELLA protein, a negative regulator of GA response (Ikeda et al. 2001; Sun 2010).
The role of GA in shoot apex development is poorly understood. However, several studies have suggested that GA plays a role in meristem function. In Arabidopsis, the DELLA protein acts as a negative regulators of GAs and limits inflorescence meristem size by directly upregulating cyclin-dependent kinase inhibitor (KRP2) in RZ (Serranomislata et al. 2017). In rice, GA2ox gene expression in RZ, which decreases following floral transition and limits the access of bioactive GAs in SAM (Sakamoto et al. 2001). The expression pattern of the GA2ox gene at the shoot apex has been reported in Arabidopsis, and it is similar to that in rice (Jasinski et al. 2005). Furthermore, the root length of GA2ox2- overexpressing lines was shorter than that of wild type in Arabidopsis due to the lower number of root meristem cells than the wild type (Li et al. 2017b).
The role of GA in carpel formation is almost completely unknown. Several lines of evidence suggest that bioactive GA can promote carpel formation, resulting in an increase in the number of locules (Liu and Li 2012; Sawhney 1983). For example, studies on tomatoes suggest that exogenous GA3 can increase locule number, while paclobutrazol (the inhibitor of GA biosynthesis), can decrease locule number (Liu and Li 2012; Sawhney 1983). Here, we demonstrate the main role of GA in carpel formation in tomato through the application of GA and the subsequent physiological experiments, and provide an in-depth understanding of the transcriptional mechanisms underlying its effects.
Materials and methods
Plant materials and treatments
The multi-locule tomato variety ‘MLK1’ has a larger fruit with 10 or more locules was grown in a greenhouse (28℃/15℃, day/night) with natural solar radiation and the humidity was approximately 60% (Li et al. 2017a). Seedlings with two to three true leaves (pre-flower bud differentiation) were sprayed with GA1 (100 µM), GA3 (100 µM), GA4 (100 µM), GA7 (100 µM), PAC (100 µM), and H2O as a control, every two days for three times per treatment. Sampling after one week of treatment, that is, the carpel primordium formation period For the GA4, PAC, and H20 treatments, shoot apexes (each approximately 5 mm) were removed from plants for RNA sequencing (RNA-Seq) and quantification of phytohormones.
Phenotypic analysis of tomato fruit
Four mature fruits of the first inflorescences from 10 tomato plants of the GA1, GA3, GA4, GA7, PAC and H2O treatments were used to analyze the locule number. Furthermore, the H2O, GA4, and PAC treatments and their phenotypic characteristics were recorded.
RNA isolation and RNA-Seq
RNA isolation, sequencing library construction and Illumina sequencing were performed by Novogene Company (Beijing, China). Total RNA was isolated from the shoot apex samples (three biological replicates) using the TRIzol reagent (Invitrogen, USA). RNA degradation and contamination were monitored on 1% agarose gels. RNA purity was determined using NanoPhotometer® spectrophotometer (IMPLEN, ca., USA). RNA concentration was measured using the Qubit® RNA Assay Kit on Qubit® 2.0 Flurometer (Life Technologies, ca., USA). RNA integrity was assessed using the RNA Nano 6000 Assay Kit on the Bioanalyzer 2100 system (Agilent Technologies, ca., USA).
A sequencing library was generated using the NEBNext® Ultra™ RNA Library Prep Kit for Illumina® (NEB, USA) following manufacturer’s recommendations. The index codes were added to attribute sequences of each sample. mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. The cleaved RNA fragments were transcribed into first-strand cDNA using reverse transcriptase, and then second-strand cDNA was synthesized using DNA polymerase I and RNase H. The fragments were ligated to sequencing adaptors and the library preparations were sequenced on an Illumina HiSeq 4000 platform to generate 150 bp paired-end reads (Lei et al. 2018).
