Abstract
Transposable elements (TEs) account for nearly half (44 %) of the human genome. However, their overall activity has been steadily declining over the past 35–50 million years, so that <0.05 % of TEs are presumably still “alive” (potentially transposable) in human populations. All the active elements are retrotransposons, either autonomous (LINE-1 and possibly the endogenous retrovirus ERVK), or non-autonomous (Alu and SVA, whose transposition is dependent on the LINE-1 enzymatic machinery). Here we show that a lineage of the endogenous retrovirus ERVE was recently engaged in ectopic recombination events and may have at least one potentially fully functional representative, initially reported as a novel retrovirus isolated from blood cells of a Chinese patient with chronic myeloid leukemia, which bears signals of positive selection on its envelope region. Altogether, there is strong evidence that ERVE should be included in the short list of potentially active TEs, and we give clues on how to identify human specific insertions of this element that are likely to be segregating in some of our populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The vast majority (~90 %) of TE insertions in the human genome correspond to retrotransposons (Lander et al. 2001), whose proliferation involves reverse transcription of an RNA intermediate into a cDNA that integrates in the host cell DNA. A small part of retrotransposons (1.5 %) results from the endogenization of retroviruses that once infected the germ line of our ancestors, generally at least 50 million years ago (Mya) (Bannert and Kurth 2006; Lander et al. 2001). Upon integration into the host genome as a provirus, ERV insertions are thereafter vertically transmitted following Mendelian rules. Altogether, the human genome contains ~200,000 ERV insertions (Lander et al. 2001), distributed among 3 classes and at least 30 families (Bannert and Kurth 2006; Gifford and Tristem 2003). Most of these insertions are solitary long terminal repeats (LTRs), produced by recombination between the flanking LTRs of a full-length provirus (Lander et al. 2001). The rest of them, with very few exceptions (de Parseval and Heidmann 2005), correspond to “fossils” that many Mya lost their capacity to transpose (Bannert and Kurth 2006; Feschotte and Gilbert 2012), although they may occasionally be transcribed (Seifarth et al. 2005) and translated with relevant consequences for their hosts (Cordaux and Batzer 2009; Jern and Coffin 2008; Lee et al. 2008). Among these families, only ERVK (HML-2) is considered likely to be still active in humans, based on the finding of several human-specific copies (~7 % of all the insertions of this family), either fixed or still segregating in our species (Buzdin 2007; Mills et al. 2007; Jha et al. 2011; Shin et al. 2013). However, not a single active ERVK copy has been found so far, although its artificial reconstruction from existing members has been possible (Dewannieux et al. 2006; Lee and Bieniasz 2007).
ERVE is a C-type endogenous retrovirus, whose prototypic element (M10976) is a 8.8 kb defective provirus bounded by LTRs of 495 nt, with overall 40 % amino acid similarity with Moloney murine leukemia virus in the gag and pol regions (Repaske et al. 1985). Results from Southern blotting analyses (Repaske et al. 1983; Steele et al. 1984) and BLAST screening of human DNA databases, suggest that the human genome contains up to 50 full-length insertions of this family, mostly confined to CpG- and Alu-rich early replicating regions, preferentially near breakpoints and fragile sites (Taruscio et al. 2002). Perhaps due in part to this bias, ERVE insertions contribute to a disproportionately large fraction of all retroelement-derived promoter or enhancer sequences of cellular genes (Bannert and Kurth 2004). On the other hand, although this family has been repeatedly associated to several forms of cancer and autoimmune diseases (Dolei 2006; Ogasawara et al. 2000; Piotrowski et al. 2005; Prusty et al. 2008; Takahashi et al. 2008; Voisset et al. 2008), its precise role in the pathogenesis remains to be elucidated. In 2002 a complete genome of a “novel” retrovirus isolated from blood cells (genomic DNA) of a patient with chronic myeloid leukemia was submitted to GenBank (HCML-ARV, AF499232—R. Z. Xu and S. Zheng, direct submission; see also AY208746 for the corresponding retroviral RNA). It has all the features of a potentially active provirus, bounded by two LTRs and with apparently intact coding sequences of all retroviral genes: gag, encoding the internal structural protein of the retrovirus, with matrix (MA), capsid (CA), and nucleocapsid (NC) domains; pro, encoding a protease (PR); pol, encoding a polyprotein with reverse transcriptase (RT), ribonuclease H (RH), and integrase (IN) domains; and env, encoding the complex that interacts with cellular receptors, with surface (SU) and transmembrane (TM) domains (standardized nomenclature proposed by Leis et al. 1988). HCML-ARV was later identified as a member of the ERVE family (Prusty et al. 2008); interestingly, its complete sequence was not detected in the published human genome, although part of it was apparently localized on chromosome 2q37. The potential of HCML-ARV to reveal new aspects of the evolution of ERVs connected with their possible involvement in human pathogenesis, moved us to carry out a genome-wide analysis that unveiled several salient features of the retroelement, such as molecular signatures of recombination with relatively old proviral insertions at the human genome and of positive selection on the sequence encoding its envelope region.
