Abstract
Genomic in situ hybridization (GISH) was used to investigate the genomic relationships among some newly collected species of genus Setaria. Previous work identified that S. viridis and S. adhaerens carry genomes A and B, respectively. GISH patterns obtained in this report clearly distinguished the genome of S. grisebachii from the known genomes A and B, and indicated its new genomic constitution which we suggest to name genome C of the Setaria genus. The two sets of chromosomes of tetraploid S. queenslandica hybridized well with the A genome of S. viridis indicating its autotetraploid nature. This is the first autotetraploid identified in the Setaria genus, which should be classified into the primary A genome gene pool rather than the tertiary gene pool as previously classified. GISH patterns did not distinguish the genome of S. leucopila from the A genome of S. viridis and S. italica, suggesting its close relation with foxtail millet. Strong hybridization signals were observed when S. adhaerens genomic DNA was used as probe to hybridize the chromosomes of diploid S. verticillata, inferring its B genome nature. Combined with morphological observation and previous work, we deduce that diploid S. verticillata and S. adhaerens are probably taxonomically the same species with different names.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Setaria Beauv. belongs to the tribe Paniceae, subfamily Panicoideae and family Poaceae in the grass family. There are about 125 species worldwide occurring in tropical, sub-tropical and temperate regions. Seventy four species originate from Africa, America has 25 native species, and others are distributed in Eurasia (Hubbard 1915; Rominger 1962). Foxtail millet (S. italica (L.) Beauv.), which is one of the oldest cereal crops contributed to human civilization not only in the past but still used as a staple food in China and India, belongs to this group of species. Some species in genus Setaria, such as foxtail millet, S. leiantha Hackel (Argentinean foxtail millet) and S. splendida Stapf var. splendida are cultivated for grazing and hay in America, Africa and Southern Asia (Wanous 1990). The genus also has members of the worst weed species that are deleterious to world agriculture management, such as S. viridis (L.) Beauv. (green foxtail), S. faberi Herrm. (giant foxtail), S. verticillata (L.) Beauv. (bristly foxtail) and S. glauca (Weigel) Hubb. (yellow foxtail) (Dekker 2003).
The nuclear genome of Setaria spp. typically has a complement of nine chromosomes in its monoploid form (Avdulov 1931) and different ploidy levels were identified in the genus including diploid (2n = 2x = 18, S. viridis, S. italica, S. adhaerens (Forssk.) Link ex Chiov., S. grisebachii Fourn. ex Hemsl., S. leucopila K. Schum., and S. verticillata (L.) Beauv.), tetraploid (2n = 4x = 36, S. verticillata, S. faberi, S. pumila (Poir.) Roem. et Schult., and S. queenslandica Domin), hexaploid (2n = 6x = 54, S. verticillata and S. pumila) and octoploid species (2n = 8x = 72, S. pumila and S. geniculata) (Avdulov 1931; Brown 1948; Gupta and Singh 1977; Dekker 2003; Wang et al. 2007). Observations drawn from interspecific hybridization, karyological studies, and genomic in situ hybridization (GISH) indicated that foxtail millet and its putative wild ancestor S. viridis, the green foxtail, carry the A genome, and S. adhaerens carries the B genome. Both S. verticillata and S. faberi, which are tetraploid species, are composed of A and B genomes (Li et al. 1945; Harlan and de Wet 1971; Darmency and Pernes 1987; Benabdelmouna et al. 2001). The genome composition of many other wild species is still unidentified and needs to be reliably established, which is useful in foxtail millet breeding and related genetic researches.
GISH offers a powerful tool for investigating genome composition and genetic relationship among species. It has been successfully used to identify closely related species in Triticeae (Anamthawat-Jónsson et al. 1990) and to investigate genomic relationship between Thinopyrum intermedium and Th. ponticum (Chen et al. 1998). It is a direct and visual method of distinguishing different genomes (Mukai et al. 1993). This technique has been applied in Setaria species genomic composition study (Benabdelmouna et al. 2001), and successfully used for genomic constitution study of Oryza and Brassica species (Li et al. 2001; Hasterok and Maluszynska 2005).
