Abstract
Gliomas are aggressive brain tumors characterized by uncontrolled cell proliferation. FAM64A, a cell cycle-related gene, has been found to promote cell proliferation in various tumors, including gliomas. However, the regulatory mechanism and clinical significance of FAM64A in gliomas remain unclear. In this study, we investigated FAM64A expression in gliomas with different grades and constructed FAM64A silenced cell lines to study its functions. Our results demonstrated that FAM64A was highly expressed in glioblastoma (P < 0.001) and associated with a poor prognosis (P < 0.001). Expression profiles at the single-cell resolution indicated FAM64A could play a role in a cell-cycle-dependent way to promote glioma cell proliferation. We further observed that FAM64A silencing in glioma cells resulted in disrupted proliferation and migration ability, and increased cell accumulation in the G2/M phase (P = 0.034). Additionally, TGF-β signaling upregulates FAM64A expression, and SMAD4 and FAM64A co-localize in high-grade glioma tissues. We found FAM64A knockdown inhibited TGF-β-induced epithelial-mesenchymal transition in glioma. Our findings suggest that FAM64A could serve as a diagnostic and therapeutic target in gliomas.






Similar content being viewed by others
Data Availability
The datasets used and analyzed during the current study are available from Gliovis (http://gliovis.bioinfo.cnio.es/) and GEO database (https://www.ncbi.nlm.nih.gov/geo/). Analysis scripts are available from the corresponding author on reasonable request.
Abbreviations
- TGF-β :
-
Transforming growth factor b
- GBM:
-
Glioblastoma
- LGG:
-
Lower grade glioma
- TCGA:
-
The cancer genome atlas
- scRNA-seq:
-
Single cell RNA-seq
- IDH:
-
Isocitrate dehydrogenase
- PLK:
-
Polo-like kinase
- TME:
-
Tumor microenvironment
- EMT:
-
Epithelial-mesenchymal transition
- TMZ:
-
Temozolomide
- MGMT:
-
O6-methylguanine-DNA methyltransferase
References
Archangelo LF, Greif PA, Maucuer A, Manceau V, Koneru N, Bigarella CL, Niemann F, dos Santos MT, Kobarg J, Bohlander SK (1833) Saad ST (2013) The CATS (FAM64A) protein is a substrate of the kinase interacting stathmin (KIS). Biochim Biophys Acta 5:1269–1279. https://doi.org/10.1016/j.bbamcr.2013.02.004
Bellail AC, Hunter SB, Brat DJ, Tan C, Van Meir EG (2004) Microregional extracellular matrix heterogeneity in brain modulates glioma cell invasion. Int J Biochem Cell Biol 36(6):1046–1069. https://doi.org/10.1016/j.biocel.2004.01.013
Bronner SM, Merrick KA, Murray J, Salphati L, Moffat JG, Pang J, Sneeringer CJ, Dompe N, Cyr P, Purkey H, Boenig GL, Li J, Kolesnikov A, Larouche-Gauthier R, Lai KW, Shen X, Aubert-Nicol S, Chen YC, Cheong J, Crawford JJ, Hafner M, Haghshenas P, Jakalian A, Leclerc JP, Lim NK, O’Brien T, Plise EG, Shalan H, Sturino C, Wai J, Xiao Y, Yin J, Zhao L, Gould S, Olivero A, Heffron TP (2019) Design of a brain-penetrant CDK4/6 inhibitor for glioblastoma. Bioorg Med Chem Lett 29(16):2294–2301. https://doi.org/10.1016/j.bmcl.2019.06.021
Butler A, Hoffman P, Smibert P, Papalexi E, Satija R (2018) Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol 36(5):411–420. https://doi.org/10.1038/nbt.4096
