Abstract
HIF-α transcription factors, as key master regulators of oxygen homeostasis, constitute a subgroup of the large bHLH-PAS transcription factor family and have been identified in many vertebrates. Although the amino acid sequences of bHLH-PAS domain are conserved, the physiological and pathological roles of this family are variable. They also have different patterns of expression. It is possible that the HIF-α copies have been retained as a consequence of adaptive amino acid replacements or relaxed selective constraint which have conferred subtle changes in function after duplications. Phylogenetic analysis indicated that at least two major duplications had occurred early in the vertebrate lineages. Analyses of the ratios of nonsynonymous/synonymous substitution rates revealed that relaxation of selective constraints might play important roles over evolutionary time and shape variation in some members of the family. The coefficients of functional divergence (θ) estimated between pairwise comparisons of gene groups from HIF-1α, HIF-2α, and HIF-3α indicated statistically significant site-specific shift of evolutionary rates between them, suggesting that altered functional constraints may have taken place at some amino acid residues after their duplications. Moreover, we also mapped sites identified to have been relaxed from purifying selection onto the three-dimensional structure of human HIF-2α. Overall, our study demonstrated that the functional diversity of HIF-αs members may be caused by relaxed negative selection on the N-terminal transactivation domains after HIF-αs duplications, which recruited new partners leading to functional specificity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hypoxia is defined as a reduction in oxygen amount available to a cell, tissue, or organism and it can arise in a variety of developmental, physiological, and pathological states. The decline in oxygen level can cause alteration in gene transcription or may result in post-translational modifications of proteins, leading to changes in cell metabolism. One of the pivotal mediators of the cellular response to hypoxia is the hypoxia-inducible factors (HIFs).
HIFs are transcription factors belonging to the basic helix-loop-helix (bHLH)-containing PER-ARNT-SIM (PAS) domain protein family. There are three known members of this family (HIF-1, HIF-2, and HIF-3), and all are α/ß heterodimeric proteins. HIF-1α was the first factor to be cloned and is the best understood isoform (Wang and Semenza 1995). Since then, the discovery of vertebrate HIF genes has grown rapidly. Now HIF proteins have been identified in several different organisms, including zebrafish, chicken, mouse, rat, and human. In normoxia, the α-subunit is ubiquitously expressed but is rapidly degraded through interaction with the VHL ubiquitin ligase complex (Maxwell et al. 1999). In hypoxic conditions, it is stabilized through multiple mechanisms that are transcriptional, post-transcriptional, as well as post-translational (Maxwell et al. 1999; Jaakkola et al. 2001; Gustafsson et al. 2005). Despite HIF-1α, its close homologues (HIF-2α and HIF-3α) have the high sequence identity (more than 67% identity in the bHLH, PAS, and transactivation domains), and all of them play roles in sensing oxygen levels; punctual amino acid replacement at key structural domains of the respective proteins may have evolved to different expression patterns and physiology roles. HIF-1α expresses in most cell types, whereas HIF-2α shows a more restricted pattern of expression. Mice deficient in HIF-1α die in utero by embryonic day 10, with embryos exhibiting poor vascularization (Ryan et al. 1998), while HIF-2α knockout mice display impairment of angiogenesis, dramatic cardiovascular defects, and failure of neural tube closure caused by mesenchymal cell death (Tian et al. 1998; Peng et al. 2000; Compernolle et al. 2002). Moreover, placentas from mice lacking both HIF-1α and HIF-2α exhibit defective placental vascularization and aberrant cell fate adoption (Cowden Dahl et al. 2005). Interestingly, hypoxia does not affect mRNA levels of HIF-1α and HIF-2α, but HIF-3α increased after 2-h hypoxia (Heidbreder et al. 2003). The function and expression pattern of HIF-3α and how hypoxia increases mRNA level of HIF-3α remain relatively unknown (Gu et al. 1998), but it was proposed that HIF-3α serves as an inhibitor of the HIF pathway (Makino et al. 2001). Although differential expression may contribute to these differences, recent studies have demonstrated that each gene appears to have distinct transcriptional targets (Iyer et al. 1998a; Hu et al. 2003; Raval et al. 2005). These transcriptional target genes are involved in erythropoiesis, angiogenesis, metastasis, energy metabolism, cell cycle arrest, differentiation, and apoptosis/proliferation (Carmeliet et al. 1998; Ryan et al. 1998; Iyer et al. 1998a; Patel and Simon 2008). Therefore, HIF-α transcription factors also contribute significantly to both normal physiology and tumorigenesis.