De novo assembly and functional annotation
Raw reads were cleaned by removing adapters and low-quality sequences. The reference genome index was constructed using Bowtie v2.2.3 and paired-end clean reads were compared with the reference genomes using TopHat v2.0.12. Then, use HTSeq v0.6.1 was used to calculate the read numbers mapped to each gene. Finally, the FPKM of each gene was calculated based on the length of the gene and read number mapped to this gene.
Differential expression analysis was performed using DESeq R package (1.18.0). The resulting P values were adjusted using the approach of Benjamini and Hochberg’s to control the false discovery rate. Using FDR < 0.05 as the threshold, genes with significantly different expression levels were identified.
Gene Ontology and Kyoto Encyclopedia of Genes and Genomes enrichment analysis
Gene Ontology (GO) enrichment analysis was performed on differentially expressed genes (DEGs) using GOseq R package. The DEGs were significantly enriched to GO terms with a P-value less than 0.05. We used KOBAS software to detect the statistical enrichment of differentially expressed genes in the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways (http://www.genome.jp/kegg/).
Quantification of phytohormones
Approximately 1 g of shoot apex was sampled from GA4, PAC, and H2O (with three biological replications) were used in the analysis. Phytohormones were extracted using a mixture of 2-propanol:H2O:concentrated HCl (2:1:0.002, v/v/v). According to the method of Pan et al. with some modifications, the phytohormones were quantified using a liquid chromatography-mass chromatography system (Agilent 1290, AB company Qtrap6500; USA) (Pan et al. 2010b).
Statistical analysis
Statistical analyses were performed using SPSS 22 Software (SPSS Inc., USA). The mean values were further indicated at P ≤ 0.05(*) using Duncan’s multiple range test for significant difference analysis. All calculations and the figures were generated using MS Excel 2016 (Redmond, WA, USA).
Results
Exogenous application of different bioactive GAs resulted in changes in the locule number in tomato
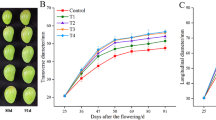
“MLK1”, a multi-locule tomato line, has an average locule number of 13.95. After exogenous bioactive GA1, GA3, GA4, and GA7 treatments, the average locule number increased to 15.93, 15.58, 16.78, and 16.48, respectively. Furthermore, the number of locule significantly increased after GA4 and GA7 treatments (P < 0.05). Among them, GA4 effect was the most obvious, while PAC reduced the locule number to 11.98 (Fig. 1a; Table S1). In addition to increasing the number of locules, GA4 was associated with malformation (Fig. 1b).
Fruit morphology of application of GA1, GA3, GA4, GA7, paclobutrazol (PAC), and H2O, as a control. (a) Quantification and comparison of locule numbers of GA1 (100 µM), GA3 (100 µM), GA4 (100 µM), GA7 (100 µM), PAC (100 µM), and H2O treatments. (b) The effect of GA4, PAC, and H2O treatments on locule number and fruit morphology. The error bars represent the standard errors. Different lowercase letters represent significant differences (p < 0.05, Duncan’s multiple range test). Scale bar: 2 cm
Overview of mRNA sequencing data
To better understand the underlying molecular mechanism of GA effect on locule number, we selected the GA4 treatment for the following portion of this experiment, considering that it caused the most significant change in locule number. We used RNA-Seq to profile the transcriptomes of the GA4, PAC, and H2O treatments in tomato. We obtained 47,293,388, 45,645,392 and 44,261,929 clean reads with three biological repeats from the GA4, PAC, and H2O treatment control libraries, respectively. The mapping rate for each library was above 93%. Additionally, we obtained 45,126,099, 43,209,618 and 41,986,486 uniquely mapped reads, respectively, for further analysis (Table 1). Using comparative analysis, we identified 438 and 67 significantly DEGs from the GA4 and PAC treatments, respectively; among these DGEs, 177 were GA-induced and 261 were GA-repressed, while 43 were PAC-induced and 24 were PAC-repressed (Fig. 2a, Table S2). Furthermore, 26 genes were differentially expressed between GA4 vs. CK and PAC vs. CK (Fig. 2b).
