Abstract
The association between CYP2D6 genotype and outcome in breast cancer patients treated with adjuvant tamoxifen remains controversial. We assessed the influence of comprehensive versus limited CYP2D6 genotype in the context of tamoxifen adherence and co-medication in a large cohort of 618 patients. Genotyping of 33 CYP2D6 alleles used two archival cohorts from tamoxifen-treated women with invasive breast cancer (Dundee, n = 391; Manchester, n = 227). Estimates for recurrence-free survival (RFS) were calculated based on inferred CYP2D6 phenotypes using Kaplan–Meier and Cox proportional hazard models, adjusted for nodal status and tumour size. Patients with at least one reduced function CYP2D6 allele (60%) or no functional alleles (6%) had a non-significant trend for worse RFS: hazard ratio (HR) 1.52 (CI 0.98–2.36, P = 0.06). For post-menopausal women on tamoxifen monotherapy, the HR for recurrence in patients with reduced functional alleles was 1.96 (CI 1.05–3.66, P = 0.036). However, RFS analysis limited to four common CYP2D6 allelic variants was no longer significant (P = 0.39). The effect of CYP2D6 genotype was increased by adjusting for adherence to tamoxifen therapy, but not significantly changed when adjusted for co-administration of potent inhibitors of CYP2D6. Comprehensive genotyping of CYP2D6 and adherence to tamoxifen therapy may be useful to identify breast cancer patients most likely to benefit from adjuvant tamoxifen.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Despite the advent of aromatase inhibitors, tamoxifen remains an important drug in the endocrine treatment of patients with oestrogen receptor-positive (ER-positive) breast cancer [1]. Tamoxifen is a pro-drug metabolised to potent anti-oestrogenic metabolites 4-hydroxy-N-desmethyl tamoxifen (endoxifen) [2] and 4-hydroxy tamoxifen [3], predominantly by the enzyme CYP2D6. Genetic variants of CYP2D6 can result in a range of enzyme activities from ultra-rapid to absent function with approximately 7% of Caucasian women homozygous for non-functional CYP2D6 alleles [4].
The effectiveness of tamoxifen may be influenced by factors including CYP2D6 metabolizer genotype [5], adherence to treatment [6], co-medications which may inhibit the conversion of tamoxifen to active metabolites including endoxifen [7] and other mechanisms of molecular resistance [8].
Some studies have suggested that women with one or two non-functional variants of CYP2D6 have a worse clinical outcome compared to women with normal CYP2D6 activity, when treated with tamoxifen in the adjuvant setting [9–13]. Others have failed to confirm an association [14], or controversially, suggested women with non-functional CYP2D6 alleles receiving tamoxifen have a better outcome [15, 16]. Such conflicting evidence may reflect the relatively small study sizes, disparate patient populations and the range of CYP2D6 alleles determined [5]. A number of recent reviews have stated that routine testing of CYP2D6 genotype is still not yet established in the evaluation of women with ER-positive breast cancer [1, 17–19].
Adherence to tamoxifen may be poor and declines over time [6], influenced by side effects from the drug [20]. Furthermore, poor adherence to adjuvant tamoxifen for breast cancer has been associated with worse survival [6]. However, no previous studies of the association between CYP2D6 genotype and breast cancer outcome have examined this crucial covariate.
A number of co-medications are potent CYP2D6 inhibitors, including antidepressant drugs such as paroxetine and fluoxetine [7], which have been prescribed to treat hot flushes associated with tamoxifen use [21]. Adjustment for CYP2D6 inhibitors in some reports has highlighted the significance of combining genetic and environmental contributors to tamoxifen response [22, 23], but data are inconsistent in establishing the influence of potent CYP2D6 inhibitors on tamoxifen response in general [24–26]. Indeed, recent evidence suggests that only concomitant use of paroxetine with tamoxifen is associated with increased breast cancer-specific mortality [25].
We sought to determine the effect of the extent of CYP2D6 genotyping, adherence to tamoxifen and co-administration of CYP2D6 inhibitors on breast cancer recurrence in patients treated with adjuvant tamoxifen.
