Abstract
Transposable elements (TEs) are mobile genes with an inherent ability to move within and among genomes. Theory predicts that TEs proliferate extensively during physiological stress due to the breakdown of TE repression systems. We tested this hypothesis in Schistosoma mansoni, a widespread trematode parasite that causes the human disease schistosomiasis. According to phylogenetic analysis, S. mansoni invaded the new world during the last 500 years. We hypothesized that new world strains of S. mansoni would have more copies of TEs than old world strains due to the physiological stress associated with invasion of the new world. We quantified the copy number of six TEs (Saci-1, Saci-2 and Saci-3, Perere-1, Merlin-sm1, and SmTRC1) in the genome and the transcriptome of old world and new world strains of S. mansoni, using qPCR relative quantification. As predicted, the genomes of new world parasites contain significantly more copies of class I and class II TEs in both laboratory and field strains. However, such differences are not observed in the transcriptome suggesting that either TE silencing mechanisms have reactivated to control the expression of these elements or the presence of inactive truncated copies of TEs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genes were originally conceived as static entities confined to specific loci along the chromosomes. The discovery of transposable elements (TEs) in eukaryotes challenged that concept by revealing a previously unknown mechanism for the production of genetic and genomic variation (Hua-Van et al. 2011). TEs are mobile DNA sequences with an inherent ability to move within and among genomes (Pritham 2009). This mobility allows them to exert control over their local genetic environments. Based on their mode of transposition, TEs are grouped into two major categories. Class I elements (retrotransposons; copy and paste) transpose through an RNA intermediate whereas Class II elements (DNA transposons; cut and paste) transpose directly as DNA (Pritham 2009).
TEs change the evolutionary trajectory of genomes in many ways. TE activity can (a) facilitate diversification of the genome by the emergence of new genes; (b) modify gene expression patterns; and (c) promote chromosomal rearrangements that ultimately increase the adaptability of the species (Oliver and Greene 2009). On the other hand, excessive TE proliferation can (a) decrease organismal fitness by mutagenic effects of insertions; (b) increase metabolic costs associated with transcription; and (c) induce translation and chromosomal damage due to ectopic recombination (Oliver and Greene 2009; Nuzhdin 1999). Thus, genomes have evolved mechanisms to repress the uncontrolled proliferation of TEs. These mechanisms include epigenetic processes such as RNAi, a transcriptional and post-transcriptional gene silencing system that relies on short, non-coding RNAs (Blumenstiel 2011).
Nonetheless, physiological stress can activate TEs and disrupt epigenetic silencing, thus leading to TE proliferation (Capy et al. 2000; Slotkin and Martienssen 2007; Zeh et al. 2009). For instance, in Drosophila simulans, developmental temperature can affect both somatic and germline TE excision rates (Chakrani et al. 1993; Garza et al. 1991). Similarly, the invasion of new habitats may challenge genomes and lead to TE activation. For example, the increased TE copy number in D. melanogaster compared to D. simulans could be associated with its worldwide colonization (Vieira and Biémont 2004). TE proliferation has also been observed in colonizing populations of Drosophila subobscura and Arabidopsis lyrata (García Guerreiro et al. 2008; Lockton et al. 2008), presumably due to the physiological stresses associated with the invasion of new habitats. Such TE proliferations may facilitate the invasion of novel habitats through genome dynamism.
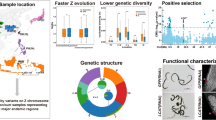
Herein, we analyze TE proliferations in the human parasite Schistosoma mansoni, which has recently invaded new habitats and thus may be exposed to novel environmental and evolutionary stressors. The parasites of the genus Schistosoma are the causative agents of schistosomiasis, a chronic and debilitating human disease of significant medical importance, contributing to 200 million cases in the tropics (Brunette 2014; Chitsulo et al. 2000). Eggs produced by adult parasites become lodged in viscera, provoking granulomatous reactions and portal hypertension (Lambertucci et al. 2000; Fig. 1). Six Schistosoma species are known to infect humans; one of the most common is S. mansoni, the etiological agent of intestinal schistosomiasis (Crompton 1999; Brindley and Hotez 2013). In fact, S. mansoni shows the most widespread distribution, spanning both old world and new world. Intestinal schistosomiasis is found in 54 countries, including the Arabian Peninsula, sub-Saharan Africa, Caribbean islands and South America (Chitsulo et al. 2000; Brunette 2014).
Life cycle of Schistosoma mansoni. Adult male and female worms inhabit definitive vertebrate hosts, mating and producing eggs. The eggs are released from the host through feces, hatch into miracidia in freshwater and infect snail intermediate hosts. The cercariae released from intermediate hosts infect vertebrate hosts by skin penetration (Figure modified from Wijayawardena et al. 2013).
The diversification of the S. mansoni lineage has intrigued both parasitologists and molecular biologists. Molecular evidence indicates that the genus Schistosoma originated in Asia (Snyder and Loker 2000; Lockyer et al. 2003; Morgan et al. 2005) and descendants of this lineage then moved to Africa approximately 12–19 million years ago, possibly by means of a widespread mammalian migration (Lockyer et al. 2003). African lineages underwent extensive radiation and gave rise to S. mansoni about 0.3–0.43 million years ago, concomitant with the emergence of modern humans (Morgan et al. 2005). From Africa, S. mansoni likely invaded the new world through the Atlantic slave trade within the last 500 years and quickly became established in the new world (Morgan et al. 2005; Files 1951, Fletcher et al. 1981, Desprès et al. 1993). This rapid colonization of the new world may have been facilitated by its genomic structure.