Single copy genes are thought to evolve conservatively because of strong negative selective pressures. Gene duplications produce a redundant gene copy and thus release one or both copies from negative selection pressure (Hughes and Criscuolo 2008). There are a number of models for the fate of duplicate gene that predict functional differentiation of paralogs based on protein sequence or regulatory divergence (Kimura 1979; Force et al. 1999). Currently, three major fates of genes after duplications are supposed to be the neofunctionalization (Goodman et al. 1975), subfunctionalization (Force et al. 1999), and pseudogenization. Thus, duplications are thought to be an important precursor of functional divergence (Lynch et al. 2006). Here, we are interested in the specific role that natural selection might play in the evolutionary history of HIF-α gene family.
Previous work has demonstrated that selection plays an important role in shaping HIF-1α structure, function, and regulation (Iyer et al. 1998b), and evolutionary rate in teleost HIF-1α is faster than in mammalian HIF-1α (Rytkonen et al. 2008). Yet, a wide-ranging study correlating its molecular evolution with structural biology in vertebrate HIF-α gene family remains unclear. In this study, we made efforts in (1) studying the evolutionary history of the HIF-α gene family, (2) evaluating the changes in selection pressures following duplications, and (3) homology modeling to understand its process of diversification. Our work may provide some clues for future investigations into HIF-αs as well as the mechanism of angiogenesis, embryogenesis, and tumorigenesis.
Materials and methods
Data collection and alignment
PSI-BLAST and TBLASTN searches were made with E value <10e-5 against the protein databases or unfinished genome sequencing projects at the National Center for Biotechnology Information Database (http://www.ncbi.nlm.nih.gov) and Ensembl database (http://www.ensembl.org) by using protein sequences of the three human HIF-α proteins (HIF-1α, HIF-2α, and HIF-3α). In this study, a total of 20 vertebrate species genomes were used to identify potential homologues (amino acid identity was above 25% over a stretch of 200 amino acids). These species were chosen to provide a range of phylogenetic distances. They were Homo sapiens (human), Macaca mulatta (macaque), Callithrix jacchus (marmoset), Pan trogolodytes (chimpanzee), Gorilla gorilla (gorilla), Pongo pygmaeus (orangutan), Mus musculus (mouse), Rattus norvegicus (rat), Vicugna pacos (alpaca), Sus scrofa (pig), Bos Taurus (cow), Canis familiaris (dog), Oryctolagus cuniculus (rabbit), Gallus gallus (chicken), Taeniopygia guttata (zebra finch), Xenopus tropicalis (frog), Danio rerio (zebrafish), Gasterosteus aculeatu (stickleback), Oryzias latipes (medaka), and Tetraodon nigroviridis (pufferfish). After the removal of expressed sequence tags and shorter splice variants, the initial data set (HIF-αs) included 50 sequences from 20 species. An additional sequence (NM_075607) from the Caenorhabditis elegans genome was served as an out-group and aided in determining the earliest diverging of HIF-αs. Accession numbers of all sequences used in the study were available as supplementary material (Table S1), together with information about the species from which they were obtained.
The protein sequences of the initial data set and complete data set (including 50 HIF-αs and 1 out-group sequence) were aligned by MUSCLE (Edgar 2004) independently using default parameter settings. In order not to introduce bias, manual alignment editing was minimized. Accurate nucleotide alignments of sequences were obtained with PAL2NAL, which is a program to construct multiple codons alignments from matching amino acid sequences (Suyama et al. 2006).
