Abstract
The locus responsible for branched spike in a branched spike mutant of Triticum monococcum L. (2n = 2x = 14, AmAm genome) and soft glume in Triticum sinskajae Filat. et Kurkiev (2n = 2x = 14, AmAm genome) were mapped by genotyping F2 populations using microsatellite markers. Phenotypic analysis in the cross T. sinskajae PI 418587/a branched spike mutant KT3-24 confirmed that both characters were under control of a recessive allele at a single locus, and they were linked with 26.6 cM. The branched head in T. monococcum (bh m) locus was located on chromosome 2AmS and the marker Xgwm122 flanked the bh m gene distally. Soft glume locus in T. sinskajae was allelic to the soft glume (sog) locus in mm09, a soft glume mutant of T. monococcum. The sog locus was linked with Xwmc644 distally. In the F2 hybrids of T. monococcum #252/PI 418587 and T. monococcum KT 3-21/PI 418587, sog was linked with Xgwm71. The gene fg which determines a false glume was also located on chromosome 2AmS and the recombination between sog and fg (1.6 cM) was obtained in F2 hybrid of KT 3-21/PI 418587.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
One of the ways to improve wheat yield is to increase grain number per spike and per unit area, and this would be partially depend on spike morphology. The wheat spike is composed of many spikelets arranged to form two opposite rows along the main axis. The spikelet is composed of several florets jointed at the axis alternatively on opposite sides. In normal spike, a single spikelet arises from each rachis node. Wheats with supernumerary spikelet are evidently a genetically heterogeneous group, where more than one spikelet arises per rachis node (Peng et al. 1998; Klindworth et al. 1990a, b). A frequently type of supernumerary spikelet is the branched form of Triticum turgidum L. (2n = 4x = 28, genome BBAA) where the extra spikelets grow from secondary spike rachides and are named genuine branching (syn. ‘turgidum type of branching’). The genes for this type of branching were mapped earlier (Klindworth et al. 1990a, b). Although the expression of the branched form was modulated by a number of environmental factors (Sharman 1944; Pennell and Halloran 1983), the branched form was recessive (Pennell and Halloran 1983; Klindworth et al. 1990a; Peng et al. 1998; Martinek and Bednar 2001). In tetraploid wheat, Klindworth et al. (1990a) suggested that branched form was conditioned by a single major gene (bh) on chromosome 2A in concert with an unknown number of minor genes. Haque et al. (2012) indicated that the bh gene was linked to microsatellite loci on chromosome 2AS. Martinek and Bednar (2001) developed winter hexaploid wheat germplasm characterized by genetically stable multirow spike (mrs), which is controlled by a single recessive gene. Dobrovolskaya et al. (2009) mapped the mrs1 gene on chromosome 2DS using microsatellite markers. The involvement of the homoeologous group 2 chromosomes in the control of branched form has been noted in both tetraploid and hexaploid wheat (Klindworth et al. 1997, 1990a, b; Peng et al. 1998). Muramatsu (2009) presumed that the homoeologous group 2 chromosomes are involved in a genetic system determining the number of spikelets per rachis node in the tribe Triticeae.