In our previous study, we have studied three diploid (S. grisebachii, S. leucopila, and S. verticillata) and one tetraploid (S. queenslandica) species (Wang et al. 2007), which are morphologically close to both the A and B genome species of the genus, but their genome compositions are not investigated. In this study, we used GISH technique to identify the genome constitution of these species, which will be useful in establishment of phylogenetic relationship of Setaria group of species.
Materials and methods
Plant materials
Wild and cultivated species of Setaria used in this study and their characteristics, are listed in Table 1. Seeds were germinated on moistened filter paper at 25°C in Petri dishes. Root tips of about 2 cm in length were cut and treated with 8-hydroxyquinoline for 2 h to accumulate metaphases. All treatments were performed at room temperature in dark. After rinsing with distilled water, the root tips were then fixed in freshly prepared 1:3 glacial acetic acid and ethanol (100%) for at least 24 h and stored at 4°C until use. Seeds of each species were grown, germinated in autoclaved soil and transferred into plastic pots in an open air field. Young leaves of plants in an age at 5- to 6-leaves stage were used to isolate DNA for probes in GISH analysis.
Chromosome preparation
The fixed root tips were rinsed with distilled water and a single root tip was transferred in a drop of 45% acetic acid onto a clean slide before gentle squashing. The slides were then scanned for good chromosome spreads at prophase or metaphase stages with phase-contrast microscope and the selected slides were stored at −80°C until use.
Isolation and labeling of genomic DNA
Total genomic DNAs of five diploid species, S. viridis, S. italica, S. adhaerens, S. grisebachii, and S. leucopila, were extracted from young leaves using the method of Li et al. (1998) with some modification. About 1 μg of the total genomic DNA from each sample was labeled with digoxigenin-11-dUTP using a nick translation kit (Roche Company) following the instructions provided by the manufacturer.
GISH analysis
GISH was carried out following Bisht and Mukai (2001) with some modifications. The hybridization mixture contained 50% formamide, 10% dextran sulfate, 2× SSC, 3 ng/μl of DNA probe, and 1 ng/μl salmon sperm DNA. Chromosomal DNA on the slides was denatured by immersion in 70% formamide in 2× SSC for 3 min at 80°C. After removing the cover with a razor blade, the slides were rapidly dehydrated in an ethanol series (70%, 95%, and 100%) for 3 min each at −20°C. The hybridization mix was denatured for 10 min at 95°C and then immediately chilled on ice for 10 min, and 20 μl of probe mix was applied to each slide. DNA–DNA in situ hybridization was carried out overnight in a moist chamber at 37°C. Following hybridization the slides were washed with high stringency in 2× SSC at 32°C, 20% formamide in 2× SSC at 32°C, 20% formamide in 0.1× SSC at 32°C for 5 min each, and 2× SSC and 4× SSC for 5 min each at room temperature, the slides were placed in the BSA (Bovine serum albumin) blocking solution (5% BSA-2× SSC) for 15 min at 37°C. To ensure the washing stringency being strong enough for a certified result we also carried out slide washing following Bennett et al. (1992) for some slides, which was carried out in 2× SSC at room temperature, 50% formamide in 2× SSC at 42°C, 2× SSC at 42°C, and 2× SSC at room temperature, for 10 min each. According to Schwarzacher et al. (1989), these conditions allow hybridization between DNA–DNA duplexes sharing approximately 80% sequence homology.
Hybridization signals were detected with anti-Digoxigenin-Rhodamine. The slides were washed twice in 4× SSC for 5 min each at room temperature. The chromosomes were finally counterstained with 4′-6-diamidio-2-phenylindole (DAPI). Chromosome preparations were visualized using an Olympus epifluorescence microscope with appropriate filters, and photos were taken digitally.
