Summary
Background Increased serum levels of soluble interleukin-2 (IL-2) receptor alpha (sIL-2Rα) are an indicator of poor prognosis in patients with B-cell non-Hodgkin lymphoma (NHL). By binding to IL-2, sIL-2Rα upregulates Foxp3 expression and induces the development of regulatory T (Treg) cells. Methods To inhibit the binding of IL-2 to sIL-2Rα with the goal of suppressing the induction of Foxp3 and decreasing Treg cell numbers, we developed peptides by structure-based computational design to disrupt the interaction between IL-2 and sIL-2Rα. Each peptide was screened using an enzyme-linked immunosorbent assay (ELISA), and 10 of 22 peptides showed variable capacity to inhibit IL-2/sIL-2Rα binding. Results We identified a lead candidate peptide, CMD178, which consistently reduced the expression of Foxp3 and STAT5 induced by IL-2/sIL-2Rα signaling. Furthermore, production of cytokines (IL-2/interferon gamma [IFN-γ]) and granules (perforin/granzyme B) was preserved in CD8+ T cells co-cultured with IL-2–stimulated CD4+ T cells that had been pretreated with CMD178 compared to CD8+ cells co-cultured with untreated IL-2–stimulated CD4+ T cells where it was inhibited. Conclusions We conclude that structure-based peptide design can be used to identify novel peptide inhibitors that block IL-2/sIL-2Rα signaling and inhibit Treg cell development. We anticipate that these peptides will have therapeutic potential in B-cell NHL and other malignancies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Interleukin (IL-2) is a T-cell growth factor that plays a key role in T-cell homeostasis. The IL-2 receptor (IL-2R) consists of 3 different subunits: alpha, beta, and gamma; however, IL-2Rα can be cleaved by proteolysis to produce a truncated and soluble form of IL-2Rα (sIL-2Rα). It has been shown that serum sIL-2Rα levels are elevated and correlate with poor survival in a variety of cancer types [1,2,3,4].
Regulatory T (Treg) cells are the gateway to immune function and typically regulate immune cell activation. While a variety of mechanisms are involved in inducing Treg cells, IL-2 is essential in the development of Treg cells. Treg cells cannot survive or expand in the absence of IL-2 either in the thymus or in the periphery. IL-2 generally induces T-cell differentiation and promotes a regulatory phenotype. Once activated via the IL-2R, a cascade of events in T cells initiate the signal transducer and activator of transcription 5 (STAT5) and forkhead box p3 (Foxp3) activation, which appear to function as important regulators of this immunologic pathway and promote the development and function of Treg cells [5,6,7].
In non-Hodgkin lymphoma (NHL), we have found that increased numbers of intratumoral Treg cells are present and suppress immune function in the tumor microenvironment [8]. We have also shown that serum levels of soluble sIL-2Rα are increased in lymphoma patients and correlate with an inferior clinical outcome [9, 10]. Shed by activated T cells that express membrane-bound IL-2Rα, sIL-2Rα contributes to the upregulation of Foxp3 expression in CD4+ T cells. Soluble IL-2Rα promotes the development of Treg cells by binding to IL-2 and the IL-2/IL-2Rα–complex signals via STAT5 [11, 12]. We hypothesized that blocking the binding of IL-2 to sIL-2Rα would prevent the induction of a regulatory T-cell phenotype. This would be associated with downregulation of STAT5 and Foxp3 expression, and subsequent promotion of T-cell effector function, including the upregulation of cytokines such as tumor necrosis factor alpha (TNF-α) and interferon gamma (INF-γ), and secretion of cytotoxic granules, including granzyme B and perforin [13].