Triticum monococcum L. (2n = 2x = 14, genome AmAm), commonly known as einkorn wheat, was widely cultivated during the pioneering human farming activities in the Fertile Crescent. Its Am genome is closely related to the Au genome of Triticum urartu Tumanian ex Gandilyan (2n = 2x = 14, genome AuAu), which was the A genome progenitor of the polyploid wheat species. T. urartu has been the dominant A genome donor of the most important polyploid wheat species, T. turgidum L., Triticum timopheevii (Zhuk.) Zhuk. (2n = 4x = 28, genome AAGG), and common wheat Triticum aestivum L. (2n = 6x = 42, genome BBAADD). In contrast, T. monococcum has only been the Am genome donor of Triticum zhukovskyi Menabde et Ericz. (2n = 6x = 42, genome AmAmAAGG) (Dvorak et al. 1993; Dubcovsky et al. 1996; Baum and Bailey 2004). Although the Am genome is under-represented in hexaploid wheat, the exploitation of genetic diversity in T. monococcum and discovery of novel variant alleles may provide opportunities for further wheat genetic improvement (Valkoun 2001). Several orthologous genes are confirmed and mapped in T. monococcum. The genes for black glume (Bg) and pubescent glume (Hg) were located on chromosome 1Am (Dubcovsky et al. 1996; Goncharov et al. 2007; Jing et al. 2007). Singh et al. (2007) located the blue aleurone (Ba) gene on chromosome 4Am. Morphological mutants in T. monococcum were induced by Multani et al. (1992) and Dr. K. Yamashita (http://www.shigen.nig.ac.jp/wheat/komugi/strains/). Kosuge et al. (2011) located the chlorina mutant gene on chromosome 7Am using a Yamashita’s mutant. The branched spike mutant was developed Dr. K. Yamashita. Triticum sinskajae Filat. et Kurkiev (2n = 2x = 14, genome AmAm) was discovered in accession K-20970 of T. monococcum during the analysis of the accessions collected in the coastal zones of Turkey by Prof. Zhukovskii (Filatenko and Kurkiev 1975). Filatenko and Kurkiev (1975) believe that T. sinskajae is a natural naked mutant of T. monococcum. Kuspira et al. (1989) found semi-compact spike and soft glume (free-threshing) of T. sinskajae were tightly linked. A false glume (gene symbol, fg), a glume-like structure, is located between the glume and lemma of the outer florets of each spikelet. False glume was also closely linked to the gene for soft glume/semi-compact spike. Goncharov et al. (2007) found that fissile inner (flower) glume (gene symbol, fig) was linked with semi-compact spike. It seems that fissile inner (flower) glume is a synonym of “false glume”. Taenzler et al. (2002) mapped the gene for soft glume (sog) of T. sinskajae on chromosome 2AmS using AFLP markers. Sood et al. (2009) also mapped the soft glume (sog) locus of a soft glume mutant of T. monococcum on chromosome 2AmS.
Molecular markers involving mapping genes of agricultural importance are widely applied. Numerous polymorphic microsatellite markers have been integrated into a genetic framework map of wheat (Röder et al. 1998a, b; Song et al. 2005; Torada et al. 2006). We applied microsatellite markers to map the genes for branched spikes and soft glumes in T. monococcum.
Materials and methods
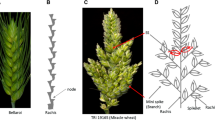
Five accessions of T. monococcum and one accession of T. sinskajae were available for the experiment. Heading of T. monococcum #252 was early. T. monococcum KT 3-24 was a mutant with branched spike developed by Dr. K. Yamashita. T. monococcum KT 3-21 has the chlorina phenotype. The spike of T. monococcum mm09 has a soft glume and simultaneously semi-compact spike determined by the gene sog located on chromosome 2AmS (Sood et al. 2009). T. sinskajae PI 418587 has soft and semi-compact spike. T. monococcum PI 584654 is a breeding line with soft spike and semi-compact spike, and is a bulk of equal quantities of seed from 34 F5 lines selected from the cross, T. monococcum/T. sinskajae (Vallega 1996). In the present study, we used the progenies of a single spike of PI 584654. Figure 1 shows the spike features of all parental accessions (Fig. 1a–f), and spikelet morphology of KT 3-21 and PI 418587 (Fig. 1g–i).
