Abstract
We have designed the user-friendly COS software with the intent to improve estimation of the probability of a family carrying a deleterious BRCA gene mutation. The COS software is similar to the widely-used Bayesian-based BRCAPRO software, but it incorporates improved assumptions on cancer incidence in women with and without a deleterious mutation, takes into account relatives up to the fourth degree and allows researchers to consider an hypothetical third gene or a polygenic model of inheritance. Since breast cancer incidence and penetrance increase over generations, we estimated birth-cohort-specific incidence and penetrance curves. We estimated breast and ovarian cancer penetrance in 384 BRCA1 and 229 BRCA2 mutated families. We tested the COS performance in 436 Italian breast/ovarian cancer families including 79 with BRCA1 and 27 with BRCA2 mutations. The area under receiver operator curve (AUROC) was 84.4 %. The best probability threshold for offering the test was 22.9 %, with sensitivity 80.2 % and specificity 80.3 %. Notwithstanding very different assumptions, COS results were similar to BRCAPRO v6.0.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Women with deleterious mutations in BRCA genes face an increased risk of breast cancer (BC) and ovarian cancer (OC), and often develop cancer at a young age. Estimates of the lifetime cumulative risk (penetrance) of BC associated with BRCA mutations range from about 80 % in earlier studies on high risk families [1–5], which were subject to ascertainment bias, to around 50–60 % in population-based or prospective studies [6–13]. Antoniou et al. [14] assembled information from 22 studies on families of index cases with BRCA1 or BRCA2 mutations, unselected for family history, and estimated a cumulative BC risk (0–70 years) of 65 % for those with BRCA1 mutations and 45 % for those with BRCA2 mutations. A meta-analysis of 10 studies that corrected for ascertainment bias showed the following cumulative risks to age 70 for mutation carriers: BC risk of 57 % for BRCA1 and 49 % for BRCA2, and OC risk of 40 % for BRCA1 and 18 % for BRCA2 [15].
Several genetic risk assessment methods are available to estimate the probability of BRCA mutation in individuals, in order to select them for molecular diagnosis [16]. Empirical methods, usually based on the number of BC and OC cases in the family, age at diagnosis, and occurrence in subsequent generations, ignore data from unaffected relatives [17–20], and may therefore grossly overestimate the probability of the mutation in large families with few affected members. By contrast, methods based on genetic models consider information from all relatives, whether they are affected or not. Berry et al. [21] and Parmigiani et al. [22] developed a method (BRCAPRO software), based on Bayes’ theorem, which required data on all first and second degree relatives of the family proband, and incorporated as prior probabilities incidence rates in the US population, allele mutation frequencies and penetrances estimated from studies in families with several BC or OC cases [4, 22, 23]. Recently, the BRCAPRO, available at http://bcb.dfci.harvard.edu/bayesmendel/brcapro.php, has been improved to include third degree relatives.
In the European case-only study (COS) on the role of gene-environment interactions in the development of BC in young women [24–27], we wanted to classify women developing BC before age 40 according to the probability that they carried a BRCA mutation, as estimated from family history. COS was carried out in seven countries with BC incidence ranging from 50 to 100 per 100,000 women per year (Estonia, France, Germany, Israel, Italy, Scotland and Slovenia) [28]. In such a context, the application of a Bayesian model requires country-specific assumptions of sporadic BC and OC incidence; however, BC incidence has increased over generations, both in the general population [29] and in mutation carriers [14, 30–32], and single age-specific incidence and penetrance curves do not accurately describe the disease risk in subsequent generations.
We have developed a computer program (COS software) to estimate the probability of carrying a deleterious BRCA mutation when incidence and penetrance are increasing over generations. The software is based on the same Bayesian logic as BRCAPRO, it is able to evaluate all third degree relatives, parents’ cousins and grandparents’ siblings (fourth degree), and it allows researchers to incorporate a hypothetical third BRCA gene to reduce overestimates of BRCA mutation probability due to the presence of other genes or gene combinations with similar penetrance.
Antoniou et al. [33] also developed a model (BOADICEA), available at http://ccge.medschl.cam.ac.uk/boadicea/, that takes into account a polygenic component beyond BRCA1 and BRCA2 and the increasing incidence of BC and OC over subsequent birth cohorts. BOADICEA also takes into account the incidence of other cancers, such as prostate and pancreas cancer [34].
Roudgari et al. [35] showed that the Scotland specific COS software compared favorably with BOADICEA in terms of sensitivity and AUROC (area under receiving operator curve), while BOADICEA showed better specificity.
We present here a new version of the COS software based on improved penetrance estimates of both BC and OC and compare its performance with BRCAPRO and BOADICEA.
Materials and methods
Study subjects and genetic testing
The present study is based on data from high risk families attending the Medical Genetics Unit of Milan National Cancer Institute (INT). Genetic counseling and testing was offered to all eligible families using widely accepted criteria based on the number of cases and ages at diagnosis (Table 1) [36, 37].