Materials and methods
Data mining
ERVE sequences from the human genome were retrieved by means of an automated nucleotide and protein sequence homology combined iterative approach (Bartolome et al. 2009). The sequence of the internal region of a full potentially active ERVE element (AF499232) was used as query, and the human genome reference sequence (GRCh37/hg19) as subject. All the sequence hits that displayed ≥60 % nucleotide homology over at least 80 % of the length of the query were regarded as full-length insertions of the ERVE family. Nucleotide sequence alignments were initially obtained with MUSCLE (Edgar 2004) and subsequently corrected by hand with the aid of Bioedit (Hall 1999). ERVE copies bearing long deletions at particular retroviral genes were not used for the corresponding phylogenetic analyses; also, when two insertions appeared to be products of segmental duplication and not of independent transposition, only one of them was kept for further analyses (Table S1). Map positions of the different insertions were obtained with the BLAT alignment tool (Kent 2002) of UCSC Genome Bioinformatics, using genome assembly Feb. 2009 (GRCh37/hg19), and do not include the LTRs.
Phylogenetic analysis
Reconstructions of the phylogenetic relationships among ERVE insertions were carried out with MEGA (Tamura et al. 2007). For neighbor-joining, distances were computed under the maximum composite likelihood model, with the pairwise deletion option; the reliability of the inferred tree was evaluated using the bootstrap resampling technique (1,000 replicates). For maximum parsimony, searches were conducted by close neighbor interchange (level = 1), with initial tree by random addition (20 replications), using all positions; a composite tree was finally obtained using a majority-rule consensus of equally most parsimonious trees.
Evolutionary and statistical analysis
Evidence of positive selection in the phylogenetic tree of ERVE was assessed by maximum likelihood ratio tests (Yang 1998), using the Codeml program in the PAML package. The program produced log likelihood values and maximum likelihood estimates of parameters for each retroviral gene, under different hierarchical models of d N /d S heterogeneity among lineages. To test whether a given model fits the data significantly better than the preceding one in the hierarchy, a likelihood ratio test was carried out, consisting of comparing the corresponding 2Δl (twice the increment in log likelihood) with a χ2 distribution with df = df2 − df1, where df2 and df1 represent the number of free parameters of the models. To assess the potential confounding effect of biased gene conversion on the molecular signature of positive selection, we compared the distribution of strong (S) and weak (W) substitutions for the different genes of HCML-ARV, after their alignment with the reconstructed ancestral sequences of lineage VIIa, with the correspondent distribution obtained from a reference database of alignments of 10,238 human genes to their orthologues in chimpanzee and macaque (Berglund et al. 2009), using Fisher’s exact tests (1-tail).
Characterization of recombination events
Analysis of mosaicism was carried out using SimPlot (Lole et al. 1999). This method is based on the distribution of phylogenetic signals supporting alternative tree topologies among four taxa: the putative mosaic sequence, one representative of each of the two “parental” lineages, and a known outgroup. We used the neighbor-joining method based on the Kimura two-parameter distance model, with a transition/transversion bias of 2.0, after generating 1,000 bootstrap replicates. SimPlot calculates and graphically plots the phylogenetic identity (bootscanning) of the query sequence to the “parental” sequences, in a sliding window of 200 nucleotides that we programmed to move along the nucleotide alignment in steps of 20.
Protein structure prediction
Predicted structures of Env protein domains were obtained using the Protein Homology/analogY Recognition Engine (Kelley and Sternberg 2009). From the ranking of possible templates obtained with the engine, the first hit—the crystal structure of FLV receptor-binding domain from Feline leukemia virus (PDB: d1lcsa)—was chosen as the best template (20 % id; E-value, 9.2e−14; estimated precision, 100 %). Visualization was accomplished with FirstGlance in Jmol (http://molvis.sdsc.edu/fgij).