Global analysis of genes expression in response to exogenous GA4 and PAC. a Volcano plot of differentially expressed genes with the cutoff (fold-change > 0.5 and FDR < 0.01). Colors of blue and green represent up- and down-regulated genes, respectively. b Comparison of expression gene numbers among GA4 vs. CK and PAC vs. CK using Venn diagram
Overall identification and functional annotation of DGEs
Tables 1 and 2 show the top 20 GO terms distributed by number of sequences of upregulated and downregulated DEGs, respectively, which were categorized into biological processes, molecular functions and cellular components; the most enriched terms were “catalytic activity”, “binding”, “metabolic process”, “single-organism process”, and “membrane” (Table S3). The KEGG analysis showed that these DEGs are involved in “Phenylpropanoid biosynthesis”, “Diterpenoid biosynthesis”, “Plant hormone signal transduction”, “Starch and sucrose metabolism” and others (Table 2).
GA biosynthetic genes in response to exogenous GA4
From the KEGG enrichment analysis, we found that the genes associated with “Diterpenoid biosynthesis” were significantly overrepresented upon GA4 and PAC treatments. In “GA biosynthetic process”, ent-copalyl diphosphate synthase, ent-kaurenoic acid oxidase 2 (SlKAO2) and gibberellin 20-oxidase 4 (SlGA20ox4), were down-regulated upon exogenous GA4 treatment. Additionally, SlGA20ox4 was up-regulated upon PAC treatment. Finally, Gibberellin 2-oxidase 2 (SlGA2ox2), which encodes GA-inactivating enzyme, was up-regulated upon exogenous GA4 treatment. In this study, we observed that the repression of GA biosynthesis-related genes may be due to the negative feedback regulation of endogenous GA biosynthesis in response to the exogenous GA4 treatment (Fig. 3).
Analysis of genes and CK and auxin levels upon exogenous GA4 and PAC applications
From the KEGG enrichment analysis, we found that the genes associated with ‘Plant hormone signal transduction’ were also significantly overrepresented upon GA4 treatment. For example, the auxin signaling pathway changed in response to the application of exogenous GA4; the expression of SlARF9, SlARF18, SlIAA7, and SAUR32 was up-regulated and only GH3 was down-regulated upon exogenous GA4 treatment in the shoot apex of tomato. Moreover, the CK signaling pathway was inhibited by exogenous GA4; the cytokinin receptor Histidine kinase 2 (SlHK2) and type-A response regulator2 (SlRRA2), which is a negative regulator of cytokinin signaling, were reduced upon exogenous GA4 treatment (Fig. 4c).
Analysis of genes and level related to CK and auxin upon exogenous GA4 and PAC applications. a The expression profile of auxin and CK biosynthesis genes. b The contents of IAA and tZ after GA4 and PAC treatment. c Gene expression pattern of auxin and CK signal transduction related genes after GA4 treatment. Detailed information on each gene and its expression level is listed in Supplementary Table 4
Furthermore, auxin and CK levels were measured, and we found that the indole-3-acetic acid (IAA) content after GA4 treatment reached 5.87 ng g−1, which was 2.08 times higher than that of the control with 2.73 ng g−1, while the PAC treatment showed no significant changes (Fig. 4b). Moreover, the content of trans-Zeatin-riboside (tZ) was 0.176 ng g−1 in the control treatment, and after exogenous application of GA4 significantly reduced to 0.09 ng g−1, it increased to 0.22 ng g−1 after PAC treatment (Fig. 4b).
Furthermore, the expression profile of auxin and CK biosynthesis-related genes was analyzed. We found that auxin synthesis-related genes YUC6 and YUC8 were upregulated after exogenous GA4 application, and the expression of auxin-induced protein was upregulated (Fig. 4a). Meanwhile, cytokinin synthesis-related genes adenosinephosphate-isopentenyltransferase (IPT) and LONELY GUY (LOG) were down-regulated after GA4 application (Fig. 4a).
