Abstract
Glycosylation is a very important post-translational modification involved in various cellular processes, such as cell adhesion, signal transduction and immune response. Urine is a rich source of glycoproteins and attractive biological fluid for biomarker discovery, owing to its availability, ease of collection, and correlation with pathophysiology of diseases. Although the urinary proteomics have been explored previously, the urinary glycoproteome characterization remains challenging requiring the development and optimization of analytical and bioinformatics methods for protein glycoprofiling. This study describes the high confident identification of 472 unique N-glycosylation sites covering 256 urinary glycoproteins. Besides, 202 unique N-glycosylation sites were identified in low molecular weight endogenous glycopeptides, which belong to 90 glycoproteins. Global site-specific characterization of the N-linked glycan heterogeneity was achieved by intact glycopeptide analysis, revealing 303 unique glycopeptides most of them displaying complex/hybrid glycans composed by sialic acid and fucose. These datasets consist in a valuable resource of glycoproteins and N-glycosylation sites found in healthy human urine that can be further explored in different disorders, in which the N-linked glycosylation may be aberrant.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glycosylation is a very important post-translational modification, which occurs in approximately half of all mammalian proteins [1]. Glycoproteins is involved in a number of physiological processes, including protein folding and trafficking [2], immune response [3] and cell-cell and cell-matrix interaction [4, 5]. In cancer, there is increasing evidence pertaining to the role of glycosylation in tumour formation and metastasis [6–8]. It was demonstrated that glycoproteins have aberrant glycosylation patterns in malignancy, producing differential occupancy of glycosylation sites or variability in attached glycan structures [9–11].
Recent developments in MS-based technologies have enabled large-scale characterization of glycoproteins [12, 13]. Generally, glycoproteomic strategies rely on the analysis of released glycans or deglycosylated peptides [14–16]. Although the removal of glycans simplifies the identification of glycosylation sites by LC–MS/MS, limited information correlating glycan structural heterogeneity to the protein attachment site are obtained.
Recently, novel strategies were developed to enrich and analyze intact N and O-linked glycopeptides, for example, using metabolic labeling of the glycoproteome combined with chemical enrichment using an isotopic recoding affinity probe [17], or chemoenzymatic method for glycopeptide enrichment on solid phase [18]. Alternatively, several MS dissociation methods have been applied to study intact glycopeptides such as collision induced dissociation (CID), high-energy collision dissociation (HCD) and electron capture/transfer dissociation (ECD/ETD) [19–21]. Despite the recent advances, the detection of intact glycopeptides often requires increased sample quantities and an enrichment step, especially because of the glycan heterogeneity and the low ionization efficiency of glycoconjugates [22]. Besides, the computational identification via database searching are still challenging due to the heterogeneity and nontemplated nature of glycopeptides. Collectively, these challenges have hindered widespread use of intact glycopeptides analysis in the glycoproteome field.
Hydrophilic interaction chromatography (HILIC) has been used increasingly to separate and enrich glycopeptides and shows some advantages over other enrichment techniques such as broad specificity towards different glycan structures, good solubility of polar samples and greater compatibility with electrospray MS [12]. Hagglund et al. explored the use of HILIC followed by partial deglycosylation to identify N-glycosylation sites in complex biological samples [23] and Mysling improved the HILIC-glycopeptides enrichment efficiency by modifying the TFA-containing mobile phases [24]. This approach was also applied in the study of intact glycopeptides and glycoproteins in different biological matrices [21, 25–30].
Urine is a rich source for glycoproteins derived from renal- [31] and distal organs [32], and has been considered as an attractive biologic fluid for disease biomarkers discovery, since it can easily be collected continuously and noninvasively [33, 34]. Although urinary proteomics approaches have been applied in biomarker discovery studies of several disorders [32, 35–37], only a few studies aimed at identifying urinary glycoproteins [38, 39]. Moreover, information about site-specific glycan-peptide micro-heterogeneity is still not well explored.
Considering the urinary glycoproteomic studies that performed site-specific glycosylation analyses, Saraswat et al. [40] reported 51 N-glycosylation sites belonging to 37 glycoproteins in urinary exosomes and Halim et al. [41] showed 58 N- and 63 O-linked glycopeptides from 53 glycoproteins in the urine from one healthy person.
This study presents a simple and straightforward methodology without extensive sample preparation processes and pre-fractionation, to comprehensively characterize N-linked glycosylation sites of urinary glycoproteins as well as low molecular weight (<10 kDa) endogenous glycopeptides. Besides, by analyzing intact glycopeptides, high confident site-specific characterization of the N-linked glycans microheterogeneity was reported.
Material and methods
Urine sample collection and protein and peptide extraction
Twenty ml of urine (first morning void) was collected from a healthy male consenting individual in two different days and stored on ice prior to processing. The healthy volunteer was 35 years old, had no medical history, and did not take any prescription medicines. The samples were centrifuged at 3000 g for 10 min at 4 °C. The cleared supernatant was further filtered using 0.22 μm filter (Millipore, Billerica, MA). Twenty milliliters of filtered urine was concentrated using 10 kDa cutoff filters (Millipore, Billerica, MA). The concentrated urine, that is, the retentate, was stored immediately at −20 °C until further use. The fraction which passed through the 10 kDa cutoff filter was also collected and the peptides (<10 kDa) were extracted and concentrated using hydrophilic − lipophilic-balanced (HLB) solid phase (SPE; Waters).
Protein digestion and desalting
The proteins were reduced by addition of DTT to a final concentration of 10 mM and incubation for 30 min. The proteins were alkylated prior to digestion by the addition of IAA to a final concentration of 40 mM and incubation for 30 min in the dark at room temperature. To quench the reaction, DTT was added to a final concentration of 5 mM. Trypsin (1:50, w/w) was added, and the mixture incubated overnight at 37 °C. The reaction was stopped with 0.4 % TFA and desalted with hydrophilic − lipophilic-balanced (HLB) solid phase extraction (SPE; Waters).