Human samples
Two sets of samples were used in this study. One consisted of 18 anonymous DNA samples from chronic myeloid leukemia patients of the Hematological Diseases Service of the Complexo Hospitalario Universitario de Santiago de Compostela (Spain). The second was the HGDP-CEPH Human Genome Diversity Panel, which includes DNA samples from 1,063 individuals representing 52 world populations (Cann et al. 2002), and is available from the Foundation Jean Dausset-CEPH in Paris.
PCR analysis and DNA sequencing
The nucleotide sequence of HCML-ARV (AF499232) and 49 other full length ERVE element insertions identified by us in the human genome were aligned with the aid of MUSCLE (Edgar 2004) and corrected by hand in Bioedit (v. 7.1.3.0) (Hall 1999). The resulting alignment was used to identify sequence motifs exclusive for HCML-ARV, which allowed us to design PCR primers specific for this insertion: ERVE-F, 5′AGC TTC ATC AAC CCA CAA GG 3′, beginning at nucleotide 3,632 (see alignment in Fig. S2); and ERVE-R, 5′ GCT GCT TTT CGA GCG TCA 3′, beginning at nucleotide 4,035. This pair of primers should amplify a 398 bp sequence corresponding to positions 4,591–4,988 in AF499232. Aliquots of every four samples of the HGDP-CEPH panel were combined in pooled samples (266 pools in total). PCR reactions were performed using FastStart High Fidelity PCR System (Roche Diagnostics) under manufacturer’s conditions. An initial denaturation step (95 °C for 3 min) was followed by 35 cycles (denaturation at 95 °C for 1′; annealing 68 °C for 1′; elongation at 72 °C for 1′), and a final elongation step at 72 °C for 1′. A PCR assay with a lower annealing temperature (59 °C), using an anonymous human DNA sample, amplified a fragment of an ERVE insertion different from AF499232 (it did not present any of its distinctive sequence motifs; data not shown). As a result of the PCR amplification process, the 5′ and 3′ ends of this amplicon are exact matches of the two primers. This amplicon was used to set positive PCR controls. PCRs were performed in 96-well plates, including 90 pooled samples, three positive, and three negative controls randomly placed in each plate.
URLs
- MUSCLE: :
- PAML: :
- SimPlot: :
- UCSC Genome Bioinformatics, Blat Search Human Genome Tool: :
- PHYRE: :
- Jmol project: :
Results and discussion
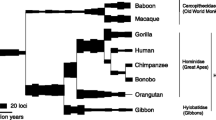
The few existing studies on the evolutionary history of ERVE (Taruscio et al. 2002; Yi and Kim 2006, 2007) reported no human-specific insertions, but exhaustive analyses of primate genome databases through efficient methods of retroviral insertion identification have not been conducted for this family yet. Using human genome build GRCh37/hg19, we obtained a set of 49 proviral nucleotide sequences (Table S1), whose phylogeny was reconstructed independently for each of several retroviral gene regions: gag, pro, rt (encoding RT), rh (encoding RH), in (encoding IN), and env. We also included in our analysis the sequence of HCML-ARV and an insertion from Pan troglodytes (P_022319), for reasons that will soon become apparent. Irrespectively of the gene or the method of reconstruction (neighbor-joining or maximum parsimony), ERVE copies can be classified into at least eight subfamilies (I through VIIb, according to rules described in López-Sánchez et al. 2005; Fig. 1; supplementary Fig. S1), grouped into two major clusters, deeply rooted in the phylogeny (I–II–III vs IV–V–VI–VIIa–VIIb). The second cluster contains previously characterized ERVE members (Taruscio et al. 2002; Yi and Kim 2007), whereas the first one contains sequences submitted to Repbase as HERV-E(a) (Smit 2008). Sequences of subfamily VIIb are part of a recently described giant chimeric composite transposon (DA Type II/III, Ji and Zhao 2008), that coamplified ERVH and ERVE variants (Lindeskog et al. 1998) most intensely in an interval that extends from 14 to 7 Mya (Li et al. 2009). Using average nucleotide distances within this subfamily as a rough yardstick, it