Data from 384 BRCA1 and 229 BRCA2 mutated families were used to calculate penetrance. An independent set of 436 consecutive families (those recruited between 2004 and 2008) was used for model validation. This set included 79 BRCA1 and 27 BRCA2 families, and 330 families tested negative for BRCA1 and BRCA2 genes. Families with variant of uncertain significance were excluded from the study.
BRCA gene mutation testing was performed either by denaturing high performance liquid chromatography (DHPLC) or by direct sequencing or by a combination of both methods examining all coding exons and corresponding splice sites of both genes. Individuals who tested negative at these analyses were investigated for the occurrence of large genomic rearrangements by multiple ligation-dependant probe amplification (MLPA), using commercially available kits (MRC-Holland). Families were considered as BRCA1 or BRCA2 mutation positive when genetic variants fulfilling one of the following criteria were ascertained. (a) Variants generating a premature stop codon (PTC), including nonsense mutations, small out-of frame insertions/deletions, splicing mutation confirmed by in vitro functional analyses, and large genomic rearrangements, with the exception of those introducing a PTC at or downstream BRCA2 codon 3326; (b) base pair changes, confirmed splicing mutations and genomic deletions leading to the loss of the translation start point; (c) confirmed splicing mutations and genomic deletions leading to the in-frame loss of exonic region coding for functional protein domains; (d) variants at the nearly invariant GT and AT dinucleotides at the 5′ and 3′ intron ends, which are predicted to affect mRNA splicing, even if not experimentally verified; (e) missense mutations and small in-frame deletions classified as pathogenic by multi-factorial probability based models [38]; (f) missense mutations affecting the highly conserved cysteine residues of the RING-finger domain of the BRCA1 protein. Families were considered as BRCA mutation negative when neither one of the above described genetic alterations nor variants of uncertain significance were ascertained.
All participants in this study signed an informed consent, approved by INT Ethic Committee, to the use of their biological samples and data for research purposes.
Mutation probability model
As shown by Parmigiani et al. [20] the probability P(M|H) of a person carrying a deleterious mutation (M) in a cancer gene, given a family history of cancer (H), is given by the Bayes theorem:
where f is the empirically-determined mutation prevalence. The expression implies that the probability P(M|H) depends on the penetrance in women with a mutation (M = 1) and on the incidence in women without a deleterious mutation (M = 0). The model is fully described in “Appendix 1”.
Model development
Estimation of incidence in women without deleterious BRCA mutations
We estimated country-specific general population BC incidence by birth cohort from cause-specific mortality data [39] and population-based cancer survival data [40, 41], using a mathematical model of the relationship between incidence, survival, mortality, and prevalence [42]. In this model, incidence was a polynomial function, with age, diagnosis period, and birth cohort as covariates. We produced BC incidence estimates for Estonia, Slovenia and Scotland, which were checked against national cancer registry data. We also produced estimates for Italy, France, and Germany, which were checked against registry data covering only part of the national populations [41]. In the present study, we are using the figures for Italy, where the cumulative lifetime BC risk increased from about 2–3 % for the women born at the turn of the 20th century, to 8–9 % for those born in the 1940 s, without any further increase subsequently [29, 43]. These estimates approximate the expected BC incidence in women carrying normal BRCA alleles. The only exception is the incidence in young women, where corrections are required, because a substantial proportion of cases in young women are attributable to BRCA mutations (see “Appendix 2”). Since male BC is rare, we used cross-sectional incidence curves estimated by pooling together 15 years of data from Italian cancer registries, without attempting any cohort-specific estimates.
As in Europe the change of OC incidence over 20th century generations has been much less dramatic than the change of BC [44], we tentatively incorporated into the software country-specific, cross-sectional age-specific incidence data from the cancer registries [28].
Incidence in women with a deleterious BRCA mutation
In order to estimate BC and OC penetrance in women with a deleterious BRCA mutation, we used pedigrees from 384 families with a BRCA1 mutation and 229 with a BRCA2 mutation. We also used 330 families which tested negative for both BRCA1 and BRCA2 to estimate the penetrance of a hypothetical third BRCA gene. For each of these sets, we simulated various age and birth cohort specific incidence and penetrance curves for BC and OC with the following model
where Ir is a 6-parameter curve given by the linear interpolation between incidence rates at ages 28, 35, 42, 49, 60, 75, and G is a 3-parameter curve given by the linear interpolation between the 1900, 1930 and 1960 cohorts. Ir is kept constant after age 75 and decreases exponentially before age 28, and G is kept constant before 1900 and after 1960.
We generated four functions (BC incidence, BC penetrance, OC incidence, OC penetrance), choosing for each function the set of 9 parameters that maximize the following
which expresses the probability of observing the histories of all families given the mutation in the probands. For the computation we used the conjugate gradient maximization method [45].
However, such functions could overestimate the incidence and penetrance curves, because families with numerous cases tend to be selected in preference to those with few cases. To partially correct for ascertainment, we assumed that the families would not have been selected for genetic testing before the occurrence of cancer in the proband. We therefore did not take into account the probands’ cancers.