Glycopeptide enrichment using HILIC
The samples were reconstituted in 100 μl of 80 % (v/v) acetonitrile, 10 % dimethyl sulfoxide (DMSO) and 1 % (v/v) trifluoroacetic acid. Peptides were loaded onto an in-house PolyHYDROXYETHYL A™ HILIC resin (PolyLC Inc) packed onto a C8 disk (Empore) in a p200 pipette tip. The flow-through and wash fraction (80 % (v/v) acetonitrile and 1 % (v/v) trifluoroacetic acid) were collected and analyzed by LC-MS/MS. The enriched glycopeptides were eluted with 50 μL 0.1 % (v/v) TFA followed by 50 μL of 25 mM NH4HCO3 and finally 50 μL of 50 % (v/v) acetonitrile. The three eluted fractions were combined and dried by vacuum centrifugation.
PNGase F deglycosylation
An aliquot of the HILIC-enriched glycopeptides (from protein trypsin digestion and endogenous glycopeptides) was resuspended in 50 mM ammonium bicarbonate, pH 8.0 and deglycosylated with 500 U of PNGase F (New England Biolabs, Ipswich, MA) for 12 h at 37 °C. After incubation, the peptides were dried by vacuum centrifugation and reconstituted in 50 μl of 0.1 % formic acid prior to LC-MS/MS analysis.
Mass spectrometry analysis
The resulting peptide mixture was analyzed on a LTQ Velos Orbitrap mass spectrometer (Thermo Fisher Scientific) coupled with LC-MS/MS by an EASY-nLC system (Thermo Fisher Scientific) through a nanoelectrospray ion source. Sample concentration and desalting were performed online using a pre-column (2 cm; 100 μm ID; 5 μm C18-A1; Thermo). Separation was accomplished on Acclaim™ PepMap™ 100 C18 column (10 cm; 75um ID; 3um C18-A2; Thermo) using a linear gradient of A and B buffers (buffer A: A = 0.1 % formic acid; Buffer B = 95 % ACN, 0.1 % formic acid) from 1 % to 50 % buffer B over 60 min at a flow rate of 0.3 μL/min to elute peptides into the mass spectrometer. Columns were washed and re-equilibrated between LC–MS/MS experiments. Mass spectra were acquired in the positive-ion mode over the range m/z 400–1200 at a resolution of 30,000 (full width at half-maximum at m/z 400) and AGC target >1 × e 6. The 20 most intense peptide ions with charge states ≥2 were sequentially isolated to a target value of 15,000 and isolation width of 2 and fragmented in the linear ion trap using low-energy CID (normalized collision energy of 35 %) with activation time of 10 ms. For intact glycopeptides analysis, the 7 most intense peptide ions with charge states ≥2 were sequentially isolated to a target value of 15,000 and isolation width of 2 and fragmented in the linear ion trap using high-energy HCD (normalized collision energy of 35 %) with activation time of 0.1 ms. Dynamic exclusion was enabled with an exclusion size list of 500, exclusion duration of 15 s, and a repeat count of 1.
Analysis of intact N-glycopeptides
Intact glycopeptide spectra from each sample were searched against database using Byonic software v2.6.46 (Protein Metrics, http://www.proteinmetrics.com/) [42]. Searches were performed with the following fixed modifications: precursor mass tolerance of 10 ppm, product ion mass tolerance of 0.02 Da, carbamidomethylation of Cys, and fully trypsin specific cleavage with a maximum of two missed cleavages. Searches were also conducted with the following variable modifications: oxidation of methionine (15.994 Da), deamidation NQ (+0.984) and glycosylation of Asn with N-glycan database (309 mammalian no sodium) available within Byonic. The list of glycoproteins identified in the formerly N-linked peptides generated after PNGase F treatment fraction were used as protein database. All searches were filtered to <1 % false discovery rate (FDR) at protein level and 0 % at peptide level [43]. Moreover Byonic software calculates two-dimensional posterior error probability (PEP 2D), meaning that the protein identity (dimension one) and quality of peptide-spectrum match are taken into account for a correct sequence assignment. The probabilities are computed by the usual target/decoy method with the decoys being reversed protein sequences [44]. Extracted ion chromatograms using the HexNAc and NeuAc oxonium ions were performed using the Xcalibur software (Thermo). Manual inspection was performed in the MS/MS spectra of all intact glycopeptides identified.
Protein identification and glycosylation site analysis
For protein identification and glycosylation site analysis, raw files were imported into MaxQuant version 1.5.2.8 [45] and the database search engine Andromeda [46] was used to search MS/MS spectra against a database composed of the Uniprot Human Protein Database (release April 15, 2015; 45,185 entries) with a tolerance level of 4.5 ppm for MS and 20 ppm and 0.5 Da for MS/MS in HCD and CID mode, respectively. Enzyme specificity was set to trypsin with a maximum of two missed cleavages. For the database search of N-formerly endogenous glycopeptides no enzyme was selected and for the N-formerly endogenous glycopeptides treated with trypsin, semi-specific free N-terminus digestion with a maximum of two missed cleavages was considered. Carbamidomethylation of cysteine (57.021 Da) was set as a fixed modification, and oxidation of methionine (15.994 Da), deamidation NQ (+0.984 Da) and protein N-terminal acetylation (42.010 Da) were selected as variable modifications. All identifications were filtered in order to achieve a FDR of 1 %.
After excluding peptides identified as potential contaminants or in reverse database, we manually filtered the peptides containing the deamidation sites within the glycosylation motif NxS/T/C. Motif-X algorithm was used to identify sequence motifs from the endogenous N-glycopeptides, using the ipi.HUMAN.fasta database as background. Only motifs with p < 10E6 were considered.
In order to evaluate the chemical (spontaneous) deamidation rate, we also performed database search using MaxQuant in the HILIC-enriched glycopeptide fraction without treatment with PNGase F (intact glycopeptides fraction) [47].
To determine biological processes and tissue protein localization statistically over-represented in a given protein list InnateDB [48] and Enrichr [49] software were used.
