Abstract
Cellular plasticity and therapy resistance are critical features of pancreatic cancer, a highly aggressive and fatal disease. The pancreas, a vital organ that produces digestive enzymes and hormones, is often affected by two main types of cancer: the pre-dominant ductal adenocarcinoma and the less common neuroendocrine tumors. These cancers are difficult to treat due to their complex biology characterized by cellular plasticity leading to therapy resistance. Cellular plasticity refers to the capability of cancer cells to change and adapt to different microenvironments within the body which includes acinar-ductal metaplasia, epithelial to mesenchymal/epigenetic/metabolic plasticity, as well as stemness. This plasticity allows heterogeneity of cancer cells, metastasis, and evasion of host’s immune system and develops resistance to radiation, chemotherapy, and targeted therapy. To overcome this resistance, extensive research is ongoing exploring the intrinsic and extrinsic factors through cellular reprogramming, chemosensitization, targeting metabolic, key survival pathways, etc. In this review, we discussed the mechanisms of cellular plasticity involving cellular adaptation and tumor microenvironment and provided a comprehensive understanding of its role in therapy resistance and ways to overcome it.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Pancreatic cancer is a highly aggressive form of cancer that affects the pancreas, a gland located in the abdomen that produces digestive enzymes and hormones. Pancreatic cancer is often difficult to detect in its early stages due to the lack of noticeable symptoms. Late diagnosis and early metastasis are the main cause attributed to its low survival rate [1]. Despite advances in surgical techniques and chemotherapy, pancreatic cancer remains a key public health issue with a 5-year survival rate of about 12% [2]. There has been minimal advancement in the clinical prognosis of this devastating disease over the past three decades [3]. It is expected to become the second most prevalent cause of cancer-related deaths in the next 10 to 15 years [4]. Surgical removal is the only potentially curative option, but it is only feasible for a small percentage of patients due to the advanced stage at diagnosis. Chemotherapy and radiation therapy are commonly used for advanced disease, but they often provide limited success in extending the life of patients. As per the most recent classification of tumors by World Health Organization (WHO), there are two primary categories of cancer found in the pancreas: (1) malignant epithelial tumors and (2) neuroendocrine neoplasms [5]. The first one encompasses ductal adenocarcinoma, acinar cell carcinoma, pancreatoblastoma, and solid pseudopapillary neoplasm. In contrast, the latter consists of functioning and non-functioning neuroendocrine tumors, neuroendocrine carcinoma, and mixed neuroendocrine-non-neuroendocrine neoplasms [5]. Pancreatic cancer typically denotes the more aggressive form known as pancreatic ductal adenocarcinoma (PDAC), which comprises approximately 90% of all cases involving exocrine cells [6,7,8,9].
Pancreas develops from the gut endoderm, one of the three primary germ layers that form in vertebrates following gastrulation. This process involves a sequence of well-coordinated signaling events and transcriptional regulatory cascades that ultimately shape it into its mature form [10]. Developing pancreatic tissue receives cues from their macro- and microenvironment results in a systematic differentiation pattern. Such ability of pancreatic tissues to adapt their development in response to their environment, regardless of their inherent genetic makeup, is referred to as the phenotypic plasticity. This highly plastic nature allows pancreatic cells to repair themselves in case of injury via transdifferentiation to progenitor like cells. However, in the presence of underlying mutation in certain genes such as Kirsten rat sarcoma (KRAS) could lead to acinar to ductal metaplastic (ADM) conversion instead of tissue repair. This inherent cellular plasticity could potentially influence not only the initial development of tumors but also the progression of these tumors, including early-stage invasion, metastasis, and metastatic organotropism [11,12,13,14].
This review includes an in-depth analysis of molecular mechanisms involved in cellular plasticity, stemness, and therapy resistance in pancreatic cancer. The review also highlights the current therapeutic strategies for pancreatic cancer, including chemotherapy and radiation therapy. The role of cancer stem cells (CSCs) in therapy resistance has been examined, and the potential for targeting cellular plasticity and stemness as a means of overcoming therapy resistance is discussed.
2 Cellular plasticity in pancreatic cancer
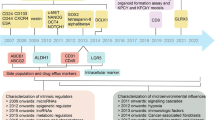
Cellular plasticity arises from various causes; however, in the context of cancer, cellular plasticity denotes the ability of cancer cells to endure dynamic changes, allowing them to acclimatize to various environmental conditions (Fig. 1) [15]. This plasticity reprograms their molecular and cellular characteristics usually without genetic mutations which enables cancer cells to survive, proliferate, and evade treatment strategies, contributing to tumor growth, metastasis, and resistance to therapy [15, 16]. Cellular plasticity in cancer involves several key aspects such as phenotypic heterogeneity, phenotypic switching, therapeutic resistance, metastasis, and microenvironment adaptation [17,18,19,20,21,22]. Cancer cells within a single tumor can exhibit diverse characteristics, including differences in gene expression, cell surface markers, and metabolic profiles [23,24,25,26,27,28]. This heterogeneity is thought to arise from reversible epigenetic changes, and microenvironmental influences [29]. Cancer cells can switch between different cellular states, often resembling different stages of cell development or differentiation. This plasticity allows them to adopt traits that aid in tumor growth, invasion, and resistance to treatment [15, 30]. For example, cancer cells might transition between epithelial and mesenchymal states, a phenomenon known as epithelial-mesenchymal transition (EMT), which can promote metastasis [31, 32].
Cellular plasticity can contribute to the development of resistance to various cancer therapies, including chemotherapy, targeted therapy, and immunotherapy [33,34,35,36]. Cancer cells that undergo phenotypic changes can escape the effects of treatments that were originally effective against a different cellular state. Plasticity is closely linked to the potential of cancer cells to disseminate or spread to distant organs. Cells that acquire more invasive and migratory characteristics can separate from the primary tumor, travel through the circulatory system, and establish secondary tumors in distant locations [37,38,39,40]. Moreover, cancer cells can adapt to the unique microenvironment of tumors, which can be characterized by low oxygen levels (hypoxia), nutrient deprivation, and inflammation. Cellular plasticity allows cancer cells to survive and thrive in these challenging conditions. Cellular plasticity is also associated with the concept of CSCs, which are a subpopulation of cancer cells with characteristics similar to stem cells. These cells have the capability of self-renewal and differentiation into different cell types within the tumor, contributing to tumor growth and heterogeneity [41].
Understanding and targeting cellular plasticity in cancer is a complex challenge in cancer research and therapy. It requires a deeper understanding of the molecular mechanisms that drive these transitions and the development of strategies to prevent or reverse these changes, ultimately leading to more effective treatments and improved patient outcomes. Pancreatic cancer is characterized by a high degree of cellular plasticity leading to tumor growth, metastasis, disease progression, and therapy resistance in various ways as discussed below.
2.1 Acinar-ductal metaplasia
Highest plastic characteristics of acinar cells in the adult pancreas allow its transdifferentiation to ductal progenitor-like cells called acinar-to-ductal metaplasia (ADM) [42]. ADM is a notable plastic feature that enables regeneration of the pancreas after injury or inflammation [43]. Ironically, this metaplastic cell acts as a precursor of pancreatic intraepithelial neoplasia (PanIN) leading to PDAC which is evident from in vivo findings and lineage tracing studies [44,45,46,47]. Multiple factors including KRAS hyperactivity and transforming growth factor (TGF)-α/β have been implicated for such transformation [48,49,50,51,52,53]. Transdifferentiation of acinar cells into duct-like structures is frequently observed in 3D cultures in the presence of oncogenic KRAS, cytokines, epidermal growth factor receptor (EGFR) activating factors, etc. [48, 50, 54,55,56]. Research also showed that Krüppel-like factor 4 (KLF4) exhibits increased expression within acinar cells and is essential for the process of acinar-to-ductal metaplasia [57]. A recent study published in 2020 showed the involvement of interleukin-22 (IL-22) in the plasticity of acinar cells that promote pancreatic tumor formation in mice [42].
The ADM is a natural reversible mechanism which becomes irreversible when acinar cells acquire KRAS mutation or under the influence of continuous exposure of growth factors. It has been demonstrated that oncogenic KRAS suppresses acinar genes including basic helix-loop-helix family member A15 (MIST1) and carboxypeptidase A1 (CPA1) and expresses ductal genes for cytokeratin 19 (KRT19) and mucin 1 (MUC1) and the overexpression of pancreatic and duodenal homeobox 1 (PDX1) and Sry-related high-mobility group box 9 (SOX9) [48, 56]. Oncogenic KRAS acts through phosphoinositide 3-kinase (PI3K) subunit p110α via extracellular signal-regulated kinases (ERK1/2) signaling as well as protein kinase B (PKB/AKT). On the other hand, it has been demonstrated that PTF1A, in collaboration with various transcription factors, plays a crucial role in preserving the mature acinar identity but inhibits KRAS-induced tumorigenesis (Fig. 2) [58,59,60,61].
Generation of pancreatic preneoplastic ductal cells from acinar cells. Acinar cells harboring oncogenic KRAS can convert to preneoplastic ductal cells with the help of PI3K p110. The process can be inhibited by mature acinar cell maintaining factor PTF1A. Acinar cells also can have similar conversion through a process known as acinar-ductal metaplasia or ADM which is promoted by KLF4 overexpression. The figure is created in BioRender.com
A serine/threonine kinase protein kinase D1 (PRKD1) plays a crucial role in converging oncogenic and wild-type KRAS signaling. PRKD1 upregulates SOX9 and PDX1 via neurogenic locus notch homolog protein 1 (NOTCH1) activation [50, 54]. PRKD1 also links oncogenic KRAS signaling by activating NF-κB and increasing mitochondrial reactive oxygen species (ROS) [50]. Lineage tracing experiment in the animal model has shown that oncogenic KRASG12D transdifferentiation pushes the cells towards PanIN1A, PanIN1B, and PanIN2 [62]. Further progression involves wild-type KRAS (EGFR signaling), inflammation, and additional gene mutations [49, 50, 63,64,65,66,67,68]. The remarkable flexibility of acinar cells, essential for their regenerative capacity following pancreatic injury, renders them susceptible to enduring transdifferentiation into PanIN cells upon exposure to oncogenic stress.