Patients and methods
Patients
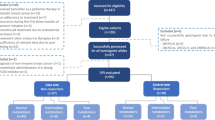
CYP2D6 genotyping was performed on samples from 618 women with ER-positive breast cancer who were prescribed 20 mg tamoxifen daily, for an intended 5 years, as adjuvant therapy. Subjects were from two geographically distinct cohorts (Fig. 1).
Cohort 1 comprised 391 Caucasian women with stage I, II or III breast cancer from Dundee, UK who had frozen primary breast tissue collected prospectively between 1997 and 2007 and updated clinical and pathological data. Matched peripheral blood lymphocyte DNA was available for 133 patients; 228 women were post-menopausal and had received adjuvant tamoxifen monotherapy. Ethical approval was obtained from the Tayside Tissue Bank Local Research Ethics Committee.
For adherence data, conducted under Caldicott Guardian approval, the Community Health Index number allowed linkage of health-related datasets providing a unique resource combining information on dispensed prescribing with detailed clinical data at the individual patient level [6]. Tamoxifen adherence was calculated as previously described as cumulative exposure, and patients with an adherence index less than 80% were deemed to have ‘low adherence’ [6].
Complete co-medication data were also recorded in this cohort and focused on the use of potent CYP2D6 inhibitors including fluoxetine, paroxetine, quinidine and bupropion (http://medicine.iupui.edu/clinpharm/COBRA/TamoxifenGuide.pdf).
Cohort 2 was composed of 227 Caucasian women with stage I, II or III breast cancer from Manchester, UK who had frozen primary breast tissue collected prospectively between 1989 and 1998; 180 of these women were post-menopausal and had received adjuvant tamoxifen monotherapy. Clinical data were collected by a comprehensive retrospective case note review to supplement prospectively collected data. The study was approved by the Trafford and Salford Research and University of Manchester Ethics Committees.
Clinical data were blinded to the laboratory investigators and data sets merged only on completion of the study.
DNA isolation and CYP2D6 genotyping
Breast tissue from Dundee was macro-dissected immediately post-operatively, and tumour was snap frozen in liquid nitrogen before storage at −80°C. Total genomic DNA was isolated using MagAttract® DNA Mini M48 Kit (Qiagen Ltd, Crawley, West Sussex, UK) on a BioRobot M48 according to the manufacturer’s protocol. Tumour specimens from Manchester patients were snap frozen in liquid nitrogen and preserved in OCT medium at −80°C. DNA was extracted by the EZ1 Qiagen robot according to manufacturer’s protocol.
Due to limited published data on the concordance of genotyping results between blood and tumour tissue [11], we undertook CYP2D6 genotyping in matched samples of tumour tissue and peripheral blood lymphocytes from 133 cases in cohort 1.
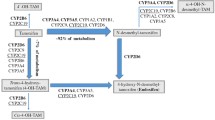
CYP2D6 genotyping was performed using the AmpliChip CYP450 Test (Roche Molecular Systems, Inc, Pleasanton, CA). The AmpliChip CYP450 Test queries 29 CYP2D6 polymorphisms to identify 33 different alleles, including a variety of gene duplications. Each allele can be assigned to one of four phenotypic categories according to its associated enzyme function: poor metabolizer (PM), intermediate metabolizer (IM), extensive metabolizer (EM) and ultra-rapid metabolizer (UM).
The predicted CYP2D6 phenotype was deduced for individual CYP2D6 alleles (http://www.cypalleles.ki.se), and CYP2D6 genotypes were classified into seven phenotypic categories using the multiple alleles, ranked from the lowest to the highest level of enzymatic function: low, PM/PM; intermediate, PM/IM, PM/EM, IM/IM and IM/EM; and high, EM/EM and EM/UM.
Statistical analysis
The association between CYP2D6 genotype and breast cancer outcomes was assessed by collapsing the seven phenotype groups described above into three functional categories: Extensive/ultra-rapid (EM/EM and EM/UM), intermediate (EM/IM, EM/PM, IM/IM and IM/PM) and poor (PM/PM) metabolism to allow comparison with recent key publications [11, 27].