The spike features of Triticum monococcum accessions (left to right; a T. monococcum #252, b T. monococcum KT 3-21, c T. monococcum KT 3-24, d T. sinskajae PI 418587, e T. monococcum mm09 and f T. monococcum PI 584654) and spikelets of g T. monococcum KT 3-21 and h, i T. sinskajae PI 418587. Arrow in spikelet i indicates the false glume, a glume-like structure located between the glume and lemma of the outer florets of each spikelet of PI 418587
To construct the linkage map of the genes for branched spike (bh), soft glume/semi-compact spike (sog) and false glume (fg) and to check allelic relationships, we developed seven F 2 hybrids, PI 418587/KT 3-24, PI 584654/KT 3-24, #252/PI 418587, KT 3-21/PI 418587, mm09/#252, mm09/PI 418587 and mm09/PI 854654. The progenies the F2 recombinants of KT 3-21/PI 418587 were grown to observe in the F3 generation. All F 2 populations and F3 progenies were grown in the experimental field at the College of Agriculture, Ibaraki University, Japan.
Microsatellite mapping
Genomic DNA was extracted from seedling leaves from individuals of three F2 populations, KT 3-24/PI 418587, #252/PI 418587 and KT 3-21/PI 418587, according to Dellaporta et al. (1983). To map the gene, we used wheat microsatellite markers located on chromosome 2A (Röder et al. 1998a, b; Song et al. 2005; Torada et al. 2006). The PCR conditions, electrophoresis of PCR products and detection of amplified fragments were performed according to Kosuge et al. (2008). Multipoint linkage values were calculated using Map Manager QTX (Manly et al. 2001). Minimum LOD scores of 3.0 were used to develop the linkage map. The software calculated genetic distances in centiMorgans (cM) by applying the Kosambi (1944) mapping function.
Results
Table 1 shows the segregation of branched spike, soft/semi-compact spike and false glume in seven F2 hybrids. The spike branching phenotype segregated in a ratio of 3 normal: 1 branched in the two F2 hybrids, PI 418587/KT 3-24 and PI 584654/KT 3-24. For the soft/semi-compact spike, F2 hybrids of T. sinskajae PI 418587/KT 3-24, #252/PI 418587 and KT 3-21/PI 418587 segregated 3 hard/monococcum type: 1 soft/semi-compact type. F2 hybrids of PI 584654/KT 3-24 also segregated 3 hard/monococcum type: 1 soft/semi-compact type. The F2 hybrid of mm09/#252 segregated 3 hard/monococcum type: 1 soft/semi-compact type. The F2 hybrids of mm09/PI 418587 and mm09/PI 584654 did not segregate for spike type indicating that the genes for semi-compact of spike in these parents were allelic. For false glume, the F2 hybrid of KT 3-21/PI 418587 segregated 3 absent: 1 present. We did not obtain any plants without false glume in the F2 hybrids of mm09/PI 418587 and mm09/PI 584654, indicating allelic gene conditioned false glume in these parents.
Linkage relationship among branched spike, soft/semi-compact spike and false glume
Table 2 shows the linkage relationship among branched spike, soft/semi-compact spike and false glume in three F2 hybrids. The F2 hybrids of PI 418587/KT 3-24 and PI 584654/KT 3-24 indicated the linkage relationships between sog and bh. The calculated recombination rates were 0.267 and 0.243, respectively. We found three recombinants in the F2 of KT 3-21/PI 418587. Joint segregation for semi/soft compact spike and false glume was 94 hard/monococcum type without false glume: 2 hard/monococcum type with false glume: 1 soft/semi-compact type without false glume: 25 soft/semi-compact type with false glume, and it did not fit a 9:3:3:1 ratio (χ2 = 88.944, df = 3, p < 0.05). The calculated recombination rate between sog and fg was 0.026 (Table 2). The genotypes of two recombinants with hard/monococcum type spike and false glume were supposed to be Sog–fgfg, wheareas the genotype of a recombinant, soft/semi-compact spike plant without false glume, was supposed to be sogsogFg. We confirmed the recombined types in F3 progenies of three recombinants.
Microsatellite mapping of bh m, sog and fg genes
It was known that the gene for soft glume (sog) was located on the chromosome 2AmS by Taenzler et al. (2002) and Sood et al. (2009). In the F2 of PI 418587/KT 3-24, 20 polymorphic markers on chromosome 2Am were used to assign the position of the spike branching gene (bh m, branched head in T. monococcum) and soft glume (sog). The linkage map was divided into two fragments (Fig. 2). The major fragment indicated the linkage relationships among bh m, sog and markers. The gene bh m was linked with Xgwm122 marker distally, whereas sog was linked with Xwmc644. The map indicated that the genes bh m and sog were located on chromosome 2AmS.