2.2 Epithelial-mesenchymal plasticity
Epithelial cell plasticity refers to the dynamic and flexible nature of these cells to undergo various changes in their characteristics, behaviors, and functions, especially mesenchymal phenotype. In normal circumstances, epithelial cells maintain their specific identity and functions within tissues, but in cancer, they can exhibit a remarkable degree of plasticity which allows them to acquire traits contributing to the heterogeneity and aggressiveness of the tumor. Changes in tumor cell motility and invasiveness often align with shifts in epithelial plasticity [69].
In pancreatic cancer, epithelial plasticity is modulated by several factors. A recent study published in 2019 reported a significant role of TANK-binding kinase 1 (TBK1) in the regulation of epithelial plasticity in KRAS-mutant PDAC towards enhanced invasion and metastasis. The TBK1 activity was found to be induced by receptor tyrosine kinase AXL in a RAS-Ras-related protein Ral-B (RALB)-dependent manner [70]. The study also found accumulation of fibrillar collagen in TBK1 genetically modified KrasLSL−G12D/+ Cdkn2aLox/Lox Ptf1aCre/+ (KIC) mice which is a hallmark of pancreatic cancer promoting EMT [70,71,72]. On the contrary, metadherin (MTDH), a single-pass transmembrane protein associated with cell proliferation in embryogenesis shown to reverse EMT or promote MET process of PDAC cells to facilitate metastatic colonization by downregulating twist-related protein 1 (TWIST1) and upregulating E-cadherin. The study also performed chromatin immunoprecipitation (ChIP) sequence analysis and revealed paired related homeobox 1A (PRRX1A) as an upstream regulator of MTDH [73, 74]. In pancreatic neuroendocrine neoplasm, EMT plasticity is shown to increase as the tumor progresses. Venugopal et al. demonstrated 17 EMT markers including CD24, CD44, DAXX, and vimentin that are significantly increased in the tissue of grade 3 pancreatic neuroendocrine tumors (PanNETs) which were 9 and 11 in grade 1 and grade 2 PanNETs, respectively [75]. Higher expression of vimentin along with E-cadherin loss is also evident in PanNETs which is associated with a poor prognosis [76]. Such overexpression of EMT markers in PanNETs could be associated with CSC marker doublecortin-like kinase 1 (DCLK1) [77].
Recent research conducted by Reichert et al. in 2018 delved into the mechanisms controlling epithelial plasticity and cell fate decisions between liver and lung colonization or metastatic organotropism in PDAC [13, 78]. They focused on the role of p120 catenin (P120CTN), a protein involved in E-cadherin binding and stabilization. Through experiments using PDAC mouse models, they found that intact or mono-allelic P120CTN expression favored liver metastasis, while bi-allelic gene deletion shifted metastatic burden towards the lungs. One allele of P120CTN was sufficient for stable E-cadherin in liver metastasis, but lung metastases lacked both proteins, suggesting non-MET-capable epithelial character. Injections of PDAC cells with different P120CTN status into mouse organs confirmed these results, indicating that epithelial plasticity is not the sole determinant of metastatic potential. Why different organs prefer diverse cell phenotypes remain unclear. Understanding such molecular mechanisms is vital for advancing therapies in pancreatic cancer. Emerging evidence suggests complex epithelial plasticity involving partial EMT (P-EMT), where cells exhibit both phenotypes and exhibit collective migration [79].
The plasticity of epithelial cells in pancreatic cancer is a complex phenomenon driven by genetic mutations, epigenetic changes, and interactions with the tumor microenvironment. This plasticity not only contributes to the aggressiveness of the disease but also poses challenges for targeted therapies, as cancer cells can adapt to treatments and develop resistance through these dynamic transitions. Understanding and targeting these plasticity mechanisms hold promise for developing more effective strategies to combat pancreatic cancer. Exploring metastatic organotropism and developing genetic manipulation tools could offer novel PDAC therapeutic avenues and possibly improving patient outcomes.
2.3 Epigenetic plasticity
It is recognized that PDAC originates from the collaboration of genetic and epigenetic reprogramming occurrences [80]. Genetically, PDAC exhibits lower heterogeneity in comparison to other types of cancers, and the prevalent KRAS mutation alone is insufficient to drive neoplastic transformation without the presence of inflammation or additional mutations [66, 81, 82]. It has been demonstrated that oncogenic KRAS collaborates with inflammation to initiate significant chromatin remodeling that stimulates the onset of tumor [83,84,85]. Nonetheless, the mechanism through which KRAS-mediated plasticity leads to the emergence of neoplastic lineages and facilitates their subsequent progression into invasive disease remains unclear. Very recently in 2023, Burdziak and colleagues demonstrated how plasticity develops during the initial phases of tumor formation by evaluating epigenetic landscape in PDAC [86]. Cellular plasticity is primarily established at the chromatin level, involving expansions or contractions in the array of transcriptional programs accessible to a specific cell [87, 88]. Cells with pronounced plasticity, like stem cells, frequently exhibit an open chromatin stage [89, 90]. Utilizing single-cell genomics (RNA-seq and ATAC-seq), computational methods, and functional perturbation in autochthonous genetically engineered mouse models, it was shown that certain KRAS mutant cell population in response to inflammation exhibits high epigenetic plasticity scores. These adaptable cellular states have an abundance of open chromatin in the vicinity of genes responsible for cell–cell communication, encompassing ligands, and cell-surface receptors. This implies an elevated inclination for interacting with the surrounding microenvironment. This group has identified a reciprocal interaction between inflammation-driven epithelial and immune cell states, which involves IL-33, a factor previously linked to pancreatic tumorigenesis [86]. These findings reveal a distinct tissue remodeling program specific to neoplasia, which could potentially be utilized for pancreatic cancer intervention.
2.4 Metabolic plasticity
Metabolic plasticity in pancreatic cancer refers to the tumor’s ability to adapt and switch between different metabolic pathways in order to sustain its growth, survival, and proliferation under varying conditions [91]. Pancreatic cancer is known for its aggressive nature and resistance to conventional treatments, partially attributed to its ability to rewire its metabolic processes [92,93,94]. Metabolic reprogramming, an emerging hallmark of PDAC, includes aerobic glycolysis, oxidative phosphorylation (OXPHOS), anti-oxidative stress, autophagy, glutaminolysis, lipogenesis, and lipolysis [95,96,97,98]. PDAC cells achieve metabolic plasticity both intrinsically and extrinsically. KRAS activation and mutations in tumor suppressor genes such as p53 act as an intrinsic factor and induce atypical mitochondrial metabolism and enhance glycolysis, with alterations in glutamine and lipid metabolism. In PDAC, KRAS rewires glucose metabolism by upregulating glucose uptake and glycolytic enzyme expression, including glucose exporter type 1 (GLUT1), hexokinase 1/2 (HK1/2), phosphofructokinase (PFK), and lactate dehydrogenase A (LDHA). These changes are regulated by c-MYC proto-oncogene (c-MYC) and hypoxia-inducible factor 1-alpha (HIF-1α). Autophagy induced by KRAS also supports glycolysis [99]. KRAS promotes anabolic metabolism, directing glycolysis intermediates towards the pentose phosphate pathway (PPP) non-oxidative arm for nucleotide biosynthesis [100]. Glutaminolysis, regulated by KRAS, is crucial for detoxification and biosynthesis [100,101,102]. Additionally, KRAS enhances glucose flux via the hexosamine biosynthesis pathway (HBP), which bridges glycolysis and glutaminolysis. PDAC cells use a noncanonical KRAS-induced glutamine pathway to maintain redox balance and support cell growth by increasing the nicotinamide adenine dinucleotide phosphates (NADPH/NADP +) ratio [98].
The acidic and oxygen/nutrient deprivation act as extrinsic factors and promote cancer cells to reprogram metabolism. The stromal cells, especially cancer-associated fibroblasts (CAFs) and immunocytes, play a significant role in metabolic reprogramming. Both intrinsic and extrinsic factors exert independent and synergistic influences on metabolic adaptability. Enhancing our comprehension of the metabolic characteristics of PDAC will play a crucial role in advancing innovative approaches for diagnosis and treatment [98].
Tumor cells and stromal cells engage in metabolic signaling to prioritize cancer cells for glucose utilization during the scarcity of nutrients. Stromal cell metabolism has been relatively underexplored, despite their prevalence in tumors, accounting for as much as 80–90% in desmoplastic pancreatic cancer [103, 104]. Cancer cells with various phenotypes use various energy sources through multiple metabolic pathways. Hypoxic cancer/stromal cells (e.g., CAFs) express higher levels of GLUT1 and utilize transported glucose for lactate production. Aerobic cancer cells receive lactate shuttle through communication between transporters monocarboxylate transporter 1 (MCT1) and MCT4, a phenomenon known as reverse Warburg effect [98].
Stromal CAFs are commonly identified to stimulate tumor advancement and metastasis [105, 106]. These CAFs undergo metabolic reprogramming, use more glucose, and secrete excess lactate mimicking the phenotype related to the Warburg effect [98, 107]. CAFs also communicate with cancer cells via exosomes in nutrient-stressed conditions [108]. Specialized CAFs called pancreatic stellate cells (PSCs) can swap between quiescent and activated phenotypes [109]. Upon activation, PSCs may release ECM molecules, intensifying hypoxia-induced metabolic reprogramming of the cancer cells [109, 110].
Targeting the metabolic plasticity of pancreatic cancer cells has emerged as a potential therapeutic strategy. Metabolic symbiosis consequently triggers a condition of “fibroblast addiction” in both primary and metastatic tumor cells, revealing what appears to be a significant vulnerability [111]. However, developing effective therapies requires a deep understanding of the complex metabolic rewiring that occurs in pancreatic cancer cells [98].