Figure 1a–d show estimates of the cumulative risks of developing BC and OC in cohorts of women with a deleterious BRCA1 or BRCA2 mutation born in 1900, 1920, 1940, and 1960. For comparison, Fig. 1a–d also show the penetrance curves estimated by the meta-analysis by Chen and Parmigiani [15]. In the women carrying a BRCA1 mutation, estimates of cumulative risk of BC to age 70 increased from 35 % in the women born in 1900 to 58 % in the women born in 1960. The corresponding penetrances for BRCA2 mutation carriers were 38 and 63 %. OC 70-year penetrance estimates also increased dramatically from 9 to 47 % for BRCA1 and from 7 to 26 % for BRCA2. These penetrances are within the literature ranges for BRCA mutations [14], and closely similar to recently estimated cumulative risks in large US population-based studies [46, 47].
Breast and ovarian cancer penetrance estimates: cumulative risk for successive birth cohorts of women carrying a BRCA1 or BRCA2 mutation. The risk curve derived by Chen and Parmigiani [15] is also shown. a BRCA1, breast cancer, b BRCA2, breast cancer. c BRCA1, ovarian cancer, d BRCA2, ovarian cancer
Incidence in eligible women who tested negative for BRCA mutations
We also estimated the penetrance curves for BC and OC in the 330 eligible families which tested negative for BRCA1 and BRCA2 mutations. The method of penetrance estimation was the same as the method used for mutated families. These estimates were incorporated into the model as if these families carried a deleterious mutation in a third hypothetical BRCA gene (BRCA3). Due to eligibility criteria, in fact, these families have a higher incidence than the general population, whether due to genetic and/or environmental factors, which must be taken into account. If the BRCA1/BRCA2 negative families had the same incidence as the general population, the assumption of a BRCA3 condition would not be necessary to discriminate BRCA1 and BRCA2 families from the other families in the eligible set. We are aware that such a hypothetical BRCA3 gene is unlikely to exist, and that the excess incidence in these families is more likely due to the interaction of several low-penetrance alleles, or to moderate/high penetrance mutations in a number of distinct non-BRCA genes [48]. However, for our purposes the real nature of the trait is irrelevant.
Allele frequencies
Allele frequencies can be set manually in the COS software. The allele frequencies for the Italian population are not known. We tentatively set the default frequencies at 0.0006 for BRCA1, and 0.0002 for BRCA2, as is widely used [49]. This BRCA1/BRCA2 ratio is consistent with the ratio of BRCA1 and BRCA2 mutations in our validation data set (79/27). However, we simulated the effect of considering other frequencies. Increasing the BRCA2/BRCA1 ratio up to 1, as suggested by recent allele frequency estimates [50, 51]. did not materially change the results.
Claus et al. [52] estimated that the genotype frequency of all mutated BC genes is 0.0033, that is to say allele frequency 0.00165. We used these estimates to hypothesize a BRCA3 allele frequency of 0.00165 − 0.0008 = 0.00085.
Software use
See “Appendix 3”.
Model evaluation
Performance of COS and comparison with other models
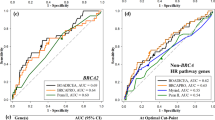
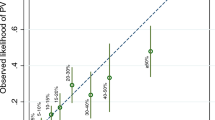
We examined the performance of COS and other programs in predicting mutations in a validation set of 436 Italian high-risk families. We calculated the area under the receiver operator characteristic curve (AUROC). ROC curves are generated by plotting sensitivity against 1-specificity, considering various probability levels as separating negative from positive predictions. The closer the AUROC is to 1, the better the software is at estimating correct outcomes. We calculated 95 % confidence intervals (CIs) for AUROC according to DeLong et al. [53]. We used the best probability threshold obtained from ROC curve to compute sensitivity and specificity. In the ROC curve, where each point is obtained by calculating the sensitivity and the specificity for a defined threshold, the best threshold is the point that minimizes the quantity (1-specificity)2 + (1-sensitivity)2 [48].
We also computed the Brier score (mean of the squared difference between each probability prediction and the genetic test outcome, which can be either 1 when positive or 0 when negative). A Brier score of 0 indicates perfect prediction; the worst possible score is 1.
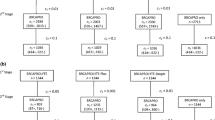
We then compared the AUROC, the Brier score, the sensitivity and the specificity obtained by the COS software with those obtained by BRCAPRO v5.1, BRCAPRO v6.0 and BOADICEA v3.0 (Fig. 2).
Each software has a specific threshold that maximizes his discrimination power. Therefore, to compare the validity of different software it is necessary to use their specific best thresholds. Otherwise, choosing a single threshold, it may occur that a software shows a better performance only because his best threshold is the nearest to the chosen one. As medical genetics units usually refer to a specific threshold, e.g. 10 % to select families for genetic testing we also show how the sensitivity and specificity change as a function of the chosen threshold (Fig. 3).
Results and discussion
We estimated the penetrance of BRCA mutation in a large set of Italian families attending genetic counseling and eligible to BRCA testing (Fig. 1). Our results could be useful for genetic daily practice in high risk families, helping geneticists and women in the complex decision process about preventive options. Overall, our penetrance estimates are in line with the ones previously published; however, they allow a more accurate personal risk definition for women belonging to different birth cohorts.