Expression profiles of TFs upon exogenous GA4 and PAC applications
GA4 and PAC induced or suppressed several TFs in the shoot apex of tomato: 31 and 23 TF genes were up- regulated and down-regulated upon exogenous GA4 applications, while 6 and 3 were up-regulated and down-regulated upon exogenous PAC applications (Table 3). Furthermore, MYB(6), ERF(4), zinc finger(4), Homeobox-leucine zipper (2), ARF(2), NAC(2), IAA(1), MADS-box(1), and TCP(1) were the upregulated TF family members, while MYB(4), YABBY (2), MADS-box(2), Y subunit A-3(2), bHLH (2), BEL1-like(2), GAI(1), and ARR(1) were downregulated TF family members upon exogenous GA4 applications. Upon exogenous PAC application, ERF(1), bHLH(1), WRKY(1), LUX(1), MADS-box(1), and MYB(1) were the upregulated TF family members, while TCP(2), and BTB/POZ(1) were the downregulated TF family members. Among them, the expression of TCP and MYB significantly changed under GA4 and PAC application, and the trend was opposite (Table 3).
Validation of DEGs by qRT-PCR
To verify the precision and reproducibility of the transcriptome data, SlARF9, SlARF18, SlIAA7, SlGH3, SlHK2, SlRRA2, SlCRCa, SlCRCb, SlTCP16, SlTCP17, SlNAC1, SlNAC2, SlWUS, SlCLV3 and SlTAG in the shoot apex were selected for qRT-PCR analysis upon GA4 and PAC treatments to confirm RNA-Seq data. The qRT-PCR assay results were consistent with the transcriptional data (Fig. 5; Table S5).
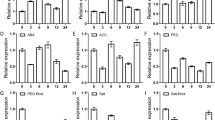
qRT-PCR analysis of SlARF9, SlARF18, SlIAA7, SlGH3, SlHK2, SlRRA2, SlCRCa, SlCRCb, SlTCP16, SlTCP17, SlNAC1, SlNAC2, SlWUS, SlCLV3 and SlTAG in the shoot apex upon GA4 and PAC treatments. The data are the mean values corresponding to three independent experiments. The error bars represent the standard errors. Different lowercase letters represent significant differences (p < 0.05, Duncan’s multiple range test)
Discussion
Domestication of tomato locule number
Cultivated tomato plants produce fruit as much as 1000 times larger than those of their wild progenitors, fruit size is the primary characteristic of commercial tomato varieties and thus an important goal of tomato domestication (Cong et al. 2008b; Lippman and Tanksley 2001; Tanksley 2004). The development of extreme fruit size has been traced to two QTLs, referred to as locule-number (lc) and fascinated (fas) that change SlWUS or SlCLV3 expression, respectively. Silencing SlWUS on an lc mutant genetic background with an RNA interference strategy resulted in smaller flowers and fruit than those of the wild-type plants, with decreased locule number(Li et al. 2017a). When the expression of SlCLV3 is decreased, the locule number increases significantly(Xu et al. 2015). In our RNA-Seq and qRT-PCR data, the expression of SlWUS and SlCLV3 was slightly increased after GA4 and PAC applications. Additionally, the DELLA protein has been reported to restrict inflorescence meristem function in Arabidopsis by directly upregulating the cell cycle inhibitor KRP2 in the underlying rib meristem, without affecting the canonical WUSCHEL-CLAVATA meristem size regulators (Serranomislata et al. 2017). Therefore, it was suggested that GA regulated the tomato locule number via a pathway other than the WUSCHEL-CLAVATA pathway. Furthermore, SlTAG regulates different processes of reproductive development in tomato which involve the identity and development of carpels and the ripening of fruits(Gimenez et al. 2010; Itkin et al. 2009; Pan et al. 2010a). In our RNA-Seq and qRT-PCR data, the expression of SlTAG was not affected by GA4 treatment. Therefore, our results indicate that GA4 application increased the locule number in tomatoes without relying on changes in the expression level of SlWUS, SlCLV3, and SlTAG.