Daneman R, Prat A (2015) The blood-brain barrier. Cold Spring Harb Perspect Biol 7(1):a020412. https://doi.org/10.1101/cshperspect.a020412
DeWire M, Fuller C, Hummel TR, Chow LML, Salloum R, de Blank P, Pater L, Lawson S, Zhu X, Dexheimer P, Carle AC, Kumar SS, Drissi R, Stevenson CB, Lane A, Breneman J, Witte D, Jones BV, Leach JL, Fouladi M (2020) A phase I/II study of ribociclib following radiation therapy in children with newly diagnosed diffuse intrinsic pontine glioma (DIPG). J Neurooncol 149(3):511–522. https://doi.org/10.1007/s11060-020-03641-2
Friedman HS, Kerby T, Calvert H (2000) Temozolomide and treatment of malignant glioma. Clin Cancer Res 6(7):2585–2597
Fu M, Zhang J, Li W, He S, Zhang J, Tennant D, Hua W, Mao Y (2021) Gene clusters based on OLIG2 and CD276 could distinguish molecular profiling in glioblastoma. J Trans Med. https://doi.org/10.1186/s12967-021-03083-y
Geeleher P, Cox N, Huang RS (2014) pRRophetic: an R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS One 9(9):e107468. https://doi.org/10.1371/journal.pone.0107468
Gratchev A (2017) TGF-beta signalling in tumour associated macrophages. Immunobiology 222(1):75–81. https://doi.org/10.1016/j.imbio.2015.11.016
Hanahan D (2022) Hallmarks of cancer: new dimensions. Cancer Discov 12(1):31–46. https://doi.org/10.1158/2159-8290.CD-21-1059
Hao Y, Baker D, Ten Dijke P (2019) TGF-β-mediated epithelial-mesenchymal transition and cancer metastasis. Int J Mol Sci. https://doi.org/10.3390/ijms20112767
Hashimoto K, Kodama A, Honda T, Hanashima A, Ujihara Y, Murayama T, Nishimatsu SI, Mohri S (2017) Fam64a is a novel cell cycle promoter of hypoxic fetal cardiomyocytes in mice. Sci Rep 7(1):4486. https://doi.org/10.1038/s41598-017-04823-1
He L, Liu Y, Lai W, Tian H, Chen L, Xie L, Liu Z (2020) DNA sensors, crucial receptors to resist pathogens, are deregulated in colorectal cancer and associated with initiation and progression of the disease. J Cancer 11(4):893–905. https://doi.org/10.7150/jca.34188
Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, Weller M, Kros JM, Hainfellner JA, Mason W, Mariani L, Bromberg JE, Hau P, Mirimanoff RO, Cairncross JG, Janzer RC, Stupp R (2005) MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med 352(10):997–1003. https://doi.org/10.1056/NEJMoa043331
Hitomi M, Deleyrolle LP, Mulkearns-Hubert EE, Jarrar A, Li M, Sinyuk M, Otvos B, Brunet S, Flavahan WA, Hubert CG, Goan W, Hale JS, Alvarado AG, Zhang A, Rohaus M, Oli M, Vedam-Mai V, Fortin JM, Futch HS, Griffith B, Wu Q, Xia CH, Gong X, Ahluwalia MS, Rich JN, Reynolds BA, Lathia JD (2015) Differential connexin function enhances self-renewal in glioblastoma. Cell Rep 11(7):1031–1042. https://doi.org/10.1016/j.celrep.2015.04.021
Iser IC, Pereira MB, Lenz G, Wink MR (2017) The epithelial-to-mesenchymal transition-like process in glioblastoma: an updated systematic review and in silico investigation. Med Res Rev 37(2):271–313. https://doi.org/10.1002/med.21408
Jiang Y, Zhou C, Gao Q, Yin ZQ, Wang J, Mu H, Yan J (2020) FAM64A promotes osteosarcoma cell growth and metastasis and is mediated by miR-493. J Oncol 2020:2518297. https://doi.org/10.1155/2020/2518297
