Abstract
Bacterial diversity and aerobic catabolic competence of dioxin-degrading bacterial strains isolated from a polluted soil in the tropics were explored. Isolation of bacteria occurred after 12 months of consecutive enrichment, with dioxin congeners serving as the only sources of carbon and energy. Seventeen strains that were isolated were subsequently screened for dioxin metabolic competence. Among these isolates, five had unique amplified ribosomal DNA restriction analysis (ARDRA) patterns out of which two exhibiting good metabolic competence were selected for further investigation. The two strains were identified as Bacillus sp. SS2 and Serratia sp. SSA1, based on their 16S rRNA gene sequences. Bacterial growth co-occurred with dioxin disappearance and near stoichiometric release of chloride for one ring of the chlorinated congeners. The overall percentage removal of dibenzofuran (DF) by strain SS2 was 93.87%; while corresponding values for 2,8-dichlorodibenzofuran (2,8-diCDF) and 2,7-dichlorodibenzo-p-dioxin (2,7-diCDD) were 86.22% and 82.30% respectively. In the case of strain SSA1, percentage removal for DF, 2,8-diCDF and 2,7-diCDD were respectively 98.9%, 80.97% and 70.80%. The presence of two dioxin dioxygenase catabolic genes (dxnA1 and dbfA1) was investigated. Only the dbfA1 gene could be amplified in SS2 strain. Results further revealed that strain SS2 presented higher expression levels for the alpha-subunit of DF dioxygenase (dbfA1) gene during growth with dioxins. The expression level for dbfA1 gene was higher when growing on DF than on the other chlorinated analogs. This study gives an insight into dioxin degradation, with the catabolic potential of strains SS2 and SSA1 (an enteric bacterium) within the sub-Sahara Africa. It further shows that dioxin catabolic potential might be more prevalent in different groups of microorganisms than previously believed. Few reports have demonstrated the degradation of chlorinated congeners of dioxins, particularly from sub-Saharan African contaminated systems.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
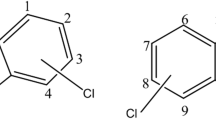
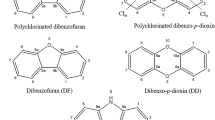
Dioxins (polychlorinated dibenzo-p-dioxins/dibenzofurans—PCDD/Fs), is a group of two-ring aromatic compounds having one to eight chlorine atoms. PCDD/Fs have 210 congeners, out of which 30 are considered significant toxins based on the chlorine atoms position and numbers on the aromatic rings (Peng et al. 2013). PCDD/Fs are produced unintentionally as unwanted by-products of various combustion and anthropogenic activities except for scientific research (Schecter et al. 2006; Klees et al. 2015). In addition, they are hydrophobic which makes them absorb onto mineral surfaces and organic matter in soils particles (Rose 2014). Such contamination has caused ecosystem imbalance in soil and reduction of microorganisms and change in abundance of functional genes which are vital to many ecological processes (Sun et al. 2018).
The application of chemical remediation strategies to decommission dioxin-polluted soils is very expensive (Kulkarni et al. 2008; Jeon et al. 2016). The use of microbial metabolic potentials as a cleanup alternative is gaining wide acceptance not only due to its cost-effectiveness, but also because of its environmental plasticity (Chen et al. 2016). Interestingly, degradation of dioxins by microorganisms has been studied, even though such reports are relatively scanty in literature. In addition, microorganisms with dioxins metabolic functionality are very scarce, especially in respect of highly chlorinated congeners due to the rarity of the gene pool in the environment (Saibu et al. 2020). Hence, very few organisms capable of dioxin degradation especially chlorinated congeners, have been isolated (Saibu et al. 2020).
Microbial degradation of dioxins is often demonstrated with growth on non-substituted dibenzofuran (DF) or dibenzo-p-dioxin (DD) as a classic substrate for dioxin compounds. Previous studies have shown that this compound is biodegraded through two catabolic routes namely, angular and lateral dioxygenation pathways (Fig. 1). The former pathway is the most common pathway encountered in the environment. This is because the initial single-step angular dioxygenation destroys the planar structure of dioxins, a factor responsible for its toxicity (Nojiri et al. 2002). Molecular oxygen attacks positions 4,4a to yield a cis-dihydrodiol that is spontaneously rearomatized to produce 2,2′,3-trihydroxybiphenyl (Jin et al. 2006; Xu et al. 2006; Saibu et al. 2020). The Meta-cleavage of this ring structure by 2,2′,3-trihydroxybiphenyl dioxygenase yields 2-hydroxy-6-oxo-6-(2′-hydroxyphenyl)-hexa-2,4-dienoate (HOHPDA) which is afterwards hydrolyzed by HOHPDA hydrolase to salicylic acid as depicted in Fig. 1. Both salicylic acid and catechol can be further oxidized to non-aromatic ring compounds via the gentisic acid or catechol ortho-pathway respectively before they are channeled into the TCA cycle for final metabolism. In lateral dioxygenation pathway, dihydroxylation occurs at the carbon–carbon bonds in 1, 2 or 3,4 positions to generate 1,2-hydroxydibenzofuran or 2,2′,3-trihydroxydiphenyl ether which is eventually meta-cleaved to 2-oxo-4-(3′-hydroxybenzofuran-2′-yl)-but-3-enoic acid (Fig. 1). This product is subsequently converted to salicylic acid or catechol (Mohammadi and Sylvestre 2005; Jin et al. 2006; Li et al. 2009; Ali et al. 2019).