3 Stemness in pancreatic cancer
Stemness is the combination of the ability of a cell to replicate its lineage, to differentiated cells, and to interconnect with its environment to maintain homeostasis in proliferation and regeneration [112]. CSCs are polymorphic, capable of differentiated and self-renewable subpopulations of tumors, which are responsible for resistance to chemotherapy and radiation therapy. CSCs in the pancreas are conducive to help in the development of PDAC. Over the past two decades, numerous specific biomarkers have been demonstrated within the realm of CSCs, such as CD44, CD24, ESA, and CD133, while other surface markers continue to be recognized as distinct features of this tumor cell population and would be found to describe elsewhere [113,114,115]. Pancreatic CSCs together with other cells, such as PSCs, CAFs, and mesenchymal stem cells, form a highly desmoplastic tumor microenvironment (TME). Understanding the CSCs of pancreatic cancer can guide the design of potent therapies against PDAC. In PDAC, it has been demonstrated that fluorescence-activated cell sorted (FACS) CD44 + /CD24 + /ESA + cells act as CSC subpopulation [113]. The study confirmed that this subpopulation could recapitulate the primary tumors in in vivo model systems [113].
The flow cytometric isolation of the secondary tumors further confirmed that the frequencies of constituent subpopulations closely resembled those initially identified during the cell sorting process of the primary resected neoplasm. Another distinct characterization of pancreatic CSCs was isolated by Hermann et al. who employed selection marker CD133 for the stem cell subpopulation [116]. Through serial passaging experiments of the CD133 + subpopulation, they demonstrated the cells’ capability to regenerate pancreatic tumor and undergo differentiation, mirroring the characteristics CD44 + /CD24 + /ESA + subpopulation mentioned above. Further investigation of pancreatic CSCs which are CD133 and C-X-C chemokine receptor type 4 (CXCR-4) positive showed its highly migratory potential towards the CXCR4 ligand chemokine SDF-1α. The inhibition of migration by anti-CXCR4 antibodies affirms the involvement of CXCR4 on the migratory nature of CSCs [116].
In addition to the investigation of pancreatic CSCs in the human cell line, researchers have created a PDAC murine model by inducing pluripotent stem cells (iPSCs), which can differentiate and transfer into progenitor cells under certain conditions [117]. It was observed that cell signaling within the niche, functioning akin to the TME, not only influenced the differentiation of CSCs but also conferred them with the ability for self-renewal [117]. Based on those novel findings, Calle et al. developed a second-generation PDAC murine model using a PDAC cell line-condition medium. Under the exposure of those mediums, iPSCs converted to CSCs, representing the upregulation of the receptor tyrosine-protein kinase ErbB-2 (ERBB2/3) cell signaling pathway in the initiation of pancreatic cancer cells. Inhibition of ERBB2/3 chemically and genetically generated three promising properties: (1) less tumorigenicity and stemness in CSCs, (2) abolishing tube formation and self-renewal ability, and (3) downregulating CD24 levels with activated mitogen-activated protein kinase kinase/ERK (MEK/ERK) cascade [118]. To further understand the formation of CSCs, Hassan et al. elucidated the abnormal overexpression of cyclin D2 (CCND2), collagen type IV alpha 1/2 chain (COL4A1/2), and actinin alpha 3 (ACTN3), which are associated with cancer aggressiveness. These genes were linked to oxidative phosphorylation and glycolysis, indicating the metabolic regulations in pancreatic CSCs [119].
Recently, CSCs have been identified in PanNETs which are marked by aldehyde dehydrogenases (ALDH) or CD90. These markers are involved in several key signaling pathways such as Src, Notch, Hedgehog, Wnt/β-catenin, EGFR, mammalian target of rapamycin (mTOR), and signal transducers and activators of transcription 3 (STAT3). Further study revealed cell surface markers CD47 and CD73 as the vulnerability of ALDH + PanNETs CSCs [120,121,122].
4 Cellular plasticity and therapy resistance in pancreatic cancer
Pancreatic tumors are highly heterogeneous, containing a mix of different cell populations with varying characteristics [123, 124]. Cellular plasticity allows some cancer cells to switch between different states, such as epithelial and mesenchymal morphologies [11, 125, 126]. Cellular plasticity can lead to changes in cellular metabolism, allowing cancer cells to rely on different energy sources or bypass metabolic vulnerabilities targeted by therapies [127]. Cellular plasticity contributes to the Darwinian evolution of cancer. Treatment exerts selective pressure on the tumor, favoring the survival and expansion of cells with plasticity-related resistance traits. Over time, this can lead to the emergence of highly therapy-resistant cell populations. Cellular plasticity can also promote tumor recurrence after initial treatment success. Residual cancer cells with plasticity-driven resistance traits can lie dormant and later reinitiate growth [128]. Additionally, plasticity plays a role in the formation of metastases, contributing to the spread of cancer cells to distant sites. Generalized plasticity-mediated therapy resistances are illustrated in Fig. 3.
Cellular plasticity–mediated therapy resistance in pancreatic cancer. FOLFIRINOX and gemcitabine both can generate quasi-mesenchymal or CSC type subpopulation that can lead to therapy resistance. Genetic events such as GATA6 loss also can induce cellular plasticity and can suppress certain MHC Class I genes resulting in less immune cell infiltration in the tumor. Extracellular vesicle-mediated signaling can enhance the communication between CSCs and non-CSC which also cause therapy resistance. PDAC, pancreatic ductal adenocarcinoma; CSC, cancer stem cells; MHC, major histocompatibility complex. The figure is created in BioRender.com
4.1 Role of cellular adaptation
PDAC has been categorized into two main subtypes based on transcriptional profiling: classical epithelial (E) type and quasi-mesenchymal (QM) type [129,130,131,132]. Epithelial cells tend to be more sensitive to chemotherapy, while mesenchymal cells are often more resistant. The phenotypic switching of cancer cells can occur in response to treatment, promoting survival of the resistant cell population. Recent evidence suggests that these subtypes exist on a spectrum, implying a potential for interconversion between them. It has been demonstrated that the FOLFIRINOX chemotherapy causes both E and QM PDAC to shift more towards the QM state, both in cell lines and patient tumors. Such QM plastic shift enables PDAC to be intrinsically resistant to chemotherapy [129]. Similarly, Kloesch and colleagues observed distinct clusters in PDAC cells through RNAseq analysis [133]. They identified cell lines expressing GATA binding protein 6 (GATA6) as epithelial clusters, whereas those lacking GATA6 as mesenchymal clusters. The study further demonstrated that the absence of GATA6 promotes cellular plasticity and evasion of the immune system. GATA6-positive cell lines exhibited an enrichment of genes related to “apical junctions” in gene set enrichment analysis (GSEA). In contrast, GATA6-deficient cells displayed an enrichment of the EMT gene set and a reduction of major histocompatibility complex (MHC) class I genes H2-d1 and H2-k1 expression. The loss of GATA6 was linked to a reduction in the infiltration of CD8α + T-cells into the tumor, suggesting that GATA6 loss may trigger immune evasion, resulting in resistance to immunotherapy [133].
4.2 Role of cancer stem cells
Cellular plasticity can endow cancer cells with stem cell-like properties, such as self-renewal and ability to differentiate [134]. CSCs, which arise from this plasticity, are often more resistant to therapies due to their ability to repair DNA damage and evade cell death mechanisms [135,136,137,138]. Therapy resistance and drug evasion in pancreatic cancer’s CSCs can be attributed to two distinct mechanisms: (1) interactions between CSCs and the TME and (2) cell signaling pathways involved in CSCs. The transmission of cellular components directly from one cell to another through cell fusion, exosomes, and tunneling nanotubes has been proposed as alternative mechanisms for intercellular communication. The direct exchange of cellular elements, such as proteins, DNA, messenger RNA (mRNA), microRNA (miRNA), long noncoding RNA (lncRNA), and organelles, between cells is a pivotal factor in the modulation of cancer cell tumorigenicity and metastatic capabilities within the TME [139,140,141,142]. Jang et al. discovered that gemcitabine chemotherapy could potentially induce intercellular transfer of molecules within the TME, leading to the reprogramming of pancreatic cancer cells. This reprogramming process enhances their tumorigenic potential and self-renewal capabilities and ultimately therapeutic resistance [143]. The phenotypes represent the upregulation of CD24 and CD44, which are the important markers of CSCs. Interestingly, the CD44 + subpopulation was found to exhibit an increased invasiveness and metastasis; on the other hand, CD24 + cells are characterized by their elevated aldehyde dehydrogenase activity. These CSCs preserve their characteristics through reprogramming themselves which involves epigenetic modulations and intercellular biomolecular interactions. Therefore, inhibition of CD44 is considered as a potential strategy for overcoming the emergence of refractoriness and recurrence in pancreatic cancer [143].
4.3 Role of metabolism
It has been demonstrated that metabolic plasticity confers resistance to erlotinib through the increased expression of glucose-6-phosphate dehydrogenase (G6PD). Pancreatic cancer cells enhance the expression of pentose phosphate pathway (PPP) including G6PD and protect cells from ROS by increasing NADPH/NADP + ratio. Mechanistically, cancer cells elevate the expression of the inhibitor of differentiation (ID1), which subsequently increases G6PD levels, leading to a modified metabolic profile and resistance to erlotinib. Inhibition of PPP using 6-aminonicotinamide (6AN) and siRNA-mediated silencing experiments confirmed such finding [144].