The genomes of three major human infecting schistosomes have been sequenced recently (Berriman et al. 2009; Protasio et al. 2012; Zhou et al. 2009; Young et al. 2012). Interestingly, 40–50 % of the schistosome genome consists of transposable elements (TEs). The estimated copy numbers of TEs in the genome of S. mansoni varies between 50 (e.g. SINBAD class I LTR TE; Thomas et al. 2010) and 20, 000 copies (e.g. SR1, class I non-LTR TE; Thomas et al. 2010). Several families of both class I and class II TE sequences have been identified in S. mansoni, most of them actively transcribed (Protasio et al. 2012; Thomas et al. 2010). For instance, LINE elements of S. mansoni show evidence of recent bursts of transposition that were not paralleled in the conspecific S. japonicum, possibly associated with the move from Asia to Africa and establishment and consequent speciation (Venancio et al. 2010). Given its phylogeography, physiological stress associated with the invasion of new habitats may have also shaped the TE repertoire of S. mansoni (Venancio et al. 2010). Herein, we test this hypothesis by performing qPCR to compare relative copy numbers of both class I and class II TEs in new and old world strains. Our null hypothesis is that there is no difference in relative copy numbers between new and old world strains. However, if increased TE content in S. mansoni results from physiological stress associated with the invasion of the new world, then we expect the new world strains of S. mansoni to have an increased copy number of TEs compared to the old world strains.
Materials and methods
Parasite strains
In total, four new world strains and two old world strains of S. mansoni were used in our study. Either single (adults or cercariae) or multiple (adults and cercariae) life cycle stages were used from each strain (Table 1). We obtained PR-1 and NMRI strains from the Biomedical Research Institute (Rockville, MD). The NMRI strain originated from eggs isolated from Puerto Rican school children in the early 1940’s whereas the PR-1 strain was collected from infected snails in Arecibo, Puerto Rico, in 1950 (Table 1; Lewis et al. 1986). The LE strain originated from Belo Horizonte, Minas Gerais, Brazil, in 1965 from an infected human patient. We also used a Brazilian field strain from Americaninhas (AMH) first isolated in 2005 from human fecal samples (Thiele et al. 2008). The hatched miracidia of AMH isolate were passed through Biomphalaria glabrata snails and Balb/cj mice to harvest the adult worms. Our old world strains included EGY-1, collected from infected humans in 1978, Qualhum, Egypt (Fletcher et al. 1981) and a Kenyan field isolate (KY: cercariae obtained from infected snails Biomplalaria pfeifferi, in 2013 from Mwea, Kenya).
All laboratory strains were maintained in the recommended snail host/strain combinations (Lewis et al. 1986, Fletcher et al. 1981). At 4–6 weeks after infection by miracidia, Biomphalaria snails were exposed to fluorescent light for 2 h to release the cercarial stage of the parasite from the snail (Fig. 1). BALB/cJ mice (n = 3–6) were infected with approximately 120 cercariae each, using the tail emersion method modified from Oliver and Stirewalt (1952). After 6–8 weeks, mice were humanely euthanized with CO2 and dissected to obtain adult worms and eggs. Adult worms were extracted from hepatic portal and mesenteric veins. The hatched eggs were used to infect the snails to maintain the parasite life cycle. All animal use followed protocols approved by the Purdue Animal Care and Use Committee (PACUC).
DNA extraction and cDNA synthesis
We used male parasites as they are considerably larger than the females, yielding sufficient quantity of good quality DNA. DNA was extracted from 30 fresh individual male worms using standard proteinase K digestion followed by organic extraction (Sambrook and Russell 2001). Extracted DNA samples were stored at −20 °C until PCR. Cercariae were collected from pooled samples, by shedding ~100 infected snails per strain. QiaAmp® (Qiagen) DNA extraction kits were used to extract DNA from single cercariae. Individual cercariae were sexed using the primers designed for the female-specific w1 repeat (Grevelding 1995). Only male cercariae were used in our analysis (n = 30 per strain), to avoid sex bias.
TRIzol® reagent (Invitrogen) was used to extract RNA from adult male worms according to the manufacturer’s instructions. As single worms yielded insufficient RNA, pooled worms were used. Each pool contained 5 adult worms, and 20 such pools per strain were used in the analysis (i.e. 20 biological replicates per strain; corresponding to strains NMRI, PR-1, LE and EGY-1). The quantity and quality of RNA was analyzed using a spectrophotometer (NanoDrop 3000, ThermoFisher Scientific) and by 1 % agarose gel electrophoresis followed by ethidium bromide staining. All RNA samples were treated with DNase I (New England Biolabs) prior to cDNA synthesis to remove trace amounts of contaminating genomic DNA. First strand cDNA synthesis was performed using the ClonTech SMART cDNA library construction kit with a modified CDS III/3′ cDNA synthesis primer (5′ TAGAGGCCGAGGCGGCCGACATGTTTTGTTTTTTTTTCTTTTTTTTTTVN 3′) (Zhu et al. 2001). A control first strand synthesis reaction was performed without reverse transcriptase to eliminate second strand amplification products that would be non-specific, i.e. generated from residual DNA contamination in the RNA samples. ClonTech PCR Advantage II polymerase mix was used for cDNA second-strand synthesis, using the following thermo cycling parameters: initial denaturation at 95 °C for 10 min followed by 40 cycles of 95 °C for 15 s and 56 °C for 1 min. Water was substituted in place of cDNA template as a negative control for second strand synthesis.
qPCR relative quantification
A single transposable element can span more than 1,000 base pairs and carry multiple genes that enable its excision and proliferation (Pritham 2009). PCR-based estimation of TE copy number can be challenging because amplification efficiency decreases as the template length increases (Huber et al. 2009). Thus, we designed primers corresponding to 100–150 bases of specific TE genes (Table 2; DeMarco et al. 2004; DeMarco et al. 2006; Feschotte 2004). All primers were designed to have optimum binding at 56 °C, so that minimum effort was required in primer optimizing (IDT® Oligo Analyzer). Six representative of the various categories of TEs in S. mansoni genome (Table 2) were selected for copy number analysis; these include four class I elements (Saci-1, Saci-2, Saci-3, Perere-1), and two class II elements (Merlin-sm1 and SmTRC1). All of these elements are known to be transcriptionally active (DeMarco et al. 2004; DeMarco et al. 2006; Feschotte 2004). To determine if the use of different genes has a significant effect on copy number estimation, two primers sets were designed for Saci-3, targeting the RNaseH gene and Protease gene (Table 2). Primer/target selection was based on primers having the same amplification efficiency as our internal control gene, GAPDH (see qPCR relative quantification method below).