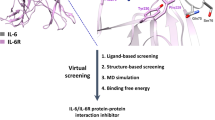
Importantly, there is interest in developing peptide drugs, particularly to block protein-protein interactions like those between IL-2 and sIL-2Rα. Moreover, a compelling case can be made for using physics-based and data-based computational methods to enable the structure-based design of novel peptide drug candidates. In this study, we therefore used a structure-based design and computational tools to design blocking peptides to inhibit the binding of IL-2 to sIL-2Rα and tested the capacity of the peptides to regulate the development of Treg cells and modulate T-cell function.
Methods
Development of the IL-2/sIL-2Rα-competitive binding assay
In order to demonstrate inhibition of binding of IL-2 to sIL-2Rα, we developed an enzyme-linked immunosorbent assay (ELISA)–based assay to determine the binding capacity between these 2 molecules. Briefly, a 96-well plate was coated with sIL-2Rα protein (PeproTech) at a concentration of 500 ng/mL in standard coating bicarbonate/carbonate buffer (100 mM). Following overnight incubation, the plate mixture was washed to remove the excess protein. Next, IL-2 protein (1 nM) was added to the plate and incubated for 1 h. After washing, biotinylated IL-2 antibody (PeproTech) was added to detect the bound IL-2 and incubated for 30 min, then washed. Next, avidin-HRP conjugate (PeproTech) was added to the wells, incubated for 30 min, then washed. Finally, 1-Step Turbo TMB-ELISA Substrate Solution (Thermo Fisher Scientific) was added to each well for 15 min, then stopped with 0.18 M H2SO4. The plate was analyzed for IL-2 expression using SpectraMax190 (Molecular Devices) and read at absorbance 450 nm. Each peptide was run in triplicate at concentrations of 0.0, 0.1, 1.0, and 10 μM. An IL-2 expression curve was calculated using the absorbance reading. The ELISA was used to create a competitive assay between IL-2 and each of the peptides to assess whether they inhibited binding between IL-2 and sIL-2Rα. An IL-2 antibody (MAB202; R&D Systems) was added as a positive control at a final concentration of 75 ng/μl. Each peptide (100 mM) was mixed with IL-2 (1 nM), incubated for 30 min at room temperature (RT), and then added to an IL-2Rα-coated plate. The mix was incubated for 1 h at RT and the solution was gently removed. The plates were washed 3 times with cell-washing buffer and developed and read using the same conditions as in the previous experiment.
Flow cytometry for STAT5
Cells were fixed and permeabilized with reagents from a Foxp3 staining kit (BioLegend). Cells were then stained with fluorochrome-conjugated antibody against Foxp3 plus fluorochrome-conjugated anti-CD4 antibody for 30 min and analyzed by flow cytometry. Acquired data were analyzed by using FlowJo (Tree Star, Ashland, OR). Phosphorylation of STAT5 was determined by using flow-based intracellular staining following the instructions described by the manufacturer (BD Biosciences, San Jose, CA). Briefly, freshly-enriched CD4+ T cells were incubated with IL-2, sIL-2Rα, or both, in combination for 30 min in a 37 °C water bath. Cells were subjected to fixation and permeabilization, stained with fluorochrome-conjugated STAT5 antibody, and analyzed by flow cytometry. The cells were stained using anti-pSTAT5-Alexa647 (BD, cat. #612599) and anti-CD4 PE (BD, cat. #347327) for flow cytometry.
Screening peptides by pSTAT5 activation
To assess whether interference by the peptides inhibited IL-2–induced STAT5 phosphorylation, normal peripheral blood T cells were selected using negative-sort T-cell enrichment beads from StemCell Technologies, Cambridge, MA (#19155). Controls for this study included enriched T cells with no stimulation or blocking peptides, as well as cells treated with IL-2 only, cells treated with sIL-2α only, cells treated with a combination of IL-2 and sIL-2α, and cells treated with a control blocking antibody for IL-2. The testing to measure inhibition included addition of the peptide, IL-2, and sIL-2Rα in RPMI full media. The cells were incubated with a mixture of the peptide, IL-2, and sIL-2Rα for 60 min at 37 °C. To stop the reaction, pre-warmed Cytofix Buffer (BD, cat. #554655) was added and incubated for 10 min at 37 °C. Finally, the cells were permeabilized with Perm Buffer III (BD, cat. #558050) and incubated on ice for 15 to 30 min to enable intracellular pSTAT5 antibody staining.