In PDAC, metabolic rewiring prioritizes anabolic processes that provide the essential cellular components required for unrestricted cancer cell proliferation. The activation of KRAS, along with autophagy and macropinocytosis, empowers PDAC cells to adjust to varying nutrient and oxygen levels. This remarkable capacity for adaptation has been identified as a factor contributing to the profound resistance to therapeutic interventions [145]. Autophagy bestows therapy resistance (chemotherapy, radiation) by enhancing cellular plasticity via supplying metabolites. Macropinocytosis works in a similar way and utilizes lysosomes to release metabolites to the cells. Consequently, multiple clinical trials (NCT01978184, NCT01128296) are currently in progress to evaluate the efficacy of hydroxychloroquine (HCQ) treatment in PDAC. Initial findings suggest the potential for enhanced outcomes when HCQ is incorporated into chemotherapy regimens for localized PDAC [146]. Plasticity of PDAC cells allows reactivation of ERK thus confers resistance to inhibitors that target upstream molecules in the RAS/RAF signaling pathway. Hence, directly targeting ERK may represent a more efficacious treatment approach [147, 148].
In metabolically rewired cells, fatty acid synthase (FASN) facilitates condensation reactions, while converting acetyl-CoA into malonyl-CoA via acetyl-CoA carboxylase. A recent investigation has revealed a connection between elevated FASN expression and gemcitabine resistance. Combining the FASN inhibitor orlistat with gemcitabine produces a synergistic response [149, 150]. Resistance to gemcitabine has also been observed to be linked to HIF1-dependent metabolic adaptation. It has been shown that by enhancing pyrimidine biosynthesis through the interplay of HIF1 with MUC1, PDAC tumors acquire resistance to gemcitabine [151]. The plastic nature of PDAC cells demonstrated resistance to glutaminase (GLS) inhibitor CB-839 when treated as a single agent. Despite cancer cells relying on GLS, their metabolic adaptability enables them to rewire alternative mechanisms for glutamate acquisition, leading to therapy resistance [152].
4.4 Other factors
The tumor microenvironment contributes to cellular plasticity and therapy resistance [36, 153]. Interactions with stromal cells, immune cells, and extracellular matrix components can induce plasticity in cancer cells [154]. The microenvironment can also provide protective niches that shield cancer cells from the effects of therapies. In addition, targeted therapies often aim to block specific signaling pathways driving cancer growth. However, cancer cells can switch to alternative pathways or bypass the blocked pathways through plasticity, rendering the targeted therapy ineffective [127]. EMT is a critical process associated with cellular plasticity [155, 156]. During EMT, cancer cells lose epithelial appearances (adhesion, polarity) and gain mesenchymal traits (motility, invasiveness) [157]. This transition can lead to increased resistance to chemotherapy and other targeted therapies.
Extracellular vehicles (EVs) contribute to communication between CSCs and non-CSCs in pancreatic tumors. Those EVs undergo conformational changes under exposure to chemotherapy or hypoxia that might thus represent the underlying mechanism that cancer cells can adapt to the changing environment thereby supporting cancer progression [158]. Moreover, the study revealed that CSC-derived agrin (AGRN)-positive EVs not only regulate Yes1-associated transcriptional regulator (YAP) cell signaling but also influence the AGRN/HIPPO axis, establishing a communication link between CSCs and non-CSCs. This observation demonstrated that the cross-linkage interaction routes inside the tumor milieu support CSC growth and resistance in PDAC [158].
Several cell signaling pathways participate in enhancing stemness of CSCs after radiotherapy and chemotherapy. Damage-associated molecular pattern (DAMP) molecule high mobility group box 1 (HMGB1) was considered as a mediator of crosstalk between the post-radiotherapy necrotic environment and resting CSCs. The binding of HMGB1 to toll-like receptor 2 (TLR2) causes inflammation that could further upregulate self-renewal potential in CD133 + pancreatic CSCs and downregulate WNT/β-catenin signaling results in therapeutic resistance [159]. Another interesting finding is the overexpression of ubiquitin-specific peptidase 22 (USP22) in cisplatin-resistant pancreatic CSCs [160]. Downregulation of USP22 could re-sensitize CSCs to cisplatin and influence WNT/β-catenin signaling. Furthermore, alterations in heat shock transcription factor 1 (HSF1), forkhead box M1 (FOXM1), and mitogen-activated protein kinase 14 (p38) have been documented as successful strategies to reverse resistance and mitigate stemness in pancreatic CSCs [161,162,163].
Understanding the mechanisms underlying cellular plasticity and its contributions to therapy resistance is crucial for developing more effective treatment strategies. Targeting pathways involved in plasticity, combining therapies to tackle multiple phenotypic states, and disrupting interactions between cancer cells and the microenvironment are potential approaches to overcome therapy resistance in pancreatic cancer.
5 Therapeutic strategies targeting cellular plasticity in pancreatic cancer
5.1 Reprogramming cancer cells
Therapeutic strategies targeting cellular plasticity in pancreatic cancer typically aim to disrupt or reverse the processes that allow cancer cells to change their characteristics and adapt to different microenvironments. The idea of reprogramming cancer cells into a benign counterpart is not a novel, and the use of all-trans retinoic acid in promyelocytic acute myelogenous leukemia is considered as an early example. However, reprogramming solid tumors is not common. In 2015, Kim and colleagues provided the initial transcriptional proof of PDAC cells undergoing reprogramming into acinar cells. Quantitative PCR confirmed the elevated expression of acinar cell lineage genes, such as serine protease 2 (PRSS2), chymotrypsin-like elastase 3 (CELA3), CPA2, and MIST1, in three distinct PDAC cell lines but not ductal genes. Moreover, chromatin immunoprecipitation revealed enhanced the basic helix-loop-helix transcription factor E47 binding at the promoter of MIST1. In vivo assessment of tumor development showed that the presence of E47 in Panc1 cells led to a notable reduction in tumor growth. Additionally, the study presented evidence of trypsinogen-positive staining in the tumor originating from PDAC/E47 cells, suggesting a persistent acinar cell reprogramming effect of E47 on Panc1 cells in animals [164]. There are a number of additional therapeutic strategies targeting cellular plasticity in PDAC. For example, the activation of urokinase plasminogen activator surface receptor (uPAR) serves as a strong unfavorable predictor in the prognosis of PDAC. uPAR and mutated KRAS collaborate to transition the tumor from a quiescent epithelial state to an active mesenchymal one, potentially elucidating the grim prognosis associated with high uPAR levels in PDAC. Interestingly, this active mesenchymal state appears to be more susceptible to gemcitabine. Therefore, strategies aimed at targeting either KRAS or uPAR should take into account the possibility of this tumor-escaping mechanism (Fig. 4) [165]. In fact, mutant KRAS specially KRASG12D has been targeted in PDAC successfully and recently reviewed by Wei et al. [166].
Targeting cellular plasticity to overcome therapy resistance in pancreatic cancer. Current strategies involve cellular reprogramming, chemosensitization, pathway targeting specifically mTOR, and metabolic targeting. Red arrow indicates upregulation of genes. SHH, sonic Hedgehog; CSC, cancer stem cells; OXPHOS, oxidative phosphorylation; bHLH, basic helix-loop-helix. The figure is created in BioRender.com
5.2 Targeting mTOR pathway
The mammalian target of rapamycin (mTOR) inhibitor was the first recognized therapeutic strategy for pancreatic CSCs. The traditional mTOR inhibitor rapamycin can reduce the viability of CD133 + CSCs, which previously correlated with increased metastasis and served as a poor prognosis biomarker in cancer cells. Matsubara et al. found that treatment with mTORC1/2 inhibitor KU-0063794 inhibits the downstream signaling of mTORC1 and decreased CD133 + CSC’s viability. Surprisingly, the inhibitor did not affect AKT phosphorylation [167]. This discovery was interpreted as an implication of mTORC1 signaling in sustaining the formation of CSCs in the pancreas [167]. Furthermore, the mTOR1/2 inhibitor was found to have an impact on the Hedgehog (HH)/Gli cascade and the Sonic Hedgehog (SHH) pathway, offering a potential means to eliminate pancreatic CSCs and their stemness properties [168]. Due to SHH’s link with enhancing stemness and invasiveness in pancreatic cancer, there has been interest in pre-clinical studies to explore the potential of combining inhibitors targeting SHH pathway with chemotherapy (Fig. 4). However, as reported by de Jesus-Acosta et al., this combination approach did not succeed in altering the stromal components of CSCs and did not achieve the intended results in the group of patients under investigation [169].
5.3 Targeting epigenetic modulator
Histone deacetylase (HDAC) inhibitor trichostatin A was reported to re-sensitize chemotherapy-resistant pancreatic CSCs by regulating them epigenetically. HDAC inhibitor, domatinostat, has been indicated to re-sensitize PDAC under the gemcitabine/taxol combination treatment in vitro and in vivo [170]. Domatinostat not only impedes the activity of FOXM1, a key player in stemness and DNA repair, but also governs cellular and mitochondrial oxidative stress in PDAC. However, the underlying mechanism of HDAC inhibitors still need further investigation [170].
5.4 Targeting cellular metabolism
Targeting cellular metabolism has the potential to diminish stemness in PDAC. Cepeloa et al. demonstrated that the mitochondria-targeted antioxidant MitoQ had the capacity to inhibit mesenchymal PDAC cells and mitigate the expression of stemness markers [171]. Pancreatic CSCs that are positive for CD133 and CD44 are shown to rely on oxidative phosphorylation (OXPHOS) and thus sensitive to OXPHOS inhibitors such as oligomycin and associated with tumor regression in vivo [172]. Such metabolic (OXPHOS) vulnerability also has been successfully targeted by metformin in PDAC models (Fig. 4) [173, 174]. These discoveries indicate that focusing on the reliance on metabolic reprogramming in cells undergoing plasticity could yield favorable outcomes [175, 176].
