Abstract
A bacterium Raoultella sp. X1, based on its 16S rRNA gene sequence, was isolated. Characteristics regarding the bacterial morphology, physiology, and genetics were investigated with an electron microscopy and conventional microbiological techniques. Although the isolate grew and degraded dimethoate poorly when the chemical was used as a sole carbon and energy source, it was able to remove up to 75% of dimethoate via co-metabolism. With a response surface methodology, we optimized carbon, nitrogen and phosphorus concentrations of the media for dimethoate degradation. Raoultella sp. X1 has a potential to be a useful organism for dimethoate degradation and a model strain for studying this biological process at the molecular level.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Organophosphorus (OP) pesticides, consisting of hundreds of OP compounds, are among the most widely used insect control agents. The chemistry, mechanism, and classification of OP pesticides have been thoroughly reviewed in recent years (Chambers et al. 2001; Costa 2006; Singh and Walker 2006). While both agriculture and our daily life are indebted to OP pesticides, their adverse effects in environments are increasingly evident. Exposure to OP pesticides could cause both acute and long-term effects (Costa 2006). While the acute effects of OP exposure are surely life-threatening by direct poisoning, the long-term effects have been the focus of most research efforts. For example, OP pesticides could damage mammalian peripheral and central nervous system by inhibiting acetylcholinesterase, a process termed organophosphate-induced delayed neuropathy (OPIDN) (Costa 2006). Moreover, studies in recent years suggested that OP pesticides may be even more threatening to human as teratogens and carcinogens (Chambers et al. 2001; Costa 2006; De Silva et al. 2006; Singh and Walker 2006).
Transformation of organic pesticides to non-toxic matters is mostly done biologically, especially by microorganisms, although physicochemical approaches are also significant (Singh and Walker 2006). In general, microbial degradation of organic pesticides is realized in two types of mechanisms: catabolism and co-metabolism. The first type covers either pure microbial isolates or mixed cultures capable of utilizing specific pesticides as a sole source of carbon (Richins et al. 2000; Horne et al. 2002; Siddavattam et al. 2003; Barton et al. 2004; Singh et al. 2004). Mixed microbial cultures are independently defined because their individual components are unable to use the chemicals as an energy source (Shelton and Somich 1988; Singh et al. 2003). Co-metabolism refers to transformation of a non-growth substrate in the presence of an obligated growth substrate. In many cases, some of the key enzymes and cofactors responsible for transformation of the non-growth substrate and utilization of the obligated growth substrate are shared. Co-metabolism can be optimized by maintaining the proper ratio of growth to non-growth substrates (Jung and Park 2005; Kan and Deshusses 2006; Sui et al. 2006; Claus et al. 2007). Microbial co-metabolism of organic pollutants has drawn a great deal of attention because of its potential for high transformation rates, wide applicability to a broad range of compounds, and a requirement for an inexpensive and widely available primary growth substrate (Kim et al. 2004; Frascari et al. 2005). In recent years, extensive studies on microbial co-metabolism of chloric organic pollutants have been carried out, greatly promoting our understanding of the process (Kim et al. 2000, 2006; Sun et al. 2000). However, little is known about co-metabolism of OP pesticides although Girbal et al. reported that the highest demeton-S-methyl biotransformation rate by Corynebacterium glutamicum was achieved in the presence of fructose (Girbal et al. 2000).
Dimethoate (O, O-dimethyl S-methylcarbamoylmethyl phosphorodithioate, C5H12NO3PS2), a typical organophosphorus pesticide, is generally metabolized by reducing the P = S group in the molecule. Although many bacteria were found to be able to degrade dimethoate (Deshpande et al. 2001; Liu et al. 2001a; Jiang et al. 2007; Debmandal et al. 2008), only two enzymes responsible for dimethoate degradation have been so far identified and purified from Aspergillus niger ZHY256, and Klebsiella sp. strain F51-1-2 (Liu et al. 2001a; Jiang et al. 2007). As one of mid-toxicity OP pesticides, dimethoate has been widely used against pests on rice, potato, cabbage, and cotton with a yearly output of 10 kt in China. As an unfortunate result, a huge amount of the chemical has accumulated in soil and water bodies. In the present study, a strain capable of degrading dimethoate via co-metabolism was isolated from soil samples and characterized phylogenetically and physiologically. Using a response surface methodology (RSM), three nutritional factors influencing growth and degradation activities of the strain were examined systematically Thus, the present study establishes the bacterial degradation of dimethoate via co-metabolism and suggests its possible application in environmental remediation or wastewater treatment.