Results
Genomic relationship of the diploid Setaria species
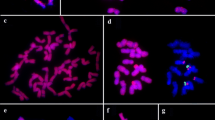
Genome identification of the three newly identified diploid species, S. grisebachii, S. leucopila, and S. verticillata was carried out using the known A genome (S. viridis) and B genome (S. adhaerens) DNAs separately as probes without blocking DNA. Total genomic DNA from S. viridis or S. adhaerens was digoxigenin labeled and used as probe on metaphase chromosomes from the target species. When genomic DNA from S. viridis was hybridized to chromosome preparations of S. leucopila, all its chromosomes had strong signals in the pericentromeric regions. The terminal regions were faintly labeled for most chromosomes (Fig. 1a, b). The same hybridization pattern was obtained in the reciprocal experiment using S. leucopila genomic DNA as probe on chromosome preparations of S. viridis (Fig. 1c, d). When the probe prepared from total genomic DNA of S. italica was hybridized on chromosomes of S. leucopila, the same results as in S. viridis experiments were obtained (data not shown). In GISH using S. viridis genomic DNA as probe, weak hybridization signals or no signals were observed on chromosome preparations of S. grisebachii (Fig. 1e) and diploid S. verticillata (data not shown).
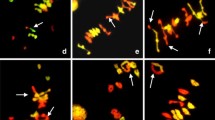
GISH patterns obtained on different Setaria species. a The metaphase chromosomes of S. leucopila were counterstained blue with DAPI. b The same metaphase cell hybridized with the probe of S. viridis digoxigenin-labeled genomic DNA (red). c DAPI counterstained metaphase cell from S. viridis. d The same metaphase cell hybridized with the probe of S. leucopila digoxigenin-labeled genomic DNA. A strong and total painting on nearly entire length of all chromosomes of S. leucopila was observed. e GISH was carried out using S. viridis genomic DNA as probe hybridizing on chromosome preparations of S. grisebachii, and chromosomes of this species showed weak signals as indicated by arrows. f Genomic DNA of S. adhaerens was applied to S. grisebachii chromosomes, only a few weak signal points were observed by dispersed pattern. g Chromosomes of S. leucopila hybridized with S. adhaerens genomic DNA as probe also showed faint signal, only two major spots signals were detected (arrowhead shows the two signals). h Metaphase of diploid S. verticillata probed with S. adhaerens total genomic DNA. A strong and total painting on the entire length of all the chromosomes was observed. i B genome of S. adhaerens genomic DNA was used as probe, very weak signals of pointed distribution were observed on a few chromosomes of S. queenslandica. j The metaphase of S. queenslandica was counterstained with DAPI. k The same metaphase cell hybridized with the genomic DNA of S. viridis, showing that 36 chromosomes of S. queenslandica were hybridized thoroughly with the S. viridis A genome. l The chromosomes of tetraploid S. queenslandica were counterstained blue with DAPI. m The same metaphase cell hybridized with genomic DNA of S. viridis (red). All the chromosomes were hybridized with strong signals. n Chromosomes of tetraploid S. verticillata hybridized with genomic DNA of S. grisebachii, some dispersed and faint spots signals were detected. Bar 5 μm
Total genomic DNA of S. adhaerens, which carries the Setaria B genome, was also used as a probe to identify the three target species, S. grisebachii, S. leucopila and diploid S. verticillata. When genomic DNA of S. adhaerens was applied to S. grisebachii chromosomes, only a few weak signals were observed by dispersed pattern (Fig. 1f), indicating the genome of S. grisebachii was distinct from the B genome. Chromosomes of S. leucopila hybridized with S. adhaerens genomic DNA as probe also showed faint signal or no signal at all (Fig. 1g). A complete different pattern was obtained when diploid S. verticillata was used as the target for probes prepared from total genomic DNA of S. adhaerens, all the chromosomes of diploid S. verticillata were strongly labeled almost entirely (Fig. 1h), indicating diploid S. verticillata carries the B genome of Setaria.