5.5 Targeting intracellular kinases
Inhibiting intracellular kinases could potentially impede the formation of CSCs. For instance, the multi-kinase inhibitor sorafenib has been demonstrated to repress PDAC CSCs which is evident from reduced tumor sphere formation, aldehyde dehydrogenase 1 (ALDH1) activity, and tumor-initiating capacity [177]. Targeting of overactive mitogen-activated protein kinase (MAPK) or WNT/β-catenin signaling along with standard chemotherapy suppresses CSCs and shows significant potential in eradicating recurrence. Moreover, targeting c-Jun N-terminal kinases (JNK) pathway increased PDAC CSC sensitivity to 5-fluorouracil (5-FU) or gemcitabine [178]. Sensitization to gemcitabine also can be enhanced by phytochemicals such as resveratrol by downregulating zinc finger E-box-binding homeobox 1 (ZEB1), zinc finger protein SNAI1 (SNAIL), zinc finger protein SNAI2 (SLUG), homeobox transcription factor nanog (NANOG), POU class 5 homeobox 1 (POU5F1/OCT4), and ATP-binding cassette (ABC) transporters, which are associated with CSC maintenance [179].
5.6 Other targets
The RNA polymerase II-associated factor 1 (PAF1) and Yes-associated protein 1 (YAP1) proteins engage in different cellular processes, comprising transcriptional regulation, stemness, and cell growth. Targeting of the PAF1/YAP1 axis is considered as an interesting strategy to regulate stemness. C3, an inhibitor of YAP1, can decrease the association of the PAF1/YAP1 complex with the SOX9 promoter, leading to a reduction in stemness and mitigating the progression of PDAC cells [180]. Notably, Zhang and colleagues demonstrated that aspirin possesses the capability to target PDAC CSCs by diminishing their self-renewal capacity and inducing apoptosis in an in vivo setting [181]. Likewise, the correlation was observed when aiming to target CD44 or impede ABC transporters with verapamil, resulting in a reduced frequency of CSCs and a restoration of sensitivity to gemcitabine [182]. Targeting CSC by phytochemical is another emerging strategy and was recently reviewed by Patil et al. [114]. Furthermore, targeting CSC-specific surface markers, signal transducer, and activator of transcription 3 (STAT3) signaling and PI3K/mTOR signaling by monoclonal antibodies or small molecule inhibitors is another effective strategy to overcome resistance developed by cellular plasticity [114].
A preclinical evaluation of galunisertib demonstrated its capability to modulate the characteristics of the mouse pancreatic cancer cell line KPC-M09. It achieved this by inhibiting the TGF-β-induced reduction of E-cadherin. Galunisertib was additionally effective in restoring the functional capabilities of both CD8 + T cells and NK cells, which had been inhibited by TGF-β in vitro [183]. In Phase II clinical investigations, galunisertib exhibited certain levels of effectiveness against tumors in patients diagnosed with PDAC [184]. These investigations, along with others, have offered compelling proof that innovative medications can influence or revert tumor cell plasticity and EMT, potentially making tumor cells more responsive to immune-mediated cell death [36]. An improved strategy for addressing PDAC involves simultaneously targeting of CSCs and non-CSCs. In this regard, salinomycin, an antibiotic in combination with gemcitabine, has demonstrated remarkable effectiveness in PDX models [185]. Given that research has demonstrated the enhanced effectiveness of a combined treatment regimen incorporating both chemotherapy and a CSC-inhibitor compared to single-agent therapy in a typically resistant form of pancreatic cancer, it is clear that targeting CSCs should be an essential component of the comprehensive treatment approach. Thus, there have been suggestions for inventive strategies aimed at restoring the sensitivity of CSCs. These include the use of a blend of medications that target ABC transporters, signaling pathways related to stemness, DDR machinery, immune checkpoint proteins, desmoplasia, and fibrosis, as well as metabolic reprogramming.
6 Conclusions and future directions
Cellular plasticity is a fundamental driver of therapy resistance in PDAC. The ability of cancer cells to switch between different states and adapt to changing conditions poses a significant challenge in treatment. Understanding the underlying molecular mechanisms of cellular plasticity and its role in therapy resistance provides a foundation for developing innovative therapeutic strategies. As we uncover more about the intricacies of this process, the potential for novel therapeutic interventions becomes increasingly promising, offering renewed hope for better outcomes in pancreatic cancer treatment. Though pancreatic cancer involves both ductal adenocarcinoma and neuroendocrine neoplasms, cellular plasticity–mediated therapy resistance is mostly studied in ductal adenocarcinoma.
Further exploration of the molecular mechanisms underlying cellular plasticity and therapy resistance is crucial. Identifying specific molecular targets within the pathways associated with plasticity could lead to the development of targeted therapies that disrupt these processes. Considering the complexity of cellular plasticity, combination therapies that target both primary tumor cells and the microenvironment may be more effective in overcoming therapy resistance. This could involve a combination of targeted therapies, immunotherapies, and therapies targeting the tumor microenvironment. Research efforts should focus on identifying biomarkers that can predict the development of therapy resistance through cellular plasticity. This would enable early intervention and personalized treatment strategies. Utilizing single cell sequencing and other advanced technologies such as spatial profiling can provide insights into the heterogeneity of cell populations within tumors, shedding light on the dynamics of cellular plasticity and its role in therapy resistance. This could involve targeting specific signaling pathways or immune cell interactions. Investigating strategies to manipulate the tumor microenvironment to prevent the emergence of therapy-resistant phenotypes is a promising avenue.
References
Mizrahi, J. D., et al. (2020). Pancreatic cancer. Lancet, 395(10242), 2008–2020.
Siegel, R. L., et al. (2023). Cancer statistics, 2023. CA: A Cancer Journal for Clinicians, 73(1), 17–48.
Hidalgo, M., et al. (2015). Addressing the challenges of pancreatic cancer: Future directions for improving outcomes. Pancreatology, 15(1), 8–18.
Schober, M., et al. (2014). Desmoplasia and chemoresistance in pancreatic cancer. Cancers (Basel), 6(4), 2137–2154.
WHO Classification of Tumours Editorial Board. (2019). Digestive System Tumours: WHO Classification of Tumours, 5th ed. Vol. 1.
Stewart, B. W., et al. (2014). World Cancer Report 2014: World Cancer Reports.
Pishvaian, M. J., & Brody, J. R. (2017). Therapeutic implications of molecular subtyping for pancreatic cancer. Oncology (Williston Park), 31(3), 159–66. 168.
Fitzgerald, T. L., et al. (2008). Changing incidence of pancreatic neoplasms: A 16-year review of statewide tumor registry. Pancreas, 37(2), 134–138.
Siegel, R. L., et al. (2022). Cancer statistics, 2022. CA: A Cancer Journal for Clinicians, 72(1), 7–33.
Gittes, G. K. (2009). Developmental biology of the pancreas: A comprehensive review. Developmental Biology, 326(1), 4–35.
Rhim, A. D., et al. (2012). EMT and dissemination precede pancreatic tumor formation. Cell, 148(1–2), 349–361.
Farrell, A. S., et al. (2017). MYC regulates ductal-neuroendocrine lineage plasticity in pancreatic ductal adenocarcinoma associated with poor outcome and chemoresistance. Nature Communications, 8(1), 1728.
Reichert, M., et al. (2018). Regulation of epithelial plasticity determines metastatic organotropism in pancreatic cancer. Developmental Cell, 45(6), 696-711 e8.
Crawford, H. C., Pasca di Magliano, M., & Banerjee, S. (2019). Signaling networks that control cellular plasticity in pancreatic tumorigenesis, progression, and metastasis. Gastroenterology., 156(7), 2073–2084.
Shen, S., & Clairambault, J. (2020). Cell plasticity in cancer cell populations. F1000Res, 9, 635.
Yuan, S., Norgard, R. J., & Stanger, B. Z. (2019). Cellular plasticity in cancer. Cancer Discovery, 9(7), 837–851.
Rambow, F., Marine, J. C., & Goding, C. R. (2019). Melanoma plasticity and phenotypic diversity: Therapeutic barriers and opportunities. Genes & Development, 33(19–20), 1295–1318.
Qin, S., et al. (2020). Emerging role of tumor cell plasticity in modifying therapeutic response. Signal Transduction and Targeted Therapy, 5(1), 228.
Kemper, K., et al. (2014). Phenotype switching: Tumor cell plasticity as a resistance mechanism and target for therapy. Cancer Research, 74(21), 5937–5941.
Gupta, P. B., et al. (2019). Phenotypic plasticity: Driver of cancer initiation, progression, and therapy resistance. Cell Stem Cell, 24(1), 65–78.
Zhuang, X., Zhang, H., & Hu, G. (2019). Cancer and microenvironment plasticity: Double-edged swords in metastasis. Trends in Pharmacological Sciences, 40(6), 419–429.
Smigiel, J. M., et al. (2019). Cellular plasticity and metastasis in breast cancer: A pre- and post-malignant problem. Journal of Cancer Metastasis and Treatment, 5, 47.
Contreras-Trujillo, H., et al. (2021). Deciphering intratumoral heterogeneity using integrated clonal tracking and single-cell transcriptome analyses. Nature Communications, 12(1), 6522.
Li, M., et al. (2020). An algorithm to quantify intratumor heterogeneity based on alterations of gene expression profiles. Communications Biology, 3(1), 505.
Hinohara, K., & Polyak, K. (2019). Intratumoral heterogeneity: More than just mutations. Trends in Cell Biology, 29(7), 569–579.
Sun, X. X., & Yu, Q. (2015). Intra-tumor heterogeneity of cancer cells and its implications for cancer treatment. Acta Pharmacologica Sinica, 36(10), 1219–1227.
Nabi, K., & Le, A. (2021). The intratumoral heterogeneity of cancer metabolism. Advances in Experimental Medicine and Biology, 1311, 149–160.
Xiao, Z., Dai, Z., & Locasale, J. W. (2019). Metabolic landscape of the tumor microenvironment at single cell resolution. Nature Communications, 10(1), 3763.
Lawson, D. A., et al. (2018). Tumour heterogeneity and metastasis at single-cell resolution. Nature Cell Biology, 20(12), 1349–1360.
da Silva-Diz, V., et al. (2018). Cancer cell plasticity: Impact on tumor progression and therapy response. Seminars in Cancer Biology, 53, 48–58.