can be estimated that the two major clusters of ERVE initiated their divergence not sooner than 58 Mya. HCML-ARV (abbreviated to HCML in Fig. 1) is included in subfamily VIIa. It should be borne in mind that AF499232 reports a genomic DNA with the characteristic LTRs of a proviral insertion, but it says nothing about the integration site, and not knowing whether there is a target site duplication or not, we can not be certain whether this copy resulted from a new insertion or from a chromosome rearrangement involving other proviral copies of the genome. However, the source of the sequence definitely indicates that it is endogenous, not a replication-competent exogenous virus. In addition, the relatively low identity (97.2 %) between its flanking LTRs clearly precludes the possibility of the copy being the result of a recent new insertion unique to one patient. Besides HCML, the subfamily VIIa includes a most peculiar sequence, H_022319, which contains the putative “part” of HCML-ARV that had been previously detected in the human genome (Prusty et al. 2008). Actually, H_022319 corresponds to an 8.1 kb defective provirus that inserted on chromosome 2 (2q37, map positions 232,265,423–232,273,554, including the 495nt LTRs, in human genome build 37.1) before the split of gibbons and great apes, 18–20 Mya. All orthologous copies in man, chimpanzee, gorilla, orangutan and gibbon share a diagnostic long deletion of most of the in region. The orthologous copy in P. troglodytes is on chromosome 2b (8.4 kb extending from 237,619,424 to 237,627,894 in genome assembly CGSC 2.1/panTro2), denoted as P_022319 in our phylogenetic trees, and shows several private differences and a few shared stop codons and small indels with H_022319 (see alignment in supplementary Fig. S2). The phylogenetic analysis of env (Fig. 1) places HCML-ARV as the closest relative of these two sequences. However, as shown in supplementary Fig. S1, there is a lack of congruence in the phylogenetic signal of the different retroviral regions on this respect. A plausible explanation for this finding is supplied in Fig. 2, which depicts the results of a bootscan test of recombination, showing that the human sequence H_022319 is most likely to be the product of ectopic recombination between the original ERVE, residing in 2q37, and a proviral insertion of HCML-ARV located elsewhere in the genome, similarly to what has been already reported for ERVK (Hughes and Coffin 2005). Three double crossovers would have left a small part of gag, nearly the whole pro, and the majority of env as the only remnants of the original ERVE insertion. According to the present day divergence between H_022319 and HCML-ARV (0.002, excluding sequence tracts from the original insertion), that recombination event dates back to approximately 0.4 Mya. What is more, the close similarity between the 5′ and 3′ flanking LTRs of H_022319 and the corresponding ones reported in AF499232 (four differences out of 987 nucleotide positions) strongly suggests that the recombination could have taken place precisely with this proviral insertion of HCML-ARV. The possibility that there is a primate ERVE sequence, which could be the source of HCML-ARV, seems to be extremely unlikely. Exhaustive searches of primate genomes using different regions from HCML-ARV as queries consistently produce the highest scores with either H022319 or the cluster formed by H022319 and P022319 (Fig. 2), with a single exception. The sequence comprising the end of RH and most of IN (4022–4693 in the alignment shown in Fig. S2), which is missing at H022319 and all its primate orthologs, only differs by two indels (otherwise, 100 % nucleotide identity) from H172358, the human locus of a proviral insertion on chromosome 17 (see also Genbank report AB062274) that we only share with chimpanzees. This strongly indicates that HCML-ARV originated in the human lineage, and that it was involved in ectopic recombination events with two other different ERVE insertions.