Pathways for metabolism of dibenzofuran through angular and lateral dioxygenation by Janibacter terrae strain XJ-1 as proposed by Jin et al. (2006) with some modifications with respect to gene expression data and metabolites encountered in strain SS2 as presented in this study. Dibenzofuran-4,4a-dioxygenase (i), 4,4a-dihydroxy-4-hydro-dibenzofuran dehydrogenase (ii), 2,2′,3-trihydroxybiphenyl dioxygenase (iii), 2-hydroxy-6-oxo-6-(2′-hydroxyphenyl)-hexa-2,4-dienoic acid hydrolase (iv), dibenzofuran-1,2-dioxygenase (v), 1,2-dihydroxy-1,2-dihydrodibenzofuran dehydrogenase (vi), 1,2-dihydroxydibenzofuran dioxygenase (vii), 2-hydroxy-4-(3′oxo-3′H-benzofuran-2′yliden)but-2-enoic acid hydrolase (viii), salicylic acid-5-hydroxylase (ix), salicylic acid-1-hydroxylase (x) and, catechol-2,3-dioxygenase (xi). The dbfA1 gene for expression of protein mediating the production of compounds 1 and 2 was well expressed in strain SS2 while chlorinated metabolites 3 and 4 (2-chlorohydroxymuconic acid) were recovered in the culture fluids all indicating angular pathway in the organism
Studies have shown that the lateral dioxygenation of DF occurs when DF serves as a secondary carbon source (co-metabolism) during bacterial growth using biphenyl and other aromatic compounds as the main carbon sources (Becher et al. 2000; Chang 2008). Perhaps due to relative chemical structural similarity, some aromatic compound-utilizing bacteria have been documented to cometabolize DF through angular and lateral catabolic pathways (Yamazoe et al. 2004; Mohammadi and Sylvestre 2005; Li et al. 2009; Le et al. 2014). Those with capacity to metabolize dioxins as carbon and energy sources are reported to belong mainly to the phyla Proteobacteria and Actinobacteria (Hong et al. 2004; Gai et al. 2007; Jaiswal et al. 2011; Peng et al. 2013; Le et al. 2014; Lin et al. 2014). There are few documented reports on Gram positive bacteria in the phylum Firmicutes (Hong et al. 2000). However, with recent advancement in molecular tools, Bacillus is now being implicated as one of the dominant groups of bacteria during degradation studies of dioxins (Chen et al. 2016).
Among the most studied microorganisms with respect to the number of congeners transformed, elucidation of dioxin catabolic pathways as well as their use as prospective candidates for remediation of contaminated soils are: Sphingomonas wittichii strain RW1, Terrabacter sp. strains DBF63 and DPO360 and Pseudomonas sp. strain CA10 (Saibu et al. 2020). Dioxin catabolic genes and the electron transport components have been characterized in these strains. Among the degradative genes, the initial angular dioxygenase genes, dbfA and dxnA play a crucial role, because it enables activation of the aromatic skeleton and cleaving of the ether linkage destroying and detoxifying the planar structure of the molecule (Nojiri and Omori 2002). Sequence analysis of the dbfA gene from strain DBF63 showed less than 40% similarity with, the dxnA gene, of strain RW1 implying distant ancestral relationship but similar functions (Noumura et al. 2004).
Despite existing body of knowledge on microbial metabolism of dioxins, degradation of this group of chemicals, whether chlorinated or not, is yet to be demonstrated for sub-Saharan African countries and Nigeria in particular where open incineration of wastes is a routine. Here we report first time, to best of our knowledge, the isolation of bacterial species able to utilize DF as the sole carbon and energy source from a historically contaminated soil. The goal was to evaluate the extent of utilization of its chlorinated analogues and also to deduce the genes involved in the initial ring cleavage. Our study shows evidence of dioxins catabolic capabilities in very unusual bacterial strains not commonly described for degradation of xenobiotic compounds, thus suggesting that such catabolic functions may be widespread in diverse bacterial groups than previously thought.
Materials and methods
Chemicals
Dibenzofuran (98% purity), 2,2,4,4,6,8,8-heptamethylnonane (HMN), analytical grade (99–100%) of 2,8-dichlorodibenzofurans (2,8-diCDF) and 2,7-dichlorodibenzo-p-dioxins (2,7-diCDD) were obtained from Sigma-Aldrich (St. Louis, MO, USA) and Accustandard (New Haven, CT, USA).
Soil sampling analysis
Soil samples were randomly taken from a depth ranging from 0 to 20 cm in six different stations from an incinerator waste dump site, Cele, Lagos, Nigeria (6.5090° N, 3.3243° E). The samples were kept in sterile ziplock bags and transported on ice to the laboratory for further analysis. This site was selected because of prolonged and indiscriminate burning of municipal waste in the premises and because it was considered one of the major sources of dioxins in the area.
The soil physico-chemistry was evaluated in Chemistry Department, University of Lagos using standard analytical protocols (AOAC 1990). Soil organic matter was monitored using the ashing procedure (Storer 1984) while organic compounds were evaluated using US EPA method (1996).
Enrichment and isolation of dioxins-degrading bacteria
Enrichment procedure was done in 250-ml conical flasks containing 100 ml of chloride-free minimal salt (MS) medium, as previously described (Kim and Picardal 2001). The medium was supplemented with a trace elements solution containing (per liter): 50 mg MnCl2·2H20, 190 mg CoCl2·6H2O, 2 mg CuSO4·5H2O, 24 mg NiSO4·6H2O, 1.1 mg FeSO4·7H2O, 18 mg NaMoO4·2H2O, 42 mg ZnCl2·7H2O, 300 mg H3BO3 as well as a mixture of vitamins containing (per liter): folic acid 0.02 mg, thiamine-HCL 0.05 mg, riboflavin 0.05 mg, biotin 0.02 mg, nicotinic acid 0.05 mg and vitamin B12 0.001 mg. The growth medium containing the trace elements and vitamins was supplied with 150 mg l−1 DF, 100 mg l−1 each of 2,8-diCDF and 2,7-diCDD. The medium was inoculated with 1 g of soil sample, sealed with cotton plug and subsequently incubation at 28 ± 2 °C on a rotary shaker programmed at 150 rpm for 30 days. Thereafter, the enriched cultures (10% v·v−1) were transferred to a freshly prepared MS medium and incubated under the same conditions for 30 days. One percent inoculum of the enrichment culture was used for subsequent transfers. The transfer was repeated eleven times, after which dioxins-utilizers were obtained through the spray plate technique. Briefly, enriched culture was inoculated on MS agar and dioxins, dissolved in acetone was sprayed on the agar uniformly (Kim and Picardal 2000) and allowed to vent off prior incubation. Typical colonies obtained were further screened for dioxins utilization.