Thiery, J. P. (2002). Epithelial-mesenchymal transitions in tumour progression. Nature Reviews Cancer, 2(6), 442–454.
Polyak, K., & Weinberg, R. A. (2009). Transitions between epithelial and mesenchymal states: Acquisition of malignant and stem cell traits. Nature Reviews Cancer, 9(4), 265–273.
Farmer, P., et al. (2009). A stroma-related gene signature predicts resistance to neoadjuvant chemotherapy in breast cancer. Nature Medicine, 15(1), 68–74.
Shibue, T., & Weinberg, R. A. (2017). EMT, CSCs, and drug resistance: The mechanistic link and clinical implications. Nature Reviews. Clinical Oncology, 14(10), 611–629.
Byers, L. A., et al. (2013). An epithelial-mesenchymal transition gene signature predicts resistance to EGFR and PI3K inhibitors and identifies Axl as a therapeutic target for overcoming EGFR inhibitor resistance. Clinical Cancer Research, 19(1), 279–290.
Horn, L. A., Fousek, K., & Palena, C. (2020). Tumor plasticity and resistance to immunotherapy. Trends in Cancer, 6(5), 432–441.
Baccelli, I., et al. (2013). Identification of a population of blood circulating tumor cells from breast cancer patients that initiates metastasis in a xenograft assay. Nature Biotechnology, 31(6), 539–544.
Aktas, B., et al. (2009). Stem cell and epithelial-mesenchymal transition markers are frequently overexpressed in circulating tumor cells of metastatic breast cancer patients. Breast Cancer Research, 11(4), R46.
Micalizzi, D. S., et al. (2009). The Six1 homeoprotein induces human mammary carcinoma cells to undergo epithelial-mesenchymal transition and metastasis in mice through increasing TGF-beta signaling. The Journal of Clinical Investigation, 119(9), 2678–2690.
Kong, D., Hughes, C. J., & Ford, H. L. (2020). Cellular plasticity in breast cancer progression and therapy. Frontiers in Molecular Biosciences, 7, 72.
Ayob, A. Z., & Ramasamy, T. S. (2018). Cancer stem cells as key drivers of tumour progression. Journal of Biomedical Science, 25(1), 20.
PerusinaLanfranca, M., et al. (2020). Interleukin 22 signaling regulates acinar cell plasticity to promote pancreatic tumor development in mice. Gastroenterology, 158(5), 1417-1432 e11.
Quilichini, E., et al. (2019). Pancreatic ductal deletion of Hnf1b disrupts exocrine homeostasis, leads to pancreatitis, and facilitates tumorigenesis. Cellular and Molecular Gastroenterology and Hepatology, 8(3), 487–511.
Tanaka, M., et al. (2012). International consensus guidelines 2012 for the management of IPMN and MCN of the pancreas. Pancreatology, 12(3), 183–197.
Strobel, O., et al. (2007). Beta cell transdifferentiation does not contribute to preneoplastic/metaplastic ductal lesions of the pancreas by genetic lineage tracing in vivo. Proceedings of the National Academy of Sciences of the United States of America, 104(11), 4419–4424.
Grippo, P. J., et al. (2003). Preinvasive pancreatic neoplasia of ductal phenotype induced by acinar cell targeting of mutant Kras in transgenic mice. Cancer Research, 63(9), 2016–2019.
Tuveson, D. A., et al. (2006). Mist1-KrasG12D knock-in mice develop mixed differentiation metastatic exocrine pancreatic carcinoma and hepatocellular carcinoma. Cancer Research, 66(1), 242–247.
Liou, G. Y., et al. (2013). Macrophage-secreted cytokines drive pancreatic acinar-to-ductal metaplasia through NF-kappaB and MMPs. Journal of Cell Biology, 202(3), 563–577.
Logsdon, C. D., & Ji, B. (2009). Ras activity in acinar cells links chronic pancreatitis and pancreatic cancer. Clinical Gastroenterology and Hepatology, 7(11 Suppl), S40–S43.
Liou, G. Y., et al. (2016). Mutant KRas-induced mitochondrial oxidative stress in acinar cells upregulates EGFR signaling to drive formation of pancreatic precancerous lesions. Cell Reports, 14(10), 2325–2336.
Hezel, A. F., et al. (2008). Pancreatic LKB1 deletion leads to acinar polarity defects and cystic neoplasms. Molecular and Cellular Biology, 28(7), 2414–2425.
Sandgren, E. P., et al. (1990). Overexpression of TGF alpha in transgenic mice: Induction of epithelial hyperplasia, pancreatic metaplasia, and carcinoma of the breast. Cell, 61(6), 1121–1135.
Liu, J., et al. (2016). TGF-beta1 promotes acinar to ductal metaplasia of human pancreatic acinar cells. Science and Reports, 6, 30904.
Liou, G. Y., et al. (2015). Protein kinase D1 drives pancreatic acinar cell reprogramming and progression to intraepithelial neoplasia. Nature Communications, 6, 6200.
Means, A. L., et al. (2005). Pancreatic epithelial plasticity mediated by acinar cell transdifferentiation and generation of nestin-positive intermediates. Development, 132(16), 3767–3776.
Shi, G., et al. (2013). Maintenance of acinar cell organization is critical to preventing Kras-induced acinar-ductal metaplasia. Oncogene, 32(15), 1950–1958.
Wei, D., et al. (2016). KLF4 is essential for induction of cellular identity change and acinar-to-ductal reprogramming during early pancreatic carcinogenesis. Cancer Cell, 29(3), 324–338.
Baer, R., et al. (2014). Pancreatic cell plasticity and cancer initiation induced by oncogenic Kras is completely dependent on wild-type PI 3-kinase p110alpha. Genes & Development, 28(23), 2621–2635.
Wu, C. Y., et al. (2014). PI3K regulation of RAC1 is required for KRAS-induced pancreatic tumorigenesis in mice. Gastroenterology, 147(6), 1405–16 e7.
Payne, S. N., et al. (2015). PIK3CA mutations can initiate pancreatic tumorigenesis and are targetable with PI3K inhibitors. Oncogenesis, 4(10), e169.
Hill, R., et al. (2010). PTEN loss accelerates KrasG12D-induced pancreatic cancer development. Cancer Research, 70(18), 7114–7124.
Kopp, J. L., et al. (2012). Identification of Sox9-dependent acinar-to-ductal reprogramming as the principal mechanism for initiation of pancreatic ductal adenocarcinoma. Cancer Cell, 22(6), 737–750.
Ardito, C. M., et al. (2012). EGF receptor is required for KRAS-induced pancreatic tumorigenesis. Cancer Cell, 22(3), 304–317.
Ji, B., et al. (2009). Ras activity levels control the development of pancreatic diseases. Gastroenterology, 137(3), 1072–82. 1082 e1-6.
Navas, C., et al. (2012). EGF receptor signaling is essential for k-ras oncogene-driven pancreatic ductal adenocarcinoma. Cancer Cell, 22(3), 318–330.
Guerra, C., et al. (2011). Pancreatitis-induced inflammation contributes to pancreatic cancer by inhibiting oncogene-induced senescence. Cancer Cell, 19(6), 728–739.
Guerra, C., et al. (2007). Chronic pancreatitis is essential for induction of pancreatic ductal adenocarcinoma by K-Ras oncogenes in adult mice. Cancer Cell, 11(3), 291–302.
Liou, G. Y., et al. (2015). Mutant KRAS-induced expression of ICAM-1 in pancreatic acinar cells causes attraction of macrophages to expedite the formation of precancerous lesions. Cancer Discovery, 5(1), 52–63.
Krebs, A. M., et al. (2017). The EMT-activator Zeb1 is a key factor for cell plasticity and promotes metastasis in pancreatic cancer. Nature Cell Biology, 19(5), 518–529.
Cruz, V. H., et al. (2019). Axl-mediated activation of TBK1 drives epithelial plasticity in pancreatic cancer. JCI Insight, 5(9), e126117.
Aguilera, K. Y., et al. (2014). Collagen signaling enhances tumor progression after anti-VEGF therapy in a murine model of pancreatic ductal adenocarcinoma. Cancer Research, 74(4), 1032–1044.
Shintani, Y., et al. (2008). Collagen I-mediated up-regulation of N-cadherin requires cooperative signals from integrins and discoidin domain receptor 1. Journal of Cell Biology, 180(6), 1277–1289.
Suzuki, K., et al. (2017). Metadherin promotes metastasis by supporting putative cancer stem cell properties and epithelial plasticity in pancreatic cancer. Oncotarget, 8(39), 66098–66111.
Jeon, H. Y., et al. (2010). Expression patterns of astrocyte elevated gene-1 (AEG-1) during development of the mouse embryo. Gene Expression Patterns, 10(7–8), 361–367.
Venugopal, A., et al. (2022). EMT molecular signatures of pancreatic neuroendocrine neoplasms. International Journal of Molecular Sciences, 23(21), 13645.
Zhou, B., et al. (2021). High vimentin expression with E-cadherin expression loss predicts a poor prognosis after resection of grade 1 and 2 pancreatic neuroendocrine tumors. BMC Cancer, 21(1), 334.
Ikezono, Y., et al. (2017). Pancreatic neuroendocrine tumors and EMT behavior are driven by the CSC marker DCLK1. Molecular Cancer Research, 15(6), 744–752.
Adamska, A., & Falasca, M. (2018). Epithelial plasticity is crucial for pancreatic cancer metastatic organotropism. Annals of Translational Medicine, 6(Suppl 1), S53.
Aiello, N. M., et al. (2018). EMT subtype influences epithelial plasticity and mode of cell migration. Dev Cell, 45(6), 681-695 e4.
Storz, P. (2017). Acinar cell plasticity and development of pancreatic ductal adenocarcinoma. Nature Reviews. Gastroenterology & Hepatology, 14(5), 296–304.