MotifX (http://motif-x.med.harvard.edu/) and Sequence2Logo (http://www.cbs.dtu.dk/biotools/Seq2Logo/) were used to create the sequence motifs for endogenous glycopeptides. For MotifX analysis, peptide sequences were centered at the glycosylation site. For Sequence2Logo analysis, peptide sequences with six aminoacids were loaded.
Results and Discussion
Herein, we employed a simple and straightforward method to characterize the N-linked glycosylation sites of glycoproteins and endogenous glycopeptides in human urine. For that, human urine was collected and concentrated using ultrafiltration. After protein digestion, glycopeptides were enriched using HILIC and N-glycans were removed by PNGase F treatment. The filtrate (peptides <10 kDa) was subjected to solid phase extraction, followed by HILIC enrichment and deglycosylation with PNGase F. Intact glycopeptides were also analyzed in order to site-specifically characterize the N-linked glycosylation heterogeneity. The glycoproteomic workflow is summarized in Fig. 1.
Schematic workflow for preparation of urinary proteins and endogenous peptides for site-specific N-linked glycosylation analysis. The glycopeptides were enriched using polyLC-HILIC affinity chromatography. The glycans were released through treatment with PNGase F and the retained glycopeptide fractions as well as the non-modified peptides (flow through) were analyzed by LC-MS/MS employing CID fragmentation. The retained glycopeptides fraction was also directly analyzed by LC-MS/MS employing HCD fragmentation to site-specific characterization of the glycan microheterogeneity
Overview of the urinary glycoproteome
Considering the formerly N-linked peptides generated after PNGase F treatment, 1092 redundant and 472 unique peptides were identified within the NxS/T/C motif (where X is not proline), covering 532 unique glycosylation sites from 256 glycoproteins (Fig. 2a and Supplemental Table S1 and S2). Considering the unmodified peptides or the peptides with deamidation modification in N or Q outside the NxS/T/C motif, we identified 422 redundant peptides. Therefore, using this strategy we observed a percentage of glycopeptides enrichment of about 72 % (1092/1514).Using the fraction of HILIC-enriched peptides without PNGase F treatment, we searched for peptides bearing a spontaneously deamidated asparagine within the N-linked glycosylation motif, however no peptides were found following this criteria.
Overview of the urinary proteome and glycoproteome. a Number of glycoproteins identified considering the formerly N-linked glycosylated peptides generated after PNGase F treatment (glycoproteome); number of proteins identified in the non-modified peptide fraction that did not bind to HILIC (proteome); number of total proteins identified the human urine considering the glycoproteome and proteome. The two biological replicates (urine collected at separate points over two different day) were considered to the total number of proteins identified in each fraction. b Distribution of N-glycoproteins (glycoproteome) identified in the urine collected at separate points over two different days. c Distribution of proteins (proteome) identified in the urine collected at separate points over two different days. d Overlap between the proteins identified in the glycoproteome and proteome fraction. e Percentage of proteins with signal peptides and transmembrane domains, considering the proteins identified in the glycoproteome or proteome fraction. f Venn diagram of the comparisons of the observed glycoproteome with those published by Saraswat et al. [40] and Halim et al. [41]
The overlap of glycoproteins identified in two different days of urine collection was about 49 % (126 glycoproteins) (Fig. 2b). The non-modified peptides that did not bind to HILIC (considered here as the urine proteome) were also analyzed by LC-MS/MS, resulting in 391 proteins identified with 57 % overlap (223 proteins) between the two different days (Fig. 2c and Supplemental Table S3). In total, 562 proteins were identified (Fig. 2a); however, the glycoproteome fraction brought out 171 exclusive proteins (Fig. 2d). A higher proportion of proteins displaying transmembrane regions (36 %) and signal peptides (70 %) were observed in the glycoproteome-enriched fraction compared to the proteome fraction (24 % and 50 %, respectively) (Fig. 2e and Supplemental Table S2 and S3); thus secreted and cell surface proteins can be more efficiently identified using glycopeptide enrichment strategies. The top ten most significant biological processes overrepresented in the glycoproteome database were: cell adhesion, negative regulation of endopeptidase activity, extracellular matrix organization, angiogenesis, platelet degranulation, regulation of proteolysis, receptor-mediated endocytosis, platelet activation, blood coagulation and heterophilic cell-cell adhesion (Supplemental Table S4). To assess the localization and distribution of the glycoproteins in human tissues we performed enrichment analysis using Human Gene Atlas [50] and Human Proteome Map [51] data-set libraries available within Enrichr software. The glycoproteins identified in the urine of a male individual had a significant enrichment localization relative to the total number of genes (Human Gene Atlas) or proteins (Human Proteome map) identified in each tissue, for liver (p = 0.004, gene names: TF, BTD, AMBP, APOH, LCAT, HRG, EPHA1, KNG1, CPN2), adipocyte (p = 0.01, gene names: LRP1, FZD4, MXRA8, TIMP1, FSTL1), smooth muscle (p = 0.04, gene names: MRC2, HEG1, QSOX1, TIMP1, FSTL1, FBN1), kidney (p = 0.03, gene names: CUBN, EGF, DPEP1, FOLR1) (Humana Gene Atlas, Supplemental Fig. 1a) and adult prostate tissue (p = 0.0004, gene names: CDH2, KLK1, CADM2, AXL, QPCT, DPEP1, SERPINI1, CPZ, NEO1, PVRL1, KLK11) (Human Proteome Map, Supplemental Fig. 1b). These results suggest that the urinary glycoproteome may be a good biofluid to study disorders related not only to urological disease but also other tissues.