Amplified 16S Ribosomal DNA Restriction Analysis (ARDRA) and identification of isolates
The isolates obtained in the enrichment were subjected to ARDRA to select unique isolate clones. DNA extraction was done on 17 isolates using the Genome DNA Kit (Qiagen Inc., Germantown, MD, USA). The amplification reaction was performed using regular 16S rRNA PCR universal primers 27F and 1401R. The PCR was prepared in 50 µl reaction mixtures: 20.75 µl of PCR grade H2O (Thermo Fisher Scientific), 25 µl master mix (4390939 Applied Biosystems), 0.25 µl BSA (0.1 ng µl−1) (Thermo Fisher Scientific), 0.5 µl of 0.2 µM of each primer and 3 µl of DNA template. The PCR conditions used had an activation time for the master mix for 10 min at 95 °C, 35 cycles at 95 °C for 30 s, 55 °C for 40 s, 72 °C for 1 min and 7 min of primer extension at 72 °C.
The amplicon was purified using the QIAquick PCR purification kit (Qiagen) as described in the kit. The reaction mixture was done in 20 µl; 10 µl PCR product (after purification), 1 µl of Hha I (20 µ ml−1), 1 µl of Msp I (20 µ ml−1), 2 µl of buffer 4 (New England Biolab) 1X, 0.2 µl BSA (0.1 ng µl−1) (Thermo) and 5.8 µl nuclease free water. The mixture was incubated at 37 °C for 3 h and a denaturation step (65 °C) for 20 min. Separation of restriction fragments was carried out on a 3% agarose gel at 60 V for 3 h. All gels were run in 1X TAE stained with SYBR safe dye (Invitrogen). The gel was visualized using a UV-transilluminator with a built-in camera. The gel image analysis for clustering was done using the PyElph software to identify identical clones and selected one of each for sequencing.
The same PCR product used for the ARDRA digestion was also cleaned using Qiaquick PCR Purification Kit (Qiagen), and sequenced using Miseq as previously described (Nguyen and Rodrigues 2018). Sequenced data were compared to GenBank database using the BLAST algorithm. A phylogenetic tree was generated using Mega software, version X with diverse species of bacteria (Saitou and Nei 1987; Kumar et al. 2018). The 16S rRNA gene sequences were deposited in GenBank database and are available under MH714859—MH714865 accession numbers.
Screening of best isolates and preliminary growth studies
Selected bacterial isolates were grown in triplicates in MS medium with DF at 150 mg l−1. The growth was monitored by turbidity with a 96 wells microplate reader (Biotek). The following procedure was programmed into the reader: set temperature at 30 °C, plate shaking at medium speed, and plate read every 30 min for 24 h at wavelength 600 nm. Selected unique clones were screened for growth on DF as the only carbon source inoculated with bacterial cells in phosphate buffer solution at pH 7.2. Growth was assayed by observing OD reads from the Biotek machine. The best two isolates (most turbid) were selected for further growth kinetics, biodegradation and gene expression studies.
Growth and utilization of dioxin congeners
Selected isolates were grown in LB medium, centrifuged (10 min, 10,000×g), washed twice with sodium phosphate buffer (pH 8.0) and resuspended in MS medium to obtain an OD600 of 0.2–0.3. Dioxin utilization was conducted in conical flasks containing MS medium which was supplemented with 150 mg l−1 of DF. DF was dissolved in HMN, a non-degradable carrier. Preliminary investigation of the cells with this solvent alone showed that it did not serve as a carbon source or impacted the cell viability. Therefore, HMN was used to dissolve the dioxins to increase mass transfer in the culture medium. Other flasks were supplied with select substituted congener (2,8-diCDF and 2,7-diCDD) at 100 mg l−1. All the flasks (set up) were incubated at 28 ± 2 °C in an incubator shaker at 150 rpm. Control flasks were inoculated with heat-killed microorganisms. Cultures were sampled periodically and concentrations of residual dioxin congeners and chloride released were determined respectively by GC–MS and spectrophotometrically. Cell densities were evaluated by standard plate count technique. Samples for residual substrate concentration were centrifuged at 7000 rpm for 10 min and clarified using 0.2 µm PTFE filter and thereafter stored at − 20 °C for analysis.
GC/MS and chloride ion assay
Residual dioxins in the medium were extracted three times with 20 ml of hexane: acetone (1:1) solvent system in a 250-ml separatory funnel. The content was forcibly agitated for 5 min while venting the funnel to reduce excess vapor pressure build-up. The organic layer was decanted gently while sodium sulphate and silica gel column were used for drying and clean-up respectively. Extract was concentrated using a rotary evaporator and reconstituted in hexane, then analyzed on an Agilent (7820A) gas chromatograph fitted with inert mass spectrophotometer (triple axis detector). Analysis was done on an electron-impact ionization mode at 70 eV with ion source temperature of 230 °C, quadruple temperature of 150 °C and transfer line temperature of 300 °C. Ion acquisition was via Scan mode (m/z 50 to 500 amu at 2.0 s) and selective ion mode (SIM). One microliter of the hexane extract injected in splitless mode was carried through the 30 m HP-5 capillary column coated with 5% (phenyl)-methylsiloxane (0.32 mm in diameter and 0.25 µm film thickness). The injector temperature was maintained at 250 °C while the detector was kept at 300 °C. The carrier gas was helium, and was set at a constant flow rate of 1 ml min−1 at an initial pressure of 1.715 psi and 37.758 cm sec·−1 average velocity. The chromatograph was initially programmed at 130 °C for 2 min, then ramped to 200 °C at 10 °C min−1 for 16 min, then to 265 °C at 5 °C min−1 for 7 min and finally to 300 °C at 5 °C min−1 and held for 3 min.
Concentration of inorganic chloride eliminated into the culture medium was determined spectrophotometrically. Silver nitrate (650 µl of 65%) was added to cell suspension (5 ml), thoroughly mixed and allowed to stand for 5 min. The mixture was filtered through 0.2 µm filter. The supernatant (2.26 ml) was transferred into a cuvette and 670 µl of 0.1 N AgNO3 was added. Incubation was done in the dark for 6 min after which AgCl was measured at 546 nm. Chloride was measured using the linear calibration curves (1 mM) as a reference point.