Gidekel Friedlander, S. Y., et al. (2009). Context-dependent transformation of adult pancreatic cells by oncogenic K-Ras. Cancer Cell, 16(5), 379–389.
Morris, JPt., et al. (2010). Beta-catenin blocks Kras-dependent reprogramming of acini into pancreatic cancer precursor lesions in mice. Journal of Clinical Investigation, 120(2), 508–20.
Alonso-Curbelo, D., et al. (2021). A gene-environment-induced epigenetic program initiates tumorigenesis. Nature, 590(7847), 642–648.
Del Poggetto, E., et al. (2021). Epithelial memory of inflammation limits tissue damage while promoting pancreatic tumorigenesis. Science, 373(6561), eabj0486.
Li, Y., et al. (2021). Mutant Kras co-opts a proto-oncogenic enhancer network in inflammation-induced metaplastic progenitor cells to initiate pancreatic cancer. Nature Cancer, 2(1), 49–65.
Burdziak, C., et al. (2023). Epigenetic plasticity cooperates with cell-cell interactions to direct pancreatic tumorigenesis. Science, 380(6645), eadd5327.
Flavahan, W. A., Gaskell, E., Bernstein, B. E. (2017). Epigenetic plasticity and the hallmarks of cancer. Science, 357(6348), eaal2380.
Dawson, M. A. (2017). The cancer epigenome: Concepts, challenges, and therapeutic opportunities. Science, 355(6330), 1147–1152.
Xie, W., et al. (2013). Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell, 153(5), 1134–1148.
Gifford, C. A., et al. (2013). Transcriptional and epigenetic dynamics during specification of human embryonic stem cells. Cell, 153(5), 1149–1163.
Vander Heiden, M. G., Cantley, L. C., & Thompson, C. B. (2009). Understanding the Warburg effect: The metabolic requirements of cell proliferation. Science., 324(5930), 1029–33.
Olivares, O., et al. (2017). Collagen-derived proline promotes pancreatic ductal adenocarcinoma cell survival under nutrient limited conditions. Nature Communications, 8, 16031.
Guillaumond, F., et al. (2013). Strengthened glycolysis under hypoxia supports tumor symbiosis and hexosamine biosynthesis in pancreatic adenocarcinoma. Proceedings of the National Academy of Sciences of the United States of America, 110(10), 3919–3924.
Lyssiotis, C. A., & Kimmelman, A. C. (2017). Metabolic interactions in the tumor microenvironment. Trends in Cell Biology, 27(11), 863–875.
Sousa, C. M., & Kimmelman, A. C. (2014). The complex landscape of pancreatic cancer metabolism. Carcinogenesis, 35(7), 1441–1450.
Perera, R. M., & Bardeesy, N. (2015). Pancreatic cancer metabolism: Breaking it down to build it back up. Cancer Discovery, 5(12), 1247–1261.
Blum, R., & Kloog, Y. (2014). Metabolism addiction in pancreatic cancer. Cell Death & Disease, 5(2), e1065.
Liang, C., et al. (2016). Metabolic plasticity in heterogeneous pancreatic ductal adenocarcinoma. Biochimica et Biophysica Acta, 1866(2), 177–188.
Bryant, K. L., et al. (2014). KRAS: Feeding pancreatic cancer proliferation. Trends in Biochemical Sciences, 39(2), 91–100.
Ying, H., et al. (2012). Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell, 149(3), 656–670.
Slawson, C., Copeland, R. J., & Hart, G. W. (2010). O-GlcNAc signaling: A metabolic link between diabetes and cancer? Trends in Biochemical Sciences, 35(10), 547–555.
Stincone, A., et al. (2015). The return of metabolism: Biochemistry and physiology of the pentose phosphate pathway. Biological Reviews of the Cambridge Philosophical Society, 90(3), 927–963.
Neesse, A., et al. (2011). Stromal biology and therapy in pancreatic cancer. Gut, 60(6), 861–868.
Casazza, A., et al. (2014). Tumor stroma: A complexity dictated by the hypoxic tumor microenvironment. Oncogene, 33(14), 1743–1754.
Kalluri, R., & Zeisberg, M. (2006). Fibroblasts in cancer. Nature Reviews Cancer, 6(5), 392–401.
Xing, Y., et al. (2015). Metabolic reprogramming of the tumour microenvironment. FEBS Journal, 282(20), 3892–3898.
Yoshida, G. J. (2015). Metabolic reprogramming: The emerging concept and associated therapeutic strategies. Journal of Experimental & Clinical Cancer Research, 34, 111.
Zhao, H., et al. (2016). Tumor microenvironment derived exosomes pleiotropically modulate cancer cell metabolism. eLife, 5, e10250.
Jaster, R. (2004). Molecular regulation of pancreatic stellate cell function. Molecular Cancer, 3, 26.
Sada, M., et al. (2016). Hypoxic stellate cells of pancreatic cancer stroma regulate extracellular matrix fiber organization and cancer cell motility. Cancer Letters, 372(2), 210–218.
Lisanti, M. P., Martinez-Outschoorn, U. E., & Sotgia, F. (2013). Oncogenes induce the cancer-associated fibroblast phenotype: Metabolic symbiosis and “fibroblast addiction” are new therapeutic targets for drug discovery. Cell Cycle, 12(17), 2723–2732.
Aponte, P. M., & Caicedo, A. (2017). Stemness in cancer: Stem cells, cancer stem cells, and their microenvironment. Stem Cells International, 2017, 5619472.
Li, C., et al. (2007). Identification of pancreatic cancer stem cells. Cancer Research, 67(3), 1030–1037.
Patil, K., et al. (2021). The plasticity of pancreatic cancer stem cells: Implications in therapeutic resistance. Cancer and Metastasis Reviews, 40(3), 691–720.
Di Carlo, C., Brandi, J., & Cecconi, D. (2018). Pancreatic cancer stem cells: Perspectives on potential therapeutic approaches of pancreatic ductal adenocarcinoma. World Journal of Stem Cells, 10(11), 172–182.
Hermann, P. C., et al. (2007). Distinct populations of cancer stem cells determine tumor growth and metastatic activity in human pancreatic cancer. Cell Stem Cell, 1(3), 313–323.
Nair, N., et al. (2017). A cancer stem cell model as the point of origin of cancer-associated fibroblasts in tumor microenvironment. Science and Reports, 7(1), 6838.
Calle, A. S., et al. (2016). A new PDAC mouse model originated from iPSCs-converted pancreatic cancer stem cells (CSCcm). American Journal of Cancer Research, 6(12), 2799–2815.
Hassan, G., et al. (2022). Different pancreatic cancer microenvironments convert iPSCs into cancer stem cells exhibiting distinct plasticity with altered gene expression of metabolic pathways. Journal of Experimental & Clinical Cancer Research, 41(1), 29.
Gaur, P., et al. (2011). Identification of cancer stem cells in human gastrointestinal carcinoid and neuroendocrine tumors. Gastroenterology, 141(5), 1728–1737.
Krampitz, G. W., et al. (2016). Identification of tumorigenic cells and therapeutic targets in pancreatic neuroendocrine tumors. Proceedings of the National Academy of Sciences of the United States of America, 113(16), 4464–4469.
Katsuta, E., et al. (2016). CD73 as a therapeutic target for pancreatic neuroendocrine tumor stem cells. International Journal of Oncology, 48(2), 657–669.
Truong, L. H., & Pauklin, S. (2021). Pancreatic cancer microenvironment and cellular composition: Current understandings and therapeutic approaches. Cancers (Basel), 13(19), 5028.
Ramon, Y. C. S., et al. (2020). Clinical implications of intratumor heterogeneity: Challenges and opportunities. Journal of Molecular Medicine (Berlin, Germany), 98(2), 161–177.
Lecharpentier, A., et al. (2011). Detection of circulating tumour cells with a hybrid (epithelial/mesenchymal) phenotype in patients with metastatic non-small cell lung cancer. British Journal of Cancer, 105(9), 1338–1341.
Dongre, A., & Weinberg, R. A. (2019). New insights into the mechanisms of epithelial-mesenchymal transition and implications for cancer. Nature Reviews Molecular Cell Biology, 20(2), 69–84.
Fendt, S. M., Frezza, C., & Erez, A. (2020). Targeting metabolic plasticity and flexibility dynamics for cancer therapy. Cancer Discovery, 10(12), 1797–1807.
Venkatesan, S., et al. (2017). Treatment-induced mutagenesis and selective pressures sculpt cancer evolution. Cold Spring Harbor Perspectives in Medicine, 7(8), a026617.
Porter, R. L., et al. (2019). Epithelial to mesenchymal plasticity and differential response to therapies in pancreatic ductal adenocarcinoma. Proceedings of the National Academy of Sciences of the United States of America, 116(52), 26835–26845.
Bailey, P., et al. (2016). Genomic analyses identify molecular subtypes of pancreatic cancer. Nature, 531(7592), 47–52.
Collisson, E. A., et al. (2011). Subtypes of pancreatic ductal adenocarcinoma and their differing responses to therapy. Nature Medicine, 17(4), 500–503.
Moffitt, R. A., et al. (2015). Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nature Genetics, 47(10), 1168–1178.
Kloesch, B., et al. (2022). A GATA6-centred gene regulatory network involving HNFs and DeltaNp63 controls plasticity and immune escape in pancreatic cancer. Gut, 71(4), 766–777.
Thankamony, A. P., et al. (2020). Cancer stem cell plasticity - a deadly deal. Frontiers in Molecular Biosciences, 7, 79.
Castelli, V., et al. (2021). The great escape: The power of cancer stem cells to evade programmed cell death. Cancers (Basel), 13(2), 328.
Ciardiello, C., Leone, A., & Budillon, A. (2018). The crosstalk between cancer stem cells and microenvironment is critical for solid tumor progression: The significant contribution of extracellular vesicles. Stem Cells International, 2018, 6392198.
Ye, J., et al. (2014). The cancer stem cell niche: Cross talk between cancer stem cells and their microenvironment. Tumour Biology, 35(5), 3945–3951.