To determine the copy number of TE genes, qPCR relative quantification was performed using StepOnePlus™ Real Time PCR system (Applied Biosystems) and Power SYBR Green PCR master mix, according to the manufacturer’s instructions (Applied Biosystems). GAPDH (Glyceraldehyde Phosphate DeHydrogenase) was used as the internal control gene (separate primers for DNA and cDNA; Messerli et al. 2006). At the beginning of each assay, five serial dilutions of the template (DNA and cDNA separately) were used in the real-time PCRs on both target and reference genes to assess efficiency and to confirm that the amplification efficiencies of both genes were equal (Schmittgen and Livak 2008). qPCR mixtures consisted of 1 × SYBR Green, 3.6 μM of each primer and 1 ng of template. The thermal profile for all genes was as follows; 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 56 °C for 1 min. A melting curve analysis was conducted from 50 to 90 °C with 0.5 °C increases per cycle for a total of 80 cycles to insure there was no mis-annealing or contaminated cDNA in the sample. Each reaction was performed in three replicates and alongside a no-template negative control to rule out contamination (see Doyle et al. 2011). Samples with Ct values higher than the negative control and multiple Tm peaks (indicative of contamination) were removed from the analysis.
Statistical analysis
The fold differences of TE copy number in the genome and the transcriptome were calculated by the comparative Ct method, with the formula 2−ΔΔCt (Schmittgen and Livak 2008).
Subsequently, the fold differences of each TE were compared between the old world and new world strains (separately for adults and cercariae). TE copy number between the field and laboratory strains were also compared to identify possible TE dynamics associated with the demographic history of the population (Fold difference = 2−ΔΔCt = 2−[(Ct TE gene, Laboratory strain−Ct GAPDH, Laboratory strain)−(Ct TE gene, Field strain−Ct GAPDH, Field strain)]). The significance was determined using non-parametric Mann Whitney U test (equal variance) and Kolmogorov–Smirnov test (unequal variance). Bonferroni corrections were used to adjust for multiple comparisons (Bland and Altman 1995).
We expected more abundant and/or transcriptionally active TEs to show a greater difference in copy number between old world and new world parasites compared to rare/inactive TEs. Pearson correlation was used to quantify the relationship between TE abundance and the difference in copy number between old world and new world parasites (i.e. estimated copy number of the TE in the genome x the average difference in copy number between the new world and old world parasites). TE abundance and activity data were adapted from DeMarco et al. (2004). All statistics were calculated using R software release 3.1.1 (www.r-project.org).
Results
TE copy number in genomic DNA
We used an array of old and new world laboratory and field strains of the parasite S. mansoni to quantify TEs. As some field strains were available only as adults or cercariae, we conducted separate comparisons for each life cycle stage. The genomes of our new world parasites (both adults and cercariae) carried more copies of both class I and class II TEs than the old world parasites for both lab and field strains (Figs. 2, 3). This difference was significant in the adults (Strains NMRI, PR-1, LE, AMH, EGY-1; Kolmogorov–Smirnov test Bonferroni corrected p < 0.05; Supplementary Table S1), yet mostly non-significant in the cercariae (Strains NMRI, PR-1, LE, EGY-1, KY; five out of the seven comparisons non-significant; Supplementary Table S1).
Class I TE copy number difference in genomic DNA of old world and new world S. mansoni cercariae (male cercariae, n = 20, field and laboratory strains NMRI, PR1, LE, EGY1, KY) and adults (adult worm DNA, n = 30 per strain, field and laboratory strains NMRI, PR1, LE, AMH, EGY1). The x axis indicates the strains while the y axis represents the fold difference in TE copy number compared to the internal control GAPDH. The level of significance is indicated by asterisks (bonferroni corrected p < 0.05: *; p < 0.005: **; p < 0.0005: ***). The error bars represent standard errors. The difference in copy number is significant in adults. In cercariae, only Saci-1 and Saci-2 TE genes are significantly different between new world and old world strains
Class II TE copy number difference in genomic DNA of old world and new world S. mansoni strains (Cercariae DNA n = 20 per strain; Adult worm DNA n = 30 per strain; comparison includes both field and laboratory strains, as in Fig. 1). The x-axis indicates the strains whereas the y-axis represents the fold difference in TE copy number compared to the internal control GAPDH. The level of significance is indicated by asterisks (bonferroni corrected p < 0.05: *; p < 0.005: **; p < 0.0005: ***). The error bars represent standard errors. The difference in class II TE copy number is significant between adult samples of old world and new world strains
When only laboratory strains (NMRI, PR-1, LE, EGY-1) were compared, new world adults still had significantly more copies of class I and class II TEs than the old world adults (Kolmogorov–Smirnov test Bonferroni corrected p < 0.05; Supplementary Figure S1; Supplementary Table S2). In cercariae of the laboratory strains, the TE copy number difference was non-significant, but the new world strains still had more copies than the old world strains (Supplementary Figure S1; Supplementary Table S2).
Overall, class I elements were more abundant in the genomes of new world strains than were class II elements (Figs. 2, 3). The correlation between the estimated TE abundance and the difference in copy number between old world and new world adult parasites was moderate (correlation coefficient = 0.44; Supplementary Figure S2). However, we observed a strong correlation (correlation coefficient = 0.80; Supplementary Figure S3) between TE activity (estimated copy number of the TE in the genome*estimated transcriptional activity of the TE) and the average difference in copy number between the new world and old world parasites. Saci-1, a retrotransposon present in 70–700 genomic copies, has the highest transcriptional activity in adults and shows the highest copy number difference between old world and new world parasites (Supplementary Figure S3; DeMarco et al. 2004). Perere, an abundant retrotransposon with relatively low transcriptional activity shows the second largest difference between the new world and the old world adult parasites. SmTRC1, the TE with the lowest copy number exhibits the lowest copy number difference between the strains (Supplementary Figure S3; DeMarco et al. 2004). We identified no significant correlation between genomic TE abundance and difference in copy number between old world and new world cercariae (data not shown).
TE expression
We used cDNA synthesized from laboratory strains to calculate the differential expression of TEs at the transcriptome level. Unlike the genome, the transcriptome data revealed no consistent significant difference between the old and new world strains (Supplementary Table S3). Only class II Merlin-sm1 TE had a significantly higher expression level in the new world samples (Kolmogorov–Smirnov test Bonferroni corrected p < 0.05; Supplementary Table S3).