Phylogenetic analysis
The full alignment of 51 sequences was used for the phylogenetic analysis. Tree searches were made with Bayesian analysis in MrBayes 3.1.2 (Ronquist and Huelsenbeck 2003) and maximum likelihood (ML) method implemented in Phyml v2.4.4 (Guindon and Gascuel 2003). In the Bayesian analysis, a general time reversible model with a proportion of invariant sites and gamma distributed among-site rate variation (GTR + I + G) was used in the DNA sequences, as selected by Akaike information criterion (AIC) in MrModeltest 2.3 (Nylander 2004). The robustness of the tree was assessed by bootstrapping (500 replicates) in Phyml. The data were partitioned by codon position (for the HIF-α genes), and we ran five million generation Bayesian Markov chain Monte Carlo analyses with four separate chains (sampling every 1,000 generations), with the first 500,000 generations discarded as burn-in. Stationarity was assumed when the average standard deviation of split frequencies dropped to less than 0.01. After discarding the first 500,000 generations as burn-in, posterior probabilities were calculated and reported on a 50% majority rule consensus tree of the post-burn-in sample.
Analyses of positive selection
We estimated the selective pressures acting on coding regions by applying a phylogenetic-based maximum likelihood method. The relevant parameters were estimated using codeml program implemented in the PAML package version 4.4 (Yang 2007), for example the branch lengths and the ratio of the nonsynonymous (d N) to synonymous substitution rates (d S), ω = d N/d S. The ratio of nonsynonymous to synonymous substitution rates (d N/d S or ω) was used to evaluate the selective regimes operating at the molecular level, expecting d N/d S = 1 for neutrality, d N/d S < 1 for purifying selection, and d N/d S > 1 for positive selection.
In this study, we used two different kinds of codon models. The first kind comprises the so-called site-specific models (Yang 2002; Wong et al. 2004) and was designed to detect positive selection among sites (codons). The second kind, the branch-site models, has the same objective but exclusively in particular lineages of a phylogeny, allowing the d N/d S ratio to vary across sites and across lineages and detecting codon-specific positive selection in preselected branches of a given tree regardless of the average d N/d S ratio (Yang and Nielsen 2002; Zhang et al. 2005). Model comparisons were made using likelihood ratio tests (LRTs), which require nested hypotheses of models being compared. Degrees of freedom are equal to the number of extra parameters in the alternative hypothesis.
We used two pairs of site models to test for individual residues under positive selection: M1a (neutral) versus M2a (positive selection) and M7 (beta) versus M8 (beta + ω). LRTs were applied to test for presence of positively selected amino acid sites. When a signature of positive selection according to the LRTs showed, the Bayes empirical Bayes (BEB) method (Nielsen and Yang 1998; Wong et al. 2004) was used to identify individual codons with d N /d S > 1. These models do not account for molecular rate shifts or positive selection in particular lineages, thus averaging over all sequences in the phylogeny. Therefore, positive selection is difficult to detect with these models. In contrast, branch-site models have more power to detect positive selection on specific lineages because they allow for branches prespecified in the phylogeny to have a class of sites with ω > 1 (model A). In a test using branch-site model A, the branch being tested for positive selection is called the foreground branch, and the other branches in the tree are called the background branches. The LRT is used to evaluate whether significant positive selection happens on the foreground branch. Using branch-site models, we tested branches of main lineages before and after important duplication events during the evolution of HIF-αs genes, which represent potential points for the evolution of possible newly acquired roles in the oxygen homeostasis following these duplications. Branches assigned as ‘‘foreground’’ were indicated with arrows in the Bayesian phylogeny.
Amino acid sequence analyses
A statistical framework modeling the functional divergence was implemented by the DIVERGE program (Gu and Vander Velden 2002), to estimate the coefficient of functional divergence (θ). This coefficient is an indicator of the level of type I functional divergence caused by an evolutionary process resulting in either different functional constraints or in a site-specific evolutionary rate shift between two duplicate genes (Gu 1999; Gu and Vander Velden 2002). The null hypothesis of θ = 0 means that the evolutionary rate is virtually the same between duplicate genes at each site. If the null hypothesis was rejected, statistical evidence for shifts in evolutionary rates or in altered functional constraints was provided.
Modeling and structure analysis
Genes identified by the above-explained methods to evolve under altered selective constraints were used to search for homologous sequences in the PDB database of protein structures (http://www.rcsb.org/pdb/home/home.do) using Blastp (Altschul et al. 1990, 1997). The Rasmol software (http://rasmol.org/) was used for all structural manipulations and highlighting the relevant amino acid replacements identified in the evolutionary analyses.