The validation set included 436 consecutive eligible index cases (416 females and 20 males): 320 women developed BC (median age 39 years, range 21–80 years), 28 OC (median age 50.5 years, range 27–77 years), and 28 both BC and OC (BC median age 47.5 years, range 30–72 years; OC median age 55 years, range 36–76 years); 13 men developed BC (median age 56 years, range 34–73 years). In 47 families, individuals affected by BC or OC were not available for testing, thus the analysis was carried out on their closest relatives.
Among the 436 families, 319 included BC cases only (41 with one early onset BC and negative family history), 15 OC cases only (one with early onset OC and negative family history) and 102 both BC and OC cases (10 with both BC and OC in the index patient and negative family history). Overall, this set included 1171 BC cases, 135 OC cases and 53 cases with both BC and OC out of 7,580 women; 33 BC cases were recorded among 6,789 men.
In the validation set of 436 families we found 106 deleterious mutations (79 BRCA1 and 27 BRCA2) corresponding to a frequency of mutated genotype of 24.3 % (95 % CI 19.6–28.9). The COS software average estimate of mutation probabilities was 24.7 %. The Brier score was 0.125 and the AUROC 0.845 (CI95 % 0.764–0.924). The best probability threshold to discriminate the BRCA1 and BRCA2 mutated families from the non-mutated ones was 22.9 %, which corresponds to a sensitivity of 80.3 % and a specificity of 80.2 %. Table 2 shows the corresponding values obtained with BRCAPRO v5.1, BRCAPRO v6.0 and BOADICEA. COS and BRCAPRO v6.0 had almost the same performance, superior to BRCAPRO v5.1 and BOADICEA for all performance indexes. The improvement of the performance of BRCAPRO v6.0 with respect to version 5.1 is likely to be due to the fact that version 6.0 allows the software to take into account third degree relatives. As all the methods are heavily dependent on the validity of the incidence and penetrance assumptions, it is most likely that the BOADICEA assumptions do not fit the Italian population so well. BOADICEA and BRCAPRO include the information on BC phenotype (hormone receptor expression), and BOADICEA allows researchers to take into account the occurrence of other BRCA associated cancers (prostate and pancreas). We have not taken into account these phenotypes, because in our database the information was sparse and COS has not yet developed these functions.
Figure 3 shows the performance of COS, BRCAPRO 6.0 and BOADICEA as a function of the threshold chosen for selecting families for BRCA testing. For the usually chosen threshold of 10 % the performance of COS and BRCAPRO 6.0 is the same (COS sensitivity 87.7 and specificity 63.2; BRCAPRO 6.0 sensitivity 87.7 and specificity 63.3), while BOADICEA shows a much lower sensitivity (71.5) and a somewhat higher specificity (70.6). For a threshold of 20 % the performance of COS would be better than BRCAPRO 6.0.
All programs predict BRCA1 and BRCA2 mutations separately. However their ability to discriminate is limited, because the assumed penetrance of the two mutated genes is fairly similar. Therefore we are only presenting the performance indicators for both genes combined.
The BRCAPRO software developed by Berry and Parmigiani [21, 22] substantially improved previous empirical approaches based on the number and ages of affected relatives. However, using BRCAPRO up to version 5.1, some families with several young cancer cases showed low mutation probabilities, because distant relatives with cancer could not be accommodated in the software. As already mentioned, version 6.0 dramatically improved the performance (Table 2), most likely because it allows researchers to take into account more distant relatives. Nevertheless, we would have expected a better performance of the COS software with respect to BRCAPRO, because the latter does not take into account the evolving cancer incidence and penetrance over subsequent generations. Moreover, we expected an overestimation of the mutation probability by BRCAPRO, because of the absence of the third gene in the model. Nevertheless, BRCAPRO v6.0 provides good estimates, most likely because of the following factors, which lower the estimation of the mutation probability:
-
(a)
the single curve for incidence incorporated in BRCAPRO v6.0 overestimates the expected BC incidence in the women belonging to older generations, when BC incidence was low. This causes an underestimation of the mutation probability.
-
(b)
BRCAPRO incorporates very low penetrance estimates, much lower than those estimated by the Chen and Parmigiani meta-analysis of studies without ascertainment bias, with BC cumulative risks at 70 years of age = 43 % for BRCA1 and 32 % for BRCA2 (vs. 57 and 49 % in the meta-analysis), and OC cumulative risks = 30 % for BRCA1 and 15 % for BRCA2 (vs. 40 and 18 %).
-
(c)
BRCAPRO v6.0 seems to give a very low weight to contralateral BC.