Phylogenetic relationships within the ERVE family based on analysis of env gene nucleotide sequences. The displayed tree was obtained by the neighbor-joining (NJ) method, and it is rooted with provirus H_0X7643. Values above branches indicate bootstrap support (1,000 replications) for NJ; below branches, support in the consensus of equally most parsimonious trees; in both cases, only values higher than 85 % are indicated. ω 2–ω 9 = model-based maximum-likelihood estimates of nonsynonymous/synonymous rate ratios (d N /d S ) along the corresponding branches (see also Table 1)
Sliding-window bootscan analysis of the triad formed by HCML-ARV and the proviral insertions H_022319 and P_022319. Nucleotide positions (bp) of H_022319 and bootstrap values (%) are depicted on the x- and y-axis, respectively. The upper right box shows the sequences (HCML-ARV and P_022319) used for comparison to H_022319. Arrows point to bootstrap NJ phylogenetic trees drawn from indicated regions of the bootscan plot. Trees were implemented in MEGA with all the insertions from subfamilies VIIa and VIIb. Bootstrap values 75 % and greater are listed at the respective nodes
To investigate the forces that have shaped HCML-ARV, we carried out an analysis of the patterns of selection along the branches of the phylogeny of ERVE by maximum likelihood ratio tests (Yang 1998). Whenever a gene has been important for the proliferation of the retrovirus, an asymmetry is expected between internal and external branches in the ratio of nonsynonymous to synonymous substitutions per site, d N/d S (ω). The former correspond to ancestral, transiently proliferating, active sequences (presumably subject to purifying selection), and the latter to present-day specific proviral insertions (presumably fixed and defective products of neutral evolution). Thus, in principle, the significantly lower average ω estimated for the internal branches of the phylogeny of the env gene, as compared to external branches (0.314 vs. 0.933, P < 0.001, Table 1; Fig. 1), should be taken as evidence that the proliferation of the ERVE family was almost entirely due to germ-line reinfection (Belshaw et al. 2005). However, ω values for active sequences of subfamilies VIIa and VIIb (1.421 and 0.905, respectively, at Table 1) deviate significantly (P < 0.001) from this general pattern, indicating either a loss of selective constraints or the action of positive Darwinian selection. The first hypothesis is most likely to be true for subfamily VIIb (associated to giant transposons whose proliferation no longer depends on mobility between cells), but the case of VIIa seems to be entirely different, since it includes HCML-ARV, a proviral insertion with an apparently intact and functional env gene. Moreover, all the other retroviral genes seem to be intact in HCML-ARV too. This is especially remarkable, since the ratio between intact and interrupted open reading frames for retroviral loci in the human genome is extremely small (Ruprecht et al. 2008) (1.8, 0.06 and 0.4 %, for gag, pro-pol—the polyprotein including PR, RT, RH, and IN—and env, respectively). To investigate the role of selection in the evolution of all these genes, we carried out several additional likelihood ratio tests. The results are shown in Table 2, and clearly hint at the action of positive selection on gag, in and env active sequences of VIIa. However, recombination-driven biased gene conversion is known to produce spurious signals of positive selection (high ω values) (Berglund et al. 2009). This factor cannot be discarded for gag and in, which show significant W → S (weak to strong) biases in their patterns of nucleotide substitution (Table 3), but it apparently does not affect env. The predicted structure of a region extending from residues 87 to 252 of Env, identified as a receptor-binding domain, was obtained for both the reconstruction of an ancestral active copy (the consensus sequence of VIIa), and its modern derivative, HCML-ARV. As shown in Fig. 3, amino acid replacements have produced a few conspicuous differences in their secondary structure (the folding/unfolding of two short β-sheets, and the turning of a α-helix into two). It is tempting to speculate that the force behind these changes was resistance to host-encoded control mechanisms of retroviral proliferation, which may work by interfering the binding of Env proteins to cell surface receptors, often mediated, paradoxically, by the Env products of other ERVs (Arnaud et al. 2007; Best et al. 1997).
Predicted structure for part of the Env protein (165 residues) of the reconstructed consensus of subfamily VIIa of ERVE (a), compared to HCML-ARV (b). The entire protein chain is shown as a smoothed backbone trace; α-helices are shown as rockets, and β-strands as planks; random coils are white. Colour scheme adapted from the DRuMS Standardized Colour Schemes for Macromolecules (http://www.umass.edu/molvis/drums). Aminoacid differences (position followed by single letter code) between both sequences are indicated on the corresponding models
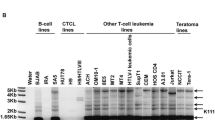
The hard prediction from this work is that the patient with chronic myeloid leukemia, wherefrom HCML-ARV DNA was obtained, carried a functional ERVE copy. Unfortunately, all our attempts to reach the authors of the submission to GenBank have failed. Diagnostic features of the HCML-ARV sequence, e.g. a private deletion of 6 nucleotides at position 448 of rh (supplementary Fig. S2), let us carry out PCR-assisted assays to detect individuals bearing this kind of insertions. Using this strategy, we failed to detect HCML-ARV either in 18 anonymous CML Spanish patients or in the 1,063 individuals of the HGDP-CEPH Human Genome Diversity Panel. This could mean that the former association between CML and the proviral insertion was fortuitous. However, there is still considerable debate over medical diagnostics of CML and related myeloid diseases (Lange and Deininger 2010; Verstovsek 2009), which advises prudence before drawing conclusions on that respect, particularly when considering the many different ways that ERVs might contribute to hematopoietic disorders (Schneider et al. 2009). Different disease-causing alleles may predominate in different populations, so that the outcome of an association study between ERVE and CML could vary between a Spanish and a Chinese sample.