Materials and methods
Chemicals
Dimethoate was purchased from the Agriculture Ministry of the People’s Republic of China. Yeast extract and agar were obtained from Difco (Shanghai, China). All other chemicals were purchased from Sigma–Aldrich (Beijing, China).
Enrichment and isolation of microorganism
A modified defined medium (DM) used in this study was composed of NaNO3 2, KH2PO4 2, KCl 0.5, MgSO4 0.5, MnSO4 0.1, BaCl2 0.2, and CaCl2 0.05 g/l (pH 6.8) (Liu et al. 2001b). DM was supplemented with 200 mg/l dimethoate (filter-sterilized), resulting in DM-D. For solid media, agar was added to a final concentration of 1.5% (w/v).
To isolate dimethoate degradation bacteria, soil samples were collected from five different sites inside a factory manufacturing dimethoate in the south central region of China (Changde, Hunan province, China). A soil sample of 5 g was suspended in 50 ml of DM-D in a 250 ml flask and incubated at 30°C in a shaker (150 rpm) for a week. The flask was then taken out and left on a bench to allow the soil particles to precipitate. A 5 ml aliquot of the aqueous phase, which contained microorganisms, was then used to inoculate a fresh 45 ml of DM-D and incubated for 7 days at 30°C in a shaker (150 rpm). After repeated twice, cultures were diluted 10-fold and spread onto DM-DY (DM-D supplemented with 0.5 g/l yeast extract) plates for isolation of single colonies.
Dimethoate degradation and tolerance test
All experiments regarding physiological characteristics of the isolate in this study were performed in triplicate unless otherwise noted. Overnight cultures in LB (150 rpm, 30°C) from single colonies were used to inoculate 50 ml of fresh LB in a 300 ml flask and the resulting culture was kept in a shaker (150 rpm) at 30°C. The cells were collected at OD600 ≈ 1.0 by centrifugation at 5,000 rpm for 10 min and washed 2 times with sterile 0.8% NaCl to remove residual nutrients. Re-suspended in 50 ml 0.8% NaCl, cells were used to inoculate DM-D in 1:50 dilution and placed in a shaker (150 rpm) for 7 days at 30°C. During incubation, the concentrations of dimethoate in cultures were monitored by the dimethoate assay. In brief, 200 μl of filtered cell-free cultures were added to 3.8 ml H2O, mixed with 1 ml 1 M NaOH and incubated at room temperature for 10 min. Following the addition of 2.5 ml HNO2 solution (prepared freshly by mixing 1 volume of 0.01 M NaNO2 with 9 volumes of 0.5 M H2SO4) and 5 min incubation, 0.5 ml 0.5% (w/v) ammonium sulfamate was added and shaken vigorously for 1 min. Subsequently, 1 ml of a mixed p-amino benzene sulfonamide (2.72% w/v) and mercury bichloride (0.02% w/v) solution was added to the resultant and incubated for 5 min. Finally, 1 ml 0.5% (w/v) N-(1-naphthyl) ethylenediamine hydrochloride was added, stabilized for 10 min, and the OD540 value of the final solution was recorded. The dimethoate concentrations of cultures were calculated out by comparison to the standard curve made with pure dimethoate. Dimethoate sensitivity assays were performed on DM-Y (DM supplemented with 0.5 g/l yeast extract) plates supplemented with various concentrations of dimethoate.
Identification of the isolate
Genomic DNA was extracted from strain X1 which showed the strongest resistance to dimethoate among all isolates with a Promega Wizard kit according to the manufacturer’s instructions (Promega, Beijing). The 16S rRNA gene was amplified with primers 16S-8F (AGAGTTTGATCCTGGCTCAG) and 16S-1537R (AAGGAGGTGATCCAGCC), and sequenced with additional primers 16S-513F (CAGCAGCCGCGGTAA) and 16S-907R (CCGTCAATTCATTTGAGTTT). The nucleotide sequence of the 16S rRNA gene of strain X1 determined in this study has been deposited in GenBank under accession number EU585749. All 16s rRNA sequences used in constructing phylogenetic tree were obtained from the Ribosomal Database (http://rdp.cme.msu.edu/). Phylogenetic analysis was performed using the Ribosomal Database Tree Builder tool as described previously (Wang et al. 2007). The resultant solution was downloaded in PostScript format and upgraded in clarification for publication.
Characterization of the isolate
The morphology of the isolate was analyzed by using scanning electron microscope (SEM) techniques. Overnight Cells on an LB plate at 30°C were washed off with phosphate-buffered saline (pH 7.0) and harvested by filtering with 0.2 μM Nuclepore Track-Etch Membrane (Whatman, Shanghai, China). The cells on the membrane were fixed in 2.5% (w/v) glutaraldehyde in 0.1 M cacodylate buffer (pH 7.2), stained by osmium tetroxide, and analyzed using a LEO 982 field emission SEM (Blair and Anderson 1998).