Allele frequencies were calculated, and tests for Hardy–Weinberg Equilibrium (HWE) were performed (http://www.r-project.org/). Genotyping analysis with limited allele coverage was based on the presence of CYP2D6 *4, *5, *10 and *41 alleles, with all other alleles detected by the AmpliChip genotyping system reassigned to the wild-type allele [10].
Patients were classified by age, menopausal status, concomitant use of adjuvant chemotherapy, node status and tumour size. For those patients missing menopausal data, menopausal status was inferred from the patient’s age at diagnosis: <45 years at diagnosis included in the pre-menopausal category, and >55 years included in the post-menopausal category. Patients aged 45–55 years were assigned to the missing data category. Based on this process, the status of 2 and 31 patients were assigned to the pre- and post-menopausal groups, respectively.
The primary planned analysis was a comparison of the hazard rate of the PM/PM group to the EM/EM group using a multivariate Cox proportional hazards models adjusted for tumour size and nodal status with relapse-free survival (RFS) as the outcome. Relapse was defined as locoregional recurrence, ductal carcinoma in situ (DCIS), distant metastases, contralateral DCIS or death due to breast cancer. A secondary analysis to estimate the difference in hazard amongst the decreased and EM groups for all patients and for the post-menopausal, tamoxifen monotherapy subgroup was also planned and performed. A set of exploratory analyses to look at subgroups that were of small size but that might give indications of interest for indicating directions for future studies, were additionally planned, such as those performed to determine the effect of adjusting for adherence and co-medication with the data available with RFS as the outcome. Estimates of hazard ratios (HRs) for the combined patient groups were obtained using a fixed-effect meta-analysis approach, where individual HR was calculated for each centre, and then combined as a mean, weighted by the inverse of the individual study variances.
The analyses were adjusted on the clinical covariates of nodal status and tumour size. These covariates were chosen based on associations seen in previous related studies [23]. Nodal status was classified as 0, 1–3 or 4+ nodes, and tumour size as < 2 cm or ≥ 2 cm. Because the proportional hazards assumption was violated for the patients with follow-up data of >12 years and because of the small number of patients with this length of follow-up, the survival analysis was performed only on the data from the first 12 years of follow time which did not grossly violate the proportional hazards assumption based on log–log plots and goodness-of-fit tests of this assumption (data not shown). All the analyses were performed in R (http://www.r-project.org/), and included the use of the package ‘survival’.
Results
Characteristics of the two cohorts
Clinical characteristics were similar between the two cohorts including mean age of disease diagnosis (60.5 vs. 63.1 years), nodal status (node negative 52% vs. 50%) and tumour size <2 cm (33% vs. 33%). However, a greater percentage of patients in Cohort 1 had undergone chemotherapy (27% vs. 4%), more patients in Cohort 1 were pre-menopausal (27% vs. 5%) and Cohort 2 had a longer median follow up (9.4 vs. 4.9 years).
When data from the two cohorts were analysed independently, there were no significant differences in CYP2D6 allele frequency or outcome (Supplementary Table 1), and thus, the two cohorts were combined for the analyses (Table 1). The AmpliChip CYP450 genotype call rate was 95.448% (671/703 samples). From 671 genotyped samples, 53 results were excluded due to incomplete clinical data, leaving 618 patient results for statistical analyses. The observed allele frequencies at the CYP2D6 locus are listed in Supplementary Table 2. Overall 60% of patients had at least one reduced function CYP2D6 allele, and 6% had no functional CYP2D6 alleles. Sensitivity analysis indicated that combining the two sets of data did not alter the inferences made from the Cox Proportional Hazards model. There were 137 recurrence events (22.2%) amongst the 618 patients during the follow-up period.
Concordance between tumour and germline genotype
Paired samples of tumour DNA and lymphocyte DNA were available from 133 patients in cohort 1. The CYP2D6 genotype concordance was 100% (data not shown) indicating that CYP2D6 genotype determination from tumour tissue accurately reflects the patients’ germline genotype.