Results and discussion
Phylogenetic analysis of the HIF-αs proteins
In this study, we compiled a data set of HIF-α genes (50 sequences in total) from 20 vertebrate species (see supplemental material Table S1) and compared conserved and variable sites between paralog group members (for example, see Fig. 1 with 6 species). This analysis identified many sites that are conserved in the same paralog group but variable between paralogs. Although the domains including bHLH, PAS, and transactivation domains are highly conserved, they exhibit polymorphism. Interestingly, HIF-αs have a higher level of DNA sequence variability in aquatic teleosts than in terrestrial tetrapods, and only 35 residues are absolutely conserved between all vertebrate HIF-αs genes in our alignment. Of these, 10 are located in bHLH domain region spanning from 168 to 220, 10 in PAS-A region from 264 to 324, and 15 in PAS-B domain region from 445 to 550 in our sequences alignment (see supplemental material Fig. S1). The N-transactivation domains (N-TAD) and oxygen-dependent degradation domain (ODD) (see supplemental material Fig. S1) are varied considerably among different species. In addition, we found the amino acids Thr (position 508 in our alignment), Gln (position 679), and Ser (position 1186) in mammals are replaced by Ala, Leu, and Arg in the fish species for HIF-1α, respectively. Also, some amino acids are conserved between all mammals but varied in teleost species for HIF-1α and HIF-2α protein sequences, for example amino acids that are located in the position 726 and 727 (overlap region of ODD and N-TAD). Further, teleost HIF-α protein sequences are 24–55 amino acids shorter than tetrapod HIF-α proteins (an average of 48 amino acids shorter for HIF-1α, 55 for HIF-2α, and 24 for HIF-3α). This is mainly due to the important deletions in fish around amino acids 439–456 and 634–658 for HIF-1α (human nomenclature), and around amino acids 577–632 for HIF-2α.
In this study, 51 sequences (one from Caenorhabditis elegans genome) were used for phylogenetic reconstruction with maximum likelihood (supplemental material Fig. S2) and Bayesian inference (BI) (Fig. 2) methods. The different reconstruction methods provided consistent topologies, especially when the major clades are considered. Exceptions are OlHIF1A and TnHIF2A sequences that are not clustered into well-defined clades in the BI method (Fig. 2). This discrepancy may be attributed to the systematic error. Our phylogenetic analysis also demonstrated that the orthologues are more related to each other from different species than to their species-specific paralogs. Most genes fell into well-defined clades supported by high Bayesian posterior probabilities (PPs) values, so the phylogeny supports our classification of HIF-αs genes into three paralog groups. HIF-1α and HIF-2α genes are sister clusters, with high PP support values (1.00 and 1.00). As sister sequences of this HIF-1α/HIF-2α clade, we recovered the clade of HIF-3α genes (1.00 PP) in a supported position (0.78 PP). From the phylogenetic analyses, we also found two subclades within clade HIF-1α. The fishes formed teleost subclade, while tetrapod subclade comprises birds and mammalian. Moreover, estimates of divergence time suggested that the tetrapods left the aquatic environment at about 360 Myr ago, and this result is quite close to the previous estimates of teleost–tetrapod splitting (400 Myr ago) (Park et al. 2008). Furthermore, we observed several species-specific gene duplications that have likely taken place in teleosts, for example one-to-many orthologous genes are present in the Tetraodon nigroviridis genome compared with other vertebrates (Fig. 2). This result is consistent with previous reports that the teleost lineage is hypothesized to have undergone additional genome duplication (Taylor et al. 2001; Taylor et al. 2003). The lost teleost orthologs, for example, only one teleost sequence was identified within HIF-2α clade, may be caused by two reasons in this study: (1) HIF-αs were truly lost from their genome; (2) incomplete genomic sequences (i.e., quality of contig sequences without finishing) may also influence reliable conclusion. The phylogenetic tree enabled us to see that an early duplication is responsible for HIF-3α and HIF-2α/HIF-1α splitting, and recent gene duplication resulted in HIF-2α and HIF-1α. As indicated above, these duplications should have occurred before speciation. Therefore, we suggest that the HIF-αs originated by duplication and divergence of a common protein at the base of the eukaryotic tree, some 700 Myr ago, and further duplication events contribute to the split between tetrapods and teleosts.