Genomic constitution of S. queenslandica
To investigate the genome constitution of tetraploid S. queenslandica, total genomic DNAs of S. viridis and S. adhaerens were used as probes on chromosome preparations of S. queenslandica, respectively. When the B genome of S. adhaerens genomic DNA was used as probe, no labeling signal or only very weak signals on few chromosomes of the species with 36 chromosomes were observed, implying that there is no B genome in S. queenslandica genomic constitution (Fig. 1i). When total genomic DNA of the Setaria A genome, S. viridis, was used as probe to test the chromosome constitution of S. queenslandica, all the 36 chromosomes were strongly hybridized with labeling signals throughout the nearly entire chromosomes (Fig. 1j, k). Using genomic DNA of S. italica as a probe, a nearly identical signal pattern was obtained (data not shown). To eliminate the possible influence of slide washing stringency on the result, using a different strong stringent washing protocol of Bennett et al. (1992), the 36 chromosomes of S. queenslandica all gave strong signals by the hybridization of S. viridis genomic DNA as probe (Fig. 1l, m). These results clearly indicate that S. queenslandica is an autotetraploid with the genome constitution of AAAA.
Identity test of the S. grisebachii genome with tetraploid species
To further understand the genome constitution of S. grisebachii, GISH analysis was carried out in tetraploid species with genomic DNAs of different diploid species as probes. When total genomic DNA of the A genome of S. viridis (AA) was used as a probe, only one set of 18 chromosomes of the tetraploid S. verticillata was labeled (data not shown). When genomic DNA of S. adhaerens (BB) was used as probe, also only 18 chromosomes showed signals. Those results were consistent with the known chromosome constitution of AABB of tetraploid S. verticillata demonstrated by Benabdelmouna et al. (2001). Using genomic DNA of S. grisebachii as a probe, no or weak hybridization signals were observed in the A and B genomes of tetraploid S. verticillata (Fig. 1n), which further indicates that the genome constitution of S. grisebachii is neither A nor B. We also hybridized genomic DNA of S. grisebachii onto the chromosomes of S. queenslandica, only very faint labeling signals was observed (data not shown).
Discussion
S. grisebachii probably carries the Setaria genome C
Compared to other genera of the grass family, the genome constitutions of Setaria are poorly understood although there are 125 species in the genus worldwide. From interspecific hybridization and karyological studies, Li et al. (1945) first suggested S. italica (foxtail millet) and its putative wild ancestor S. viridis were composed of genome A, and S. faberi Herrm. was an allotetraploid composed of A and another unknown genome. GISH by Benabdelmouna et al. (2001) indicated that chromosomes of S. adhaerens could not hybridize with the A genome of S. viridis and S. italica genomic DNA, and hybrid weakness between S. adhaerens and S. viridis also confirmed the different genomes of these two species. The genome of S. adhaerens was designated B and further analysis indicated that S. faberi and tetraploid S. verticillata were all allotetraploids, which were composed of A and B genomes (Benabdelmouna et al. 2001). In this study, genomic DNA of S. grisebachii hybridized neither to the A genome of S. viridis nor to the B genome of S. adhaerens, and reciprocal performance gave the same results. It is reasonable to assume that S. grisebachii carries a new genome, designated genome C, which is different from the known genomes A and B. This point of view is well supported by the known fact that S. grisebachii originated from Mexico, while S. viridis and S. adhaerens originated from Eurasia. Our three years field observations indicate that botanical morphology of S. grisebachii also was clearly different from S. viridis and S. adhaerens, which supports the new genome hypothesis. Our data from phylogenetic analysis of Setaria species with DNA sequences from genes of Adh1, Adh2 and Gpa1 also clearly distinguishes S. grisebachii from S. viridis and S. adhaerens (not published data), which also support the C genome constitution of S. grisebachii.
The A genome species and the first autotetraploid in Setaria
Clear understanding of genomic constitution and relationship will be important for germplasm management and breeding by use of wild resources. Probes from both S. viridis and S. italica genomic DNA hybridized well with chromosomes of S. leucopila, indicating that S. leucopila is a diploid species of the A genome. So far three A genome diploid Setaria species were identified including S. viridis, S. italica, and S. leucopila. S. viridis was thought to be the putative wild ancestor of domesticated S. italica, and the close genetic relationship of these two species was verified by interspecific hybridization and karyological studies (Li et al. 1945; Darmency and Pernes 1987), RAPD classification (Li et al. 1998), RFLP based genetic map construction (Wang et al. 1998) and GISH analysis (Benabdelmouna et al. 2001). Accumulated data made probably that these species are taxonomically the same species, which was first suggested by Harlan and de Wet (1971). Our GISH results showed that the genome of S. leucopila could not be differentiated from S. viridis and S. italica, indicating their close karyological relationship. This makes S. leucopila a valuable genetic resource for improvement of foxtail millet because of its possession of original genetic information. Morphological observation found that S. leucopila is someway intermediate between S. viridis and S. italica with fewer tillers and larger seeds than S. viridis, but the seed shattering habit indicates its wild species character. Crossability of S. leucopila with S. viridis and S. italica must be further investigated in order to determine their genetic evolution relationship.