Jiao Y, Fu Z, Li YQ, Zhang W, Liu YH (2019) Aberrant FAM64A mRNA expression is an independent predictor of poor survival in pancreatic cancer. Plos One. https://doi.org/10.1371/journal.pone.0211291
Kaminska B, Cyranowski S (2020) Recent advances in understanding mechanisms of TGF Beta signaling and its role in glioma pathogenesis. Adv Exp Med Biol 1202:179–201. https://doi.org/10.1007/978-3-030-30651-9_9
Kaminska B, Kocyk M, Kijewska M (2013) TGF beta signaling and its role in glioma pathogenesis. Adv Exp Med Biol 986:171–187. https://doi.org/10.1007/978-94-007-4719-7_9
Korsunsky I, Millard N, Fan J, Slowikowski K, Zhang F, Wei K, Baglaenko Y, Brenner M, Loh PR, Raychaudhuri S (2019) Fast, sensitive and accurate integration of single-cell data with Harmony. Nat Methods 16(12):1289–1296. https://doi.org/10.1038/s41592-019-0619-0
Liu J, Peng Y, Wei W (2022) Cell cycle on the crossroad of tumorigenesis and cancer therapy. Trends Cell Biol 32(1):30–44. https://doi.org/10.1016/j.tcb.2021.07.001
Loh CY, Chai JY, Tang TF, Wong WF, Sethi G, Shanmugam MK, Chong PP, Looi CY (2019) The E-cadherin and N-cadherin switch in epithelial-to-mesenchymal transition: signaling, therapeutic implications, and challenges. Cells. https://doi.org/10.3390/cells8101118
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW (2016) The 2016 world health organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131(6):803–820. https://doi.org/10.1007/s00401-016-1545-1
Louis DN, Perry A, Wesseling P, Brat DJ, Cree IA, Figarella-Branger D, Hawkins C, Ng HK, Pfister SM, Reifenberger G, Soffietti R, von Deimling A, Ellison DW (2021) The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol 23(8):1231–1251. https://doi.org/10.1093/neuonc/noab106
Mizuno K, Tanigawa K, Nohata N, Misono S, Okada R, Asai S, Moriya S, Suetsugu T, Inoue H, Seki N (2020) FAM64A: a novel oncogenic target of lung adenocarcinoma regulated by both strands of miR-99a (miR-99a-5p and miR-99a-3p). Cells. https://doi.org/10.3390/cells9092083
Ostrom QT, Gittleman H, de Blank PM, Finlay JL, Gurney JG, McKean-Cowdin R, Stearns DS, Wolff JE, Liu M, Wolinsky Y, Kruchko C, Barnholtz-Sloan JS (2016) American brain tumor association adolescent and young adult primary brain and central nervous system tumors diagnosed in the united states in 2008–2012. Neuro Oncol 18(Suppl 1):i1–i50. https://doi.org/10.1093/neuonc/nov297
Pastushenko I, Blanpain C (2019) EMT transition states during tumor progression and metastasis. Trends Cell Biol 29(3):212–226. https://doi.org/10.1016/j.tcb.2018.12.001
Qiu X, Qin F (2021) FAM64A antagonizes tumor suppressive effects of miR-610 in neuroblastoma in vitro. J Neurosurg Sci. https://doi.org/10.23736/S0390-5616.21.05261-9
Ross JL, Vega JV, Plant A, MacDonald TJ, Becher OJ, Hambardzumyan D (2021) Tumour immune landscape of paediatric high-grade gliomas. Brain 144:2594–2609. https://doi.org/10.1093/brain/awab155
Schichor C, Albrecht V, Korte B, Buchner A, Riesenberg R, Mysliwietz J, Paron I, Motaln H, Turnšek TL, Jürchott K, Selbig J, Tonn JC (2012) Mesenchymal stem cells and glioma cells form a structural as well as a functional syncytium in vitro. Exp Neurol 234(1):208–219. https://doi.org/10.1016/j.expneurol.2011.12.033