Flow cytometry for Foxp3
Cells were mixed with the same controls as in the pSTAT5 protocol. However, Foxp3 staining required cells to be stimulated for 3 days prior to analysis. Plates were pre-coated with OKT3 and anti-CD28 and incubated for 3 days to allow the Foxp3 to be synthesized. Next, cells were prepared for analysis using a True-Nuclear™ Transcription Factor Buffer Set (BioLegend, cat. #422601) according to manufacturer protocol. Briefly, cells were fixed and permeabilized with reagents from the Foxp3 staining kit. Cells were then stained with Alexa488-conjugated antibody Foxp3 (BioLegend, cat. #320112), according to manufacturer protocol, plus PE-conjugated anti-CD4 antibody (BD, cat. #347327) for 30 min and analyzed by flow cytometry. Acquired data were analyzed by gating on the CD4 and Foxp3 cells using FlowJo (Tree Star, Ashland, OR). CD4 cells were gated and assayed for the percentage of Foxp3 within the cells.
Computational peptide drug design
CMDInventus is a physics-based, data-driven computational platform for peptide drug design. It consists of core modules for addressing biophysical and ADMET problems associated with peptide drug design. The modules can be strung together into project-specific design workflows. Key modules include CMDpeptide (peptide conformational analysis), CMDdock (protein-peptide docking), CMDdesign (combinatorial amino acid sampling along a peptide backbone), CMDboltzmann (ensemble-based, protein-peptide binding affinity prediction), CMDescore and CMDmscore (single-structure–based protein-peptide binding affinity predictions), and CMDscaffold (peptide scaffold database). CMDInventus and many of its modules have been previously described [14].
In the present study, 3 structure-based design approaches were implemented using CMDInventus. First, peptides were designed based on a wild-type (WT) motif derived from IL-2 helix 59–72 (LEEELKPLEEVLNL). Second, peptides were designed based on a small IL-2 peptide WT interface motif comprising residues 42–45 (FKFY). In both approaches, CMDdesign was used to generate motif analogues through systematic side-chain scanning [15]. Analogues were then scored and ranked using CMDescore, CMDmscore, and CMDboltzmann. In the case of the helix-motif approach, a total of 8 peptides were synthesized and tested (CMD161-CMD168) (including the WT sequence motif). In the case of the FKFY-motif approach, a total of 7 novel peptide analogues were synthesized and tested (CMD176-CMD182). Third, peptides were designed de novo using a disulfide-cyclized all D-amino acid scaffold (XCXXC) “interaction” approach. In this approach, key IL2-Rα anchor residue interaction sites were identified [11, 16], and CMDdock and CMDboltzmann were used to generate cyclized analogues that best satisfied all interaction constraints (Table 1). Ultimately, 7 de novo–designed cyclic analogs (CMD169-CMD175) were synthesized and tested. All peptides were purchased from the University of Missouri, Molecular Interactions Core Facility (Dr Fabio Gallazzi, http://micore.missouri.edu). All peptides were synthesized using Fmoc/OtBu protocols, purified to >95% purity (RP-HPLC), and analyzed for expected molecular weight using ESI-MS. See Table 1 for peptide sequence information and characterization details.