According to Sood et al. (2009) and Haque et al. (2012), the Xgwm71 marker was a pivotal marker franking sog and bh loci. However, in the cross PI 418587/KT3-24, Xgwm71 was not polymorphic between parents. Hence, we mapped the sog gene further using two F2 hybrids of #252/PI 418587 and KT 3-21/PI 418587, paying attention to the Xgwm71 marker franking the sog gene. In the F2 of #252/PI 418587, the sog gene was bracketed by the markers Xgwm558 and Xgwm71 (Fig. 2) at distances of 5.0 and 5.2 cM respectively. In the F2 of KT 3-21/PI 418587, the sog gene was linked with Xgwm71 distally, and the distance between sog and Xgwm71 was 15.3 cM. Recombination between sog and fg (1.6 cM) was obtained in the F2 of KT 3-21/PI 418587.
Discussion
In the present study, the segregation behavior of the branched spike in T. monococcum indicated that it is controlled by a single major gene, bh m. This result coincided with Klindworth et al. (1997) and Haque et al. (2012) in T. durum. Shitsukawa et al. (2009) hypothesized that the branched spike phenotype is caused by the transformation of the floret meristem into a spikelet-like meristem. Spike architecture influences the processes of pollination and seed formation, and thus plays an important role in the determination of grain number in spike. The branched spike allows the formation of more spikelets per spike, thereby increasing the spike sink capacity and can influence then harvest index of the plant. T. monococcum, commonly known as einkorn wheat coined from the German expression of ‘one grain’, generally has one grain per spikelet, Triticum monococcum L. subsp. aegilopoides (Link) Thell. also has one grain per spikelet. The thaoudar type of T. monococcum subsp. aegilopoides (syn. Triticum boeoticum Boiss. subsp. thaoudar (Hausskn.) E. Schiem.) and T. urartu has two grains per spikelet. It indicated that introduction of bh m gene concerted with increasing spikelet and floret number in spike is essential for increasing yield potential. Multirow spike has also been considered as a usable donor for increasing the spikelet and seed number per spike (Martinek and Bednar 2001; Dobrovolskaya et al. 2009; Martinek et al. 2011). This is important for enhancing the crop yield potential, since wheat yield is generally thought to be sink-limited (Wang et al. 1998). The development of genetic resources which increase number of reproductive organs is particularly desirable (Miralles and Slafer 2007; Reynolds et al. 2005). It is interesting that Dobrovolskaya et al. (2009) mapped the gene for multirow spike (mrs1) on chromosome 2DS. The location of bh m was found on chromosome 2AmS. The similar homoeoallelic position of both genes responsible for supernumerary spikelet expression suggests that these genes may be orthologous loci.
We found that the naturally occurring sog mutation in T. sinskajae and the artificially induced sog mutation in T. monococcum were allelic. The sog mutation was associated with ‘false glume’ formation in the mutants. Observation by Kuspira et al. (1989), Goncharov et al. (2007) and the present study indicated mutations occurred in the genomic region of the soft glume-false glume complex. In T. monococcum, Multani et al. (1992) induced a liguleless mutant, and Dr. K. Yamashita (http://www.shigen.nig.ac.jp/wheat/komugi/strains/) produced a non-glossy mutant. The liguleless and non-glossy mutants may be useful produce a more precise linkage map of chromosome 2Am because those genes should be orthologous with the lg and W I loci of tetraploid and hexaploid wheat.