Variation in copy number estimates between genes
We estimated the copy number of Saci-3 using two different TE genes (RNaseH and Protease) to determine whether the amplicon has a significant effect on TE copy number estimation. Both RNaseH and Protease data indicate greater copy numbers of Saci-3 in new world strains (Fig. 2e–h), and our calculations indicate that estimates are not significantly affected by the TE gene assayed (\({\text{Fold difference}} = 2^{{ - \Delta \Delta {\text{Ct}} }} = 2^{{ - [\left( {{\text{Ct}}Saci{\text -}3,{\text{ RNaseH{-}Ct GAPDH}}} \right) - ({\text{Ct}}Saci{\text -}3,{\text{ protease{-}Ct GAPDH}})]}}\); Kolmogorov–Smirnov test p > 0.05).
TE copy number in laboratory versus field strains
Laboratory and field parasites are subjected to differential evolutionary pressures and therefore could exhibit differences in their genetic makeup (Fletcher et al. 1981; Stohler et al. 2004). Therefore, we calculated the TE copy number differences between laboratory and field strains in two different ways. First, we compared TE copy number between laboratory and field strains of geographically proximate sampling sites (i.e. Brazilian field strain vs. Brazilian lab strain and Egyptian lab strain vs. Kenyan field strain) to reduce the effects of differential evolutionary history while identifying such disparities. Second, we compared the pooled new world adults of laboratory strains (NMRI, PR1 and LE) against the new world field strain (AMH) to identify significant differences in TE copy number, despite the geographical distance. Our analysis indicates that the TE copy numbers are different between well-established laboratory strains and field collected parasites in both old and new world samples (Table 3). The differences were significant in all adult worm comparisons, but only marginally significant in a single cercarial comparison (Table 3). Interestingly, the direction of change was the same in all the comparisons, except in Perere and SmTRC1 of cercariae EGY-1 versus KY. Specifically, both adult and cercarial field samples seem to carry more copies of Saci-1 (RNase H), Saci-2 (RNase H), Saci-3 (RNase H), Saci-3 (Protease) and Merlin-sm1 (Transposase). The comparisons were significant in adults (Table 3). In both adult worm comparisons, four out of seven significant comparisons indicated more TE copies in the field strains. Despite these differences, both field and laboratory samples showed the same overall trend (i.e. new world parasites have more TE copies), when they were analyzed separately for old world vs. new world (Figs. 2, 3; Supplementary Figure S1; Supplementary Tables S1–S2).
Discussion
Schistosoma mansoni is one of the most notorious human parasites due to its pathogenicity and widespread distribution (Brunette 2014). Phylogenetic analyses suggest that S. mansoni invaded the new world during the trans-Atlantic slave trade (Morgan et al. 2005; Files 1951; Fletcher et al. 1981; Desprès et al. 1993) and quickly became established (Dejong et al. 2001). This suggests that S. mansoni must have adapted rapidly to the new world, where it no doubt faced novel intermediate and definitive hosts (Dejong et al. 2001, 2003), different environmental/climatic regimes, and dissimilar suites of pathogens, predators, and competitors. Beyond such ecological factors, genomic factors such as TE proliferation might have helped the parasite to adapt to new habitats (Venancio et al. 2010). We used qPCR quantification to test the idea that TE repertoires differ between parasites from established versus new habitats, by calculating the copy number of TEs in new and old world strains of S. mansoni relative to a control gene (GAPDH). Our results indicate that new world strains of S. mansoni do indeed carry more TE copies than old world strains. The trend is consistent in both cercarial and adult samples, albeit only significant in adults.
Both class I and class II elements are more abundant in the genomes of new world strains than in old world strains. Class I TEs, which copy and paste via an RNA intermediate, create new copies of themselves and thus the population of elements in a genome can grow rapidly. As a consequence, large genomes often consist primarily of class I retrotransposons (Venancio et al. 2010; Kidwell 2002). Conversely, class II elements increase their copy numbers through indirect mechanisms that rely on the host machinery such as DNA replication and double-strand break repair (Engels et al. 1990; Ros and Kunze 2001). Although not extensively as class I, class II DNA transposons can also proliferate due to stress (Naito et al. 2009; Feschotte et al. 2003).
We were surprised to observe a disparity in TE copy number between adult and cercarial life history stages (Figs. 2, 3). One possible explanation for this difference is that mitotic chromosomal recombination is often observed in schistosome cercariae (Bayne and Grevelding 2003; Semyenova et al. 2007), despite their asexual proliferation within the snail host (Fig. 1). Intra- and inter-chromosomal recombination events result in cercariae having differential numbers of repetitive elements (Rinaldi et al. 2012; Bayne and Grevelding 2003). Mitotic recombination can produce populations that are more diverse within hosts than among hosts (Korsunenko et al. 2012). TEs can facilitate ectopic recombination events, and could be associated with increased TE expression in cercariae (DeMarco et al. 2004). Another possible explanation for the disparity in TE copy number between life history stages is the use of parasites from different generations, as parasites with complex life cycles undergo several bottlenecks that can produce differences in genetic and genomic structure (Stohler et al. 2004).
Despite the differences in genomic copy number, there was no significant difference in the expression of most TEs between new world and old world strains. There are at least three possible explanations for this: pooling, bias, and TE repression. Pooling is often carried out in transcriptome analysis when insufficient RNA is available from individuals or individual tissues (Marinov et al. 2014). However, pooling can reduce the likelihood of identifying true biological variation in TE gene expression (Jolly et al. 2005) and thus cannot be discounted. However, there are also biological reasons that might explain the discordance in copy number variation between the genome and transcriptome. One of these is bias, whereby truncated TE copies are much more likely to become fixed in populations than full length copies—yet not necessarily more likely to be more highly expressed—because of purifying selection against active full length copies (Tollis and Boissinot 2013).