Results
Design of sIL-2Rα-blocking CMD peptides
To investigate whether blocking sIL-2Rα binding to IL-2 inhibits the development of Treg cells, we designed 22 peptide compounds to inhibit the binding of IL-2 and IL-2Rα using a structure-based computational design (Fig. 1 and Table 1). Three basic design approaches were taken: 1) design around a 14 residue IL-2 helical motif comprised of residues 59–72 (LEEELKPLEEVLNL) (Fig. 1a), 2) design around a small IL-2 interface motif made of residues 42–45 (FKFY) (Fig. 1b), and 3) use a de novo design of disulfide-cyclized peptides (XCXXC) to optimize interactions with key IL-2Rα residues (Fig. 1c). The helical motif-based design resulted in the identification of 8 candidate peptides (CMD161-CMD168), while the FKFY motif-based design resulted in the identification of 7 predicted peptides (CMD176-CMD182) and the de novo design identified 7 prospective interference peptides (CMD169-CMD175).
Design of sIL-2Rα-blocking CMD peptides. Images showing peptide structure. Each panel represents the computer simulation image of 1 of the 3 peptide backbones designed to block/interfere the IL-2/IL-2Rα binding. The 3 backbones derived form the following peptide templates: a, LEELKPLEEVLNL helix; b, Interferance-based peptide WCRFC; and c, FKFY peptide
Identification by ELISA of CMD peptides that interfere with IL-2Rα/IL-2 binding
All 22 peptides were evaluated for sIL-2Rα/IL-2 binding interference using an ELISA assay. This assay was repeated a minimum of 3 times in triplicate to evaluate possible candidate proteins that either blocked IL-2 binding with sIL-2Rα or blocked sIL-2Rα binding sites, therefore inhibiting binding of IL-2. Due to assay variability, some peptides were tested up to 5 times. All peptides showing any decrease in IL-2 expression were selected for cell-based evaluation. As shown in Fig. 2, while some displayed no inhibition, many peptides, from all 3 design classes, exhibited variable IL-2-binding inhibition, including CMDs 161, 162, 164, 166, and 168 of the helix motif group (range, 7.7%–23.1%), CMD-169 and 171 of the de novo interference group (range, 8.4%–11.8%), and CMDs 178, 179, and 181 from the interface motif group (range, 22.4%–41.3%).
Identification by ELISA of CMD peptides that interfere with IL-2Rα/IL-2 binding. Graph showing binding IL-2 level in sIL-2Rα coated plate in the presence of at different concentrations of CMD peptides assayed by ELISA. Ten peptides, each with 4 concentrations, incubated with IL-2, prior to adding to the coated sIL-2Rα plate. The binding IL-2 level was measured by ELISA
Effect of CMD peptides on STAT5 phosphorylation and Foxp3 expression in T cells
To further confirm blockade of IL-2/sIL-2Rα binding, we assessed whether peptide blockade of IL-2/sIL-2Rα affected STAT5 phosphorylation and Foxp3 expression. To do this, we pre-incubated sIL-2Rα and IL-2 in the presence or absence of peptides. We then cultured freshly isolated T cells in the presence or absence of the IL-2/sIL-2Rα mixture alone or with the addition of potential blocking peptides identified by ELISA. T cells treated with IL-2 or sIL-2Rα alone were used as controls. The STAT5 phosphorylation and Foxp3 expression were measured using flow-based assays. As shown in Fig. 3, while IL-2 substantially induced STAT5 phosphorylation in T cells as expected, the pre-incubated peptide-IL-2/sIL-2Rα mix variably suppressed IL-2–induced phosphorylation. Several peptides, including CMDs 162, 176, and 178, exhibited greater suppression than other peptides (Fig. 3). Similarly, Foxp3 expression was inhibited in T cells treated with the peptide-IL-2/sIL-2α mix. As shown in Fig. 3, while some peptides displayed modest inhibition, others, including CMDs 178, 179, and 182, exhibited substantial inhibition of Foxp3 expression in T cells.