Although many Setaria species are tetraploid, most of their genome constitutions are not clear except for two allotetraploid species composed of AABB, S. faberi and tetraploid S. verticillata (Benabdelmouna et al. 2001). In this research, the two sets of chromosomes of S. queenslandica all hybridized well with S. viridis and S. italica genomic DNA, indicating that S. queenslandica probably is an autotetraploid composed of AAAA. Using different washing protocols with high stringency, multiple independent hybridizations of chromosome preparations of S. queenslandica with genomic DNA of S. viridis as probe all gave the same result (Fig. 1j–m), which eliminated the possibility of an inaccurate result generated by not enough washing stringency and ensured the autotetraploid AAAA genome constitution of S. queenslandica. But we do not know whether the two sets of A genome in S. queenslandica are identical or in some degree identical, or in other words whether its genome constitution is AAAA or AAA1A1. Further experiments with DNA sequence data need to be done to clarify this question. Because artificially made autotetraploid lines of S. italica from foxtail millet cultivars all suffer from certain degree of seed setting sterility (Ahanchede et al. 2004), which made polyploid breeding in foxtail millet not successful, the finding of a naturally formed fertile autotetraploid might be helpful for polyploid breeding in foxtail millet.
The Setaria genus was previously classified into three gene pools. The primary gene pool is composed of S. viridis, S. italica and its close relatives (Harlan and de Wet 1971; Jusuf and Pernès 1985). The secondary gene pool is composed of more related wild species, including S. adhaerens, S. verticillata, and S. faberi. The remaining wild Setaria species were classified into the tertiary gene pool, which includes S. queenslandica (Zangré et al. 1992; Panaud 2006). Our experiment in this report clearly identified the close genetic relationship of S. queenslandica with S. viridis and S. italica. Hence, it is reasonable to classify S. queenslandica into the primary gene pool, which is the only natural autotetraploid species in Setaria so far identified.
Diploid S. verticillata and S. adhaerens are taxonomically the same species
Our GISH results showed that the genome of diploid S. verticillata and S. adhaerens were closely related and could not be separated. In morphology, there was no distinct difference between the two species; they are all with thin and fragile stems, and the same structure of panicle and spikelet. Based on GISH experiments and morphology observation, we deduce that diploid S. verticillata and S. adhaerens are probably taxonomically the same species with different names. Using GISH experiment Benabdelmouna et al. (2001) identified that the A genome of tetraploid S. verticillata comes from S. viridis and its B genome from S. adhaerens. From isoenzyme analysis and cytogenetical studies with F1 hybrid between S. italica (4n) and tetraploid S. verticillata, Wu and Bai (2000) deduce that S. viridis and diploid S. verticillata were the ancestors of tetraploid S. verticillata. The results of these two studies strongly support our conclusion that diploid S. verticillata and S. adhaerens are taxonomically the same species.
Because of its worldwide distribution and broad genetic variation, the A genome species of S. viridis was thought to be the primary gene pool of the Setaria genus (Harlan and de Wet 1971; Zangré et al. 1992; Panaud 2006). The diploid species S. verticillata or S. adhaerens had probably evolved from the A genome S. viridis or from an unknown diploid Setaria ancestor. Two allotetraploid species, S. verticillata, and S. faberi, were probably descended from the hybrid genome duplication between two diploid species of the A and B genome after the evolution of the B genome species. Successful hybridization between S. viridis and S. adhaerens (Benabdelmouna et al. 2001) strongly supports this point of view.