Sepulveda-Sanchez JM, Gil-Gil M, Alonso-Garcia M, Vaz Salgado MA, Vicente E, Mesia Barroso C, Rodriguez Sanchez A, Duran G, De Las PR, Munoz-Langa J, Velasco G, Hernandez-Lain A, Hilario A, Navarro Martin M, Benavides M, Oleaga L, Cantero Montenegro D, Ruano Y, Sanchez-Gomez P, Martin-Soberon MC, Morales-Llombart R, Pachon V, Pineda E (2020) Phase II trial of palbociclib in recurrent retinoblastoma-positive anaplastic oligodendroglioma: a study from the spanish group for research in neuro-oncology (GEINO). Target Oncol 15(5):613–622. https://doi.org/10.1007/s11523-020-00754-6
Singh N, Miner A, Hennis L, Mittal S (2021) Mechanisms of temozolomide resistance in glioblastoma - a comprehensive review. Cancer Drug Resist 4(1):17–43. https://doi.org/10.20517/cdr.2020.79
Soukupova J, Malfettone A, Bertran E, Hernández-Alvarez MI, Peñuelas-Haro I, Dituri F, Giannelli G, Zorzano A, Fabregat I (2021) Epithelial-mesenchymal transition (EMT) induced by TGF-β in hepatocellular carcinoma cells reprograms lipid metabolism. Int J Mol Sci. https://doi.org/10.3390/ijms22115543
Stanic J, Carta M, Eberini I, Pelucchi S, Marcello E, Genazzani AA, Racca C, Mulle C, Di Luca M, Gardoni F (2015) Rabphilin 3A retains NMDA receptors at synaptic sites through interaction with GluN2A/PSD-95 complex. Nat Commun 6:10181. https://doi.org/10.1038/ncomms10181
Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, Taphoorn MJ, Belanger K, Brandes AA, Marosi C, Bogdahn U, Curschmann J, Janzer RC, Ludwin SK, Gorlia T, Allgeier A, Lacombe D, Cairncross JG, Eisenhauer E, Mirimanoff RO (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352(10):987–996. https://doi.org/10.1056/NEJMoa043330
Vitucci M, Hayes DN, Miller CR (2011) Gene expression profiling of gliomas: merging genomic and histopathological classification for personalised therapy. Brit J Cancer 104(4):545–553. https://doi.org/10.1038/sj.bjc.6606031
Wang D, Hu A, Peng H, Li D, Zhang L (2021) Tumor-promoting function of PIMREG in glioma by activating the beta-catenin pathway. 3Biotech 11(8):380. https://doi.org/10.1007/s13205-021-02922-5
Wen PY, Packer RJ (2021) The 2021 WHO classification of tumors of the central nervous system: clinical implications. Neuro Oncol 23(8):1215–1217. https://doi.org/10.1093/neuonc/noab120
Wen NY, Guo BF, Zheng HW, Xu LB, Liang H, Wang Q, Wang D, Chen XY, Zhang SN, Li Y, Zhang L (2019) Bromodomain inhibitor jq1 induces cell cycle arrest and apoptosis of glioma stem cells through the VEGF/PI3K/AKT signaling pathway. Int J Oncol 55(4):879–895. https://doi.org/10.3892/ijo.2019.4863
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z, Feng T, Zhou L, Tang W, Zhan L, Fu X, Liu S, Bo X, Yu G (2021) ClusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2(3):100141. https://doi.org/10.1016/j.xinn.2021.100141
Xu J, Lamouille S, Derynck R (2009) TGF-beta-induced epithelial to mesenchymal transition. Cell Res 19(2):156–172. https://doi.org/10.1038/cr.2009.5
Yao ZC, Zheng XC, Lu ST, He ZX, Miao YT, Huang HH, Chu XW, Cai CQ, Zou F (2019) Knockdown of FAM64A suppresses proliferation and migration of breast cancer cells. Breast Cancer 26(6):835–845. https://doi.org/10.1007/s12282-019-00991-2