GA induced multi-locule fruit formation
GA plays a vital role in regulating the locule number in tomatoes. For example, previous research found that application of exogenous gibberellin acid (GA3) increased tomato locule number, while PAC application the number (Liu and Li 2012), both of which are consistent with our findings (Fig. 1). Many pathways involved in plant responses to exogenous GA may be conserved in most plants. For instance, the GA signaling pathway crosstalks with the CK and auxin signaling pathway, which is consistent with that in other previously well-studied plant species (Nemhauser et al. 2006; Zhang et al. 2018). Moreover, several lines of evidence suggest that ARR, a negative regulator of cytokinin signaling has an important function in the meristem (Buechel et al. 2010; Giulini et al. 2004; Zhao et al. 2010). In our data, the expression of SlRRA2 was reduced upon exogenous GA4 treatment suggesting that GA action in carpel development is achieved by reducing the expression of SlARR2. Auxin is a positive regulator of plant lateral organ initiation, and the inhibition of polar auxin transport blocks leaf formation in the vegetative tomato meristem, resulting in pin-like naked stems (Reinhardt et al. 2000). In our data, the expression of SlARF9, SlIAA7, and SlSAUR32 was up-regulated upon exogenous GA4 treatment in the shoot apex of tomato. However, auxin regulates the size of the meristem and the number of floral organs, which has to be further validated. Furthermore, exogenous GA rapidly induced more lignin condensation and reduced photosynthesis in moso bamboo seedlings (Zhang et al. 2018). Similarly, in our shoot apex transcriptomics experiments, we found that KEGG enrichment pathways also included “Phenylpropanoid biosynthesis” and “Photosynthesis-antenna proteins” (Table 2). However, its regulatory function on meristem remains to be studied.
Comparative global analysis of TFs potentially involved in the regulation of locule number
The plant life cycle involves many biological processes, such as cell division and differentiation, embryonic development, seed germination, reproductive growth, and flowering, which are regulated by complex transcriptional networks. Fifty-four differentially expressed TFs were identified upon exogenous GA4 treatment during floral bud differentiation. Some of those differentially expressed TFs are related to hormone and stress responses, such as MYBs (Li et al. 2015; Prabu and Prasad 2012; Zhang et al. 2012), ERFs (Mizoi et al. 2012; Sharma et al. 2010), ARFs (Guilfoyle et al. 1998; Jain and Khurana 2009) and RRA (Buechel et al. 2010; Zwack and Rashotte 2015), and others directly regulate lateral organs development. For example, SlCRCa and SlCRCb are a part of the YABBY gene family, and participate in the control of polarity of the lateral organs (Lee et al. 2005). Additionally, SlTCP16 is a member of the SlTCP gene family, which is involved in leaf development and axillary shoots formation (Parapunova et al. 2014). Lastly, NAC transcription factors promote abiotic stress tolerance and lateral root formation in transgenic soybean plants (Hao et al. 2011).
Conclusions
To reveal how GA4 regulates the locule number in tomato, we applied exogenous GA4 and PAC to tomato seedlings, and observed that it could significantly increase and decrease the locule number, respectively. It has been reported that endogenous GA and auxin are located in the PZ region, controlling the differentiation of lateral organs, while endogenous CK is located in the CZ region, promoting the division of stem cells. In this study, we found that exogenous GA4 positively regulated auxin and negatively regulated CK content in the shoot apex of tomato. The expression of SlARF9, SlARF18, SlIAA7, and SlSAUR32 was up-regulated, while the expression of Histidine kinase 2 (SlHK2) and type-A response regulator2 (SlRRA2) was reduced upon exogenous GA4 treatment. Furthermore, we found that some genes related to lateral organ development, such as SlCRCa and SlCRCb, were down-regulated, while SlTCP16 and SlTCP17 were slightly up-regulated upon exogenous GA4 treatment. These results provide theoretical support for the molecular mechanism underlying GA-induced multi-locule fruit formation (Fig. 6).