Recurrence-free survival analyses according to CYP2D6 metabolism groups
As part of our pre-planned statistical analysis, we compared patients who were homozygous for PM CYP2D6 alleles with patients who had at least two normally functioning CYP2D6 alleles (EM). Somewhat surprisingly, we observed no breast cancer recurrences in the 27 patients comprising the PM/PM group for whom complete covariate data were available. This lack of events in this small PM/PM group may be attributed to the higher prevalence of favourable clinical characteristics including tumours <2 cm (59% vs. 34%, P = 0.01), less node-positive disease (37% vs. 45%) and fewer pre-menopausal women (15% vs. 21%) (Supplementary Table 3).
Because of their surprising nature, it was appropriate to perform a post-hoc power analysis of these results. For this data set, there was approximately 55% power to detect a HR of 1.5 in the comparison amongst PM/PM to the EM patients. Therefore, it is possible that the PM/PM group may have a greater risk of recurrence than the EM/EM group, but this study was unable to definitively test that hypothesis. Although the study was underpowered to detect anything other than a large effect of CYP2D6 on recurrence rate when this analysis was planned, because of some large effects that had been reported in the literature and because of the relevance to the rest of the planned analyses, this comparison was performed.
Another pre-planned statistical analysis compared the outcome between patients with predicted normal CYP2D6 activity (UM/EM and EM/EM patients) and patients with any reduced function alleles. This analysis, despite the lack of events in the PM group, demonstrated a reduced RFS in patients with one or more reduced function or null alleles (HR 1.52, 95% CI 0.98–2.36, P = 0.06), with adjustment for covariates (Fig. 2, Table 2).
Sub-group analysis considering only post-menopausal women who received adjuvant tamoxifen as monotherapy demonstrated that decreased metabolizers (including all the patients with one or more reduced function or null alleles) had a significantly greater relative risk of breast cancer recurrence (HR 1.96, 95% CI 1.05–3.66, P = 0.04) (Table 2) both with and without covariate adjustment for tumour size and nodal status (Fig. 3).
Comparison of the association between comprehensive versus limited CYP2D6 genotyping and RFS
Previous assessments of the relationship between CYP2D6 genotype and response to tamoxifen treatment in patients with breast cancer have considered a limited panel of CYP2D6 variant alleles, either CYP2D6*4 alone [22, 28] or a selected panel of four more common variant alleles—CYP2D6*4,*5, *10 and *41 [10, 23]. We observed in silico that re-assigning CYP2D6 genotype to a limited panel of reduced/null function alleles (CYP2D6 *4, *5, *10 and *41) compared with those classified as having normal CYP2D6 function resulted in no significant differences (Table 2). This analysis indicates that limited CYP2D6 allele coverage may result in the misclassification of some patients with reduced CYP2D6 function as EMs in keeping with recent evidence [27].
Influence of tamoxifen adherence on RFS
Tamoxifen adherence data were available on 257 patients in cohort 1, based on linkage of their complete records of prescription encashment in the community. Thirty-seven patients had adherence less than 80%, a level previously associated with an increased risk of breast cancer recurrence [6]. Two of these 37 patients had recurrence events and were EMs. Reassigning patients in cohort 1 with adherence <80% to the decreased metabolizer group (Fig. 4) resulted in the HR of this group increasing from the multivariate HR of 2.57 (P = 0.03) to a HR of 3.02 (1.07, 8.47), P = 0.04. Within the post-menopausal subset treated with tamoxifen only, reassigning patients based on adherence showed a change from initial multivariate HR of 7.14 (P = 0.06) to a HR of 5.57 (0.74, 41.77), P = 0.09. After adjusting for adherence, patients within the EM group had an extremely low breast cancer recurrence rate.
Influence of CYP2D6 inhibitors on RFS
The CYP2D6 metabolism status of patients was adjusted for co-medication by re-classifying those patients in the EM group (UM + EM) who were prescribed either fluoxetine or paroxetine, both strong inhibitors of CYP2D6, as decreased metabolizers (no patients were taking quinidine or bupropion). In the 424 patients with either available co-medication data and/or genetically decreased metabolism status, 32 (7.6%) were known to be prescribed a strong inhibitor. Of these, 14 EM patients were reclassified to decreased metabolizers based on inhibitory co-medications, and adjustment for these co-medications did not alter the HR (data not shown).