Detection and expression of dioxin dioxygenase genes by quantitative real-time PCR (qRT-PCR)
For the detection and evaluation of dioxin dioxygenase genes dxnA1 and dbfA1 expression in the isolates, cells were pre-grown separately on DF and 2,8-diCDF. Cells grown on non-inducible medium (LB broth) without DF or 2,8-diCDF served as control. The qPCR primers dxnA1 F (5′-TCATG GCTGG GTGTT CAATA-3′) and dxnA1 R (5′-CGAAA ATCAG CCCCT TGTAG-3′) (Hartmann et al. 2012) as well as dbfA1 TSY30d F (5′-TTGAAGTGGCAGGTCCCATC-3′) and TSY30d R (5′-CGATCCAATTCCAGACCCAC-3′) designed by Iida et al. (2006) were used for the amplification. Cultures were collected at exponential growth phase and centrifuged at 10,000 rpm for 5 min at 25 °C. RNeasy Mini kit (Qiagen) was used to extract the RNA and subsequent DNase I treatment for DNA clean up prior to synthesis of cDNA. cDNA was synthesized using High-Capacity cDNA Reverse Transcription kit (Applied Biosystems) with random primers according to the protocol provided by the manufacturer (Rodrigues and Tiedje 2007). RT-PCR was conducted in a Step One Plus Real-time PCR system (Applied Biosystems). Reaction mixture consist of 2X Power SYBR Green (7.5 μl) (Applied Biosystems), 0.3 μl of 0.1 ng μl−1 BSA, 0.9 μl of 5 mM primer and 1 μl of cDNA template. Gene dbfA1 optimized qPCR conditions were as follows: 95 °C for 10 min, followed by 40 cycles each at 95 °C for 15 s, 55 °C for 1 min, 72 °C for 1 min. Optimized qPCR conditions for gene dxnA1 were 95 °C for 2 min, followed by 40 cycles each cycle of 95 °C for 10 s, 58 °C for 20 s and 68 °C for 30 s. At the end of the final cycle, a melting curve was produced. The 2−ΔΔCT protocol was used to calculate gene expression (Livak and Schmittgen 2001).
The expression of the gene without dioxin congener used for calibration and normalization of the results was conducted using the 16S rRNA reference gene. The gene expression study, the 16S rRNA reference and the target genes (dxnA1 and dbfA1) were applied using the comparative critical threshold (2−ΔΔCT) quantification method. All samples were run in triplicate.
Statistical analysis
Prism 6.0 software was used for all the statistical analyses (GraphPad Software, San Diego CA, USA).
Results
Physico-chemical properties of soil sample and contaminant persistence in the soil
Investigation of the physico-chemical properties of the polluted soil revealed a slightly basic soil characterized with very high organic matter content, organic carbon and DF—a model compound of dioxin congeners (Table 1). Moisture content was low, which could be due to the high concentration of organic pollutants, causing high soil hydrophobicity and consequent reduction in the water holding capacity of the sample. The high DF concentration was a result of the incineration activities, which is one of the key sources of dioxin pollution in the environment. Also, DF strongly sorbs on to organic matter in soils and reduces mobility leading to its persistence in the soils. Mineral nutrients (Total N and P) needed for efficient growth and functionality of microorganisms in relatively large concentration, were minimal in the soil.
16S rRNA based identification of dioxin degrading bacteria
In the present soil, we were able to isolate 17 strains. To avoid duplication of clones from the enrichment and isolation process, all 17 isolates were subjected to the ARDRA analysis. A comparison of the dendrogram (Fig. 2) showed a clustering of identical strains. Out of the 17 isolates, only five were unique. A positive identification of the five isolates was established on their 16S rRNA gene sequence analysis. They were identified as Bacillus sp. SS2, Klebsiella sp. SSA7, Prevotella sp. SSA5, Bordetella sp. SSA6 and Serratia sp. SSA1.
The phylogenetic tree (Fig. 3) shows the diversity and clustering of the 5 isolates obtained in this study with previous known dioxin degraders. For instance, Bacillus sp. SS2 clustered with known dioxin degrader, Paenibacillus sp. YK5 while Serattia sp. SSA1 clustered with Serratia marcescens PSAD-9 and Klebsiella sp. HL1 which are respectively wax- and dioxins-degrading organisms. Following further screening of these five isolates for growth in DF, the strains SS2 and SSA1 showed the best ability to grow in DF (Fig. 4) and were subsequently selected for further investigation.
Phylogenetic tree based on the 16S rRNA gene sequences of dioxin-degrading bacterial isolates (with asterisks) obtained from an incinerator waste dump site and reference sequences from GenBank. The tree was created with Mega software version X using the neighbor-joining algorithm with 500 bootstrap replications; bootstrap values less than 50% are shown at the branch nodes; the scale bars represents 20% sequence divergence. The 16S rRNA gene sequence of Janibacter sp. strain XJ-1 was used as outgroup reference The GenBank accession number of all sequences used to build the tree is given in parentheses
Growth dynamics of the of five isolates (Bacillus sp. SS2, open circle; Serrattia sp. SSA1, open square; Klebsiella sp. SSA7, open triangle; Bordetella sp. SSA6, Inverted triangle; Provetella sp. SSA5, open diamond) on DF. The organisms were grown in MS medium supplemented with 100 mg l−1 of the substrate and cultivated for 24 h. No growth was observed in non-inoculate or heat-inactivated control flasks
Degradation capability of Strains SS2 and SSA1 on dioxins congeners
Metabolic competence of the isolates was tested on each of DF, 2, 8-diCDF and 2,7-diCDD as the only source of carbon as depicted in Figs. 5, 6 and 7 and summarized in Table 2. In the control flasks inoculated with heat-inactivated cells, losses due to non-biological factors did not occur and chloride elimination was negligible.