Model depicting how the exogenous GA4 mediates related gene expression in shoot apex to regulate locule number. The shoot apex is divided into three functional regions, CZ, PZ, and RZ, CK distribution in the CZ region, GA and auxin distribution in the PZ region (Shani et al. 2006). Arrows for promotion, T-shaped arrows for inhibition
References
Bowman JL, Eshed Y (2000) Formation and maintenance of the shoot apical meristem. Trends Plant Sci 5:110–115
Buechel S, Leibfried A, To JPC, Zhao Z, Andersen SU, Kieber JJ, Lohmann JU (2010) Role of A-type ARABIDOPSIS RESPONSE REGULATORS in meristem maintenance and regeneration. Eur J Cell Biol 89:279–284. https://doi.org/10.1016/j.ejcb.2009.11.016
Cong B, Barrero L, Tanksley SD (2008a) Regulatory change in YABBY-like transcription factor led to evolution of extreme fruit size during tomato domestication. Nat Genet 40:800–804
Cong B, Barrero LS, Tanksley SD (2008b) Regulatory change in YABBY-like transcription factor led to evolution of extreme fruit size during tomato domestication. Nat Genet 40:800–804
Eriksson S, Bohlenius H, Moritz T, Nilsson O (2006) GA4 Is the Active Gibberellin in the Regulation of LEAFY Transcription and Arabidopsis Floral Initiation. Plant Cell 18:2172–2181
Fleet CM, Sun T (2005) A DELLAcate balance: the role of gibberellin in plant morphogenesis. Curr Opin Plant Biol 8:77–85
Galli M, Gallavotti A (2016) Expanding the regulatory network for meristem size in. Plants Trends Genet 32:372–383
Gimenez E et al (2010) Functional analysis of the Arlequin mutant corroborates the essential role of the Arlequin/TAGL1 gene during reproductive development of tomato. PLoS ONE 5:e14427. https://doi.org/10.1371/journal.pone.0014427
Giulini A, Wang J, Jackson D (2004) Control of phyllotaxy by the cytokinin-inducible response regulator homologue ABPHYL. Nature 430 1:1031–1034
Gou J, Strauss SH, Tsai C, Fang K, Chen Y, Jiang X, Busov V (2010) Gibberellins regulate lateral root formation in populus through interactions with auxin and other hormones. Plant Cell 22:623–639
Guilfoyle TJ, Ulmasov T, Hagen G, THE ARF FAMILY OF TRANSCRIPTION FACTORS AND THEIR ROLE IN PLANT HORMONE-RESPONSIVE TRANSCRIPTION (1998) Cell Mol Life Sci 54:619–627
Hao Y et al (2011) Soybean NAC transcription factors promote abiotic stress tolerance and lateral root formation in transgenic plants. Plant J 68:302–313
Hedden P, Thomas SG (2012) Gibberellin biosynthesis and its regulation. Biochem J 444:11–25
Helliwell CA, Chandler PM, Poole AJ, Dennis ES, Peacock WJ (2001) The CYP88A cytochrome P450, ent-kaurenoic acid oxidase, catalyzes three steps of the gibberellin biosynthesis pathway. Proc Natl Acad Sci USA 98:2065–2070
Holt AL, Van Haperen JM, Groot EP, Laux T (2014) Signaling in shoot and flower meristems of Arabidopsis thaliana. Curr Opin Plant Biol 17:96–102
Ikeda A et al (2001) Slender Rice, a constitutive gibberellin response mutant, is caused by a null mutation of the SLR1 gene, an ortholog of the height-regulating gene GAI/RGA/RHT/D8. Plant Cell 13:999–1010
Itkin M, Seybold H, Breitel D, Rogachev I, Meir S, Aharoni A (2009) TOMATO AGAMOUS-LIKE 1 is a component of the fruit ripening regulatory network. Plant J 60:1081–1095. https://doi.org/10.1111/j.1365-313X.2009.04064.x
Jain MK, Khurana JP (2009) Transcript profiling reveals diverse roles of auxin-responsive genes during reproductive development and abiotic stress in rice. FEBS J 276:3148–3162