Discussion
There is a growing interest in personalized medicine and in the use of genetic testing to provide information to select specific treatments for patients with cancer. The influence of CYP2D6 polymorphisms on outcome in tamoxifen-treated, ER-positive breast cancer patients remains controversial [9–19]. The current study was conducted in routine practice and supports the association between CYP2D6 polymorphisms and clinical outcome [9–14, 22, 23], but highlights the importance of comprehensive genotyping and particularly the adverse effect of poor adherence [6, 26, 29, 30].
The failure to observe an increased recurrence risk in breast cancer patients with a PM phenotype was surprising. The lack of events in this group may be attributed to the small number of patients and, by chance, the more favourable clinical features (Supplementary Table 3) compared to the other genotype groups or simply due to random sampling effects.
Nonetheless, we observed an increased breast cancer risk amongst the much larger group of patients with decreased metabolism based on the presence of one or more hypofunctional CYP2D6 alleles. Hypofunctional alleles in our analysis included both intermediate and null alleles. By these criteria, 60% of the treated women had one or more hypofunctional alleles compared to only 6% who had two null alleles and would be predicted to PMs. However, it is important to note that different conclusions may be drawn from the data if the groups for comparison were defined differently and further study is required to establish the most appropriate classification of CYP2D6 status in relation to tamoxifen. The detrimental effect of carrying one or more hypofunctional alleles was most apparent in the post-menopausal cohort treated with tamoxifen monotherapy, who received no adjuvant cytotoxic chemotherapy. The separation of the survival curves for extensive and reduce metabolizers (Fig. 3) occurred earlier (~2.5 years) in the post-menopausal tamoxifen monotherapy group than in the entire cohort (~4 years). This mirrors the findings of Goetz et al. [22] and further confirms that the effect of CYP2D6 metabolizer status is greater in this post-menopausal patient subset. Although prior studies established the adverse effect of a CYP2D6 PM genotype on outcome [22, 23], other studies have demonstrated that individuals with one or more hypofunctional alleles have an increased risk of breast cancer recurrence when treated with tamoxifen [10, 11]. Together these data suggest that only patients with two fully functional CYP2D6 alleles experience the full clinical benefit of tamoxifen.
Many previous studies of CYP2D6 genotyping and tamoxifen have used formalin-fixed paraffin-embedded tissue [10, 22] and genotype assays, which define a limited number of alleles. The availability here of DNA from fresh-frozen tumour tissue permitted genotyping for a wider range of variant CYP2D6 alleles. The correlation between tumour and blood DNA provides justification for the use of frozen tumour DNA as source material for CYP2D6 genotyping if blood samples are not available.
The association between CYP2D6 genotype and outcome in our cohort was statistically significant only with the extended CYP2D6 allele coverage. This suggests that the method of genotyping may be critical for accurate phenotype prediction [17] and the results of prior studies of CYP2D6 genotype may have been confounded by the inadvertent mis-assignment of some patients to a normal or EM phenotype. This is consistent with previous reports that highlight the importance of broad CYP2D6 variant allele coverage [27, 31], which is critical to identify hypofunctional alleles present in certain ethnic minorities [4].
Reliable adherence data are rarely available outside the context of clinical trials [30], but was uniquely available in this study for a proportion of patients through clinical record linkage [6]. Women who are CYP2D6 EMs are more likely to have poor adherence and to discontinue tamoxifen as they experience more severe side effects related to oestrogen deprivation [20]. Thus, those patients who are most likely to benefit from tamoxifen therapy paradoxically may also have the highest incidence of side effects. Whilst symptomatic data or tamoxifen metabolite levels were not available in this study and the reasons for reduced adherence remain uncertain, whatever the cause, poor adherence enhanced the effect of reduced CYP2D6 function, emphasizing the importance of adherence on disease-free and overall survival from breast cancer [6, 26, 29, 30].