All the models share the limitations that they use penetrance estimates, and also published allele frequency estimates, which may be inaccurate. The relatively poor performance of BOADICEA on our Italian families highlights the importance of the availability of population specific incidence and/or penetrance estimates. Several approaches have been used to estimate the average penetrance associated with BRCA mutations. Earlier estimates applied the maximum-LOD-score method to multiple-case families collected for linkage studies for the identification of disease loci [3, 4]. This method provided dramatically overestimated life-long cumulative risk estimates, of the order of 80 % by age 70 for BC. A few years later, however, Struewing et al. [9], in a population based study of 5318 Ashkenazi Jews, unselected for family history, showed that the penetrance is likely to be much lower. By comparing the cancer histories of relatives of carriers and non-carriers of the three founder mutations, he estimated that the risk of BC by the age of 70 was 56 %. The major limitation of studies based on cases unselected for family history in other populations, in which mutations are almost spread all over the two genes, is their small size and, therefore, the imprecision of estimates. Antoniou et al. [14], however, pooled 22 studies, half population based and half selected on the basis of the young age of the proband. Mothers and sisters of the proband were assumed to be followed from age 20 years and were censored at the age of first cancer diagnosis: the resulting estimates for BC risk to the age of 70 years were 65 % for BRCA1 and 45 % for BRCA2. The estimated penetrance of BRCA1, however, was higher if the studied families were selected for the young age of the proband, suggesting some residual ascertainment bias (mutations associated with early age of cancer may confer higher lifetime cumulative risks). The recent meta-analysis of Chen and Parmigiani [15], who pooled population based studies and studies that corrected for ascertainment bias, gave somewhat lower estimates for BRCA1 (57 %), and similar results (49 %) for BRCA2 mutation carriers. An excellent method to estimate penetrance is to prospectively follow-up healthy mutation carriers, as was made in the EMBRACE study [13], with a 10-year follow-up of 988 mutation carriers without a previous diagnosis of breast or ovarian cancer: the average BC cumulative risks by age 70 years were estimated to be 60 % for BRCA1 and 55 % for BRCA2 carriers. These results cannot be influenced by the previous familial cancer history, but, still, may be somewhat overestimated, because in the studied families, selected for family history, high penetrance mutations may have been overrepresented. There is increasing evidence, in fact, that different mutations may confer different risk. We estimated penetrance from families selected for young age of the proband or the presence of several cases in the family, i.e. from a series biased for higher penetrance. We are confident, however, that the exclusion of the proband’s cancer from the computation provided fairly good correction of ascertainment bias. As shown in Fig. 1, in fact, our estimates are quite in line with those of Cheng and Parmigiani [15], and the increasing penetrance over subsequent birth cohorts is of the same order of the increasing relative risk estimated by Antoniou et al. [14].
Nevertheless, whatever the validity of the estimates, there is an intrinsic limitation in the discriminating power of the models, which depends on differences in phenotype incidence. When the difference in disease incidence between women with and without mutation is large, the power is high. Similarly, if penetrance is low and if it approaches the incidence of sporadic cancer, the discriminating power is reduced. It is because BC penetrance estimates for BRCA1 and BRCA2 are so similar that reliable discrimination between them is actually difficult.
The COS software is available for free on request to patrizia.pasanisi@istitutotumori.mi.it.
The COS risk prediction model demonstrated high performance indexes and can be a useful risk stratification tool in research studies. At present, however, the use of prediction models and strict probability thresholds in clinical practice protocols still present evident limitations. To select families eligible for BRCA testing, models should not be used alone and genetic counseling should always be provided.
References
The Breast Cancer Linkage (1999) Cancer risks in BRCA2 mutation carriers. J Natl Cancer Inst 91:1310–1316
Brose MS, Rebbeck TR, Calzone KA, Stopfer JE, Nathanson KL, Weber BL (2002) Cancer risk estimates for BRCA1 mutation carriers identified in a risk evaluation program. J Natl Cancer Inst 94:1365–1372
Easton DF, Bishop DT, Ford D, Crockford GP (1993) Genetic linkage analysis in familial breast and ovarian cancer: results from 214 families. The breast cancer linkage consortium. Am J Hum Genet 52:678–701
Ford D, Easton DF, Stratton M et al (1998) Genetic heterogeneity and penetrance analysis of the BRCA1 and BRCA2 genes in breast cancer families. The breast cancer linkage consortium. Am J Hum Genet 62:676–689
Narod S, Ford D, Devilee P et al (1995) Genetic heterogeneity of breast-ovarian cancer revisited. Breast cancer linkage consortium. Am J Hum Genet 57:957–958