References
Baum BR, Bailey LG (2004) The origin of the A genome donor of wheats (Triticum: Poaceae)—a perspective based on the sequence variation of the 5S DNA gene units. Genet Resour Crop Evol 51:183–196
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA minipreparation: version II. Plant Mol Biol Rep 1:19–21
Dobrovolskaya O, Martinek P, Voylokov AV, Korzun V, Röder MS, Börner A (2009) Microsatellite mapping of genes that determine supernumerary spikelets in wheat (T. aestivum) and rye (S. cereale). Theor Appl Genet 119:867–874
Dubcovsky J, Luo MC, Zhong JY, Bransteitter R, Desai A, Kilian A, Kleinhofs A, Dvorak J (1996) Genetic map of diploid wheat, Triticum monococcum L., and its comparison with maps of Hordeum vulgare L. Genetics 143:983–999
Dvorak J, Diterlizzi P, Zhang HB, Resta P (1993) The evolution of polyploid wheats—identification of the A-genome donor species. Genome 36:21–31
Filatenko AA, Kurkiev UK (1975) A new species—Triticum sinskajae A. Filat. et Kurk. Trudy po Prikl Bot, Genet i Selekts 54:239–241 (in Russian)
Goncharov NP, Kondratenko EJA, Bannikova SV, Konovalov AA, Golovnina KA (2007) Comparative genetic analysis of diploid naked wheat Triticum sinskajae and the progenitor T. monococcum accession. Russ J Genet 43:1248–1256
Haque MA, Martinek P, Kobayashi S, Kita I, Ohwaku K, Watanabe N, Kuboyama T (2012) Microsatellite mapping of the genes for semi-dwarfism and branched spike in Triticum durum Desf. var. ramosoobscurum Jakubz. “Vetvistokoloskaya”. Genet Resour Crop Evol 59:831–837
Jing H-C, Kornyukhin D, Kanyuka K, Orford S, Zlatska A, Mitrofanova OP, Koebner R, Hammond-Kosack K (2007) Identification of variation in adaptively important traits and genome-wide analysis of trait-marker associations in Triticum monococcum. J Exp Bot 58:3749–3764
Klindworth DL, Williams ND, Joppa LR (1990a) Inheritance of supernumerary spikelets in a tetraploid wheat cross. Genome 33:509–514
Klindworth DL, Williams ND, Joppa LR (1990b) Chromosomal location of genes for supernumerary spikelets in tetraploid wheat. Genome 33:515–520
Klindworth DL, Klindworth MM, Williams ND (1997) Telosomic mapping of four genetic markers in durum wheat. J Hered 88:229–232
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Kosuge K, Watanabe N, Kuboyama T, Melnik VM, Yanchenko VI, Rosova MA, Goncharov NP (2008) Cytological and microsatellite mapping of mutant genes for spherical grain and compact spikes in durum wheat. Euphytica 159:289–296
Kosuge K, Watanabe N, Kuboyama T (2011) Comparative genetic mapping of the chlorina mutant genes in genus Triticum. Euphytica 179:257–263
Kuspira J, MacLagen J, Bhambhani RN, Sadasivaiah RS, Kim N-S (1989) Genetic and cytogentic analyses of the A genome of Triticum monococcum L. V. Inheritance and linkage relationships of genes determining the expression of 12 qualitative characters. Genome 32:869–881
Manly KF, Cudmore RH Jr, Meer JM (2001) Map Manager QTX, cross-platform software for genetic mapping. Mam Genome 12:930–932
Martinek P, Bednar J (2001) Changes of spike morphology (multirow spike-MRS, long glumes-LG) in wheat (Triticum aestivum L.) and their importance for breeding. In: The proceedings of international conference ‘‘Genetic collections, isogenic and alloplasmic lines’’, Novosibirsk, Russia, pp 192–194
Martinek P, Dobrovolskaya OB, Pokorova P, Vanova M (2011) Gene resources of wheat (Triticum aestivum L.) with changed spike morphology and their utilisation. In: Pazderu K (ed) Seed and seedlings. Scientific and Technical Seminar, Prague, pp 62–69