Whilst CYP2D6 inhibitors reduce endoxifen levels [13, 21] and have been associated with worse outcome in patients treated with tamoxifen [22, 23, 25], there was no significant adverse effect of potent CYP2D6 inhibitors in this study. This may reflect the infrequent use of such medications in the two cohorts, as only a small number of patients were identified as having taken potent inhibitors. Recent evidence suggests that amongst antidepressants, also used for reducing side effects from tamoxifen, only paroxetine has a significant effect on breast cancer survival [25]. Further studies considering the effects of CYP2D6 inhibitors are required to establish the relevance of this potential drug interaction on clinical outcome.
Conclusions
The decision whether to adopt CYP2D6 testing into routine clinical practice will require analysis of large randomised controlled trial datasets and large meta-analyses such as those being conducted by the International Tamoxifen Pharmacogenetics Consortium. However, for adjuvant therapy after ER-positive breast cancer, we recommend that comprehensive genotyping should be considered and this may utilise either constitutional or tumour DNA. Furthermore, the CYP2D6 genotype needs to be interpreted in the context of the clinical setting where adherence to prescribed tamoxifen therapy may have a significant influence on disease recurrence.
Abbreviations
- CYP450:
-
cytochrome p450
- RFS:
-
recurrence-free survival
- HR:
-
hazard ratio
- CI:
-
confidence intervals
- ER:
-
oestrogen receptor
- PM:
-
poor metabolizer
- IM:
-
intermediate metabolizer
- EM:
-
extensive metabolizer
- UM:
-
ultrametabolizer
- HWE:
-
Hardy–Weinberg Equilibrium
- DCIS:
-
ductal carcinoma in situ
- DNA:
-
deoxyribonucleic acid
References
Ingle JN (2008) Pharmacogenetics and pharmacogenomics of endocrine agents for breast cancer. Breast Cancer Res 10(Suppl 4):S17
Johnson MD, Zuo H, Lee KH, Trebley JP, Rae JM, Weatherman RV, Desta Z, Flockhart DA, Skaar TC (2004) Pharmacological characterization of 4-hydroxy-N-desmethyl tamoxifen, a novel active metabolite of tamoxifen. Breast Cancer Res Treat 85:151–159
Osborne CK (1998) Tamoxifen in the treatment of breast cancer. N Engl J Med 339:1609–1618
Sachse C, Brockmoller J, Bauer S, Roots I (1997) Cytochrome P450 2D6 variants in a Caucasian population: allele frequencies and phenotypic consequences. Am J Hum Genet 60:284–295
Dezentje VO, Guchelaar HJ, Nortier JW, van de Velde CJ, Gelderblom H (2009) Clinical implications of CYP2D6 genotyping in tamoxifen treatment for breast cancer. Clin Cancer Res 15:15–21
McCowan C, Shearer J, Donnan PT, Dewar JA, Crilly M, Thompson AM, Fahey TP (2008) Cohort study examining tamoxifen adherence and its relationship to mortality in women with breast cancer. Br J Cancer 99:1763–1768
Stearns V, Johnson MD, Rae JM, Morocho A, Novielli A, Bhargava P, Hayes DF, Desta Z, Flockhart DA (2003) Active tamoxifen metabolite plasma concentrations after coadministration of tamoxifen and the selective serotonin reuptake inhibitor paroxetine. J Natl Cancer Inst 95:1758–1764
Vendrell J, Robertson KE, Ravel P, Bray SE, Bajard A, Purdie CA, Nguyen C, Hadad SM, Bieche I, Chabaud S, Bachelot T, Thompson AM, Cohen PA (2008) A candidate signature associated with tamoxifen failure in primary breast cancer. Breast Cancer Res 10:R88