Effect of CMD peptides on STAT5 phosphorylation and Foxp3 expression in T cells. Graphs showing expression of pSTAT5 (top panel) and Foxp3 (bottom panel) in T cells treated with or without IL-2, sIL-2Ra alone or in combination in the presence or absence of CMD peptides. Note: CDMD178 has reduced expression of both STAT5 and Foxp3. Four subsequent experiments showed similar blocking of pSTAT5 (mean 22% reduction ±10%) and Foxp3 (mean 33.8% reduction ±23%)
Effect of CMD peptides on T-cell function
Given the capacity of CMD peptides to reduce Treg cell development, we next wanted to know whether CMD peptide-treated CD4+ T cells that lost suppressive properties would affect the effector function of CD8+ T cells or whether effector function would no longer be suppressed. To do this, we cultured CD4+ T cells in OKT-3–coated plates plus anti-CD28 Ab in the presence or absence of the peptide IL-2/sIL-2Rα mix for 3 days. Cells treated with blocking IL-2 Ab were used as a control. Cells were then harvested and co-cultured with autologous CD8+ T cells for another 3 days, and cytokine production of CD8+ T cells was measured using intracellular cytokine staining. We chose to test CMD178, as this peptide showed consistent and substantial inhibition of IL-2/IL-2R signaling in all previous assays. As shown in Fig. 4, we first compared CD4+ cells treated with IL-2/sIL-2Rα and vehicle alone (negative control) to those treated with a control anti-IL-2 blocking antibody (positive control). These cells were then co-cultured with active CD8+ T cells, and the cytokine and granule production by the CD8+ cell were measured. As expected, cytokine expression including TNF-α and IFN-γ, as well as granule production (granzyme B and perforin) in CD8+ T cells co-cultured with the IL-2 blocking Ab-treated CD4+ T cells was upregulated. We then tested our lead peptide in the same fashion. When CD4+ T cells were treated with the CMD178 peptide plus IL-2/sIL-2Rα, the production of cytokines and granules in CD8+ T cells was further enhanced compared with IL-2 Ab plus IL-2/sIL-2Rα–treated CD4+ T cells. These results suggested that the CMD178 peptide substantially inhibits the development of Treg cells induced by IL-2/sIL-2Rα signaling (Fig. 4).
Effect of CMD peptides on T-cell function. Graph showing cytokine and granule production by CD8+ T -cells when co-cultured with CD4+ T cells and CMD178. The effect of Treg cells on CD8+ cells showed an increase in activity when compared to IL-2/sIL-2Rα controls for Granzyme B, INF-γ, Perforin and TNF-α
Discussion
IL-2 is a cytokine produced primarily by activated CD4+ T cells that plays a crucial role in inducing the development of Treg cells [17]. IL-2 mediates its effects by binding to IL-2 receptors, which can be expressed by CD4+ or CD8+ T cells. Once bound to its receptor, IL-2 starts a cascade of effects, including phosphorylation of STAT5. This activation is necessary for production of Foxp3 and it has been established that Foxp3 is required for development of Treg cells in humans. Activated Treg cells produce more IL-2 thereby further promoting their regulatory function. By blocking the binding of IL-2 to IL-2R, Foxp3 regulatory functions are limited, and regulatory pathways promoting the development and function of Treg cells are suppressed.
In B-cell NHL, IL-2 promotes Treg cells and inhibits TH17 cell development, leading to an inhibitory tumor microenvironment in lymphoma [18]. We previously reported that elevated serum sIL-2Rα levels are associated with a poor prognosis in NHL and that sIL-2Rα facilitates IL-2 signaling and induces Foxp3 expression in T cells, resulting in cells with a regulatory phenotype [12]. We showed that sIL-2Rα binds to and synergizes with IL-2 to suppress anti-tumor immunity by suppressing the proliferation and granule production by intratumoral effector T cells. These results indicate that sIL-2Rα plays an active biologic role in NHL by binding IL-2 and promoting IL-2 signaling rather than depleting IL-2 and blocking its function. Therefore, we contend that the use of novel therapies that deplete sIL-2Rα, eliminate sIL-2Rα production by depleting T cells expressing IL-2Rα, or block the binding of sIL-2Rα to IL-2 will result in considerable clinical benefit for NHL patients.