Standard protocols, including those for determining Gram reaction, catalase activities and enzymatic hydrolysis of various substrates were employed to establish the physiological and biochemical properties of strain X1 (Smibert and Krieg 1994). Growth of the isolate under normal or various stress conditions, including pH, temperature, and osmolarity by NaCl was examined in LB. Experiments were conducted in triplicate. The effects of different co-substrates on dimethoate degradation of strain X1 were assessed. Sixteen carbon sources, including starch, methionine, threonine, sorbitol, mannitol, glucose, sucrose, d-galactose, trehalose, pyruvic acid, aspartic acid, glutamic acid, sodium citrate, α-ketoglutaric acid, succinic acid, and malic acid were tested individually in liquid DM-D at a concentration of 0.5% (w/v).
To determine the degradation products of dimethoate by X1, cells grown to the late log-phase in DM with 0.5% (w/v) of sodium citrate were collected by centrifugation and resuspended in DM-D to ~1.2 of OD600. After 2 h, the supernatant of the degradation assay was applied to both two thin-layer chromatography (TLC) and colorimetric analysis as reported previously (Munnecke and Hsieh 1976; Kanagawa et al. 1980). Cell-less DM-D medium was used as the control.
Central composite rotatable design, data analysis and validation
Dynamics of dimethoate degradation and optimal concentrations of carbon, nitrogen, and phosphorus in DM-D were further studied by means of response surface methodology (RSM). To this end, a central composite design (CCD) was applied to reduce the number of experiments but not the statistical significance of the results (Cochran and Cox 1992). In this analysis, three variables with five concentration levels for the bacterial dimethoate degradation were examined by varying one variable at a time. A total of 20 experimental runs were carried out, of which 6 replicates at the center of the design were used to estimate the error of the measurements. The second-order polynomial was used to predict the response. SAS software (Version 8.2, SAS Institute, Cary, NC, USA) was used for regression and graphical analyses of the data obtained. Response surfaces were drawn as a function of two independent variables, while the third was held constant and equal to its mean value. To validate the predicted values, experiments with the predicted optimum concentrations of carbon, nitrogen, and phosphorus in DM-D were carried out.
Results
Strain isolation, physiological, and morphological characterization
The soil samples used in this study were collected from sites heavily contaminated with dimethoate, ranging from 5 to 200 μg per gram of soil. During the enrichment, the maximum cell density of the culture was approximately 0.16 (OD600) and degradation of dimethoate was observed. The percentages of degraded dimethoate (PDD) in three rounds of enrichment in 7 days were virtually the same: 25.3, 27.4, and 28.1%. Thirty individual colonies were obtained from these soil samples at the final step of the enrichment. One of these colonies, namely X1, was chosen for further identification and characterization because of its significantly stronger resistance to dimethoate (8 g/l). The strain X1 is stable in terms of its dimethoate degradation ability after repeated cultivation (~70 generations) in rich media.
Strain X1 is a rod-shaped Gram-negative bacterium, 1.5 and 0.5 μm in length and diameter, respectively (Fig. 1).
Scanning electron microscope image of Raoultella sp. X1. Cells were prepared as described in Materials and methods
Positive reactions for catalase, nitrate reductase and glucose fermentation and negative for gelatinase, amylase, oxidase and hydrogen sulfide production were observed. The optimal temperature, pH, and salt concentration for growth were 30°C, pH 6–8, and 2% (absolute concentration of NaCl), respectively. The lowest temperature which allows growth was 10°C. Growth and dimethoate degradation of X1, like other 29 isolates, was poor when dimethoate was used as the sole carbon and energy source with a maximum cell density of approximately 0.12 (OD600) and a PDD of approximately 27% over 10 day’s incubation (Fig. 2). When grown in DM-DY, X1 exhibited fast growth with a doubling time of ~40 min during the exponential phase and was able to degrade up to 28% of dimethoate in 10 days (Fig. 2).