Finally, the lack of expression differences could correspond to reactivation of TE repression systems sometime after new world colonization. We favor this idea because our data are consistent with the emerging body of evidence summarized here. Elevated transposition rates cannot be maintained for long without deleterious effects caused by TE insertions (Zeh et al. 2009). In schistosomes, hyper-methylation of TEs has been identified as a mechanism for silencing TE transcription (Geyer et al. 2011). Furthermore, schistosomes utilize an RNAi pathway to silence the expression of genes (Batista and Marques 2011; Gomes et al. 2009; Chen et al. 2010). Hao et al. (2010) identified and characterized endogenous siRNAs which were primarily derived from S. japonicum TEs. Several genes encoding core and accessory proteins that function in RNAi pathways are conserved in the genome of schistosomes and gene silencing induced by introduction of exogenous dsRNA can be readily induced (Krautz-Peterson et al. 2007). This suggests that the TEs in schistosome genomes are now under tight control to prevent TE proliferation.
Repression is an arms race that is not always effective. Consider Merlin-sm1, a short class II element with an estimated 50–500 copies in the genome (Feschotte 2004) and the only expressed TE with significantly higher copy number in the new world. This suggests that Merlin-sm1 escaped TE repression systems, continuing to actively transcribe long after the new world invasion of S. mansoni. The patchy phylogenetic distribution of Merlin-like sequences suggests the possibility that it might have spread across distantly related species (including schistosomes) through repeated horizontal transfer (Feschotte 2004).
TE proliferations are susceptible to environmental changes (Zeh et al. 2009). The link between stress and TE activation is not always straightforward, but emerging patterns point towards underlying processes (Casacuberta and Gonzalez 2013). TE activation is often a consequence of the relaxation of epigenetic control induced by environmental changes (Slotkin and Martienssen 2007; Zeh et al. 2009; Rebollo et al. 2010). According to the “epi-transposon hypothesis”, epigenetic regulatory mechanisms (e.g., RNA interference, DNA methylation, and histone modifications) help maintain genome stability by suppressing TE mobilization. However, physiological stress, as induced by climate change or the invasion of new habitats, disrupts epigenetic regulation and unleashes TEs. Such TE proliferations can also induce the expression of adjacent genes when TEs act as transcription enhancers. For instance, activation of mPing elements in rice, can upregulate the expression of nearby genes in a stress inducible manner (Naito et al. 2009). Conversely some TEs, such as TF1 in Schizosaccharomyces pombe, specifically integrate near stress-responsive genes and are activated during stress (Bowen et al. 2003).
Other processes could also lead to increased proliferation of TEs. According to the “founder event hypothesis” TE proliferation occurs due to founder effects associated with genetic drift (García Guerreiro et al. 2008; García Guerreiro and Fontdevila 2007; Vieira and Biémont 2004). The differential fixation of TE copies in colonized versus native populations of Drosophila subobscura (García Guerreiro et al. 2008) and Arabidopsis lyrata (Lockton et al. 2008) strongly suggests that demographic history can influence the fate of TEs in the genome. Such founder effects are evidenced by low nucleotide diversity and have been identified in the new world strains of S. mansoni (Morgan et al. 2005). TE dynamics are governed by transposition, selection, and drift. Purifying selection purges deleterious copies of TEs in large populations, but selection is less effective at purging TEs in small populations where genetic drift predominates (Lynch and Conery 2003). Therefore, deleterious TEs are expected to proliferate most rapidly in small populations governed by genetic drift. This scenario is consistent with the evolutionarily recent establishment of S. mansoni in the new world, where colonizers were no doubt subject to bottlenecks and founder effects. Likewise, laboratory strains are notoriously affected by inbreeding and founder events as they are often established from one or few individuals (Fletcher et al. 1981). In addition, they are more susceptible to population bottlenecks and differential selection pressures (i.e. due to the maintenance in the mouse host) than field isolates (Blank et al. 2010; Stohler et al. 2004). Our results indicate differential TE profiles in laboratory and field strains, possibly due to their different demographic histories (Table 3).
Once unleashed due to environmental stressors, TE proliferations can inadvertently foster genetic innovations required for rapid adaptation (Zeh et al. 2009). Indeed, transposable elements have the ability to generate mutations through the process of transposition, which can create ectopic rearrangements, inversions, translocations and gene duplications (Casacuberta and Gonzalez 2013). Mutations are the ultimate source of all genetic variation and serve as the raw material for evolutionary adaptations. TE-induced mutations are known to confer adaptive advantages to host genomes (Schlenke and Begun 2004; Zeh et al. 2009). For instance in Drosophila simulans, the Doc element within a P450 gene provides increased resistance to insecticides (Schlenke and Begun 2004). Following this logic, TE proliferations might have assisted colonizers such as S. mansoni in the invasion of new habitats (and new worlds).
Conclusions
Dynamic environmental conditions pose a major challenge to the survival of organisms and positive selection acts upon those genetic variants that foster organismal adaptation (Visser 2008). TEs act as “mutator loci” that can give rise to new innovations that serve as a basis for adaptation. We report one of the first studies of TE dynamics in parasites. We used genes from multiple TEs and two life cycle stages to substantiate our results. Using qPCR, we demonstrated that new world strains of S. mansoni carry more copies of TEs in their genomes than old world strains. This finding is consistent with a burst of stress-induced TE activity concomitant with the invasion of the new world by S. mansoni. Such amplifications might have positively influenced the parasite’s ability to invade new habitats, but if so the signature of TE bursts was absent in transcriptomes, perhaps due to the reactivation of TE repression systems and/or indicating the presence of inactive TE copies. Although new world strains possess more TE copies in both laboratory strain and field strain comparisons, we found that field and laboratory samples also differ in their TE repertoires. Our results demonstrate the value of studying TE dynamics at multiple hierarchical levels in order to better understand TE and genome evolution.