The list of glycoproteins identified in our study covered 17 out of 37 (46 %) glycoproteins identified in human urinary exosomes [40] and 16 out of 53 (25 %) glycoproteins identified in the previous healthy urinary glycoproteomic study [41]. However our study showed 231 exclusive glycoproteins that were not identified in the previous urinary glycoproteome studies (Fig. 2f). The modest overlap of glycoproteins identified in our study comparing with these two other studies were expected since biological as well methodological variation may produce different sets of enriched glycoproteins/glycopeptides. For example, the study by Halim et al., used hydrazine capture to enrich sialylated glycoproteins in human urine and the intact glycopeptides were analysis by CID and ECD fragmentation method. In the study by Saraswat et al., they focused on the urinary exosomes and the glycopeptides were enriched using SNA affinity and analyzed by CID-tandem MS. However, despite all these variation we could still observe common glycoproteins with these studies and demonstrate that our approach brings exclusive identifications.
Intact glycopeptides analysis
Site-specific glycan micro-heterogeneity of the urinary glycoproteins was investigated by analyzing an aliquot of the HILIC-enriched glycopeptides without prior PNGase F treatment, using high resolution LC-MS/MS and higher energy collision dissociation (HCD) (Fig. 1). Using the intense diagnostic saccharide oxonium ions such as m/z 138.06/204.09 for N-acetylhexosamine (HexNAc) and/or m/z 274.09/292.10 for N-acetylneuraminic acid (NeuAc), we compared the glycosylation profile of intact glycopeptides expressed in the human urine collected in two different days (Fig. 3a).
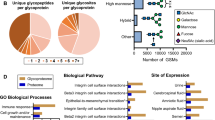
Investigation of the site-specific N-linked glycans of the human urinary glycoproteins. a Extracted ion chromatogram of HexNAc (m/z 138.06/204.09) and NeuAc (m/z 274.09 and 292.10) oxonium ions with a mass tolerance of 20 ppm. b Distribution of unique N-linked intact glycopeptides in the urine collected in two different days. c Distribution of glycoproteins covered by the identified intact glycopeptides in the urine collected in two different days. d Distribution of the number of intact glycopeptides identified according to the class (high mannose or complex/hybrid) and features
The extracted ion chromatogram for the HexNAc and NeuAc oxonium ions showed similar profile between the two different days (Fig. 3a). Interestingly, the diagnostic ion at m/z 274.091/292.10 pointed out to large amount of glycopeptides containing NeuAc in human urine.
To assess site-specific glycan composition, intact glycopeptides were identified using Byonic software. 1437 and 1038 spectra were assigned to a glycopeptide, in the first and second day, respectively (data not shown). Considering the non-glycosylated peptides, 172 and 91 were identified in the first and second day, respectively, corresponding to a percentage of glycopeptide enrichment in almost 90 %. However, for high confident identification, we filtered only the glycopeptides identified with 0 % FDR, resulting in 156 and 402 identifications in the first and second day, respectively (Supplemental Table S5). From the 156 redundant intact glycopeptides identified with 0 % FDR in the first day, all of them had PEP 2D <0.001 (less than 0.1 % chance that the match is wrong). In the second day, from 402 redundant intact glycopeptides identified with 0 % FDR, 257 had PEP 2D <0.001 and 142 had PEP 2D <0.01 (less than 1 % chance of a wrong match) and only 3 had PEP 2D <0.1 (less than 10 % chance of a wrong match). The glycopeptides identified with the lowest PEP score had 14 fragments assigned (not shown). These parameters were chosen for stringent and confident assignments [17, 52]. However, more efforts are needed to establish a method for intact glycopeptide FDR calculation.
In total, 303 unique intact glycopeptides were identified, with an overlap of 20 % between the first and second day (Fig. 3b). These glycopeptides covered a total of 60 glycoproteins, of which 30 (50 %) were identified in both days (Fig. 3c). Unfortunately, the FDR and scores only assesses the correctness of the peptide sequence [44], which means that the glycopeptide identification can be partially correct, e.g. correct peptide sequence and glycosylation site but still remain incorrect at the identified monosaccharide composition. For example, we observed that from the 303 unique intact glycopeptides, 61 showed the glycan NeuGc, which is uncommon in human. We manually inspect all MS/MS spectra to search for its oxonium ions at m/z 308/290 however we did not find these ions but the NeuAc oxonium ion instead (some examples are provided in Supplemental Fig. 3). Since the mass difference between NeuAc and NeuGc is 16 Da, we checked if the Y1 ion of these peptides also contained the addition of +16 Da. Interestingly, from the 43 peptide sequences assigned with NeuGc, we were able to find in 22 of them the Y1 + 16 in the MS/MS spectra. Therefore, the peptides identified with NeuGc have a modification of +16 Da in the peptide sequence (Supplemental Fig. 2 A-U).
Most of the glycan structures were classified as hybrid/complex; amongst them 110 glycopeptides were identified with fucose and sialic acid (36 %), 78 with only fucose (25 %), 49 with only sialic acid (16 %) and 30 (10 %) composed by only HexNAc and Hex. High mannose was observed in 35 glycopeptides (11 %) (Fig. 3d and Supplemental Table S5). Interestingly, Gao et al. showed very similar distribution of N-glycans composition in human serum, where complex/hybrid glycans containing fucose and sialic acid were the main composition, followed by complex/hybrid with only sialic acid or only fucose, undecorated complex/hybrid and in less proportion the high mannose [53]. We also performed charge state distribution analysis of the intact glycopeptides identified in each class. We observed that glycopeptides containing NeuAc presented the highest percentage of >4 charges, while high mannose glycopeptides were identified mainly with +2 and +3 charges (Supplemental Fig. 3).
The 303 unique intact glycopeptides covered to 104 unique peptide sequences. These sequences had an overlap of 58 % (60 peptides) with the peptides identified in the deglycosylated fraction, in which the peptide harbored the deamidation site in the consensus NXS/T/C motif (where X is not proline) (Supplemental Fig. 4). Using UnicarbKB database, we manually checked the possible structures for each glycan assigned to the 186 intact glycopeptides, whose peptide sequences were also identified in the formerly N-linked peptide fraction (Supplemental Table S6). From 57 glycan unique structures, 48 were found in the UnicarbKB database [54] (http://www.unicarbkb.org/) (Table 1). The glycopeptides showed in Table 1 were considered here as high confident identified glycopeptides in human urine, since they were evidenced by two different MS-based approaches and had the glycan structure annotated in a public database. Examples of annotated spectra of N-linked glycopeptides, directly from the Xcalibur program, with typical oxonium ions, glycosidic fragments and b/y-ions, are provided in Supplemental Fig. 5a-d.