Phylogenetic tree of 50 homologues of vertebrate HIF-αs. The phylogenetic tree of the HIF-α protein family was inferred by the Bayesian analyses. Posterior probabilities are labeled above branches. Caenorhabditis elegans was used as outgroup. For species name abbreviations, see supplementary Table S1. The numbers indicate the Bayesian probabilities for each phylogenetic clade. Shaded boxes denote the three lineages. Arrows indicated duplication events and used to set as foreground branches
Selection and functional diversification
Several models of molecular evolution have been proposed to account for the preservation of duplicate genes. They differ in their predictions regarding the pattern of sequence evolution following gene duplications. A previous study reported that positive Darwinian or adaptive selection might have been important in fixing specific protein residues after some of the main duplication events leading to the paralogous groups of the gene family (Martinez-Castilla and Alvarez-Buylla 2003), while Hughes et al. (2009) have reported the possible acquisition of the new function of UCP1 through neutrally evolving, relaxed constraints after duplications. Moreover, the neofunctionalization is usually supposed to include a stage of neutral evolution, with functional change only occurring by a last step including positive selection (Wagner 2008). Therefore, we tested for significant differences in the substitution rates and for positive selection by comparing nonsynonymous (d N) to synonymous (d S) substitution rate (Lynch et al. 2006) to test evolutionary patterns playing in this gene family formation.
Codon-based substitution models were used to test for site-specific selection and then address whether some sites had been fixed by positive selection. The tests contrasting the models M1a against M2a resulted in nearly identical log-likelihood scores (data not shown), suggested that the amino acid changes were neutral or under purifying selection. M1a, the parameter estimates for the least parameter rich model, describes that most sites with low ω estimates (indication of strong selective constraints), that is 87% of HIF-αs sites, were under strong purifying selection. The test with M7 and M8, which allows for beta-distributed site-specific ω ratio, also failed to detect any site under possible positive selection (data not shown). These results provided consistently strong evidence that the test does not find any positively selected sites across the whole multiple sequences alignments.
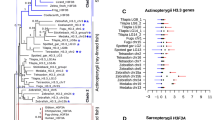
Since positive selection will likely affect a few amino acids at specific lineages on the phylogeny, models estimating ω ratios averaged over all lineages might hide the signal for positive selection. Therefore, it is reasonable to expect molecular rate shifts and positive selection after gene duplications when functional diversification processes affect one or both duplicates. In this study, we decided to assess whether positive selection had fixed certain amino acids at particular moments of HIF-αs gene evolution, especially following the main duplication events. To achieve this, we used branch-site models that detect positive selection at specific coding sites and in particular branches of the phylogenetic tree. We focused on two key duplications during the evolution of HIF-αs genes (see Fig. 2). In spite that null hypothesis of the test 2 (compare the model A to the modified model A with ω = 1 fixed) was not rejected, the lineages after duplications exhibit several positions with evidence of relaxed selection (the test 1 (compare the model A to the M1a) was significant) (Table 1); nevertheless, many relaxed amino acid positions are located in the highly variable ODD region. Though several positively selected sites (estimated ω greater than 1) were identified in previously assigned lineages by empirical Bayes estimation, a class of sites in each was identified with ω2 = 1 (Table 1). As such, relaxed functional constraint is most consistent with the molecular evolutionary analyses of the HIF-αs data. Further, our observation of strong purifying selection being the primary mode of evolution throughout the HIF-αs phylogeny is consistent with the findings of recent results on the fate of Hox and UCP genes after duplications (Lynch et al. 2006; Hughes et al. 2009). Gene duplication–specific changes in the substitution rates (type I functional divergence) might reflect the difference in evolutionary rate at amino acid sites after gene duplication (Gu 1999; Gu 2001). In this analysis, we performed analysis using the DIVERGE program (Gu and Vander Velden 2002) for all pairs of HIF-αs clusters to detect evolutionary rate pattern shifts and to identify which amino acid sites may have contributed to the functional divergence. Pairwise comparisons of gene groups from HIF-1α, HIF-2α, and HIF-3α were carried out, and the sites that are most likely to have evolved at different rates between two clusters were identified by establishing empirically a cutoff value. Three coefficients of functional divergence (θ) with standard errors and significance levels are given in Table 2. We found significant evidence of type I functional divergence for comparisons between different gene clusters, with θ varying markedly from 0.22 to 0.35. Namely, there were some amino acid sites with discrepancies in their evolutionary rates between these paralogous pairs. The amino acids responsible for the functional divergence after gene duplication can be predicted based on a site-specific profile by choosing a suitable cutoff value (Thompson et al. 2007). In this study, we used the cutoff values: Q(k) ≥ 0.67. As expected, most amino acids had very low posterior probability (PP) values and, therefore, they would not be involved in the hypothetical functional divergence (Fig. 3). Specifically, we detected five, twenty-two, and fourteen amino acid positions (with PP threshold values higher than 0.67) in the HIF-1α/HIF-3α, HIF-2α/HIF-3α, and HIF-1α/HIF-2α comparisons, respectively. Besides, eleven of these functional divergence candidate positions are also detected by LRT in codeml which may be related to relaxed selective constraint during HIF-αs duplications (indicated in bold in Table 1). These residues were mapped onto the three-dimensional structure of the Homo sapiens HIF-2α which we have modeled.