References
Batista TM, Marques JT (2011) RNAi pathways in parasitic protists and worms. J Proteomics 74:1504–1514
Bayne CJ, Grevelding CG (2003) Cloning of Schistosoma mansoni sporocysts in vitro and detection of genetic heterogeneity among individuals within clones. J Parasitol 89:1056–1060
Berriman M, Haas BJ, LoVerde PJ, Wilson RA, Dillon GP, Cerqueira GC, Mashiyama ST, Al-Lazikani B, Andrade LF, Ashton PD, Aslett MA, Bartholomeu DC, Blandin G, Caffrey CR, Coghlan A, Coulson R, Day TA, Delcher A, DeMarco R, Djikeng A, Eyre T, Gamble JA, Ghedin E, Gu Y, Hertz-Fowler C, Hirai H, Hirai Y, Houston R, Ivens A, Johnston DA, Lacerda D, Macedo CD, McVeigh P, Ning Z, Oliveira G, Overington JP, Parkhill J, Pertea M, Pierce RJ, Protasio AV, Quail MA, Rajandream M, Rogers J, Sajid M, Salzberg SL, Stanke M, Tivey AR, White O, Williams DL, Wortman J, Wu W, Zamanian M, Zerlotini A, Fraser-Liggett CM, Barrell BG, El-Sayed NM (2009) The genome of the blood fluke Schistosoma mansoni. Nature 460:352–358
Bland JM, Altman DG (1995) Multiple significance tests: the Bonferroni method. BMJ 310:170
Blank WA, Test MR, Liu SF, Lewis FA, Blanton RE (2010) Long-term genetic stability and population dynamics of laboratory strains of Schistosoma mansoni. J Parasitol 96:900–907
Blumenstiel JP (2011) Evolutionary dynamics of transposable elements in a small RNA world. Trends Genet 27:23–31
Bowen NJ, Jordan IK, Epstein JA, Wood V, Levin HL (2003) Retrotransposons and their recognition of pol II promoters: a comprehensive survey of the transposable elements from the complete genome sequence of Schizosaccharomyces pombe. Genome Res 13:1984–1997
Brindley PJ, Hotez PJ (2013) Break out: urogenital schistosomiasis and Schistosoma haematobium infection in the post-genomic era. PLoS Negl Trop Dis 7(3):e1961. doi:10.1371/journal.pntd.0001961
Brunette GW (2014) CDC health information for international travel 2014. Oxford University Press, Oxford
Capy P, Gasperi G, Biemont C, Bazin C (2000) Stress and transposable elements: co-evolution or useful parasites? Heredity 85:101–106
Casacuberta E, Gonzalez J (2013) The impact of transposable elements in environmental adaptation. Mol Ecol 22:1503–1517
Chakrani F, Capy P, David JR (1993) Developmental temperature and somatic excision rate of mariner transposable element in three natural populations of Drosophila simulans. Genet Sel Evol 25:121–132
Chen J, Yang Y, Guo S, Peng J, Liu Z, Li J, Lin J, Cheng G (2010) Molecular cloning and expression profiles of Argonaute proteins in Schistosoma japonicum. Parasitol Res 107:889–899
Chitsulo L, Engels D, Montresor A, Savioli L (2000) The global status of schistosomiasis and its control. Acta Trop 77:41–51
Crompton DWT (1999) How much human helminthiasis is there in the world? J Parasitol 85:397–403
DeJong RJ, Morgan JAT, Paraense WL, Pointier JP, Amarista M, Ayeh-Kumi PF, Babiker A, Barbosa CS, Brémond P, Pedro Canese A, de Souza CP, Dominguez C, File S, Gutierrez A, Incani RN, Kawano T, Kazibwe F, Kpikpi J, Lwambo NJ, Mimpfoundi R, Njiokou F, Noël Poda J, Sene M, Velásquez LE, Yong M, Adema CM, Hofkin BV, Mkoji GM, Loker ES (2001) Evolutionary relationships and biogeography of Biomphalaria (Gastropoda: Planorbidae) with implications regarding its role as host of the human bloodfluke, Schistosoma mansoni. Mol Biol Evol 18:2225–2239
DeJong RJ, Morgan JAT, Wilson WD, Al-Jaser MH, Appleton CC, Coulibaly G, D’Andrea PS, Doenhoff MJ, Haas W, Idris MA, Magalhães LA, Moné H, Mouahid G, Mubila L, Pointier JP, Webster JP, Zanotti-Magalhães EM, Paraense WL, Mkoji GM, Loker ES (2003) Phylogeography of Biomphalaria glabrata and B. pfeifferi, important intermediate hosts of Schistosoma mansoni in the New and Old World tropics. Mol Ecol 12:3041–3056
DeMarco R, Machado AA, Soares MB, Gargioni C, Kawano T, Rodrigues V, Madeira AMBN, Wilson RA, Menck CFM, Setubal JC, Dias-Neto E, Leite LCC, Verjovski-Almeida S (2004) Saci-1, -2, and -3 and Perere, four novel retrotransposons with high transcriptional activities from the human parasite Schistosoma mansoni. J Virol 78:967–2978
DeMarco R, Venancio TM, Verjovski-Almeida S (2006) SmTRC1, a novel Schistosoma mansoni DNA transposon, discloses new families of animal and fungi transposons belonging to the CACTA superfamily. BMC Evol Biol 6:89–102
Desprès L, Imbert-Establet D, Monnerot M (1993) Molecular characterization of mitochondrial-DNA provides evidence for the recent introduction of Schistosoma mansoni into America. Mol Biochem Parasitol 60:221–230
Doyle JM, McCormick CR, DeWoody JA (2011) The quantification of spermatozoa by real-time quantitative PCR, spectrophotometry, and spermatophore cap size. Mol Ecol Res 11:101–106
Engels WR, Johnson-Schlitz DM, Eggleston WB, Sved J (1990) High-frequency P element loss in Drosophila is homolog dependent. Cell 62:515–525
Feschotte C (2004) Merlin, a new superfamily of DNA transposons identified in diverse animal genomes and related to bacterial IS1016 insertion sequences. Mol Biol Evol 21:1769–1780
Feschotte C, Swamy L, Wessler SR (2003) Genome-wide analysis of mariner-like transposable elements in rice reveals complex relationships with Stowaway MITEs. Genetics 163:747–758
Files VS (1951) A study of the vector-parasite relationships in Schistosoma mansoni. Parasitology 41:264–269