Goetz MP, Rae JM, Suman VJ, Safgren SL, Ames MM, Visscher DW, Reynolds C, Couch FJ, Lingle WL, Flockhart DA, Desta Z, Perez EA, Ingle JN (2005) Pharmacogenetics of tamoxifen biotransformation is associated with clinical outcomes of efficacy and hot flashes. J Clin Oncol 23:9312–9318
Schroth W, Antoniadou L, Fritz P, Schwab M, Muerdter T, Zanger UM, Simon W, Eichelbaum M, Brauch H (2007) Breast cancer treatment outcome with adjuvant tamoxifen relative to patient CYP2D6 and CYP2C19 genotypes. J Clin Oncol 25:5187–5193
Schroth W, Goetz MP, Hamann U, Fasching PA, Schmidt M, Winter S, Fritz P, Simon W, Suman VJ, Ames MM, Safgren SL, Kuffel MJ, Ulmer HU, Boländer J, Strick R, Beckmann MW, Koelbl H, Weinshilboum RM, Ingle JN, Eichelbaum M, Schwab M, Brauch H (2009) Association between CYP2D6 polymorphisms and outcomes among women with early stage breast cancer treated with tamoxifen. JAMA 302:1429–1436
Ramón y Cajal T, Altés A, Paré L, del Rio E, Alonso C, Barnadas A, Baiget M (2010) Impact of CYP2D6 polymorphisms in tamoxifen adjuvant breast cancer treatment. Breast Cancer Res Treat 119:33–38
Kiyotani K, Mushiroda T, Imamura CK, Hosono N, Tsunoda T, Kubo M, Tanigawara Y, Flockhart DA, Desta Z, Skaar TC, Aki F, Hirata K, Takatsuka Y, Okazaki M, Ohsumi S, Yamakawa T, Sasa M, Nakamura Y, Zembutsu H (2010) Significant effect of polymorphisms in CYP2D6 and ABCC2 on clinical outcomes of adjuvant tamoxifen therapy for breast cancer patients. J Clin Oncol 28:1287–1293
Nowell SA, Ahn J, Rae JM, Scheys JO, Trovato A, Sweeney C, MacLeod SL, Kadlubar FF, Ambrosone CB (2005) Association of genetic variation in tamoxifen-metabolizing enzymes with overall survival and recurrence of disease in breast cancer patients. Breast Cancer Res Treat 91:249–258
Wegman P, Vainikka L, Stål O, Nordenskjöld B, Skoog L, Rutqvist LE, Wingren S (2005) Genotype of metabolic enzymes and the benefit of tamoxifen in postmenopausal breast cancer patients. Breast Cancer Res 7:R284–R290
Wegman P, Elingarami S, Carstensen J, Stål O, Nordenskjold B, Wingren S (2007) Genetic variants of CYP3A5, CYP2D6, SULT1A1, UGT2B15 and tamoxifen response in postmenopausal patients with breast cancer. Breast Cancer Res 9:R7
Lash TL, Lien EA, Sørensen HT, Hamilton-Dutoit S (2009) Genotype-guided tamoxifen therapy: time to pause for reflection? Lancet Oncol 10:825–833
Higgins MJ, Rae JM, Flockhart DA, Hayes DF, Stearns V (2009) Pharmacogenetics of tamoxifen: who should undergo CYP2D6 genetic testing? J Natl Compr Cancer Netw 7:203–213
Hoskins JM, Carey LA, McLeod HL (2009) CYP2D6 and tamoxifen: DNA matters in breast cancer. Nat Rev Cancer 9:576–586
Rae JM, Sikora MJ, Henry NL, Li L, Kim S, Oesterreich S, Skaar TC, Nguyen AT, Desta Z, Storniolo AM, Flockhart DA, Hayes DF, Stearns V, COBRA investigators (2009) Cytochrome P450 2D6 activity predicts discontinuation of tamoxifen therapy in breast cancer patients. Pharmacogenomics J 9:258–264
Jin Y, Desta Z, Stearns V, Ward B, Ho H, Lee KH, Skaar T, Storniolo AM, Li L, Araba A, Blanchard R, Nguyen A, Ullmer L, Hayden J, Lemler S, Weinshilboum RM, Rae JM, Hayes DF, Flockhart DA (2005) CYP2D6 genotype, antidepressant use, and tamoxifen metabolism during adjuvant breast cancer treatment. J Natl Cancer Inst 97:30–39
Goetz MP, Knox SK, Suman VJ, Rae JM, Safgren SL, Ames MM, Visscher DW, Reynolds C, Couch FJ, Lingle WL, Weinshilboum RM, Fritcher EG, Nibbe AM, Desta Z, Nguyen A, Flockhart DA, Perez EA, Ingle JN (2007) The impact of cytochrome P450 2D6 metabolism in women receiving adjuvant tamoxifen. Breast Cancer Res Treat 101:113–121