In the present study, we developed 22 peptides using a structure-based computational design and screened them for their efficacy to block sIL-2Rα/IL-2 binding. Two conservative IL-2 motif design approaches and an ambitious de novo design approach were explored. The motif-based design approaches sought to exploit IL-2 helix 59–72 (LEEELKPLEEVLNL) and IL-2 interface peptide fragment 42–45 (FKFY), respectively. The de novo design approach sought to identify disulfide-linked cyclic peptides (XCXXC) predicted to interfere with IL-2Rα binding by interacting favorably with key IL-2Rα anchor residues. Ultimately, the helix motif approach, the de novo interference approach, and the interface FKFY motif approach resulted in 8 peptides (CMD161–168), 7 peptides (CMD169-CMD175), and 7 peptides (CMD176-CMD182) being tested, respectively. Importantly, all 3 design approaches yielded hits in the ELISA assay and showed the capacity to block sIL-2Rα/IL-2 binding. More importantly, by blocking IL-2/sIL-2Rα binding, several CMD peptides were shown to impact T-cell function by down-regulating the development of Treg cells. Our findings are promising, as these peptides may provide candidates for developing drugs with therapeutic potential that work by reversing the suppressive tumor microenvironment.
In particular, CMD178 displayed consistently encouraging activity by blocking IL-2/sIL-2Rα binding in the ELISA assay, suppressing STAT5 phosphorylation and inhibiting Foxp3 expression in cell-based assays, and thereby increasing cytokine production by CD8+ T cells in co-culture assays. The small size of CMD178 also makes it an attractive candidate for continued development and therefore it is worth reviewing the basic design logic underlying CMD178. The design of CMD178 is based on the IL-2 FKFY interface motif. Four lines of evidence converged on the FKFY motif as a promising template for structure-based peptide-drug design. First, previous experimental work showed that F42 and Y45 mutations dramatically reduced IL-2 affinity for IL-2Rα. Similarly, K43 and F44 mutations were shown to reduce IL-2 binding to IL-2Rα [19,20,21]. Hence, the FKFY motif represents a validated IL-2/IL-2Rα binding motif. Second, analysis of the 1Z92 IL-2/IL-2Rα crystal structure shows the FKFY backbone to lack well-defined secondary structure [22]. This suggests the potential for a small peptide to adopt a similar binding conformation without having to pay a large free-energy penalty. Third, a phage display study by Liu and colleagues [23] identified a 12-residue IL-2/IL-2R blocking peptide that contained the internal sequence motif AKFH. Finally, analysis of the IL-2/IL-2Rα crystal structure revealed promising opportunities for introducing side-chain additions and substitutions into the FKFY sequence aimed at enhancing binding to IL-2Rα. Ultimately, computational sequence optimization of the FKFY motif resulted in the synthesis and testing of 7 candidate peptides (CMD176-CMD182), with 3 emerging as validated ELISA hits (CMD179, CMD181, and CMD182), and CMD178 displaying consistently promising activity across all assays.
It has been shown that Treg cells exhibit a profound impact on the immune response of lymphoma patients. Highly prevalent in NHL biopsies, Treg cells have been shown to efficiently suppress intratumoral CD4+ and CD8+ T cells, resulting in suppressed antitumor immunity. While Treg cells show a significant biological relevance in lymphoma, studies have unexpectedly found that high numbers of Treg cells predicted improved survival in NHL [24]. Further investigation has suggested that instead of numbers, the location of Treg cells in the tumor microenvironment impacts patient outcome [25]. It appears that increased numbers of Treg cells in NHL may be a surrogate for an active, but inhibited, anti-tumor immune response. Inhibiting the immunological effect of Treg cells may therefore allow effector cells to target the malignant clone.