Homology modeling
To gain a better understanding of altered functional constraints on the functional diversity, we mapped the location of sites under altered functional constraints onto the sequence logo (supplemental material Fig. S1) and three-dimensional structure. Figure 4 shows the location in the three-dimensional structure of the relevant relaxed selected amino acid positions and those inferred to contribute to type I functional divergence (with high PP values). Interestingly, we found that these sites are located in the region adjacent to the hydroxylation domains (ODD and C-TAD in supplementary Fig. S1). This suggests that sequence variation in these regions could be adaptive because it may affect the oxygen sensitivity of the protein by affecting its three-dimensional structure and the binding affinities of the oxygen-sensitive hydroxylases (Rytkonen et al. 2008). It has been reported that the ODD domain of HIF-1α plays a dual role in the context of the full-length protein: (1) it functions as a transcriptional activator that stimulates gene expression via protein–protein interactions with the basal machinery and (2) it senses changes in oxygen levels through enzymatic hydroxylation of two of its prolines, resulting in subsequent ubiquitination by the VHL-elongin B-elongin C (VCB) complex followed by degradation (Sanchez-Puig et al. 2005). The findings of Sanchez-Puig et al. suggest that HIF-1α ODD domain contained two binding sites for p53 core. The fragments comprised full-length ODD domain, and N-TAD and inhibitory domain, respectively. Previous attempt was also made to narrow down the ODD domain by dividing the ODD domain into three parts, and each of them independently conferred hypoxic induction to different degrees (Huang et al. 1998). However, previous results pose a stiff challenge to determine the precise sites responsible for O2-dependent degradation. Our work just provided some probable functional sites spread over the domain which could be mutated simultaneously to verify their functions. Moreover, N-terminal transactivation domains (aa360–aa600 for human HIF-1α) overlapping with the hydroxylation domains confer functional specificity on HIF-αs proteins (Brahimi-Horn and Pouyssegur 2007). The majority of sites under relaxed purifying selection in this region demonstrate that functional constraints are more relaxed for the region of the protein that exposed to the degradation at normoxia and transcriptional activation at hypoxia. This may have enabled the recruitment of novel interactions with proteins which potentially modify functional specificity between paralogs. Finally, some relaxed selection sites are conserved between all mammals but varied in teleost species or replaced by another conserved amino acid, for example amino acid Thr (position 508) and Gln (position 679) in mammals. We thought that this may be related to the multigenerational adaptation to separate environment for these species. Without further analyses of function, we are still unable to define the biological significance of these relaxed sites between past and present aquatic environment. Of course, we also could not exclude the important roles of the deletion of HIF-αs in teleosts. Just because the present computational models cannot take the indels into account does not mean that they would be unimportant. This pattern of more stringent negative selection in HIF-αs proteins reflects an ancient fixation of the HIF-αs pathway in response to constant oxygen supply in the vertebrate habitat. Accordingly, the acquisition of novel protein interaction partners was likely driven by relaxed negative selection on the N-terminal transactivation domains after HIF-αs duplications. These derived interactions could be those responsible for functional diversity of HIF-αs proteins in vertebrates.