Newman WG, Hadfield KD, Latif A, Roberts SA, Shenton A, McHague C, Lalloo F, Howell S, Evans DG (2008) Impaired tamoxifen metabolism reduces survival in familial breast cancer patients. Clin Cancer Res 14:5913–5938
Ahern TP, Pedersen L, Cronin-Fenton DP, Sørensen HT, Lash TL (2009) No increase in breast cancer recurrence with concurrent use of tamoxifen and some CYP2D6-inhibiting medications. Cancer Epidemiol Biomarkers Prev 18:2562–2564
Kelly CM, Juurlink DN, Gomes T, Duong-Hua M, Pritchard KI, Austin PC, Paszat LF (2010) Selective serotonin reuptake inhibitors and breast cancer mortality in women receiving tamoxifen: a population based cohort study. BMJ 340:c693
Dezentjé VO, van Blijderveen NJ, Gelderblom H, Putter H, van Herk-Sukel MP, Casparie MK, Egberts AC, Nortier JW, Guchelaar HJ (2010) Effect of concomitant CYP2D6 inhibitor use and tamoxifen adherence on breast cancer recurrence in early-stage breast cancer. J Clin Oncol 28:2423–2429
Schroth W, Hamann U, Fasching PA, Dauser S, Winter S, Eichelbaum M, Schwab M, Brauch HB (2010) CYP2D6 polymorphisms as predictors of outcome in breast cancer patients treated with tamoxifen: expanded polymorphism coverage improves risk stratification. Clin Cancer Res Jun 1. [Epub ahead of print]
Bijl MJ, van Schaik RH, Lammers LA, Hofman A, Vulto AG, van Gelder T, Stricker BH, Visser LE (2009) The CYP2D6*4 polymorphism affects breast cancer survival in tamoxifen users. Breast Cancer Res Treat 118:125–130
Partridge AH, Wang PS, Winer EP, Avorn J (2003) Nonadherence to adjuvant tamoxifen therapy in women with primary breast cancer. J Clin Oncol 21:602–606
Ruddy KJ, Partridge AH (2009) Adherence with adjuvant hormonal therapy for breast cancer. Ann Oncol 20:401–402
Borges S, Desta Z, Li L, Skaar TC, Ward BA, Nguyen A, Jin Y, Storniolo AM, Nikoloff DM, Wu L, Hillman G, Hayes DF, Stearns V, Flockhart DA (2006) Quantitative effect of CYP2D6 genotype and inhibitors on tamoxifen metabolism: implication for optimization of breast cancer treatment. Clin Pharmacol Ther 80:61–74
Acknowledgments
We thank H McBurney for technical assistance, A Ashfield, L-J Birse-Stuart-Bell and C McCowan for clinical data acquisition and the patients for donating their tissue for cancer research. The DNA extractions, clinical data gathering, data analysis and report writing were supported by Breast Cancer Research Scotland, Cancer Research-UK, NHS Tayside and the Tayside Tissue Bank (in Dundee), the National Institutes for Health Research Manchester Biomedical Research Centre and Cancer Research-UK (in Manchester) where RF is supported by an ESMO Clinical Translational Fellowship.
Conflict of interest
AM Thompson, CA Purdie and W Newman have received reimbursements and research funding, respectively, from Roche within the last 5 years. Marcel Fontecha, Grant Hillman, Andrea Johnson, Jeffrey Lawrence and Michele Nikoloff were employed by Roche Molecular Systems at the time this study was conducted. The other authors declare that they have no competing interests. The authors have full control of all the primary data and agree to external review by the Journal if requested.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Thompson, A.M., Johnson, A., Quinlan, P. et al. Comprehensive CYP2D6 genotype and adherence affect outcome in breast cancer patients treated with tamoxifen monotherapy. Breast Cancer Res Treat 125, 279–287 (2011). https://doi.org/10.1007/s10549-010-1139-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-010-1139-x