N-linked glycosylation sites analysis of urinary endogenous peptides
Low molecular weight glycopeptides that were not retained by the 10 kDa-cutoff membrane was extracted and enriched using HILIC. The formerly N-linked peptides generated after PNGase F treatment were analyzed by LC-MS/MS. We identified 125 glycosylated peptides covering 70 unique glycosylation sites within NxS/T/C motifs, from 40 glycoproteins (Fig. 4a and Supplemental Table S7).
N-glycosylation site analysis of endogenous glycopeptides (<10 kDa). a Distribution of identified formerly N-linked glycosylated peptides generated after PNGase F treatment in intact or tryptic endogenous glycopeptides. Only deamidation sites within NxS/T/C motif where x is not P were considered. b Distribution of proteins identified in the glycoproteome, proteome and glycopeptidome fraction
We further performed amino acid frequency analysis in order to search for specific protease preference cleavage sites that may be generating the endogenous glycopeptides. We found that leucine is the main amino acid occurring in the the NxT motif from the glycosylation site to the N-terminal of the peptide. On the other hand, in the NxS motif we did not observe the same preference and leucine was observed in the vicinity of the S to the C-terminal direction (Supplemental Fig. 6a and b). It was demonstrated that some metalloproteases (MMPs 1, 2, 3, 7, 9, 12, 13, 14) and ADAMs (ADAM17, ADAM10) have preferences for hydrophobic residues, such as leucine in the cleavage site [55–58]. Moreover, we found that leucine and isoleucine were the main aminoacids present at the N-terminus of the endogeneous glycopeptides (Supplemental Fig. 6c).
Since the peptide dabatabase search is compromised by large peptide sequences and no digestion mode, we performed tryptic digestion of the HILIC-enriched N-glycopeptides in order to increase the coverage of N-linked glycosylation sites and reduce false positives. Using this strategy we were able to increase the number of unique glycosylation sites to 134, which covered to 69 glycoproteins (Fig 4a and Supplemental Table S8).
In total, 204 unique N-linked glycosylation sites were identified in the human endogenous glycopeptide, covering90 glycoproteins. Interestingly, when we compared the overlap of proteins identified in the glycoproteome, proteome and glycopeptidome fraction, we observed that the glycopeptidome were able to retrieve exclusive proteins (57 out of 90) (Fig. 4b).
Chemical artifacts were searched using the intact glycopeptides fraction and the peptides bearing a spontaneously deamidated asparagine within the N-linked glycosylation motif were reported in Supplemental Table S1 and S2.
Conclusion
To our knowledge, the present dataset provides a highly confident characterization of N-linked glycosylation sites in human urinary proteins, including site-specific glycan microheterogeneity and for the first time the N-glycosylation sites from endogenous glycopeptides. This dataset, as well the methodology showed in this study, consist in a valuable resource for future MS-based glycoproteomic studies concerned with disorders in which N-linked protein glycosylation may be aberrant and an alternative approach for searching for novel candidate biomarkers in human urine.
References
Apweiler R., Hermjakob H., Sharon N.: On the frequency of protein glycosylation, as deduced from analysis of the SWISS-PROT database. Biochim. Biophys. Acta. 1473(1), 4–8 (1999)
Ferris S.P., Kodali V.K., Kaufman R.J.: Glycoprotein folding and quality-control mechanisms in protein-folding diseases. Dis. Model. Mech. 7(3), 331–341 (2014). doi:10.1242/dmm.014589
Marth J.D., Grewal P.K.: Mammalian glycosylation in immunity. Nat. Rev. Immunol. 8(11), 874–887 (2008). doi:10.1038/nri2417
Cummings R.D., Pierce J.M.: The challenge and promise of glycomics. Chem. Biol. 21(1), 1–15 (2014). doi:10.1016/j.chembiol.2013.12.010
Crocker P.R., Feizi T.: Carbohydrate recognition systems: functional triads in cell-cell interactions. Curr. Opin. Struct. Biol. 6(5), 679–691 (1996)
Pinho S.S., Reis C.A.: Glycosylation in cancer: mechanisms and clinical implications. Nat. Rev. Cancer. 15(9), 540–555 (2015). doi:10.1038/nrc3982
Fuster M.M., Esko J.D.: The sweet and sour of cancer: glycans as novel therapeutic targets. Nat. Rev. Cancer. 5(7), 526–542 (2005). doi:10.1038/nrc1649
Dube D.H., Bertozzi C.R.: Glycans in cancer and inflammation–potential for therapeutics and diagnostics. Nat. Rev. Drug Discov. 4(6), 477–488 (2005). doi:10.1038/nrd1751
Stowell S.R., Ju T., Cummings R.D.: Protein glycosylation in cancer. Annu. Rev. Pathol. 10, 473–510 (2015). doi:10.1146/annurev-pathol-012414-040438
Gilgunn S., Conroy P.J., Saldova R., Rudd P.M., O'Kennedy R.J.: Aberrant PSA glycosylation–a sweet predictor of prostate cancer. Nat. Rev. Urol. 10(2), 99–107 (2013). doi:10.1038/nrurol.2012.258
Hauselmann I., Borsig L.: Altered tumor-cell glycosylation promotes metastasis. Frontiers in oncology. 4, 28 (2014). doi:10.3389/fonc.2014.00028