Growth of bacterial strains in MS Medium supplemented with 150 mg l−1 of DF as a sole source of carbon and energy under aerobic batch conditions. Growth is shown as an increase in total viable counts (filled circle) and decrease in concentration of DF (filled square) monitored by GC–MS. In controls with heat-inactivated cells (open square), substrate was not utilized a minimal abiotic loss occurred. Data points represent means of at least two replicate determinations. In some cases, error bars were eliminated to improve clarity
Growth of bacterial strain in MS Medium supplemented with 100 mg l−1 of 2,8-diCDF as a sole source of carbon and energy under aerobic batch conditions. Growth is shown as an increase in total viable counts (filled circle), decrease in concentration of 2,8-diCDF (filled square) monitored by GC–MS and chloride released (filled triangle) determined spectrophotometrically. In controls with heat-inactivated cells, substrate (open square) was not utilized and minimal abiotic loss occurred. Data points represent means of at least two replicate determinations. In some cases, error bars were eliminated to improve clarity
Growth of bacterial strain in MS Medium supplemented with 100 mg l−1 of 2,7-diCDD as a sole source of carbon and energy under aerobic batch conditions. Growth is shown as an increase in total viable counts (filled circle), decrease in concentration of 2,7-diCDF (filled square) monitored by GC–MS and chloride released (filled triangle) determined spectrophotometrically. In controls with heat-inactivated cells, substrate (open square) was not utilized and minimal abiotic loss occurred. Data points represent means of at least two replicate determinations. In some cases, error bars were eliminated to improve clarity
Both bacterial strains utilized the concentration of DF supplied within the 120-h incubation period. The organisms grew exponentially without exhibiting any lag phase since they were acclimated to the pollutants prior to the biodegradation study (Fig. 5). Although cell increase appeared consistent in all time points, consumption of the substrate was highest during the first 24 h of growth, resulting in over 50% degradation especially in flasks inoculated with strain SS2. An additional 96 h was, however, necessary to degrade the remaining DF in the growth media. It is also noteworthy that the growth of strain SS2 produced significantly more biomass compared to SSA1. In spite of these observable differences between the two strains, they consumed nearly all the DF at the end of the experiment (Table 2). Statistical analysis indicated lack of significant difference in the degradation competence of both isolates (P < 0.05).
Although, in 2,7-diCDD incubations, degradation had a similar trend as 2,8-diCDF for both isolates, it appears the later congener is more amenable to degradation than the former. Growth of strain SS2 on 2,7-diCDD produced over three-orders-magnitude cell increase with corresponding decrease in substrate concentration and near stoichiometric chloride release for one ring. The amount of chloride eliminated is indicative of possible production of halogenated products as metabolites and more importantly, mineralization of over 40% of the initial substrate concentration (Table 2). For SSA1, biomass production and carbon utilization were much lower (Fig. 7, Table 2). In contrast to 2,8-diCDF, the most active substrate utilization was witnessed between 48 and 96 h, in which 24–26% degradation was obtained. This amount of time was necessary to obtain the highest chloride production by SSA1 (Fig. 6). Although the rate of degradation observed in 2,8-diCDF supplementation was more than twice obtained for 2,7-diCDD, mineralization rate was slightly higher in the later than the former. Interestingly, there was no significant difference (P < 0.05) irrespective of the congener utilized for carbon and the choice of strain for inoculation.
Detection of metabolites and dioxin catabolic genes and their gene expression
During examination of the culture fluids for possible production of metabolites, salicylic acid and 5-chlorosalicylic acid were identified as metabolites of DF and 2,8-diCDF respectively according to the GC/MS profile. 2-Chlorohydroxymuconic acid was the only metabolite found in flasks supplied with 2,7-diCDD. No other metabolites were recovered. The recovery of these metabolites readily suggests attack of the aromatic ring bearing chlorine substituent at position 2 and subsequent assimilation into the cellular architecture. This inference is further reinforced by the near-stoichiometric chloride released for one ring.
The detection and expression of putative dioxin dioxygenase genes dxnA1 and dbfA1 in strain SS2 was conducted using qRT-PCR primers for dxnA1 (dioxin dioxygenase, large subunit) gene of strain RW1 and dbfA1 (alpha subunit of dibenzofuran dioxygenase) of Paenibacillus sp. YK5 since both strains encode DF dioxygenases. Only dbfA1 gene was detected while dxnA1 gene was not amplified irrespective of the optimization conditions. The fold change in expression of dbfA gene was measured in different dioxin congeners.
The gene was well expressed differentially in the presence of DF and 2,8-diCDF. The fact that no expression was observed in control cells readily suggests the requirement of dioxins for induction of the gene. Moreover, the DF was a better inducer of the dbfA1 gene compared to the substituted analogue (Fig. 8). Fold change was calculated and dbfA1 gene was shown to be up-regulated 5 and 25 times higher in the 2,8-diCDF and DF grown cells, respectively, versus the control.
Relative fold induction of dbfA1 gene in SS2 cells grown with each of DF and 2,8-diCDF as a carbon source. The expression level of dbfA1 gene by real-time PCR was relative to the expression of 16S rRNA gene and normalized. The expression level in non-inducible medium control was near zero. Relative fold induction was calculated using the 2−ΔΔCT method. Data points represent means ± SD of at least two replicate determinations
Discussion
Worldwide interest in the use of microorganisms to degrade recalcitrant compounds in the environment has been on for several decades. However, the application of this approach is always limited by scanty desirable microorganisms or favorable environmental conditions. In this study, 17 bacterial isolates that had the capacity to grow on two chlorinated congeners of dioxin were isolated from a dioxin polluted soil during enrichment. These isolated strains whose colonies were surrounded by a large zone of clearance on dioxins coated MS agar plates (spray plate technique), indicated dioxin utilization.
These results readily suggest that the site had microorganisms for the target compounds. However, in the absence of adequate micronutrients (N and P) (Table 1), which are essential for growth, rate of dioxin degradation might be reduced despite available carbon and energy sources. It is possible that in the current soil, the biodegradation of dioxins by these microorganisms might be affected due to the lack of nutrients, hence the high concentration of dioxins found in the soil. Furthermore, high organic content can also be a result of sorption of the contaminants to soil particles making the contaminants unavailable for biodegradation in the soil. This correlation of organic content and contaminant availability has been previously implicated as the primary factor limiting microbial accessibility to contaminants (Kretzschmar et al. 1999; Carberry 2005). All these may somewhat clarify the persistence of dioxin in the soil despite the presence of dioxin degraders in the microbial community.