Bonadona V, Sinilnikova OM, Chopin S et al (2005) Contribution of BRCA1 and BRCA2 germ-line mutations to the incidence of breast cancer in young women: results from a prospective population-based study in France. Genes Chromosomes Cancer 43:404–413
Hopper JL, Southey MC, Dite GS et al (1999) Population-based estimate of the average age-specific cumulative risk of breast cancer for a defined set of protein-truncating mutations in BRCA1 and BRCA2. Australian breast cancer family study. Cancer Epidemiol Biomarkers Prev 8:741–747
Loman N, Bladstrom A, Johannsson O, Borg A, Olsson H (2003) Cancer incidence in relatives of a population-based set of cases of early-onset breast cancer with a known BRCA1 and BRCA2 mutation status. Breast Cancer Res 5:R175–R186
Struewing JP, Hartge P, Wacholder S et al (1997) The risk of cancer associated with specific mutations of BRCA1 and BRCA2 among Ashkenazi Jews. N Engl J Med 336:1401–1408
Risch HA, McLaughlin JR, Cole DE et al (2006) Population BRCA1 and BRCA2 mutation frequencies and cancer penetrances: a kin-cohort study in Ontario, Canada. J Natl Cancer Inst 98:1694–1706
Thorlacius S, Struewing JP, Hartge P et al (1998) Population-based study of risk of breast cancer in carriers of BRCA2 mutation. Lancet 352:1337–1339
Anglian Breast Cancer Study Group (2000) Prevalence and penetrance of BRCA1 and BRCA2 mutations in a population-based series of breast cancer cases. Br J Cancer 83:1301–1308
Mavaddat N, Peock S, Frost D et al (2013) Cancer risks for BRCA1 and BRCA2 mutation carriers: results from prospective analysis of EMBRACE. J Natl Cancer Inst 105:812–822
Antoniou A, Pharoah PD, Narod S et al (2003) Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case Series unselected for family history: a combined analysis of 22 studies. Am J Hum Genet 72:1117–1130
Chen S, Parmigiani G (2007) Meta-analysis of BRCA1 and BRCA2 penetrance. J Clin Oncol 25:1329–1333
Antoniou AC, Easton DF (2006) Models of genetic susceptibility to breast cancer. Oncogene 25:5898–5905
Claus EB, Risch N, Thompson WD (1994) Autosomal dominant inheritance of early-onset breast cancer. Implications for risk prediction. Cancer 73:643–651
Federico M, Maiorana A, Mangone L et al (1999) Identification of families with hereditary breast and ovarian cancer for clinical and mammographic surveillance: the Modena Study Group proposal. Breast Cancer Res Treat 55:213–221
Frank TS, Deffenbaugh AM, Reid JE et al (2002) Clinical characteristics of individuals with germline mutations in BRCA1 and BRCA2: analysis of 10,000 individuals. J Clin Oncol 20:1480–1490
Shattuck-Eidens D, Oliphant A, McClure M et al (1997) BRCA1 sequence analysis in women at high risk for susceptibility mutations. Risk factor analysis and implications for genetic testing. JAMA 278:1242–1250
Berry DA, Parmigiani G, Sanchez J, Schildkraut J, Winer E (1997) Probability of carrying a mutation of breast-ovarian cancer gene BRCA1 based on family history. J Natl Cancer Inst 89:227–238
Parmigiani G, Berry D, Aguilar O (1998) Determining carrier probabilities for breast cancer-susceptibility genes BRCA1 and BRCA2. Am J Hum Genet 62:145–158
Andersen TI (1996) Genetic heterogeneity in breast cancer susceptibility. Acta Oncol 35:407–410
Berrino F, Pasanisi P, Berrino J, Curtosi P, Bellati C (2002) A European case-only study on familial breast cancer. IARC Sci Publ 156:63–65
Pasanisi P, Berrino J, Fusconi E, Curtosi P, Berrino F (2005) A European case-only study (COS) on familial breast cancer. J Nutr 135:3040S–3041S
Pasanisi P, Hédelin G, Berrino J et al (2009) Oral contraceptive use and BRCA penetrance: a case-only study. Cancer Epidemiol Biomarkers Prev 18:2107–2113
Pasanisi P, Bruno E, Manoukian S, Berrino F (2013) A randomized controlled trial of diet and physical activity in BRCA mutation carriers. Fam Cancer. doi:10.1007/s10689-013-9691
Parkin D, Whelan SL, Ferlay J (2002) Cancer incidence in five continents. IARC scientific publication no. 155. IARC Press, Lyon
Capocaccia R, Verdecchia A, Micheli A, Sant M, Gatta G, Berrino F (1990) Breast cancer incidence and prevalence estimated from survival and mortality. Cancer Causes Control 1:23–29
Chang-Claude J, Becher H, Eby N, Bastert G, Wahrendorf J, Hamann U (1997) Modifying effect of reproductive risk factors on the age at onset of breast cancer for German BRCA1 mutation carriers. J Cancer Res Clin Oncol 123:272–279
Narod SA, Goldgar D, Cannon-Albright L et al (1995) Risk modifiers in carriers of BRCA1 mutations. Int J Cancer 64:394–398
Tryggvadottir L, Sigvaldason H, Olafsdottir GH et al (2006) Population-based study of changing breast cancer risk in Icelandic BRCA2 mutation carriers, 1920–2000. J Natl Cancer Inst 98:116–122
Antoniou AC, Pharoah PP, Smith P, Easton DF (2004) The BOADICEA model of genetic susceptibility to breast and ovarian cancer. Br J Cancer 91:1580–1590
Lee AJ, Cunningham AP, Kuchenbaecker KB, Mavaddat N, Easton DF, Antoniou AC (2014) BOADICEA breast cancer risk prediction model: updates to cancer incidences, tumour pathology and web interface. Br J Cancer 110:535–545
Roudgari H, Miedzybrodzka ZH, Haites NE (2008) Probability estimation models for prediction of BRCA1 and BRCA2 mutation carriers: COS compares favourably with other models. Fam Cancer 7:199–212