Conclusions
The results of this study addressed the evolutionary history of vertebrate HIF-α gene family for the first time from molecular sequence data. Our results suggested that gene duplication events trigger diversification of the HIF-α family into three groups. Statistical analyses of selective pressures indicated that relaxed selective constraint might play important roles over evolutionary time and shape variation in some members of the family. The detection of functional divergence among HIF-αs groups also confirmed that the HIF-αs members have evolved into different functional properties owing to rate shifts of a small set of amino acids after gene duplication events. At last, we mapped the location of sites under alter functional constraint onto the sequence logo and three-dimensional structure. The probably relaxed selection sites are located both inside and on the surface of the three-dimensional structure. All these studies will certainly contribute to better understand the precise role of natural selection and functional diversity of this family.
References
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Altschul SF, Madden TL, Schaffer AA et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Brahimi-Horn MC, Pouyssegur J (2007) Harnessing the hypoxia-inducible factor in cancer and ischemic disease. Biochem Pharmacol 73:450–457
Carmeliet P, Dor Y, Herbert JM et al (1998) Role of HIF-1alpha in hypoxia-mediated apoptosis, cell proliferation and tumour angiogenesis. Nature 394:485–490
Compernolle V, Brusselmans K, Acker T et al (2002) Loss of HIF-2alpha and inhibition of VEGF impair fetal lung maturation, whereas treatment with VEGF prevents fatal respiratory distress in premature mice. Nat Med 8:702–710
Cowden Dahl KD, Fryer BH, Mack FA et al (2005) Hypoxia-inducible factors 1alpha and 2alpha regulate trophoblast differentiation. Mol Cell Biol 25:10479–10491
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinform 5:113
Force A, Lynch M, Pickett FB et al (1999) Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151:1531–1545
Goodman M, Moore GW, Matsuda G (1975) Darwinian evolution in the genealogy of haemoglobin. Nature 253:603–608
Gu X (1999) Statistical methods for testing functional divergence after gene duplication. Mol Biol Evol 16:1664–1674
Gu X (2001) Maximum-likelihood approach for gene family evolution under functional divergence. Mol Biol Evol 18:453–464
Gu X, Vander Velden K (2002) DIVERGE: phylogeny-based analysis for functional-structural divergence of a protein family. Bioinformatics 18:500–501
Gu Y-Z, Moran SM, Hogenesch JB et al (1998) Molecular characterization and chromosomal localization of a third α-class hypoxia inducible factor subunit, HIF3α. Gene Expr 7:205–213
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Gustafsson MV, Zheng X, Pereira T et al (2005) Hypoxia requires notch signaling to maintain the undifferentiated cell state. Dev Cell 9:617–628
Heidbreder M, Frohlich F, Johren O et al (2003) Hypoxia rapidly activates HIF-3α mRNA expression. FASEB J 17:1541–1543
Hu CJ, Wang LY, Chodosh LA et al (2003) Differential roles of hypoxia-inducible factor 1alpha (HIF-1alpha) and HIF-2alpha in hypoxic gene regulation. Mol Cell Biol 23:9361–9374
Huang LE, Gu J, Schau M et al (1998) Regulation of hypoxia-inducible factor 1α is mediated by an O2-dependent degradation domain via the ubiquitin-proteasome pathway. Proc Natl Acad Sci U S A 95:7987–7992
Hughes J, Criscuolo F (2008) Evolutionary history of the UCP gene family: gene duplication and selection. BMC Evol Biol 8:306
Hughes DA, Jastroch M, Stoneking M et al (2009) Molecular evolution of UCP1 and the evolutionary history of mammalian non-shivering thermogenesis. BMC Evol Biol 9:4
Iyer NV, Kotch LE, Agani F et al (1998a) Cellular and developmental control of O2 homeostasis by hypoxia-inducible factor 1 alpha. Genes Dev 12:149–162
Iyer NV, Leung SW, Semenza GL (1998b) The human hypoxia-inducible factor 1alpha gene: HIF1A structure and evolutionary conservation. Genomics 52:159–165
Jaakkola P, Mole DR, Tian YM et al (2001) Targeting of HIF-alpha to the von Hippel-Lindau ubiquitylation complex by O2-regulated prolyl hydroxylation. Science 292:468–472
Kimura M (1979) The neutral theory of molecular evolution. Sci Am 241:98–100, 102, 108 passim