The successful isolation of strains SS2 and SSA1 from the contaminated soil may not be unconnected with the enrichment strategy used in this study. Consortium of selected dioxin congeners as against a non-chlorinated substrate was used to enrich and isolate the best dioxin bacterial degraders from the polluted soil. The essence of this blueprint lies with the fact that several bacteria of the microbial community would easily catalyze the non-substituted analogue while utilizing highly chlorinated congeners on the sideline. The isolation of strain SSA1, a member of the genus Serratia, is unusual since it is an enteric microorganism and is hardly implicated in degradation of persistent organic compounds. The ability of enteric bacteria to utilize DF and some selected chlorinated congeners is yet to be reported until now. However, previous work of Jaiswal and Thakur (2007) documented the degradation of DF by a strain of Serratia marcescens but not the substituted analogues. Furthermore, until now, very few reports have been given on the isolation of Bacillus species after aerobic enrichment using DF and dioxin congeners as the only sources of carbon. The ability of Bacillus sp. strain SS2 obtained from a dioxin polluted environment to degrade dioxins is also uncommon because Firmicutes is a group that is rarely implicated in the degradation of this group of chemicals. For instance, only one investigation with extracts from Geobacillus sp. UZO 3 have been shown to degrade 2,7-diCDD (Suzuki et al. 2011). Therefore, the growth of both strains on chlorinated congeners suggests that they may possess novel dioxygensaes and perhaps pathways not previously investigated. The metabolic versatility of strains SS2 and SSA1 enabling them to grow with dioxins and its chlorinated congeners confirms that the genera Bacillus and Serratia, which are phylogenetically close to previously reported DF degraders, Paenibacillus YK5 (Iida et al. 2006) and Serratia marcescens (Jaiswal and Thakur 2007), are indeed members with dioxin metabolic functionality. It is not unlikely that these strains may possess unusual and novel multifunctional dioxygenases, which is a beneficial feature for bioremediation of polluted sites containing various other persistent organic pollutants.
In a study undertaken by Jin et al. (2006), Janibacter terrae strain XJ-1 depleted almost 100 mg l−1 DF after 120 h. Similarly, Pseudomonas aeruginosa strain FA-HZ1, an organism that could barely tolerate 0.5 mM (84 mg l−1) of DF took nearly 80 h to degrade 0.1 mM (~ 17 mg l−1) of the same compound (Ali et al. 2019). In this study, however, 150 mg l−1 DF was completely utilized by both strains SS2 and SSA1 in 120 h. It is noteworthy that none of the organisms described by Jin et al. (2006) and Ali et al. (2019) could grow on chlorinated analogs of dioxins. Generally, very few bacteria have been reported to utilize chlorinated dioxins. For instance, Pseudomonas veronii PH-03 and strain RW1 have been described to degrade chlorinated dioxins (Wittich et al. 1992; Wilkes et al. 1996; Hong et al. 2004), However, in these studies chloride ion analysis was not conducted. The substantial release of chloride during growth on the chlorinated congeners by both strains did not account for all of the consumed chlorinated congeners. This suggests that only one mole of the chlorine was cleaved off the aromatic ring and the possibility that there was accumulation of unknown chlorinated products. Incomplete biomineralization of chlorinated dioxin congeners is usually noted with formation and build-up of different chlorinated intermediates (Wilkes et al. 1996). The bacterial degradation capacities on the selected congeners were in the rank of DF > 2,8-diCDF > 2,7-diCDD. This indicates that the degradation activity of the tested dioxin congeners did not depend on the positional location of the substituted chlorine atoms (steric hindrance effect of chlorine substituents at the ortho position on the electron density of the aromatic ring) since the degradation rates of both 2,7-diCDD and 2,8-diCDF for both strains did not differ significantly. This may be due to the substrate mixture used for the enrichment set up, which most likely selected organisms with capacity for both DF and chlorinated congeners. The isolation of these two bacterial strains from the same dioxin contaminated soil further indicates that extensive degradation of the compounds might have been evolving in response to dioxin exposure over a prolonged period (continuous incineration). This study has brought to focus the persistence of robust organisms with unique metabolic properties not only for chlorinated furans but also chlorinated dioxins wherein positional locations of chlorine on the aromatic nuclei does not appear affect the degradation.
Microorganisms are known to utilize several pathways to degrade a compound. In dioxin degradation, the significant step of the ring cleavage, is catalyzed by dioxygenases (such as dbfA1 and dxnA1) at the angular position. The fact that SS2 presented the gene dbfA1 suggests that it might present a degradation pathway similar to other organisms, such as Sphingomonas wittich RW1, Terrabacter sp. DBF63 and Rhodococcus sp. HA01 (Armengaud et al. 1999; Habe et al. 2001; Nojiri et al. 2002; Aly et al. 2008), which also have the same dioxygenase gene. This initial single step reaction (cleavage) destroys the planar structure of DF, which is a factor responsible for toxicity. In this study, elucidation of the metabolic pathway was not investigated. However, the dbfA1 dioxygenase appears to be operational in SS2 and suggest potential similar catabolic route as proposed in Fig. 1. In addition, the recovery of 5-chlorosalicylic acid and the non-halogenated analogue from the culture medium further reinforces that the initial hydroxylation of the dioxin aromatic ring occurs at the angular position.