Jasinski S et al (2005) KNOX action in Arabidopsis is mediated by coordinate regulation of cytokinin and gibberellin activities. Curr Biol 15:1560–1565
Liu S, Li TL (2012) Regulation effects of exogenous gibberellin acid (GA3) on the formation of tomato (Solanum lycoperscium) ovary locule and fasciated transcription. Afr J Biotechnol. https://doi.org/10.5897/ajb12.2244
Lee J, Baum SF, Alvarez JP, Patel A, Chitwood DH, Bowman JL (2005) Activation of CRABS CLAW in the nectaries and carpels of arabidopsis. Plant Cell 17:25–36
Lei Y et al (2018) Comparative analysis of alfalfa (Medicago sativa L.) leaf transcriptomes reveals genotype-specific salt tolerance mechanisms. BMC Plant Biol 18:35
Li C, Ng CKY, Fan L (2015) MYB transcription factors, active players in abiotic stress signaling. Environ Exp Bot 114:80–91
Li H et al (2017) Tomato transcription factor SlWUS plays an important role in tomato flower and locule development. Front Plant Sci 8:457
Li H et al (2017) Plant-specific histone deacetylases HDT1/2 regulate GIBBERELLIN 2-OXIDASE2 expression to control arabidopsis root meristem cell. Number Plant Cell 29:2183–2196. https://doi.org/10.1105/tpc.17.00366
Lippman Z, Tanksley SD (2001) Dissecting the genetic pathway to extreme fruit size in tomato using a cross between the small-fruited wild species Lycopersicon pimpinellifolium and L. esculentum var. Giant Heirloom Genet 158:413–422
Lozano R, Gimenez E, Cara B, Capel J, Angosto T (2009) Genetic analysis of reproductive development in tomato. Int J Dev Biol 53:1635–1648
Mizoi J, Shinozaki K, Yamaguchishinozaki K (2012) AP2/ERF family transcription factors in plant abiotic stress responses ☆. Biochem Biophys Acta 1819:86–96
Nemhauser JL, Hong F, Chory J (2006) Different plant hormones regulate similar processes through largely nonoverlapping transcriptional responses. Cell 126:467–475. https://doi.org/10.1016/j.cell.2006.05.050
Olszewski NE, Sun T, Gubler F (2002) Gibberellin signaling: biosynthesis, catabolism, and response pathways. Plant Cell 14:561–580
Pan IL, McQuinn R, Giovannoni JJ, Irish VF (2010a) Functional diversification of AGAMOUS lineage genes in regulating tomato flower and fruit development. J Exp Bot 61:1795–1806. https://doi.org/10.1093/jxb/erq046
Pan X, Welti R, Wang X (2010b) Quantitative analysis of major plant hormones in crude plant extracts by high-performance liquid chromatography–mass spectrometry. Nat Protoc 5:986–992
Parapunova V et al (2014) Identification, cloning and characterization of the tomato TCP transcription factor family. BMC Plant Biol 14:157–157
Prabu G, Prasad DT (2012) Functional characterization of sugarcane MYB transcription factor gene promoter (PScMYBAS1) in response to abiotic stresses and hormones. Plant Cell Rep 31:661–669
Reinhardt D, Mandel T, Kuhlemeier C (2000) Auxin regulates the initiation and radial position of plant lateral organs. Plant Cell 12:507–518
Sakamoto T et al (2001) Expression of a gibberellin 2-oxidase gene around the shoot apex is related to phase transition in rice. Plant Physiol 125:1508–1516
Sawhney VK (1983) The role of temperature and its relationship with gibberellic acid in the development of floral organs of tomato (Lycopersicon esculentum). Botany 61:1258–1265