Lynch VJ, Roth JJ, Wagner GP (2006) Adaptive evolution of Hox-gene homeodomains after cluster duplications. BMC Evol Biol 6:86
Makino Y, Cao R, Svensson K et al (2001) Inhibitory PAS domain protein is a negative regulator of hypoxia-inducible gene expression. Nature 414:550–554
Martinez-Castilla LP, Alvarez-Buylla ER (2003) Adaptive evolution in the Arabidopsis MADS-box gene family inferred from its complete resolved phylogeny. Proc Natl Acad Sci U S A 100:13407–13412
Maxwell PH, Wiesener MS, Chang GW et al (1999) The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 399:271–275
Nielsen R, Yang Z (1998) Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics 148:929–936
Nylander JAA (2004) MrModeltest v2. Program distributed by the author. Evolutionary Biology Centre, Uppsala University, Uppsala
Park JI, Semyonov J, Chang CL et al (2008) Origin of INSL3-mediated testicular descent in therian mammals. Genome Res 18:974–985
Patel SA, Simon MC (2008) Biology of hypoxia-inducible factor-2alpha in development and disease. Cell Death Differ 15:628–634
Peng J, Zhang L, Drysdale L et al (2000) The transcription factor EPAS-1/hypoxia-inducible factor 2alpha plays an important role in vascular remodeling. Proc Natl Acad Sci U S A 97:8386–8391
Raval RR, Lau KW, Tran MG et al (2005) Contrasting properties of hypoxia-inducible factor 1 (HIF-1) and HIF-2 in von Hippel-Lindau-associated renal cell carcinoma. Mol Cell Biol 25:5675–5686
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Ryan HE, Lo J, Johnson RS (1998) HIF-1 alpha is required for solid tumor formation and embryonic vascularization. EMBO J 17:3005–3015
Rytkonen KT, Ryynanen HJ, Nikinmaa M et al (2008) Variable patterns in the molecular evolution of the hypoxia-inducible factor-1 alpha (HIF-1alpha) gene in teleost fishes and mammals. Gene 420:1–10
Sanchez-Puig N, Veprintsev DB, Fersht AR (2005) Binding of natively unfolded HIF-1alpha ODD domain to p53. Mol Cell 17:11–21
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34:W609–W612
Taylor JS, Van de Peer Y, Braasch I et al (2001) Comparative genomics provides evidence for an ancient genome duplication event in fish. Philos Trans R Soc Lond B Biol Sci 356:1661–1679
Taylor JS, Braasch I, Frickey T et al (2003) Genome duplication, a trait shared by 22000 species of ray-finned fish. Genome Res 13:382–390
Thompson CE, Salzano FM, de Souza ON et al (2007) Sequence and structural aspects of the functional diversification of plant alcohol dehydrogenases. Gene 396:108–115
Tian H, Hammer RE, Matsumoto AM et al (1998) The hypoxia-responsive transcription factor EPAS1 is essential for catecholamine homeostasis and protection against heart failure during embryonic development. Genes Dev 12:3320–3324
Wagner A (2008) Neutralism and selectionism: a network-based reconciliation. Nat Rev Genet 9:965–974
Wang GL, Semenza GL (1995) Purification and characterization of hypoxia-inducible factor 1. J Biol Chem 270:1230–1237
Wong WS, Yang Z, Goldman N et al (2004) Accuracy and power of statistical methods for detecting adaptive evolution in protein coding sequences and for identifying positively selected sites. Genetics 168:1041–1051
Yang Z (2002) Likelihood and Bayes estimation of ancestral population sizes in hominoids using data from multiple loci. Genetics 162:1811–1823
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yang Z, Nielsen R (2002) Codon-substitution models for detecting molecular adaptation at individual sites along specific lineages. Mol Biol Evol 19:908–917
Zhang J, Nielsen R, Yang Z (2005) Evaluation of an improved branch-site likelihood method for detecting positive selection at the molecular level. Mol Biol Evol 22:2472–2479
Acknowledgments
This work is supported by the National High Technology Research and Development Program of China (863 project) (grant no. 2006AA10Z1E3), the National Natural Science Foundation of China (grant no. 30671492), and the National 973 Key Basic Research Program (grant no. 2006CB102102, 2004CB117502).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zhang, X., Wang, M., Tan, G. et al. Molecular selection and functional divergence of HIF-α proteins in vertebrates. Genetica 138, 1241–1250 (2010). https://doi.org/10.1007/s10709-010-9523-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-010-9523-3