Safa, A. R. (2016). Resistance to cell death and its modulation in cancer stem cells. Critical Reviews in Oncogenesis, 21(3–4), 203–219.
Wang, H. F., et al. (2021). Cell fusion in cancer hallmarks: Current research status and future indications. Oncology Letters, 22(1), 530.
Dai, J., et al. (2020). Exosomes: Key players in cancer and potential therapeutic strategy. Signal Transduction and Targeted Therapy, 5(1), 145.
Roehlecke, C., & Schmidt, M. H. H. (2020). Tunneling nanotubes and tumor microtubes in cancer. Cancers (Basel), 12(4), 857.
Manjunath, Y., et al. (2020). Tumor-cell-macrophage fusion cells as liquid biomarkers and tumor enhancers in cancer. International Journal of Molecular Sciences, 21(5), 1872.
Jang, G., et al. (2022). Direct cell-to-cell transfer in stressed tumor microenvironment aggravates tumorigenic or metastatic potential in pancreatic cancer. NPJ Genomic Medicine, 7(1), 63.
Sharma, N., et al. (2020). Metabolic plasticity imparts erlotinib-resistance in pancreatic cancer by upregulating glucose-6-phosphate dehydrogenase. Cancer & Metabolism, 8, 19.
Biancur, D. E., & Kimmelman, A. C. (2018). The plasticity of pancreatic cancer metabolism in tumor progression and therapeutic resistance. Biochimica et Biophysica Acta - Reviews on Cancer, 1870(1), 67–75.
Boone, B. A., et al. (2015). Safety and biologic response of pre-operative autophagy inhibition in combination with gemcitabine in patients with pancreatic adenocarcinoma. Annals of Surgical Oncology, 22(13), 4402–4410.
Van Cutsem, E., et al. (2018). Phase I/II trial of pimasertib plus gemcitabine in patients with metastatic pancreatic cancer. International Journal of Cancer, 143(8), 2053–2064.
Hayes, T. K., et al. (2016). Long-term ERK inhibition in KRAS-mutant pancreatic cancer is associated with MYC degradation and senescence-like growth suppression. Cancer Cell, 29(1), 75–89.
Nishi, K., et al. (2016). Inhibition of fatty acid synthesis induces apoptosis of human pancreatic cancer cells. Anticancer Research, 36(9), 4655–4660.
Tadros, S., et al. (2017). De novo lipid synthesis facilitates gemcitabine resistance through endoplasmic reticulum stress in pancreatic cancer. Cancer Research, 77(20), 5503–5517.
Shukla, S. K., et al. (2017). MUC1 and HIF-1alpha signaling crosstalk induces anabolic glucose metabolism to impart gemcitabine resistance to pancreatic cancer. Cancer Cell, 32(1), 71-87 e7.
Biancur, D. E., et al. (2017). Compensatory metabolic networks in pancreatic cancers upon perturbation of glutamine metabolism. Nature Communications, 8, 15965.
Khalaf, K., et al. (2021). Aspects of the tumor microenvironment involved in immune resistance and drug resistance. Frontiers in Immunology, 12, 656364.
Poltavets, V., et al. (2018). The role of the extracellular matrix and its molecular and cellular regulators in cancer cell plasticity. Frontiers in Oncology, 8, 431.
Kalluri, R., & Weinberg, R. A. (2009). The basics of epithelial-mesenchymal transition. The Journal of Clinical Investigation, 119(6), 1420–1428.
Lu, W., & Kang, Y. (2019). Epithelial-mesenchymal plasticity in cancer progression and metastasis. Developmental Cell, 49(3), 361–374.
Ribatti, D., Tamma, R., & Annese, T. (2020). Epithelial-mesenchymal transition in cancer: A historical overview. Translational Oncology, 13(6), 100773.
Ruivo, C. F., et al. (2022). Extracellular vesicles from pancreatic cancer stem cells lead an intratumor communication network (EVNet) to fuel tumour progression. Gut, 71(10), 2043–2068.
Cebrian, M. J., et al. (2016). Paradoxical role of HMGB1 in pancreatic cancer: Tumor suppressor or tumor promoter? Anticancer Research, 36(9), 4381–4389.
Li, J., et al. (2020). Tumor cell-intrinsic USP22 suppresses antitumor immunity in pancreatic cancer. Cancer Immunology Research, 8(3), 282–291.
Qian, W., et al. (2021). The EGFR-HSF1 axis accelerates the tumorigenesis of pancreatic cancer. Journal of Experimental & Clinical Cancer Research, 40(1), 25.
Huang, C., Du, J., & Xie, K. (2014). FOXM1 and its oncogenic signaling in pancreatic cancer pathogenesis. Biochimica et Biophysica Acta, 1845(2), 104–116.
Zhao, J., et al. (2022). 5-fluorouracil suppresses stem cell-like properties by inhibiting p38 in pancreatic cancer cell line PANC-1. Folia Histochemica et Cytobiologica, 60(1), 55–65.
Kim, S., et al. (2015). The basic helix-loop-helix transcription factor E47 reprograms human pancreatic cancer cells to a quiescent acinar state with reduced tumorigenic potential. Pancreas, 44(5), 718–727.
Peng, L., et al. (2023). Urokinase-type plasminogen activator receptor (uPAR) cooperates with mutated KRAS in regulating cellular plasticity and gemcitabine response in pancreatic adenocarcinomas. Cancers (Basel), 15(5), 1587.
Wei, D., et al. (2023). A small molecule with big impact: MRTX1133 targets the KRASG12D mutation in pancreatic cancer. Clinical Cancer Research, 30, 1–8.
Matsubara, S., et al. (2020). Prevention of Akt phosphorylation is a key to targeting cancer stem-like cells by mTOR inhibition. Human Cell, 33(4), 1197–1203.
Peer, E., Tesanovic, S., & Aberger, F. (2019). Next-generation Hedgehog/GLI pathway inhibitors for cancer therapy. Cancers (Basel), 11(4), 538.
Nakashima, H., et al. (2006). Nuclear factor-kappaB contributes to Hedgehog signaling pathway activation through sonic Hedgehog induction in pancreatic cancer. Cancer Research, 66(14), 7041–7049.
Roca, M. S., et al. (2022). HDAC class I inhibitor domatinostat sensitizes pancreatic cancer to chemotherapy by targeting cancer stem cell compartment via FOXM1 modulation. Journal of Experimental & Clinical Cancer Research, 41(1), 83.
Capeloa, T., et al. (2022). Inhibition of mitochondrial redox signaling with MitoQ prevents metastasis of human pancreatic cancer in mice. Cancers (Basel), 14(19), 4918.
Viale, A., et al. (2014). Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature, 514(7524), 628–632.
Bao, B., et al. (2012). Metformin inhibits cell proliferation, migration and invasion by attenuating CSC function mediated by deregulating miRNAs in pancreatic cancer cells. Cancer Prevention Research (Philadelphia, Pa.), 5(3), 355–364.
Mohammed, A., et al. (2013). Antidiabetic drug metformin prevents progression of pancreatic cancer by targeting in part cancer stem cells and mTOR signaling. Translational Oncology, 6(6), 649–659.
Lonardo, E., et al. (2013). Metformin targets the metabolic achilles heel of human pancreatic cancer stem cells. PLoS One, 8(10), e76518.
Sancho, P., et al. (2015). MYC/PGC-1alpha balance determines the metabolic phenotype and plasticity of pancreatic cancer stem cells. Cell Metabolism, 22(4), 590–605.
Rausch, V., et al. (2010). Synergistic activity of sorafenib and sulforaphane abolishes pancreatic cancer stem cell characteristics. Cancer Research, 70(12), 5004–5013.
Suzuki, S., et al. (2015). JNK suppression of chemotherapeutic agents-induced ROS confers chemoresistance on pancreatic cancer stem cells. Oncotarget, 6(1), 458–470.
Shankar, S., et al. (2011). Resveratrol inhibits pancreatic cancer stem cell characteristics in human and KrasG12D transgenic mice by inhibiting pluripotency maintaining factors and epithelial-mesenchymal transition. PLoS One, 6(1), e16530.
Ben, Q., et al. (2020). A nicotine-induced positive feedback loop between HIF1A and YAP1 contributes to epithelial-to-mesenchymal transition in pancreatic ductal adenocarcinoma. Journal of Experimental & Clinical Cancer Research, 39(1), 181.
Zhang, Y., et al. (2015). Aspirin counteracts cancer stem cell features, desmoplasia and gemcitabine resistance in pancreatic cancer. Oncotarget, 6(12), 9999–10015.
Hong, S. P., et al. (2009). CD44-positive cells are responsible for gemcitabine resistance in pancreatic cancer cells. International Journal of Cancer, 125(10), 2323–2331.
Yingling, J. M., et al. (2018). Preclinical assessment of galunisertib (LY2157299 monohydrate), a first-in-class transforming growth factor-beta receptor type I inhibitor. Oncotarget, 9(6), 6659–6677.
Melisi, D., et al. (2018). Galunisertib plus gemcitabine vs. gemcitabine for first-line treatment of patients with unresectable pancreatic cancer. British Journal of Cancer, 119(10), 1208–1214.
Zhang, G. N., et al. (2011). Combination of salinomycin and gemcitabine eliminates pancreatic cancer cells. Cancer Letters, 313(2), 137–144.
Funding
Work in the lab of Azmi AS is supported by R01CA24060701A1 and R37CA215427.
Ethics declarations
Conflict of interest
ASA received funding from Colorado Chromatography and Blackstone Therapeutics. ASA serves as a consultant for GLG and Guidepoint.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Uddin, M.H., Zhang, D., Muqbil, I. et al. Deciphering cellular plasticity in pancreatic cancer for effective treatments. Cancer Metastasis Rev 43, 393–408 (2024). https://doi.org/10.1007/s10555-023-10164-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-023-10164-5