Fletcher M, Loverde PT, Woodruff DS (1981) Genetic variation in Schistosoma mansoni enzyme polymorphisms in poulations from Africa, southwest Asia, South America, and the West Indies. Am J Trop Med Hyg 30:406–421
García Guerreiro MP, Fontdevila A (2007) The evolutionary history of Drosophila buzzatii. XXXVI. Molecular structural analysis of Osvaldo retrotransposon insertions in colonizing populations unveils drift effects in founder events. Genetics 175:301–310
García Guerreiro MP, Chávez-Sandoval BE, Balanyà J, Serra L, Fontdevila A (2008) Distribution of the transposable elements bilbo and gypsy in original and colonizing populations of Drosophila subobscura. BMC Evol Biol 8:234. doi:10.1186/1471-2148-8-234
Garza D, Medhora M, Koga A, Hartl DL (1991) Introduction of the transposable element mariner into the germline of Drosophila melanogaster. Genetics 128:303–310
Geyer KK, Rodríguez López CM, Chalmers IW, Munshi SE, Truscott M, Heald J, Wilkinson MJ, Hoffmann KF (2011) Cytosine methylation regulates oviposition in the pathogenic blood fluke Schistosoma mansoni. Nat Commun 2:424. doi:10.1038/ncomms1433
Gomes MS, Cabral FJ, Jannotti-Passos LK, Carvalho O, Rodrigues V, Baba EH, Renata GS (2009) Preliminary analysis of miRNA pathway in Schistosoma mansoni. Parasitol Int 58:61–68
Grevelding CG (1995) The female-specific W1 sequence of the Puerto Rican strain of Schistosoma mansoni. Mol Biochem Parasitol 71:269–272
Hao L, Cai P, Jiang N, Wang H, Chen Q (2010) Identification and characterization of microRNAs and endogenous siRNAs in Schistosoma japonicum. BMC Genom 11:55. doi:10.1186/1471-2164-11-55
Hua-Van A, Le Rouzic A, Boutin TS, Filee J, Capy P (2011) The struggle for life of the genome’s selfish architects. Biol Direct 6:19. doi:10.1186/1745-6150-6-19
Huber JA, Morrison HG, Huse SM, Neal PR, Sogin ML, Welch DBM (2009) Effect of PCR amplicon size on assessments of clone library microbial diversity and community structure. Environ Microbiol 11:1292–1302
Jolly RA, Goldstein KM, Wei T, Gao H, Chen P, Huang S, Colet J, Ryan TP, Thomas CE, Estrem ST (2005) Pooling samples within microarray studies: a comparative analysis of rat liver transcription response to prototypical toxicants. Physiol Genomics 22:346–355
Kidwell M (2002) Transposable elements and the evolution of genome size in eukaryotes. Genetica 115:49–63
Korsunenko A, Chrisanfova G, Lopatkin A, Beer SA, Voronin M, Ryskov AP, Semyenova SK (2012) Genetic differentiation of cercariae infrapopulations of the avian schistosome Trichobilharzia szidati based on RAPD markers and mitochondrial cox1 gene. Parasitol Res 110:833–841
Krautz-Peterson G, Radwanska M, Ndegwa D, Shoemaker CB, Skelly PJ (2007) Optimizing gene suppression in schistosomes using RNA interference. Mol Biochem Parasitol 153:194–202
Lambertucci JR, Serufo JC, Gerspacher-Lara R, Rayes AA, Teixeira R, Nobre V, Antunes CM (2000) Schistosoma mansoni: assessment of morbidity before and after control. Acta Trop 77:101–109
Lewis FA, Stirewalt MA, Souza CP, Gazzinelli G (1986) Large-scale laboratory maintenance of Schistosoma mansoni, with observations on three schistosome/snail host combinations. J Parasitol 72:813–829
Lockton S, Ross-Ibarra J, Gaut BS (2008) Demography and weak selection drive patterns of transposable element diversity in natural populations of Arabidopsis lyrata. Proc Natl Acad Sci USA 105:13965–13970
Lockyer AE, Olson PD, Ostergaard P, Rollins D, Johnston DA, Attwood SW, Horak P, Snyder SD, Le TH, Agatsuma T, McManus DP, Carmichael AC, Naem S, Littlewood DT (2003) The phylogeny of the Schistosomatidae based on three genes with emphasis on the interrelationships of Schistosoma Weinland, 1858. Parasitology 126:203–224
Lynch M, Conery JS (2003) The origins of genome complexity. Science 302:1401–1404
Marinov GK, Williams BA, McCue K, Schroth GP, Gertz J, Myers RM, Wold BJ (2014) From single-cell to cell-pool transcriptomes: stochasticity in gene expression and RNA splicing. Genome Res 24:496–510
Messerli SM, Kasinathan RS, Morgan W, Spranger S, Greenberg RM (2006) Schistosoma mansoni P-glycoprotein levels increase in response to praziquantel exposure and correlate with reduced praziquantel susceptibility. Mol Biochem Parasitol 167:54–59
Morgan JA, Dejong RJ, Adeoye GO, Ansa ED, Barbosa CS, Bremond P, Cesari IM, Charbonnel N, Correa LR, Coulibaly G, D’Andrea PS, De Souza CP, Doenhoff MJ, File S, Idris MA, Incani RN, Jarne P, Karanja DM, Kazibwe F, Kpikpi J, Lwambo NJ, Mabaye A, Magalhaes LA, Makundi A, Mone H, Mouahid G, Muchemi GM, Mungai BN, Sene M, Southgate V, Tchuente LA, Theron A, Yousif F, Zanotti-Magalhaes EM, Mkoji GM, Loker ES (2005) Origin and diversification of the human parasite Schistosoma mansoni. Mol Ecol 14:3889–3902
Naito K, Zhang F, Tsukiyama T, Saito H, Hancock CN, Richardson AO, Okumoto Y, Tanisaka T, Wessler SR (2009) Unexpected consequences of a sudden and massive transposon amplification on rice gene expression. Nature 461:1130–1134
Nuzhdin SV (1999) Sure facts, speculations, and open questions about the evolution of transposable element copy number. Genetica 107:129–137
Oliver KR, Greene WK (2009) Transposable elements: powerful facilitators of evolution. BioEssays 31:703–714
Pritham EJ (2009) Transposable elements and factors influencing their success in eukaryotes. J Hered 100:648–655