Serranomislata A, Bencivenga S, Bush M, Schiessl K, Boden SA, Sablowski R (2017) DELLA genes restrict inflorescence meristem function independently of plant height. Nat Plants 3:749–754
Shani E, Yanai O, Ori N (2006) The role of hormones in shoot apical meristem function. Curr Opin Plant Biol 9:484–489
Sharma MK, Kumar R, Solanke AU, Sharma R, Tyagi AK, Sharma AK (2010) Identification, phylogeny, and transcript profiling of ERF family genes during development and abiotic stress treatments in tomato. Mol Genet Genomics 284:455–475
Shimada A et al (2008) Structural basis for gibberellin recognition by its receptor GID. Nature 456 1:520–523
Sun T, Kamiya Y (1997) Regulation and cellular localization of ent-kaurene synthesis. Physiol Plant 101:701–708
Sun T (2010) Gibberellin-GID1-DELLA: a pivotal regulatory module for plant growth and development. Plant Physiol 154:567–570
Tanksley SD (2004) The genetic, developmental, and molecular bases of fruit size and shape variation in tomato. Plant Cell 16:S181–S189
Xu C et al (2015) A cascade of arabinosyltransferases controls shoot meristem size in tomato. Nat Genet 47:784–792
Yamaguchi S (2008) Gibberellin metabolism and its regulation. Annu Rev Plant Biol 59:225–251
Yu H, Ito T, Zhao Y, Peng J, Kumar PP, Meyerowitz EM (2004) Floral homeotic genes are targets of gibberellin signaling in flower development. Proc Natl Acad Sci USA 101:7827–7832
Zhang L, Zhao G, Jia J, Liu XW, Kong X (2012) Molecular characterization of 60 isolated wheat MYB genes and analysis of their expression during abiotic stress. J Exp Bot 63:203–214
Zhang H et al (2018) Transcriptome characterization of moso bamboo (Phyllostachys edulis) seedlings in response to exogenous gibberellin applications. BMC Plant Biol 18:125
Zhao Z, Andersen SU, Ljung K, Dolezal K, Miotk A, Schultheiss SJ, Lohmann JU (2010) Hormonal control of the shoot stem-cell niche. Nature 465:1089–1092
Zwack PJ, Rashotte AM (2015) Interactions between cytokinin signalling and abiotic stress responses. J Exp Bot 66:4863–4871
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant No. 31972397, U1708232), China Agriculture Research System (Grant No. CARS-25), and the Shenyang Scientific Research Project (18-013-0-36, RC180123).
Author information
Authors and Affiliations
Contributions
YL designed and carried out the experiments, analyzed the results, and wrote the manuscript. MS, HX, SM and BW provided scientific advice, and revised the manuscript. MQ and TL conceived the research area, provided scientific advice, and supervised the project.
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
The raw data of locule number investigation (XLSX 9.6 kb)
Table S2
The DEGs identified in GA4 vs. CK and PAC vs. CK (XLSX 89.1 kb)
Table S3
Gene Ontology analysis of DEGs in of GA4(100 µM), PAC (100 µM), and H2O treatments DEGs were annotated in three main categories: biological process, cellular component, and molecular function (DOCX 19.2 kb)
Table S4
Expression level and fold change of genes in RNA-seq (XLSX 10.8 kb)
Table S5
The raw data of qRT-PCR validation (XLSX 11.2 kb)
Rights and permissions
About this article
Cite this article
Li, Y., Sun, M., Xiang, H. et al. Transcriptome analysis reveals regulatory effects of exogenous gibberellin on locule number in tomato. Plant Growth Regul 91, 407–417 (2020). https://doi.org/10.1007/s10725-020-00614-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-020-00614-3