In summary, our results suggest that structure-based peptide design can be used to identify novel peptide inhibitors that block IL-2/IL-2Rα signaling and inhibit STAT5 and Foxp3 upregulation. These peptides could be used as new therapeutic agents to limit immune suppression by Treg cells and promote a more effective antitumor immune response in NHL.
References
Bien E, Balcerska A (2008) Serum soluble interleukin 2 receptor alpha in human cancer of adults and children: a review. Biomarkers 13(1):1–26
Gupta M, Stenson M, O'Byrne M, Maurer MJ, Habermann T, Cerhan JR, Weiner GW, Witzig TE (2016) Comprehensive serum cytokine analysis identifies IL-1RA and soluble IL-2Ralpha as predictors of event-free survival in T-cell lymphoma. Ann Oncol: Off J Eur Soc for Med Oncol/ESMO 27(1):165–172
Wang L, Liao DZ, Zhang J, Xia ZJ, Peng XW, Lu Y (2013) Clinical significance of serum soluble interleukin-2 receptor-alpha in extranodal natural killer/T-cell lymphoma (ENKTL): a predictive biomarker for treatment efficacy and valuable prognostic factor. Med Oncol 30(4):723
Jo SA, Hwang SH, Chang CL, Kim SY, Shin HJ, Chung JS, Sol MY, Lee EY (2010) Clinical relevance of elevated levels of serum soluble interleukin-2 receptor alpha (sIL-2Ralpha) in patients with non-Hodgkin's lymphoma. Korean J Lab Med 30(6):600–605
Jeffery HC, Jeffery LE, Lutz P, Corrigan M, Webb GJ, Hirschfield GM, Adams DH, Oo YH (2017) Low-dose interleukin-2 promotes STAT-5 phosphorylation, Treg survival and CTLA-4-dependent function in autoimmune liver diseases. Clin Exp Immunol 188(3):394–411
Liu W, Putnam AL, Xu-Yu Z, Szot GL, Lee MR, Zhu S, Gottlieb PA, Kapranov P, Gingeras TR, Fazekas de St Groth B, Clayberger C, Soper DM, Ziegler SF, Bluestone JA (2006) CD127 expression inversely correlates with FoxP3 and suppressive function of human CD4+ T reg cells. J Exp Med 203(7):1701–1711
Mahmud SA, Manlove LS, Farrar MA (2013) Interleukin-2 and STAT5 in regulatory T cell development and function. JAKSTAT 2(1):e23154
Ai WZ, Hou JZ, Zeiser R, Czerwinski D, Negrin RS, Levy R (2009) Follicular lymphoma B cells induce the conversion of conventional CD4+ T cells to T-regulatory cells. Int J Cancer 124(1):239–244
Binder M, O'Byrne MM, Maurer MJ, Ansell S, Feldman AL, Cerhan J, Novak A, Porrata LF, Markovic S, Link BK, Witzig TE (2017) Associations between elevated pre-treatment serum cytokines and peripheral blood cellular markers of immunosuppression in patients with lymphoma. Am J Hematol 92(8):752–758
Mir MA, Maurer MJ, Ziesmer SC, Slager SL, Habermann T, Macon WR, Link BK, Syrbu S, Witzig T, Friedberg JW, Press O, LeBlanc M, Cerhan JR, Novak A, Ansell SM (2015) Elevated serum levels of IL-2R, IL-1RA, and CXCL9 are associated with a poor prognosis in follicular lymphoma. Blood 125(6):992–998
Zorn E, Nelson EA, Mohseni M, Porcheray F, Kim H, Litsa D, Bellucci R, Raderschall E, Canning C, Soiffer RJ, Frank DA, Ritz J (2006) IL-2 regulates FOXP3 expression in human CD4+CD25+ regulatory T cells through a STAT-dependent mechanism and induces the expansion of these cells in vivo. Blood 108(5):1571–1579
Yang ZZ, Grote DM, Ziesmer SC, Manske MK, Witzig TE, Novak AJ, Ansell SM (2011) Soluble IL-2Ralpha facilitates IL-2-mediated immune responses and predicts reduced survival in follicular B-cell non-Hodgkin lymphoma. Blood 118(10):2809–2820