Zhang W, Couldwell WT, Simard MF, Song H, Lin JH, Nedergaard M (1999) Direct gap junction communication between malignant glioma cells and astrocytes. Cancer Res 59(8):1994–2003
Zhang C, Han Y, Huang H, Min L, Qu L, Shou C (2014) Integrated analysis of expression profiling data identifies three genes in correlation with poor prognosis of triple-negative breast cancer. Int J Oncol 44(6):2025–2033. https://doi.org/10.3892/ijo.2014.2352
Zhang J, Qian L, Wu JQ, Lu D, Yuan HY, Li WJ, Ying XM, Hu SF (2019) Up-regulation of FAM64A promotes epithelial-to-mesenchymal transition and enhances stemness features in breast cancer cells. Biochem Biophys Res Commun 513(2):472–478. https://doi.org/10.1016/j.bbrc.2019.03.207
Zhang JW, Fu MJ, Zhang ML, Zhang JS, Du ZG, Zhang HY, Hua W, Mao Y (2021) DDX60 is associated with glioma malignancy and serves as a potential immunotherapy biomarker. Front Oncol. https://doi.org/10.3389/fonc.2021.665360
Zheng R, Wan C, Mei S, Qin Q, Wu Q, Sun H, Chen CH, Brown M, Zhang X, Meyer CA, Liu XS (2019) Cistrome data browser: expanded datasets and new tools for gene regulatory analysis. Nucleic Acids Res 47(D1):D729–D735. https://doi.org/10.1093/nar/gky1094
Zhou Y, Ou L, Xu J, Yuan H, Luo J, Shi B, Li X, Yang S, Wang Y (2021) FAM64A is an androgen receptor-regulated feedback tumor promoter in prostate cancer. Cell Death Dis 12(7):668. https://doi.org/10.1038/s41419-021-03933-z
Zhu XL, Yang H, Zhang MY, Wu XW, Jiang L, Liu XC, Lv K (2021) YTHDC1-mediated VPS25 regulates cell cycle by targeting JAK-STAT signaling in human glioma cells. Cancer Cell Int. https://doi.org/10.1186/s12935-021-02304-0
Acknowledgements
The bioinformatic analysis was supported by the Medical Research Data Center of Fudan University.
Funding
This work was supported by the National Natural Science Foundation of China (82072784, 8210113482, 82103690), National Key R&D Program of China (2022YFC3401600), and CAMS Innovation Fund for Medical Sciences (2022-I2M-C&T-B-112). The funders had no role in study design, data collection, interpretation, or the decision to submit the work for publication.
Author information
Authors and Affiliations
Contributions
MF, JZ and LZ have contributed equally to this study. WH, WW, and YM conceived the general framework of this study and revised the manuscript. JZ, MF and LZ performed the cell experiments. MF, YF, QW, XF, LZ, and JZ analyzed data. MF and LZ interpreted the results. FF collected clinical samples. MF, LZ, and JZ wrote the manuscript. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing financial interests.
Ethics Approval
The study was conducted in accordance with the Declaration of Helsinki and approved by the Ethics Committee of Huashan Hospital (Approval number: KY2015-256).
Consent to Participate
Glioma tissues used in the current study were resected in 2020 from patients in the Department of Neurosurgery, Huashan Hospital of Fudan University. Normal brain tissues were gathered from traumatic brain injury patients. Written informed consent were obtained from all patients for use of tissue sample.