The nomenclature of S. verticillata
A wide range of chromosome complements have been reported by many literature papers with the same taxonomical name of S. verticillata, including diploid 2n = 18, tetraploid 2n = 36 and hexaploid 2n = 54, and the situation was well summarized by Dekker (2003). Whether this is due to misidentification, or a product of the complex taxonomy of the genus, is not apparent. If these reports are true, they raise important questions about intra-specific fertility, and may indicate reproductive barriers within species, and we know that polyploidization (either alloploidy or autoploidy) has played an active role in speciation and delimiting taxa in Setaria (Khosla and Sharma 1973). Auguier (1979) named the diploid form with S. verticillata and the tetraploid form with S. verticiformis Auguier and Wu and Bai (2000) supported this nomenclature. Our result, combined with the GISH result of Benabdelmouna, et al. (2001) and the isoenzyme result of Wu and Bai (2000), clearly indicate that S. verticillata and S. adhaerens are taxonomically the same species. So misidentification and different names with the synonymous species made the complex situation for some Setaria species. We suggest that diploid S. verticillata should be named S. adhaerens and the tetraploid form be kept the commonly accepted name of S. verticillata. Because we do not have the hexaploid S. verticillata sample, its genome constitution and nomenclature need to be studied in the future.
References
Ahanchede A, Poirier-Hamon S, Darmency H (2004) Why no tetraploid cultivar of foxtail millet? Genet Resour Crop Evol 51:227–230. doi:10.1023/B:GRES.0000024020.91764.8d
Anamthawat-Jónsson K, Schwarzacher T, Leith AR, Bennett MD, Heslop-Harrison JS (1990) Discrimination between closely related Triticeae species using genomic DNA as a probe. Theor Appl Genet 79:721–728. doi:10.1007/BF00224236
Auguier P (1979) Dumortier’s concepts in the genus Setaria Beauv. (Poaceae) (J). Bulletindu Jardin Botanique Natl Belgidue 49:427–433. doi:10.2307/3668095
Avdulov NP (1931) Karyo-systematische Untersuchungen der Familie Gramineen. Bull Appl Bot Suppl 43. Leningrad
Benabdelmouna A, Shi Y, Abirached-Darmency M, Darmency H (2001) Genomic in situ hybridization (GISH) discriminates between the A and the B genomes in diploid and tetraploid Setaria species. Genome 44:685–690. doi:10.1139/gen-44-4-685
Bennett ST, Kenton AY, Bennett MD (1992) Genomic in situ hybridization reveals the allopolyploid nature of Milium montianum (Gramineae). Chromosoma 101:420–424. doi:10.1007/BF00582836
Bisht MS, Mukai Y (2001) Genomic in situ hybridization identifies genome donor of finger millet (Eleusine coracana). Theor Appl Genet 102:825–832. doi:10.1007/s001220000497
Brown WV (1948) A cytological study in the Gramineae. Am J Bot 35:382–385. doi:10.2307/2437938
Chen Q, Conner RL, Laroche A, Thomas JB (1998) Genome analysis of Thinopyrum intermedium and Thinopyrum ponticum using genomic in situ hybridization. Genome 41:580–586. doi:10.1139/gen-41-4-580
Darmency H, Pernes J (1987) An inheritance study of domestication in foxtail millet using an interspecific cross. Plant Breed 99:30–33. doi:10.1111/j.1439-0523.1987.tb01146.x
Dekker J (2003) The foxtail (Setaria) species-group. Weed Sci 51:641–656. doi:10.1614/P2002-IR
Gupta PK, Singh RV (1977) Variations in chromosomes and flavonoids in Setaria Beauv. Nucleus 20:167–171
Harlan JR, de Wet JMJ (1971) Towards a rational taxonomy of cultivated plants. Taxon 20:509–517. doi:10.2307/1218252
Hasterok R, Maluszynska J (2005) FISH and GISH analysis of Brassica genomes. Acta Biol Crac Ser Bot 47:185–192