Zhang Y., Jiao J., Yang P., Lu H.: Mass spectrometry-based N-glycoproteomics for cancer biomarker discovery. Clin. Proteomics. 11(1), 18 (2014). doi:10.1186/1559-0275-11-18
Thaysen-Andersen M., Packer N.H.: Advances in LC-MS/MS-based glycoproteomics: getting closer to system-wide site-specific mapping of the N- and O-glycoproteome. Biochim. Biophys. Acta. 1844(9), 1437–1452 (2014). doi:10.1016/j.bbapap.2014.05.002
Kaji H., Saito H., Yamauchi Y., Shinkawa T., Taoka M., Hirabayashi J., Kasai K., Takahashi N., Isobe T.: Lectin affinity capture, isotope-coded tagging and mass spectrometry to identify N-linked glycoproteins. Nat. Biotechnol. 21(6), 667–672 (2003). doi:10.1038/nbt829
Morelle W., Faid V., Chirat F., Michalski J.C.: Analysis of N- and O-linked glycans from glycoproteins using MALDI-TOF mass spectrometry. Methods Mol. Biol. 534, 5–21 (2009). doi:10.1007/978-1-59745-022-5_1
Jensen P.H., Karlsson N.G., Kolarich D., Packer N.H.: Structural analysis of N- and O-glycans released from glycoproteins. Nat. Protoc. 7(7), 1299–1310 (2012). doi:10.1038/nprot.2012.063
Woo C.M., Iavarone A.T., Spiciarich D.R., Palaniappan K.K., Bertozzi C.R.: Isotope-targeted glycoproteomics (IsoTaG): a mass-independent platform for intact N- and O-glycopeptide discovery and analysis. Nat. Methods. 12(6), 561–567 (2015). doi:10.1038/nmeth.3366
Sun S., Shah P., Eshghi S.T., Yang W., Trikannad N., Yang S., Chen L., Aiyetan P., Hoti N., Zhang Z., Chan D.W., Zhang H.: Comprehensive analysis of protein glycosylation by solid-phase extraction of N-linked glycans and glycosite-containing peptides. Nat. Biotechnol. 34(1), 84–88 (2016). doi:10.1038/nbt.3403
Medzihradszky K.F., Kaasik K., Chalkley R.J.: Tissue-specific glycosylation at the glycopeptide level. Mol. Cell. Proteomics: MCP. 14(8), 2103–2110 (2015). doi:10.1074/mcp.M115.050393
Hoffmann M., Marx K., Reichl U., Wuhrer M., Rapp E.: Site-specific O-glycosylation analysis of human blood plasma proteins. Mol. Cell. Proteomics: MCP. 15(2), 624–641 (2016). doi:10.1074/mcp.M115.053546
Parker B.L., Thaysen-Andersen M., Solis N., Scott N.E., Larsen M.R., Graham M.E., Packer N.H., Cordwell S.J.: Site-specific glycan-peptide analysis for determination of N-glycoproteome heterogeneity. J. Proteome Res. 12(12), 5791–5800 (2013). doi:10.1021/pr400783j
Stavenhagen K., Hinneburg H., Thaysen-Andersen M., Hartmann L., Varon Silva D., Fuchser J., Kaspar S., Rapp E., Seeberger P.H., Kolarich D.: Quantitative mapping of glycoprotein micro-heterogeneity and macro-heterogeneity: an evaluation of mass spectrometry signal strengths using synthetic peptides and glycopeptides. J. Mass Spectrom.: JMS. 48(6), 627–639 (2013). doi:10.1002/jms.3210
Hagglund P., Bunkenborg J., Elortza F., Jensen O.N., Roepstorff P.: A new strategy for identification of N-glycosylated proteins and unambiguous assignment of their glycosylation sites using HILIC enrichment and partial deglycosylation. J. Proteome Res. 3(3), 556–566 (2004)
Mysling S., Palmisano G., Hojrup P., Thaysen-Andersen M.: Utilizing ion-pairing hydrophilic interaction chromatography solid phase extraction for efficient glycopeptide enrichment in glycoproteomics. Anal. Chem. 82(13), 5598–5609 (2010). doi:10.1021/ac100530w
Li, X., Jiang, J., Zhao, X., Wang, J., Han, H., Zhao, Y., Peng, B., Zhong, R., Ying, W., Qian, X.: N-glycoproteome analysis of the secretome of human metastatic hepatocellular carcinoma cell lines combining hydrazide chemistry, HILIC enrichment and mass spectrometry. PloS one 8(12), e81921 (2013). doi:10.1371/journal.pone.0081921
Melo-Braga M.N., Schulz M., Liu Q., Swistowski A., Palmisano G., Engholm-Keller K., Jakobsen L., Zeng X., Larsen M.R.: Comprehensive quantitative comparison of the membrane proteome, phosphoproteome, and sialiome of human embryonic and neural stem cells. Mol. Cell. Proteomics: MCP. 13(1), 311–328 (2014). doi:10.1074/mcp.M112.026898
Pompach P., Chandler K.B., Lan R., Edwards N., Goldman R.: Semi-automated identification of N-glycopeptides by hydrophilic interaction chromatography, nano-reverse-phase LC-MS/MS, and glycan database search. J. Proteome Res. 11(3), 1728–1740 (2012). doi:10.1021/pr201183w
Cheng K., Chen R., Seebun D., Ye M., Figeys D., Zou H.: Large-scale characterization of intact N-glycopeptides using an automated glycoproteomic method. J. Proteome. 110, 145–154 (2014). doi:10.1016/j.jprot.2014.08.006
Wuhrer M., de Boer A.R., Deelder A.M.: Structural glycomics using hydrophilic interaction chromatography (HILIC) with mass spectrometry. Mass Spectrom. Rev. 28(2), 192–206 (2009). doi:10.1002/mas.20195
Yu Y.Q., Gilar M., Kaska J., Gebler J.C.: A rapid sample preparation method for mass spectrometric characterization of N-linked glycans. Rapid Commun. Mass Spectrom.: RCM. 19(16), 2331–2336 (2005). doi:10.1002/rcm.2067