Janas ML, Groves P, Kienzle N, Kelso A (2005) IL-2 regulates perforin and granzyme gene expression in CD8+ T cells independently of its effects on survival and proliferation. J Immunol 175(12):8003–8010
Onoprienko LV, Mikhaleva II, Lunev VE, Nesmeianov VA, Ivanov VT (1989) Synthesis and immunochemical properties of peptides corresponding to sequences 59-72 and 25-36 of human interleukin-2. Bioorg Khim 15(7):908–921
Wang X, Rickert M, Garcia KC (2005) Structure of the quaternary complex of interleukin-2 with its alpha, beta, and gammac receptors. Science 310(5751):1159–1163
Du J, Yang H, Zhang D, Wang J, Guo H, Peng B, Guo Y, Ding J (2010) Structural basis for the blockage of IL-2 signaling by therapeutic antibody basiliximab. J Immunol 184(3):1361–1368
Boyman O, Sprent J (2012) The role of interleukin-2 during homeostasis and activation of the immune system. Nat Rev Immunol 12(3):180–190
Liao W, Lin JX, Leonard WJ (2013) Interleukin-2 at the crossroads of effector responses, tolerance, and immunotherapy. Immunity 38(1):13–25
Emerson SD, Palermo R, Liu CM, Tilley JW, Chen L, Danho W, Madison VS, Greeley DN, Ju G, Fry DC (2003) NMR characterization of interleukin-2 in complexes with the IL-2Ralpha receptor component, and with low molecular weight compounds that inhibit the IL-2/IL-Ralpha interaction. Protein Sci 12(4):811–822
Sauve K, Nachman M, Spence C, Bailon P, Campbell E, Tsien WH, Kondas JA, Hakimi J, Localization JG (1991) In human interleukin 2 of the binding site to the alpha chain (p55) of the interleukin 2 receptor. Proc Natl Acad Sci U S A 88(11):4636–4640
Thanos CD, DeLano WL, Wells JA (2006) Hot-spot mimicry of a cytokine receptor by a small molecule. Proc Natl Acad Sci U S A 103(42):15422–15427
Rickert M, Wang X, Boulanger MJ, Goriatcheva N, Garcia KC (2005) The structure of interleukin-2 complexed with its alpha receptor. Science 308(5727):1477–1480
Liu BY, Zhu P, Luo HB, Fu N (2006) screening of short peptides binding to cell surface interleukin-2 receptor alpha chain. Nan Fang Yi Ke Da Xue Xue Bao 26(7):971–974
Tzankov A, Meier C, Hirschmann P, Went P, Pileri SA, Dirnhofer S (2008) Correlation of high numbers of intratumoral FOXP3+ regulatory T cells with improved survival in germinal center-like diffuse large B-cell lymphoma, follicular lymphoma and classical Hodgkin's lymphoma. Haematologica 93(2):193–200
Hude I, Sasse S, Engert A, Brockelmann PJ (2017) The emerging role of immune checkpoint inhibition in malignant lymphoma. Haematologica 102(1):30–42
Acknowledgments
This work was supported in part by grants from the National Institutes of Health (P50 CA97274), the Leukemia & Lymphoma Society, the Landow Foundation, and the Predolin Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare no conflict of interest.
Ethical approval
The authors declared that this article does not contain any studies with human or animal participants.
Informed consent
For this type of study, formal consent is not required.
Rights and permissions
About this article
Cite this article
Price-Troska, T., Yang, ZZ., Diller, D. et al. Inhibiting IL-2 signaling and the regulatory T-cell pathway using computationally designed peptides. Invest New Drugs 37, 9–16 (2019). https://doi.org/10.1007/s10637-018-0606-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10637-018-0606-9