Protasio AV, Tsai IJ, Babbage A, Nichol S, Hunt M, Aslett MA, De Silva N, Velarde GS, Anderson TJC, Clark RC, Davidson C, Gary PG, Dillon S, Holroyd NE, LoVerde PT, Lloyd C, McQuillan J, Oliveira G, Otto TD, Parker-Manuel SJ, Quail MA, Wilson RA, Zerlotini A, Dunne DW, Berriman M (2012) A systematically improved high quality genome and transcriptome of the human blood fluke Schistosoma mansoni. PLoS Negl Trop Dis 6:e1455. doi:10.1371/journal.pntd.0001455
Rebollo R, Horard B, Hubert B, Vieira C (2010) Jumping genes and epigenetics: towards new species. Gene 454:1–7
Rinaldi G, Eckert SE, Tsai IJ, Suttiprapa S, Kines KJ et al (2012) Germline transgenesis and insertional mutagenesis in Schistosoma mansoni mediated by murine leukemia virus. PLoS Pathog 8(7):e1002820. doi:10.1371/journal.ppat.1002820
Ros F, Kunze R (2001) Regulation of activator/dissociation transposition by replication and DNA methylation. Genetics 157:1723–1733
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York
Schlenke TA, Begun DJ (2004) Strong selective sweep associated with a transposon insertion in Drosophila simulans. Proc Natl Acad Sci USA 101:1626–1631
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108
Semyenova SK, Khrisanfova GG, Korsunenko AV, Voronin MV, Beer SV, Vodyanitskaya SV, Serbina EA, Yurlova NI, Ryskov AP (2007) Multilocus variation in cercariae, parthenogenetic progeny of different species of the class Trematoda. Dokl Biol Sci 414:235–238
Slotkin RK, Martienssen R (2007) Transposable elements and the epigenetic regulation of the genome. Nat Rev Genet 8:272–285
Snyder SD, Loker ES (2000) Evolutionary relationships among the Schistosomatidae (Platyhelminthes: Digenea) and an Asian origin for Schistosoma. J Parasitol 86:283–288
Stohler RA, Curtis J, Minchella DJ (2004) A comparison of microsatellite polymorphism and heterozygosity among field and laboratory populations of Schistosoma mansoni. Int J Parasitol 34:595–601
Thiele EA, Sorenson RE, Gazzinelli A, Minchella DJ (2008) Genetic diversity and population structuring of Schistosoma mansoni in a Brazilian village. Int J Parasitol 38:389–399
Thomas M, Macias C, Alonso C, Lopez MC (2010) The Biology and Evolution of transposable elements in parasites. Trends Parasitol 26:350–362
Tollis M, Boissinot S (2013) Biology Lizards and LINEs: selection and demography affect the fate of L1 retrotransposons in the genome of the green anole (Anolis carolinensis). Genome Biol Evol 5:1754–1768
Venancio TM, Wilson RA, Verjovski-Almeida S, DeMarco R (2010) Burst of transposition from non-long terminal repeat retrotransposon families of the RTE clade in Schistosoma mansoni. Int J Parasitol 40:743–749
Vieira C, Biémont C (2004) Transposable element dynamics in two sibling species: Drosophila melanogaster and Drosophila simulans. Genetica 120:115–123
Visser ME (2008) Keeping up with a warming world; assessing the rate of adaptation to climate change. Proc Biol Sci 275:649–659
Young ND, Jex AR, Li B, Liu S, Yang L, Xiong Z, Li Y, Cantacessi C, Hall RS, Xu X, Chen F, Wu X, Zerlotini A, Oliveira G, Hofmann A, Zhang G, Fang X, Kang Y, Campbell BE, Loukas A, Ranganathan S, Rollins D, Rinaldi G, Brindley P, Yang H, Yang J, Wang J, Gasser RB (2012) Whole-genome sequence of Schistosoma haematobium. Nat Genet 44:221–225
Zeh DW, Zeh JA, Ishida Y (2009) Transposable elements and an epigenetic basis for punctuated equilibria. BioEssays 31:715–726
Zhou Y, Zheng H, Chen Y, Zhang L, Wang K, Guo J, Huang Z, Zhang B, Huang W, Jin K, Dou T, Hasegawa M, Wang L, Zhang Y, Zhou J, Tao L, Cao Z, Li Y, Vinar T, Brejova B, Brown D, Li M, Miller DJ, Blair D, Zhong Y, Chen Z, Liu F, Hu W, Wang Z, Zhang Q, Song H, Chen S, Xu X, Xu B, Ju C, Huang Y, Brindley PJ, McManus DP, Feng Z, Han Z, Lu G, Ren S, WangY GuW, Kang H, Chen J, Chen X, Chen S, Wang L, Yan J, Wang B, Lv X, Jin L, Wang B, Pu S, Zhang X, Zhang W, Hu Q, Zhu G, Wang J, Yu J, Wang J, Yang H, Ning Z, Beriman M, Wei C, Ruan Y, Zhao G, Wang S (2009) The Schistosoma japonicum genome reveals features of host–parasite interplay. Nature 460:345–351
Zhu YY, Machleder EM, Chenchik A, Li R, Siebert PD (2001) Reverse transcriptase template switching: a SMART™ approach for full-length cDNA library construction. Biotechniques 30:892–897
Acknowledgments
We sincerely acknowledge Dr. Phil LoVerde and Dr. Eric Loker for sharing the LE, EGY and Kenyan field isolates with us. We also thank Dr. Fred Lewis, Dr. Matt Tucker and the staff of the Biomedical Research Institute for providing the following reagents through BEI Resources, NIAID, NIH: (1) Schistosoma mansoni, Strain NMRI, Exposed Biomphalaria glabrata strain NMRI, NR-21962 and (2) Schistosoma mansoni, Strain PR-1, Exposed Biomphalaria glabrata strain M-line, NR-21961. We greatly appreciate the members of the Minchella and DeWoody labs for their comments on an earlier version of the manuscript. This project was funded by the Purdue University Department of Biological Sciences (DJM), Sigma Xi grants-in-aid program (BKW), and the University Faculty Scholar program (JAD).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wijayawardena, B.K., DeWoody, J.A. & Minchella, D.J. The genomic proliferation of transposable elements in colonizing populations: Schistosoma mansoni in the new world. Genetica 143, 287–298 (2015). https://doi.org/10.1007/s10709-015-9825-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-015-9825-6