Shimwell N.J., Bryan R.T., Wei W., James N.D., Cheng K.K., Zeegers M.P., Johnson P.J., Martin A., Ward D.G.: Combined proteome and transcriptome analyses for the discovery of urinary biomarkers for urothelial carcinoma. Br. J. Cancer. 108(9), 1854–1861 (2013). doi:10.1038/bjc.2013.157
Zhang H., Cao J., Li L., Liu Y., Zhao H., Li N., Li B., Zhang A., Huang H., Chen S., Dong M., Yu L., Zhang J., Chen L.: Identification of urine protein biomarkers with the potential for early detection of lung cancer. Sci. Rep. 5, 11805 (2015). doi:10.1038/srep11805
Wu J., Chen Y.D., Gu W.: Urinary proteomics as a novel tool for biomarker discovery in kidney diseases. J. Zhejiang Univ. Sci. B. 11(4), 227–237 (2010). doi:10.1631/jzus.B0900327
Thomas, C.E., Sexton, W., Benson, K., Sutphen, R., Koomen, J.: Urine collection and processing for protein biomarker discovery and quantification. Cancer epidemiology, biomarkers & prevention: a publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive Oncology 19(4), 953–959 (2010). doi:10.1158/1055-9965.EPI-10-0069
Overbye A., Skotland T., Koehler C.J., Thiede B., Seierstad T., Berge V., Sandvig K., Llorente A.: Identification of prostate cancer biomarkers in urinary exosomes. Oncotarget. 6(30), 30357–30376 (2015). doi:10.18632/oncotarget.4851
Haj-Ahmad T.A., Abdalla M.A., Haj-Ahmad Y.: Potential urinary protein biomarker candidates for the accurate detection of prostate cancer among benign prostatic hyperplasia patients. J. Cancer. 5(2), 103–114 (2014). doi:10.7150/jca.6890
Jedinak A., Curatolo A., Zurakowski D., Dillon S., Bhasin M.K., Libermann T.A., Roy R., Sachdev M., Loughlin K.R., Moses M.A.: Novel non-invasive biomarkers that distinguish between benign prostate hyperplasia and prostate cancer. BMC Cancer. 15, 259 (2015). doi:10.1186/s12885-015-1284-z
Wang L., Li F., Sun W., Wu S., Wang X., Zhang L., Zheng D., Wang J., Gao Y.: Concanavalin A-captured glycoproteins in healthy human urine. Mol. Cell. Proteomics: MCP. 5(3), 560–562 (2006). doi:10.1074/mcp.D500013-MCP200
Yang N., Feng S., Shedden K., Xie X., Liu Y., Rosser C.J., Lubman D.M., Goodison S.: Urinary glycoprotein biomarker discovery for bladder cancer detection using LC/MS-MS and label-free quantification. Clinical cancer research: an official journal of the American Association for Cancer Research. 17(10), 3349–3359 (2011). doi:10.1158/1078-0432.CCR-10-3121
Saraswat M., Joenvaara S., Musante L., Peltoniemi H., Holthofer H., Renkonen R.: N-linked (N-) glycoproteomics of urinary exosomes. [Corrected]. Mol. Cell. Proteomics: MCP. 14(2), 263–276 (2015). doi:10.1074/mcp.M114.040345
Halim A., Nilsson J., Ruetschi U., Hesse C., Larson G.: : Human urinary glycoproteomics; attachment site specific analysis of N- and O-linked glycosylations by CID and ECD. Mol. Cell. Proteomics: MCP. 11(4), M111 013649 (2012). doi:10.1074/mcp.M111.013649
Bern, M., Kil, Y.J., Becker, C.: Byonic: advanced peptide and protein identification software. Current protocols in bioinformatics / editoral board, Andreas D. Baxevanis. .. [et al.] Chapter 13, Unit13 20 (2012). doi:10.1002/0471250953.bi1320s40
Bern M., Cai Y., Goldberg D.: Lookup peaks: a hybrid of de novo sequencing and database search for protein identification by tandem mass spectrometry. Anal. Chem. 79(4), 1393–1400 (2007). doi:10.1021/ac0617013
Bern M.W., Kil Y.J.: Two-dimensional target decoy strategy for shotgun proteomics. J. Proteome Res. 10(12), 5296–5301 (2011). doi:10.1021/pr200780j
Cox J., Mann M.: MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26(12), 1367–1372 (2008). doi:10.1038/nbt.1511
Cox J., Neuhauser N., Michalski A., Scheltema R.A., Olsen J.V., Mann M.: Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10(4), 1794–1805 (2011). doi:10.1021/pr101065j
Palmisano G., Melo-Braga M.N., Engholm-Keller K., Parker B.L., Larsen M.R.: Chemical deamidation: a common pitfall in large-scale N-linked glycoproteomic mass spectrometry-based analyses. J. Proteome Res. 11(3), 1949–1957 (2012). doi:10.1021/pr2011268
Breuer K., Foroushani A.K., Laird M.R., Chen C., Sribnaia A., Lo R., Winsor G.L., Hancock R.E., Brinkman F.S., Lynn D.J.: InnateDB: systems biology of innate immunity and beyond–recent updates and continuing curation. Nucleic Acids Res. 41(Database issue), D1228–D1233 (2013). doi:10.1093/nar/gks1147
Chen E.Y., Tan C.M., Kou Y., Duan Q., Wang Z., Meirelles G.V., Clark N.R., Ma'ayan A.: Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinf. 14, 128 (2013). doi:10.1186/1471-2105-14-128
Su A.I., Wiltshire T., Batalov S., Lapp H., Ching K.A., Block D., Zhang J., Soden R., Hayakawa M., Kreiman G., Cooke M.P., Walker J.R., Hogenesch J.B.: A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl. Acad. Sci. U. S. A. 101(16), 6062–6067 (2004). doi:10.1073/pnas.0400782101