Manoukian S, Peissel B, Pensotti V et al (2007) Germline mutations of TP53 and BRCA2 genes in breast cancer/sarcoma families. Eur J Cancer 43:601–606
Radice P (2002) Mutations of BRCA genes in hereditary breast and ovarian cancer. J Exp Clin Cancer Res 21:9–12
Lindor NM, Guidugli L, Wang X et al (2012) A review of a multifactorial probability-based model for classification of BRCA1 and BRCA2 variants of uncertain significance (VUS). Hum Mutat 33:8–21
Verdecchia A, Capocaccia R, Egidi V, Golini A (1989) A method for the estimation of chronic disease morbidity and trends from mortality data. Stat Med 8:201–216
Berrino F, Capocaccia R, Esteve J et al (1999) Survival of cancer patients in Europe: the EUROCARE-2 study. IARC scientific publications no. 151, Lyon
Berrino F, Capocaccia R, Coleman P et al (2003) Survival of cancer patients in Europe: the EUROCARE-3 study. Ann Oncol 14:v1–v155
De Angelis G, De Angelis R, Frova L, Verdecchia A (1994) MIAMOD: a computer package to estimate chronic disease morbidity using mortality and survival data. Comput Methods Programs Biomed 44:99–107
Micheli A, Verdecchia A, Capocaccia R et al (1992) Estimated incidence and prevalence of female breast cancer in Italian regions. Tumori 78:13–21
Gonzalez-Diego P, Lopez-Abente G, Pollan M, Ruiz M (2000) Time trends in ovarian cancer mortality in Europe (1955–1993): effect of age, birth cohort and period of death. Eur J Cancer 36:1816–1824
Press WH, Tekolski SA, Vetterling WT, Flannery BP (1992) Numerical Recipes in C: the art of scientific computing, 2edn. Cambridge University Press, Cambridge
Chen S, Iversen ES, Friebel T et al (2006) Characterization of BRCA1 and BRCA2 mutations in a large United States sample. J Clin Oncol 24:863–871
Begg CB, Haile RW, Borg A et al (2008) Variation of breast cancer risk among BRCA1/2 carriers. JAMA 299:194–201
Eccles SA, Aboagye EO, Ali S et al (2013) Critical research gaps and translational priorities for the successful prevention and treatment of breast cancer. Breast Cancer Res 15:R92
Ford D, Easton DF, Peto J (1995) Estimates of the gene frequency of BRCA1 and its contribution to breast and ovarian cancer incidence. Am J Hum Genet 57:1457–1462
National Cancer Institute. (2013) Genetics of breast and ovarian cancer (PDQ). http://www.cancer.gov/cancertopics/pdq/genetics/breast-and-ovarian/HealthProfessional/page2
Malone KE, Daling JR, Doody DR et al (2006) Prevalence and predictors of BRCA1 and BRCA2 mutations in a population-based study of breast cancer in white and black American women ages 35–64 years. Cancer Res 66:8297–8308
Claus EB, Risch N, Thompson WD (1991) Genetic analysis of breast cancer in the cancer and steroid hormone study. Am J Hum Genet 48:232–242
DeLong ER, DeLong DM, Clarke-Pearson DL (1988) Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44:837–845
Perkins NJ, Schisterman EF (2006) The inconsistency of “optimal” cutpoints obtained using two criteria based on the receiver operating characteristic curve. Am J Epidemiol 163:670–675
Acknowledgments
The study was financed by the 6th Framework Program of the European Community and by funds from Italian citizens who allocated the 5 × 1,000 share of their tax payment in support of the Fondazione IRCCS Istituto Nazionale dei Tumori, according to Italian laws (INT-Institutional strategic projects ‘5 × 1,000’). We also thank Dr Maria Teresa Penci for help with the language and Dr Giulietta Scuvera for data collection.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix 1
We have modified the original method of Parmigiani et al. [22] in three different points. First, we have included third degree relatives plus the grandparent sibling offspring (IV degree); second, we use age and birth cohort specific incidence and penetrance; third, we have incorporated the possibility of taking into account an hypothetical third gene. We present here the formulas respecting as far as possible the notation that Parmigiani used in his article.
For each family member b, the contribution to the likelihood function depends on the age a b , the genetic status of BRCA1i, of BRCA2j, of BRCA3k, and on the disease status. As Parmigiani did, we factorize the conditional probability ρ ijk b of observing the disease history of member b, given the genetic status ijk, into two disease-specific term, one for breast cancer and one for ovary cancer (notice that at this point we can add other phenotypes).
where the meaning of indexes is: family member b, O = ovary, B = breast and
the ρ are defined in the following formula
where R ijk(a, g) is the cumulative risk at age a for the generation g and r ijk(a, g) is the age and birth cohort specific incidence.
To rewrite the formula used by Parmigiani in the case of 3 genes and for III degree relatives, we assign a label b to every family member and we define the following functions
m(b) | returns the label of the mother of b |
f(b) | returns the label of the father of b |
no(b) | returns the number of sons and daughters of b |
o l (b) l = 1, …, no(b) | returns the label of person l belonging to the offspring of b |
q(b) | returns the label of the mate of b |
ns(b) | returns the number of brothers and sisters of b |
s l (b) | l = 1, …, ns(b) returns the label of person l belonging to the sibling of b |
For instance, the label of the paternal grandmother of b will be f(m(b)).