Consent for Publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10571_2023_1348_MOESM1_ESM.tif
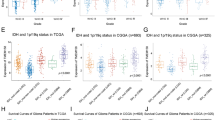
Supplementary file1 (TIF 33731 KB)— Suppl. Fig. 1 (A) Pan-cancer analysis of FAM64A. Most TCGA tumors had a higher accumulation of FAM64A than the adjacent non-tumor tissues. (B) IDH wildtype gliomas expressed higher FAM64A compared with IDH mutant gliomas in TCGA-GBMLGG cohort. (C) FAM64A in mesenchymal subtype is higher than neural and proneural subtype. (D) FAM64A expression in negative correlation with the sensitivity of BI-2536, a PLK inhibitor. Data represents the mean ± SD of triplicate samples, *P < .05, **P < .01, ***P < .001, t test was used to compare two individual groups
10571_2023_1348_MOESM2_ESM.tif
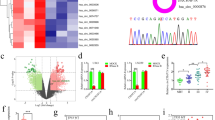
Supplementary file2 (TIF 26981 KB)— Suppl. Fig. 2 (A) Umaps of GSE131928 dataset before and after removing batch effect using harmony algorithm. And the batch effect was very apparent. (B) Cell groups (Monocytes/Macrophage/Microglia cells, Glia/Glioma cells, and T cells) were verified by classic gene markers (T cells: CD3D, CD3E, CD4; Glia/Glioma cells: OLIG2, FA2H, CNP, GFAP; Monocytes/Macrophages/Microglias: CD68, CD163, CD14, FPR1). (C) FAM64A knock-down efficiency of stable cell lines was validated by immunoblotting. ShFAM64A represented the FAM64A silencing U87 cells and shScramble is the control group. (D) shFAM64A U251 cells have lower viability after 48h and 72h compared with control groups
10571_2023_1348_MOESM3_ESM.tif
Supplementary file3 (TIF 30471 KB)— Suppl. Fig. 3 (A) Heatmap for output of copy number variation of glia/glioma cells via inferCNV algorithm. The T cell and TAMs are set as references. (B) Copy number variation (CNV) visualization of GSE131928 scRNA dataset. Color is coded for different clustering of inferCNV algorithm. CNV score for Group 18 is low. Therefore, other groups were included in the following analysis
10571_2023_1348_MOESM4_ESM.tif
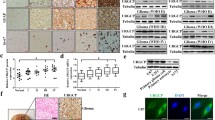
Supplementary file4 (TIF 26265 KB)— Suppl. Fig. 4 (A) Network plot of TGF-β signaling crosstalk. TAMs have a very strong interaction with FAM64A highly expressed neoplasm cells via the TGF-β signaling pathway (the directional red and green bands mean the crosstalk between TAMs and FAM64A highly expressed neoplasm cells). (B) Ligand-receptor plot of TGF-β signaling pathway. TGFB1-(TGFBR1+TGFBR2) contribute most to the signaling flow between TAMs and neoplasm cells. (C) Integrated genome browser visualization of tag density profiles for ChIP-Seq Smad2, ChIP-Seq Smad3, and ChIP-Seq Smad4. (D) FAM64A mRNA expression in control group and Smad3 knock out groups in GSE125116. (E) TGF-β concentration in WHO IV gliomas is higher than normal brain tissues, WHO II and III gliomas significantly. (F) Representative images of IHC staining for FAM64A in different pathological grades of gliomas. (G) TGF-β signaling upregulated SMAD3 and SMAD4 in HA-MG scramble controls and FAM64A silencing groups (shFAM64A#1 and shFAM64A#2). TGF-β of low concentration (10 ng/mL) could promote N-Cadherin expression. However, N-Cadherin expression decreased with the elevated TGF-β concentrations (20 ng/mL and 40 ng/mL). Control groups exhibited higher N-cadherin expression compared with FAM64A silencing groups (Paired t-test, P= 0.0171, 0.0963, respectively). The protein level of N-Cadherin, SMAD3, and SMAD4 in cells was determined by western blotting, normalized by β-TUBULIN. Data represents the mean ± SD of triplicate samples, *P < .05, **P < .01, ***P < .001, t test was used to compare two individual groups
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fu, M., Zhang, J., Zhang, L. et al. Cell Cycle-Related FAM64A Could be Activated by TGF-β Signaling to Promote Glioma Progression. Cell Mol Neurobiol 43, 2975–2987 (2023). https://doi.org/10.1007/s10571-023-01348-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10571-023-01348-2