Kim M.S., Pinto S.M., Getnet D., Nirujogi R.S., Manda S.S., Chaerkady R., Madugundu A.K., Kelkar D.S., Isserlin R., Jain S., Thomas J.K., Muthusamy B., Leal-Rojas P., Kumar P., Sahasrabuddhe N.A., Balakrishnan L., Advani J., George B., Renuse S., Selvan L.D., Patil A.H., Nanjappa V., Radhakrishnan A., Prasad S., Subbannayya T., Raju R., Kumar M., Sreenivasamurthy S.K., Marimuthu A., Sathe G.J., Chavan S., Datta K.K., Subbannayya Y., Sahu A., Yelamanchi S.D., Jayaram S., Rajagopalan P., Sharma J., Murthy K.R., Syed N., Goel R., Khan A.A., Ahmad S., Dey G., Mudgal K., Chatterjee A., Huang T.C., Zhong J., Wu X., Shaw P.G., Freed D., Zahari M.S., Mukherjee K.K., Shankar S., Mahadevan A., Lam H., Mitchell C.J., Shankar S.K., Satishchandra P., Schroeder J.T., Sirdeshmukh R., Maitra A., Leach S.D., Drake C.G., Halushka M.K., Prasad T.S., Hruban R.H., Kerr C.L., Bader G.D., Iacobuzio-Donahue C.A., Gowda H., Pandey A.: A draft map of the human proteome. Nature. 509(7502), 575–581 (2014). doi:10.1038/nature13302
Shah P., Wang X., Yang W., Toghi Eshghi S., Sun S., Hoti N., Chen L., Yang S., Pasay J., Rubin A., Zhang H.: Integrated proteomic and glycoproteomic analyses of prostate cancer cells reveal glycoprotein alteration in protein abundance and glycosylation. Mol. Cell. Proteomics: MCP. 14(10), 2753–2763 (2015). doi:10.1074/mcp.M115.047928
Gao W.N., Yau L.F., Liu L., Zeng X., Chen D.C., Jiang M., Liu J., Wang J.R., Jiang Z.H.: Microfluidic Chip-LC/MS-based Glycomic analysis revealed distinct N-glycan profile of rat serum. Sci. Rep. 5, 12844 (2015). doi:10.1038/srep12844
Campbell M.P., Peterson R., Mariethoz J., Gasteiger E., Akune Y., Aoki-Kinoshita K.F., Lisacek F., Packer N.H.: UniCarbKB: building a knowledge platform for glycoproteomics. Nucleic Acids Res. 42(Database issue), D215–D221 (2014). doi:10.1093/nar/gkt1128
Eckhard U., Huesgen P.F., Schilling O., Bellac C.L., Butler G.S., Cox J.H., Dufour A., Goebeler V., Kappelhoff R., Keller U.A., Klein T., Lange P.F., Marino G., Morrison C.J., Prudova A., Rodriguez D., Starr A.E., Wang Y., Overall C.M.: Active site specificity profiling of the matrix metalloproteinase family: Proteomic identification of 4300 cleavage sites by nine MMPs explored with structural and synthetic peptide cleavage analyses. Matrix Biol.: journal of the International Society for Matrix Biology. 49, 37–60 (2016). doi:10.1016/j.matbio.2015.09.003
Tucher J., Linke D., Koudelka T., Cassidy L., Tredup C., Wichert R., Pietrzik C., Becker-Pauly C., Tholey A.: LC-MS based cleavage site profiling of the proteases ADAM10 and ADAM17 using proteome-derived peptide libraries. J. Proteome Res. 13(4), 2205–2214 (2014). doi:10.1021/pr401135u
Prudova A., auf dem Keller U., Butler G.S., Overall C.M.: Multiplex N-terminome analysis of MMP-2 and MMP-9 substrate degradomes by iTRAQ-TAILS quantitative proteomics. Mol. Cell. Proteomics: MCP. 9(5), 894–911 (2010). doi:10.1074/mcp.M000050-MCP201
Schilling O., Overall C.M.: Proteome-derived, database-searchable peptide libraries for identifying protease cleavage sites. Nat. Biotechnol. 26(6), 685–694 (2008). doi:10.1038/nbt1408
Acknowledgments
CNPq (GP 441878/2014-8) and FAPESP (GP: 2014/06863-3), Rebeca Kawahara is supported the Capes, PNPD, and FAPESP (2015/02866-0). Joyce Saad is supported by the “Programa Unificado de Bolsas de Estudo”. The facility Biomass at CEFAP-USP is acknowledged for the analyses.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic Supplementary Material
Supplemental Figure 1
Tissue localization enrichment analysis in the list of identified urinary glycoproteins (PDF 36 kb)
Supplemental Figure 2
Manual verification of the Y1 + 16 Da in the MS/MS spectra from intact glycopeptides identified with NeuGc. The Y1 ion +16 Da is indicated by the red arrow. (PDF 248 kb)
Supplemental Figure 3
Charge state distribution of intact glycopeptides identified in each glycan composition class. The percentage was calculated based on the number of glycopeptides with charge +2, +3, +4 and >4 in each class to the total number of glycopeptides identified in each class. (PDF 15 kb)
Supplemental Figure 4
Overlap between the number of peptide sequences identified in the formerly N-linked peptides generated after PNGase F treatment and intact glycopeptides (with 0 % FDR) identified using Byonic software. (PDF 12 kb)
Supplemental Figure 5
Examples of annotated spectra for N-linked glycopeptides identified in urinary glycoproteins (PDF 62 kb)
Supplemental Figure 6
Sequence recognition motif in endogenous N-glycopeptides. N-glycosylation consensus sequence as derived using MotifX. A) MotifX analysis centered on the NxT glycosylation motif. B) MotifX analysis centered on the NxS glycosylation motif. C) Sequence2logo analysis obtained using the endogenous peptide sequences with six aminoacids length. (PDF 93 kb)
ESM 7
(XLSX 1076 kb)
Rights and permissions
About this article
Cite this article
Kawahara, R., Saad, J., Angeli, C.B. et al. Site-specific characterization of N-linked glycosylation in human urinary glycoproteins and endogenous glycopeptides. Glycoconj J 33, 937–951 (2016). https://doi.org/10.1007/s10719-016-9677-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10719-016-9677-z