Human endogenous retroviruses (HERVs) have been repeatedly associated to cancer (Ruprecht et al. 2008; Kassiotis 2014), autoimmune diseases (Balada et al. 2009; Nissen et al. 2013; Volkman and Stetson 2014), prion diseases (Lee et al. 2013), and abnormally acute reactions to different kinds of stress (Cho et al. 2008), but the evidence for their precise role in pathogenesis is still quite controversial (Voisset et al. 2008; Young et al. 2012, 2013), pivoting around the impact of variation in ERV insertion profiles on the individual susceptibility to these serious health problems. Since all human beings share the vast majority of TE insertions, the pinpointing of variants that may still be active and segregating in our populations becomes of outmost importance for developing efficient tests of association (Moyes et al. 2007). HCML-ARV, as reported in AF499232, is a proviral insertion of a human endogenous virus in a so far unknown genome location, which could be segregating in some human (Chinese?) populations. The results of this paper identify HCML-ARV as a representative of a potentially active lineage of ERVE, whose origin traces back to relatively recent genetic exchanges with different proviral copies that inserted in the genome of our ancestors several million of years ago, and that bears distinctive molecular signatures of positive selection on its envelope region which could be the key of its success in running away from the control mechanisms of retroviral proliferation encoded in our genome.
ERVE was a prolific family of endogenous retroviruses that displayed at least eight long-lasting bursts of activity in our ancestors, two of them (subfamilies VIIa and VIIb) still in course after the separation of the gorilla from our common trunk. This offers a wide-open field for recombination that should make us cautious in pronouncing its sentence of death.
References
Arnaud F, Caporale M, Varela M, Biek R, Chessa B et al (2007) A paradigm for virus–host coevolution: sequential counter-adaptations between endogenous and exogenous retroviruses. PLoS Pathog 3:e170
Balada E, Ordi-Ros J, Vilardell-Tarrés M (2009) Molecular mechanisms mediated by human endogenous retroviruses (HERVs) in autoimmunity. Rev Med Virol 19:273–286
Bannert N, Kurth R (2004) Retroelements and the human genome: new perspectives on an old relation. Proc Natl Acad Sci USA 101(Suppl 2):14572–14579
Bannert N, Kurth R (2006) The evolutionary dynamics of human endogenous retroviral families. Annu Rev Genomics Hum Genet 7:149–173
Bartolome C, Bello X, Maside X (2009) Widespread evidence for horizontal transfer of transposable elements across Drosophila genomes. Genome Biol 10:R22
Belshaw R, Katzourakis A, Paces J, Burt A, Tristem M (2005) High copy number in human endogenous retrovirus families is associated with copying mechanisms in addition to reinfection. Mol Biol Evol 22:814–817
Berglund J, Pollard KS, Webster MT (2009) Hotspots of biased nucleotide substitutions in human genes. PLoS Biol 7:e26
Best S, Le Tissier PR, Stoye JP (1997) Endogenous retroviruses and the evolution of resistance to retroviral infection. Trends Microbiol 5:313–318
Buzdin A (2007) Human-specific endogenous retroviruses. Sci World J 7:1848–1868
Cann HM, de Toma C, Cazes L, Legrand M-F, Morel V et al (2002) A human genome diversity cell line panel. Science 296:261
Cho K, Lee YK, Greenhalgh DG (2008) Endogenous retroviruses in systemic response to stress signals. Shock 30:105–116
Cordaux R, Batzer MA (2009) The impact of retrotransposons on human genome evolution. Nat Rev Genet 10:691–703
de Parseval N, Heidmann T (2005) Human endogenous retroviruses: from infectious elements to human genes. Cytogenet Genome Res 110:318–332
Dewannieux M, Harper F, Richaud A, Letzelter C, Ribet D et al (2006) Identification of an infectious progenitor for the multiple-copy HERV-K human endogenous retroelements. Genome Res 16:1548–1556