Hubbard FT (1915) A taxonomic study of Setaria and its immediate allies. Am J Bot 2:169–198. doi:10.2307/2435051
Jusuf M, Pernès J (1985) Genetic variability of foxtail millet (Setaria italica P. Beauv.). Electrophoretic study of five isoenzyme systems. Theor Appl Genet 71:385–391. doi:10.1007/BF00251177
Khosla PK, Sharma ML (1973) Cytological observations on some species of Setaria. Nucleus 26:38–41
Li HW, Li CH, Pao WK (1945) Cytological genetical studies of the interspecific cross of cultivated foxtail millet, Setaria italica (L.) Beauv., and S. viridis L. J Am Soc Agron 37:32–54
Li Y, Jia JJ, Wang Y, Wu S (1998) Intraspecific and interspecific variation in Setaria revealed by RAPD analysis. Genet Resour Crop Evol 45:279–285. doi:10.1023/A:1008600123509
Li CB, Zhang DM, Ge S, Lu BR, Hong DY (2001) Identification of genome constitution of Oryza malampuzhaensis, O.minuta, and O. punctata by multicolor genomic in situ hybridization. Theor Appl Genet 103:204–211. doi:10.1007/s001220100563
Mukai Y, Nakahara Y, Yamamoto M (1993) Simultaneous discrimination of the three genomes in hexaploid wheat by multicolor fluorescence in situ hybridization using total genomic and highly repeated DNA probes. Genome 36:489–495. doi:10.1139/g93-067
Panaud O (2006) In: Kole C (ed) Foxtail millet, genome mapping and molecular breeding in plants, volume 1 cereals and millets. Springer-Verlag, Berlin
Rominger JM (ed) (1962) Taxonomy of Setaria (Gramineae) in North America. Illinois Biological Monographs
Schwarzacher T, Leitch AR, Bennett MD, Heslop-Harrison JS (1989) In situ localization of parental genomes in a wide hybrid. Ann Bot (Lond) 64:315–324
Wang ZM, Devos KM, Liu CJ, Wang RQ, Gale MD (1998) Construction of RFLP-based maps of foxtail millet, Setaria italica (L.) P. Beauv. Theor Appl Genet 96:31–36. doi:10.1007/s001220050705
Wang YQ, Zhi H, Li W, Li HQ, Wang YF, Diao XM (2007) Chromosome number identification of some wild Setaria species. J Plant Genet Resour 8:159–164
Wanous M (1990) Origin, taxonomy and ploidy of the millets and minor cereals. Plant Var Seeds 3:99–112
Wu QM, Bai JL (2000) Cytogenetic and isoenzymic studies on foxtail millet and S. verticillata (2x) and S. verticiformis (4x). Acta Bot Boreal-Occident Sinica 20:954–959
Zangré R, Nguyen-Van E, Rherissi B, Till-Bottraud I (1992) Organisation du pool génique de Setaria italica (L.) P. Beauv. et exploitation des ressources génétiques d’espèces spontanées. In: Complexes d’espèces, flux de gènes et ressources génétiques des plantes. Lavoisier édition. BRG, Paris, pp 87–97
Acknowledgements
This research was supported by National Natural Science Foundation of China (30630045, 30471117), China National Technology Supporting Program (2006BAD02B02) and Hebei Natural Science Foundation (C2004000697, C2006000725). We thank Dr. H. Darmency for his kindly providing S. adhaerens seeds. The authors also thank Dr. Li Hongjie for his technical assistance and manuscript modification.
Author contributions
Xianmin Diao conceived and designed the experiments. Yongqiang Wang and Hui Zhi performed the experiments. Hui Zhi, Wei Li, Haiquan Li, Yongfang Wang and Zhanjing Huang contributed to reagents, materials planting and data analysis. Xianmin Diao wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Additional information
Y. Wang and H. Zhi contributed equally to this article.
Rights and permissions
About this article
Cite this article
Wang, Y., Zhi, H., Li, W. et al. A novel genome of C and the first autotetraploid species in the Setaria genus identified by genomic in situ hybridization. Genet Resour Crop Evol 56, 843–850 (2009). https://doi.org/10.1007/s10722-009-9405-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-009-9405-7