Miralles DJ, Slafer GA (2007) Sink limitations to yield in wheat: how could it be reduced? J Agric Sci 145:139–149
Multani DS, Sharma SK, Dhaliwal HS, Gill KS (1992) Inheritance of induced morphological mutants in Triticum monococcum L. Plant Breed 109:259–262
Muramatsu M (2009) A presumed genetic system determining the number of spikelets per rachis node in the tribe Triticeae. Breed Sci 59:617–620
Peng ZS, Yen C, Yang JL (1998) Chromosomal location of genes for supernumerary spikelet in bread wheat. Euphytica 103:109–114
Pennell AL, Halloran GM (1983) Inheritance of supernumerary spikelets in wheat. Euphytica 32:767–776
Reynolds MP, Pellegrineschi A, Skowmand B (2005) Sink limitation to yield and biomass: a summary of some investigations in spring wheat. Ann Appl Biol 146:39–49
Röder MS, Korzun V, Gill BS, Ganal MW (1998a) The physical mapping of microsatellite markers in wheat. Genome 41:278–283
Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998b) A microsatellite map of wheat. Genetics 149:2007–2023
Sharman BC (1944) Branched head in wheat and wheat hybrids. Nature 153:497–498
Shitsukawa N, Kinjo H, Takumi S, Murai K (2009) Heterochronic development of the floret meristem determines grain number per spikelet in diploid, tetraploid and hexaploid wheats. Ann Bot 104:243–251
Singh K, Ghai M, Garg M, Chhuneja P, Kaur P, Schnurbusch T, Keller B, Dhaliwal HS (2007) An integrated molecular linkage map of diploid wheat based on a Triticum boeoticum × T. monococcum RIL population. Theor Appl Genet 115:301–312
Song QJ, Shi JR, Singh S, Fikus EW, Costa JM, Lewis J, Gill BS, Ward R, Cregan PB (2005) Development and mapping of microsatellite (SSR) markers in wheat. Theor Appl Genet 110:550–560
Sood S, Kuraparthy V, Bai G, Gill BS (2009) The major threshability genes soft glum (Sog) and tenacious glume (Tg) of diploid and polyploidy wheat, trace their origin to independent mutations at non-orthologous loci. Theor Appl Genet 119:341–351
Taenzler B, Esposti RF, Vaccino P, Brandolini A, Effgen S, Heun M, Schäfer-Pregl R, Borghi B, Salamini F (2002) Molecular linkage map of einkorn wheat: mapping of storage-protein and soft-glume genes and bread-making quality QTLs. Genet Res Camb 80:131–143
Torada A, Koike M, Mochida K, Ogihara Y (2006) SSR-based linkage map with new markers using an intraspecific population of common wheat. Theor Appl Genet 112:1042–1051
Vallega V (1996) Registration of partially free-threshing diploid wheat germplasm. Crop Sci 36:1717
Valkoun J (2001) Wheat pre-breeding using wild progenitors. In: Bedo Z, Lang L (Eds) Book from conference: 6IWC Budapest, Hungary, Jun 05–09, 2000, Wheat in a Global Environment, Book Series Development in Plant Breeding vol 9, pp 699–707
Wang ZL, Yin YP, He MR, Cao HM (1998) Source-sink manipulation effects on postanthesis photosynthesis and grain setting on spike in winter wheat. Photosynthetica 35:453–459
Acknowledgments
We acknowledge the gift of seed of diploid wheat accessions from the National Small Grain Collection (NGSC), Aberdeen, Idaho, USA, and technical assistance Dr. M. A. Haque and Miss I. Kita. We thank Dr. D. L. Klindworth, USDA-ARS, Northern Crop Science Lab., Fargo, USA for helpful comments on our manuscript. P.M. thanks for project RO0211 of the Ministry of Agriculture of the Czech Republic.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Amagai, Y., Martinek, P., Watanabe, N. et al. Microsatellite mapping of genes for branched spike and soft glumes in Triticum monococcum L.. Genet Resour Crop Evol 61, 465–471 (2014). https://doi.org/10.1007/s10722-013-0050-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-013-0050-9