Dolei A (2006) Endogenous retroviruses and human disease. Expert Rev Clin Immunol 2:149–167
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Feschotte C, Gilbert C (2012) Endogenous viruses: insights into viral evolution and impact on host biology. Nat Rev Genet 13:283–296
Gifford R, Tristem M (2003) The evolution, distribution and diversity of endogenous retroviruses. Virus Genes 26:291–315
Hall TA (1999) BIOEDIT: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hughes JF, Coffin JM (2005) Human endogenous retroviral elements as indicators of ectopic recombination events in the primate genome. Genetics 171:1183–1194
Jern P, Coffin JM (2008) Effects of retroviruses on host genome function. Annu Rev Genet 42:709–732
Jha AR, Nixon DF, Rosenberg MG, Martin JN, Deeks SG et al (2011) Human endogenous retrovirus K106 (HERV-K106) was infectious after the emergence of anatomically modern humans. PLoS One 6:e20234
Ji X, Zhao S (2008) DA and Xiao—two giant and composite LTR-retrotransposon-like elements identified in the human genome. Genomics 91:249–258
Kassiotis G (2014) Endogenous retroviruses and the development of cancer. J Immunol 192:1343–1349
Kelley LA, Sternberg MJ (2009) Protein structure prediction on the Web: a case study using the Phyre server. Nat Protoc 4:363–371
Kent WJ (2002) BLAT—the BLAST-like alignment tool. Genome Res 12:656
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC et al (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Lange T, Deininger MW (2010) Molecular diagnostics in chronic myeloid leukemia. Expert Opin Med Diagn 4:113–124
Lee YN, Bieniasz PD (2007) Reconstitution of an infectious human endogenous retrovirus. PLoS Pathog 3:e10
Lee YK, Chew A, Phan H, Greenhalgh DG, Cho K (2008) Genome-wide expression profiles of endogenous retroviruses in lymphoid tissues and their biological properties. Virology 373:263–273
Lee Y-J, Jeong B-H, Choi E-K, Kim Y-S (2013) Involvement of endogenous retroviruses in prion diseases. Pathogens 2:533–543
Leis J, Baltimore D, Bishop JM, Coffin J, Fleissner E et al (1988) Standardized and simplified nomenclature for proteins common to all retroviruses. J Virol 62:1808–1809
Li X, Slife J, Patel N, Zhao S (2009) Stepwise evolution of two giant composite LTR-retrotransposon-like elements DA and Xiao. BMC Evol Biol 9:128
Lindeskog M, Medstrand P, Cunningham AA, Blomberg J (1998) Coamplification and dispersion of adjacent human endogenous retroviral HERV-H and HERV-E elements; presence of spliced hybrid transcripts in normal leukocytes. Virology 244:219–229
Lole KS, Bollinger RC, Paranjape RS, Gadkari D, Kulkarni SS et al (1999) Full-length human immunodeficiency virus type 1 genomes from subtype C-infected seroconverters in India, with evidence of intersubtype recombination. J Virol 73:152–160
López-Sánchez P, Costas JC, Naveira HF (2005) Paleogenomic record of the extinction of human endogenous retrovirus ERV9. J Virol 79:6997–7004
Mills RE, Bennett EA, Iskow RC, Devine SE (2007) Which transposable elements are active in the human genome? Trends Genet 23:183–191
Moyes D, Griffiths DJ, Venables PJ (2007) Insertional polymorphisms: a new lease of life for endogenous retroviruses in human disease. Trends Genet 23:326–333
Nissen KK, Laska MJ, Hansen B, Terkelsen T, Villesen P et al (2013) Endogenous retroviruses and multiple sclerosis—new pieces to the puzzle. BMC Neurol 13:111
Ogasawara H, Hishikawa T, Sekigawa I, Hashimoto H, Yamamoto N, Maruyama N (2000) Sequence analysis of human endogenous retrovirus clone 4-1 in systemic lupus erythematosus. Autoimmunity 33:15–21
Piotrowski PC, Duriagin S, Jagodzinski PP (2005) Expression of human endogenous retrovirus clone 4-1 may correlate with blood plasma concentration of anti-U1 RNP and anti-Sm nuclear antibodies. Clin Rheumatol 24:620–624
Prusty BK, zur Hausen H, Schmidt R, Kimmel R, de Villiers EM (2008) Transcription of HERV-E and HERV-E-related sequences in malignant and non-malignant human haematopoietic cells. Virology 382:37–45