We use the following notation to express multi-summation symbol
We rewrite the Parmigiani function d(m 1, m 2, f 1, f 2, n) in the case of three genes
It represents the probability of observing a determined clinical history in the offspring of b [formed by no(b) individuals] given the allele configuration (m 1, m 2, m 3) in the mother and (f 1, f 2, f 3) in the father (the temporary proband b could be either the mother or the father label). The function P[i|m, f] represents the probability to inherit i mutated alleles given m mutated alleles in the mother and f in the father following mendelian rules of gene transmission.
We can extend the d function in order to take into account two generations (sons and nephew) by defining
where \( \rho_{{q(o_{l} (b))}}^{{i_{q} j_{q} k_{q} }} \) is related to offspring mates (i.e. sons and daughter in law), if no information is available we consider ρ ijk q() = 1 for every ijk; the function P[i q , j q , k q ] represents the probability of the mate to have a certain allele configuration and it is directly related to the prevalence of the three mutations, that is
with
where f a is the allele frequency for gene BRCA a .
In case of three generations we can define the function
which incorporates the d2 function, and so on for further generations. Now we have solved how to “move down” in the family.
In order to write the functions to “move up” in the family we need to define the following function U b (o1, o2, o3) that represents the probability of observing the history of the parents and of the siblings of family member b given his gene status (o1, o2, o3)
The function P[i f , i m |o], used in the latter, represents the probability to find i f mutated alleles in the father and i m in the mother given o mutated alleles in the son or daughter. It is defined as follow using the Bayes theorem.
We have used the function ds where only the siblings of b (temporary proband) are taken into account;
It represents the probability of observing the history of siblings of b and of their offspring (up to two generations) given the genetic status of parents (m 1, m 2, m 3) and (f 1, f 2, f 3)
Note that in this case, even if the structure is the same of function d3, the label b is not referred to the parents as it was before but to temporary proband.
With the U function we can “move up” of one generation, but we want reach the great-grandparents so we need functions to do two more steps. We use U function to move from grandparents to great-grandparents and grandparents siblings; The U2 function defined by
is used to move from parents to grandparents, aunts and uncles and the function U3 defined by
is used to move from the proband to parents and proband’ s sibling.
Finally, the probability of observing the clinical family history given the genetic status (i, j, k) of the proband is
where ρ ijk p is refers to the proband personal clinical history, i m j m k m are refer to the genetic status of the proband’ s mate and ρ ijk q(p) to his history.
Finally according to the Bayes theorem the probability of observing mutation status [BRCA1, BRCA2, BRCA3] given the family history can be written
Appendix 2
The observed incidence rates in the general population ir all can be written as the sum of two or terms: the incidence rates for women carrying wild-type BRCA gene ir 0 time the prevalence of wild-type and the incidence rates ir 1 for women carrying BRCA mutation times the prevalence of mutation
where f is the allele frequency, (1 − f)2 is the wild-type prevalence and (f 2 + 2f(1 − f)) is the mutation prevalence. Because of the very low allele frequency we can approximate the latter by omitting the second order terms f 2.
In the case of three genes the latter becomes
The incidence rates for the wild-type BRCA is then
For instance we have simulated how mutation probability changes in a proband with breast cancer at different ages using respectively ir all and ir 0
Age | Mut. prob. using ir all | Mut. prob. using ir 0 |
|---|---|---|
20 | 0.452 | 0.837 |
25 | 0.243 | 0.323 |
30 | 0.141 | 0.166 |
40 | 0.071 | 0.077 |
50 | 0.033 | 0.035 |
60 | 0.028 | 0.030 |
70 | 0.015 | 0.016 |
We can note that the correction is important for younger ages when there is a considerable proportion of genetic cases.
Appendix 3
Software input
To estimate BRCA mutation probability, the COS software requires sex, current age or age at death, age at BC or OC diagnosis, age at diagnosis of second or contralateral BC, and age at any bilateral prophylactic mastectomy or oophorectomy in each family member.
The results of genetic testing can also be included (allowing researchers to calculate the mutation probability in untested family members).
These data are also required by BRCAPRO. Unlike BRCAPRO, however, the COS software additionally requires the date of birth of each family member (to allow for changing incidence and penetrance over generations).
Since data on the distant relatives of a proband are frequently incomplete, the COS software incorporates routines for the automatic imputation of missing information. These routines incorporate the following assumptions: that the interval between generations is 25 years, that BC and OC deaths occur 5 and 2 years, respectively, after diagnosis, and that life expectancy has increased from 65 years for people born before 1920 to 75 years for those born afterwards. A routine to automatically import family data from Progeny is available at http://www.progenygenetics.com/.
Rights and permissions
About this article
Cite this article
Berrino, J., Berrino, F., Francisci, S. et al. Estimate of the penetrance of BRCA mutation and the COS software for the assessment of BRCA mutation probability. Familial Cancer 14, 117–128 (2015). https://doi.org/10.1007/s10689-014-9766-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10689-014-9766-8