It is generally known that during utilization of a compound through a classified pathway, induction of other pathways may occur fortuitously as reported in Agrobacterium sp. PH-08 (Le et al. 2014) and Cupriavidus sp. strain SK-3 (Adebusoye 2017). The catabolic primers failed to amplify the catabolic genes in strain SSA1 probably because the genes are plasmid borne or the bacterium possess novel set of genes or pathways different from the previous ones. A critical review of previous studies shows that it may be difficult to design a single primer to target aromatic-ring-hydroxylating dioxygenase (ARHDO) genes in view of their functional and structural diversity. It is noteworthy that Kahl and Hofer (2003) used three primer sets and not all were successful in amplification of the ARHDO genes from 11 bacterial strains spanning 4 genera which were known to aerobically grow on predominantly bicyclic aromatic compounds. However, Jaiswal and Thakur (2007) isolated a close strain of SSA1, Serratia marscescens which followed the angular dioxygenation pathway. It is not impossible therefore, for SSA1 to follow the angular pathway judging from the recovery of 5-chlorosalicylic acid as a central metabolite. It is also a possibility that the strain can metabolize dioxins through other pathways.
Conclusion
The isolation of these bacterial strains from a Nigerian polluted soil expands our knowledge on the bacterial diversity in a peculiar geographical environment. Also, this is a pioneer report on bacteria capable of aerobic degradation of chlorinated dioxins within the tropics of the sub-Saharan African continent. These organisms grew luxuriantly and degraded dioxin congeners via angular dioxygenation as demonstrated by gene expression and detection of some metabolites in the culture fluids. The catabolic profiles and genetic attributes of these organisms show their potential importance as bioremediation candidates for clean-up of contaminated sites. Nevertheless, more study is required to fully characterize the physiology and metabolic pathways in both strains and to further demonstrate the influence of positional location of chlorine substituents on dioxin degradation.
References
Adebusoye SA (2017) Biological degradation of 4-chlorobenzoic acid by a PCB metabolizing bacterium through a pathway not involving (chloro)catechol. Biodegradation 28:37–51
Ali F, Hu H, Wang W, Zhou Z, Shah SB, Xu P, Tang H (2019) Characterization of a dibenzofuran-degrading strain of Pseudomonas aeruginosa, FA-HZ1. Environ Pollut 250:262–273
Aly HAH, Huu NB, Wray V, Junca H, Pieper DH (2008) Two angular dioxygenases contribute to the metabolic versatility of dibenzofuran-degrading Rhodococcus sp. strain HA01. Appl Environ Microbiol 74:3812–3822
Armengaud J, Timmis KN, Wittich RM (1999) A functional 4-hydroxysalicylate/hydroxyquinol degradative pathway gene cluster is linked to the initial dibenzo-p-dioxin pathway genes in Sphingomonas sp. strain RW1. J Bacteriol 181:3452–3461
AOAC (1990) Official methods of analysis. Association of Official Analytical Chemists, Washington, DC
Becher D, Specht M, Hammer E, Francke W, Schauer F (2000) Cometabolic degradation of dibenzofuran by biphenyl-cultivated Ralstonia sp. strain SBUG 290. Appl Environ Microbiol 66:4528–4531
Carberry JB (2005) Bioremediation of hydrocarbon- contaminated soils using indigenous microbes. Biores Technol 96:1049–1055
Chang YS (2008) Recent developments in microbial biotransformation and biodegradation of dioxins. J Mol Micro Biotech 15:152–171
Chen WY, Wu JH, Lin SC, Chang JE (2016) Bioremediation of polychlorinated-p-dioxins/dibenzofurans contaminated soil using simulated compost-amended landfill reactors under hypoxic conditions. J Hazard Mater 312:159–168
Gai Z, Yu B, Li L, Wang Y, Ma C, Feng J, Deng Z, Xu P (2007) Cometabolic degradation of dibenzofuran and dibenzothiophene by a newly isolated carbazole-degrading Sphingomonas sp. strain. Appl Environ Microbiol 73:2832–2838
Habe H, Chung JS, Lee JH, Kasuga K, Yoshida T, Nojiri H, Omori T (2001) Degradation of chlorinated dibenzofurans and dibenzo-p-dioxins by two types of bacteria having angular dioxygenases with different features. Appl Environ Microbiol 67:3610–3617
Hartmann EM, Badalamenti JP, Krajmalnik-Brown R, Halden RU (2012) Quantitative PCR for tracking megaplasmid-borne biodegradation potential of a model sphingomonad. Appl Environ Microbiol 78(12):4493–4496
Hong HB, Hwang SH, Chang YS (2000) Biosorption of 1,2,3,4-tetrachlorodibenzo- p -dioxin and polychlorinated dibenzofurans by Bacillus pumilus. Water Res 34:349–353
Hong HB, Nam IH, Murugesan K, Kim YM, Chang YS (2004) Biodegradation of dibenzo-p-dioxin, dibenzofuran, and chlorodibenzo-p-dioxins by Pseudomonas veronii PH-03. Biodegradation 15:303–313
Iida T, Nakamura K, Izumi A, Mukouzaka Y, Kudo T (2006) Isolation and characterization of a gene cluster for dibenzofuran degradation in a new dibenzofuran-utilizing bacterium, Paenibacillus sp. strain YK5. Arch Microbiol 184:305–315
Jaiswal PK, Kohli S, Gopal M (2011) Isolation and characterization of alkalo tolerant Pseudomonas sp. strain ISTDF1 for degradation of dibenzofuran. J Ind Microbiol Biotechnol 38:503–511
Jaiswal PK, Thakur IS (2007) Isolation and characterization of dibenzofuran-degrading Serratia marcescens from alkalophilic bacterial consortium of the chemostat. Curr Microbiol 55:447–454
Jeon JR, Murugesan K, Baldrian P, Schmidt S, Chang YS (2016) Aerobic bacterial catabolism of persistent organic pollutants - potential impact of biotic and abiotic Interaction. Curr Opin Biotechnol 38:71–78
Jin S, Zhu T, Xu X, Xu Y (2006) Biodegradation of dibenzofuran by Janibacter terrae strain XJ-1. Curr Microbiol 53:30–36