Repaske R, O’Neill RR, Steele PE, Martin MA (1983) Characterization and partial nucleotide sequence of endogenous type C retrovirus segments in human chromosomal DNA. Proc Natl Acad Sci USA 80:678–682
Repaske R, Steele PE, O’Neill RR, Rabson AB, Martin MA (1985) Nucleotide sequence of a full-length human endogenous retroviral segment. J Virol 54:764–772
Ruprecht K, Mayer J, Sauter M, Roemer K, Mueller-Lantzsch N (2008) Endogenous retroviruses and cancer. Cell Mol Life Sci 65:3366–3382
Schneider AM, Duffield AS, Symer DE, Burns KH (2009) Roles of retrotransposons in benign and malignant hematologic disease. Cellscience 6:121–145
Seifarth W, Frank O, Zeilfelder U, Spiess B, Greenwood AD et al (2005) Comprehensive analysis of human endogenous retrovirus transcriptional activity in human tissues with a retrovirus—specific microarray. J Virol 79:341–352
Shin W, Lee J, Son SY, Ahn K, Kim HS, Han K (2013) Human-specific HERV-K insertion causes genomic variations in the human genome. PLoS One 8:e60605
Smit AF (2008) HERVE_a - ERV1 endogenous retrovirus from Catarrhini. Direct submission to Repbase Update. http://www.girinst.org/repbase/index.html
Steele PE, Rabson AB, Bryan T, Martin MA (1984) Distinctive termini characterize two families of human endogenous retroviral sequences. Science 225:943–947
Takahashi Y, Harashima N, Kajigaya S, Yokoyama H, Cherkasova E et al (2008) Regression of human kidney cancer following allogeneic stem cell transplantation is associated with recognition of an HERV-E antigen by T cells. J Clin Invest 118:1099–1109
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Taruscio D, Floridia G, Zoraqi GK, Mantovani A, Falbo V (2002) Organization and integration sites in the human genome of endogenous retroviral sequences belonging to HERV-E family. Mamm Genome 13:216–222
Verstovsek S (2009) Preclinical and clinical experience with dasatinib in Philadelphia chromosome-negative leukemias and myeloid disorders. Leuk Res 33:617–623
Voisset C, Weiss RA, Griffiths DJ (2008) Human RNA “rumor” viruses: the search for novel human retroviruses in chronic disease. Microbiol Mol Biol Rev 72:157–196
Volkman HE, Stetson DB (2014) The enemy within: endogenous retroelements and autoimmune disease. Nat Immunol 15:415–422
Yang Z (1998) Likelihood ratio tests for detecting positive selection and application to primate lysozyme evolution. Mol Biol Evol 15:568–573
Yi JM, Kim HS (2006) Molecular evolution of the HERV-E family in primates. Arch Virol 151:1107–1116
Yi JM, Kim HS (2007) Molecular phylogenetic analysis of the human endogenous retrovirus E (HERV-E) family in human tissues and human cancers. Genes Genet Syst 82:89–98
Young GR, Eksmond U, Salcedo R, Alexopoulou L, Stoye JP, Kassiotis G (2012) Resurrection of endogenous retroviruses in antibody-deficient mice. Nature 491:774–778
Young GR, Stoye JP, Kassiotis G (2013) Are human endogenous retroviruses pathogenic? An approach to testing the hypothesis. BioEssays 35:794–803
Acknowledgments
We thank C. Phillips and. M. V. Lareu for kindly letting us use their copy of the HGDP-CEPH Human Genome Diversity Panel.
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10709_2014_9789_MOESM2_ESM.pdf
Figure S1. Phylogenetic relationships within the ERVE family based on the analysis of gag, pr, rt and rh gene nucleotide sequences. (PDF 1051 kb)
10709_2014_9789_MOESM3_ESM.pdf
Figure S2. Multiple nucleotide alignment of HCML-ARV with the consensus of subfamily VIIa and the orthologous copies of provirus 022319 in humans and chimpanzees. (PDF 87 kb)
Rights and permissions
About this article
Cite this article
Naveira, H., Bello, X., Abal-Fabeiro, J.L. et al. Evidence for the persistence of an active endogenous retrovirus (ERVE) in humans. Genetica 142, 451–460 (2014). https://doi.org/10.1007/s10709-014-9789-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-014-9789-y