Kahl S, Hofer B (2003) A genetic system for the rapid isolation of aromatic-ring-hydroxylating dioxygenase activities. Microbiol 149:1475–1481
Kim S, Picardal F (2000) A novel bacterium that utilizes monochlorobiphenyls and 4-chlorobenzoate as growth substrates. FEMS Microbiol Lett 158:225–229
Kim S, Picardal F (2001) Microbial growth on dichlorobiphenyls chlorinated on both rings as a sole carbon and energy source. Appl Environ Microbiol 67:1953–1955
Klees M, Hiester E, Bruckmann P, Molt K, Schmidt TC (2015) Polychlorinated biphenyls, polychlorinated dibenzo-p-dioxins and dibenzofurans in street dust of North Rhine-Westphalia, Germany. Sci Total Environ 511:72–81
Kretzschmar R, Borkovec M, Grolimund D, Elimelech M (1999) Mobile subsurface colloids and their role in contaminant transport. Adv Agron 66:121–193
Kulkarni PS, Crespo JG, Afonso CAM (2008) Dioxins sources and current remediation technologies—a review. Environ Int 34:139–153
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Le TT, Murugesan K, Nam IH, Jeon JR, Chang YS (2014) Degradation of dibenzofuran via multiple dioxygenation by a newly isolated Agrobacterium sp. PH-08. J Appl Microbiol 116:542–553
Li Q, Wang X, Yin G, Gai Z, Tang H, Ma C, Deng Z, Xu P (2009) New metabolites in dibenzofuran cometabolic degradation by a biphenyl-cultivated Pseudomonas putida strain B6–2. Environ Sci Technol 43:8635–8642
Lin WC, Chien GPC, Kao CM, Newman L, Wong TY, Liu JK (2014) Biodegradation of polychlorinated dibenzo-p-dioxins by Pseudomonas mendocina Strain NSYSU. J Environ Qual 43:349–357
Livak K, Schmittgen T (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Mohammadi M, Sylvestre M (2005) Resolving the profile of metabolites generated during oxidation of dibenzofuran and chlorodibenzofurans by the biphenyl catabolic pathway enzymes. Chem Biol 12:835–846
Nguyen HN, Rodrigues DF (2018) Chronic toxicity of graphene and graphene oxide in sequencing batch bioreactors: a comparative investigation. J Hazard Mater 343:200–207
Nojiri H, Omori T (2002) Molecular bases of aerobic bacterial degradation of dioxins: involvement of angular dioxygenation. Biosci Biotechnol Biochem 66:2001–2016
Nojiri H, Kamakura M, Urata M, Tanaka T, Chung JS, Takemura T, Yoshida T, Habe H, Omori T (2002) Dioxin catabolic genes are dispersed on the Terrabacter sp. DBF63 genome. Biochem Biophys Res Comm 296:233–240
Noumura T, Habe H, Widada J, Chung JS, Yoshida T, Nojiri H, Omori T (2004) Genetic characterization of the dibenzofuran-degrading Actinobacteria carrying the dbfA1A2 gene homologues isolated from activated sludge. FEMS Microbiol Lett 239:147–155
Peng P, Haiyan Y, Ruibao J, Li L (2013) Biodegradation of dioxin by a newly isolated Rhodococcus sp. with the involvement of self-transmissible plasmids. Appl Microbiol Biotechnol 97:5585–5595
Rodrigues DF, Tiedje JM (2007) Multi-locus real-time PCR for quantitation of bacteria in the environment reveals Exiguobacterium to be prevalent in permafrost. FEMS Microbiol Ecol 59:489–499
Rose M (2014) Environmental contaminants: dioxins, furans, and dioxin-like polychlorinated biphenyls A2. In: Motarjemi Y (ed) Encyclopedia of food safety. Academic Press, Waltham Massachusetts, pp 315–322
Saibu S, Adebusoye SA, Oyetibo GO (2020) Aerobic bacterial transformation and biodegradation of dioxins: a review. Bioresour Bioprocess 7:1–21
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schecter A, Birnbaum L, Ryan JJ, Constable JD (2006) Dioxins: an overview. Environ Res 101:419–428
Storer DA (1984) A simple high sample volume ashing procedure for determining soil organic matter. Comm Soil Sci Plant Anal 15:759–772
Sun J, Pan L, Tsang DC, Zhan Y, Zhu L, Li X (2018) Organic contamination and remediation in the agricultural soils of China: a critical review. Sci Total Environ 615:724–740
Suzuki Y, Nakamura M, Otsuka Y, Suzuki N, Ohyama K, Kawakami T, Sato K, Kajita S, Hishiyama S, Fujii T, Takahashi A, Katayama Y (2011) Novel enzymatic activity of cell free extract from thermophilic Geobacillus sp. UZO 3 catalyzes reductive cleavage of diaryl ether bonds of 2,7-dichlorodibenzo-p-dioxin. Chemosphere 83:868–872
US Environmental Protection Agency (US EPA) (1996) Method 3630, Revision B. Silica gel Cleanup. SW-846 Manual. Washington, DC, USA
Wilkes H, Wittich R, Timmis KN, Fortnagel P, Francke W (1996) Degradation of chlorinated dibenzofurans and dibenzo-p-dioxins by Sphingomonas sp. strain RW1. Appl Environ Microbiol 62:367–371
Wittich RM, Wilkes H, Sinnwell V, Francke W, Fortnagel P (1992) Metabolism of dibenzo-p-dioxin by Sphingomonas sp. strain RW1. Appl Environ Microbiol 58:1005–1010
Xu P, Yu B, Li FL, Cai XF, Ma CQ (2006) Microbial degradation of sulfur, nitrogen and oxygen heterocycles. Trends Microbiol 14:398–405
Yamazoe A, Yagi O, Oyaizu H (2004) Degradation of polycyclic aromatic hydrocarbons by a newly isolated dibenzofuran-utilizing Janibacter sp. strain YY-1. Appl Microbiol Biotechnol 65:211–221
Acknowledgements
This study was partially supported by Welch foundation award number (E-2011-20190330) and Biological and Environmental Research Science Focus Area grant: U.S. Department of Energy (Grant No. DE-AC52-06NA25396) as well as University of Lagos Central Research Committee grant.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Saibu, S., Adebusoye, S.A., Oyetibo, G.O. et al. Aerobic degradation of dichlorinated dibenzo-p-dioxin and dichlorinated dibenzofuran by bacteria strains obtained from tropical contaminated soil. Biodegradation 31, 123–137 (2020). https://doi.org/10.1007/s10532-020-09898-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-020-09898-8