Growth of X1 and time course of dimethoate degradation by X1 in various media. The cells from a single preparation were used to inoculate three different media. Growth was recorded as OD600 readings. Growth and PDD were presented by solid and dash lines, respectively. In all cases, DM-DY (♦), DM-DSC (DM-D supplemented with sodium citrate, ▲), and DM-D (●). The following controls were included in experiments but not presented in this figure: no growth of X1 was observed in DM (no carbon source); Natural degradation of dimethoate (~2%) was observed in all these tested media without cells. The data are averages from three independent cultures. Error bars were omitted for clarity
Identification of strain X1
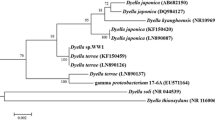
The total DNA was extracted from X1 cells grown in LB to the exponential phase for genetic analysis. The sequence of 16S rRNA gene was determined for identification. The 16S rRNA gene of X1 exhibited a 99.3% nucleotide identity to Raoultella ornithinol type strain. Based on these data, we propose that strain X1 represents a member of Raoultella genus in the enterobacterium subgroup of γ-proteobacteria, namely Raoultella sp. X1 (Fig. 3).
Phylogenetic relationship among strains in Raoultella (R) and Klebsiella (K) genus. K(R) represents that both genus names are currently used for the species. The neighbour-joining tree was prepared at Ribosomal Database (http://rdp.cme.msu.edu/) by Tree Builder with bootstraps values of 500 replicates at the branches
Selection of co-substrate
Growth and dimethoate degradation of strain X1 in DM-D supplemented with one of substrates at 5 g/l were examined and the results were summarized in Table 1. The strain was unable to utilize starch, methionine, and threonine as carbon sources for growth. While saccharides, sugar alcohols and pyruvate could be used as carbon sources to support growth of X1, they were not helpful for dimethoate degradation. Interestingly, aspartate, glutamate, sodium citrate, α-ketoglutarate, succinate and malate facilitated degradation, suggesting that the dimethoate degradation via co-metabolism by strain X1 could be exploited. The highest PDDs were observed in the presence of either sodium citrate (Fig. 2) or malate (Table 1). For the subsequent experiments, we chose sodium citrate (Na3C6H5O7) simply because it is less expensive, resulting in DM-DC medium. Based on these results and the finding of a low level of dimethoate degradation by X1 in DM-DY, we proposed that dimethoate degradation by X1 could be increased by the optimization of media.
In an attempt to determine the degradation products by X1, a degradation assay was performed with TLC and colorimetric approaches. The first step of dimethoate degradation by X1 was to break down the P–S linkage of dimethoate revealed by the P–S specific chemical in TLC. The colorimetric results further demonstrated that O, O-dimethyl phosphorothioate was present in the supernatant of the experimental assay mixture but not in that of the control assay mixture. These results suggest that the first step of the dimethoate degradation pathway of X1 appears to be the same to those in reported strains (Liu et al. 2001a; Jiang et al. 2007).
Screening of other factors influencing the dimethoate degradation
Although strain X1 required at least one of the carbon sources as co-substrate for the dimethoate degradation, other factors may either facilitate or inhibit this biological process. A series of tests were conducted to investigate effects of individual factors on the dimethoate degradation. Factors showing a significant impact on the process included temperature, pH, osmolarity by NaCl, and other nutrients. However, the influence of these environmental factors on degradation was highly correlated to the biomass (data not shown). This result suggests that the impaired degradation ability of X1 under unfavorable conditions is most likely due to poor growth. In this study, we focused our exploration on nutrients. Strain X1 was able to use dimethoate as sole nitrogen or phosphorus source evidenced by its ability to grow in DM-DSC in the absence of either NaNO3 or KH2PO4 (data not shown). However, growth rate of strain X1 was about 20% slower and the maximum cell density was reduced greatly to ~0.6 of OD600. We have showed above that dimethoate, 0.2% of NaNO3 and KH2PO4 in DM-DC as the nitrogen or phosphorus source were capable of supporting a healthy growth of strain X1. A further investigation revealed that NaNO3 and KH2PO4 in DM-DC exhibited a significant impact on dimethoate degradation only when their levels exceeded 0.2%. This observation prompted us to develop a central composite experiment for optimization of dimethoate degradation by strain X1 using DM-DSC containing 0.2% of NaNO3 and KH2PO4 as the base medium. The minimum and maximum limits of variables were given in Table 2.
Development and examination of the fitted model
A central composite experiment was designed to optimize carbon, nitrogen and phosphorus concentrations for dimethoate degradation via co-metabolism by strain X1. To this end, a total of 20 experiments were conducted to generate the experimental data (Table 2). Based on these data, a second-order polynomial equation (Eq. 1) was established to reflect the dimethoate degradation using multiple regression analysis:
where C, N and P represented the coded values of carbon, nitrogen, and phosphorus, respectively. The predicted results from this equation were also given in Table 2.
Using the standard analysis of variance (ANOVA), we found that this quadratic regression model met our expectation for optimizing our experimental parameters (Table 3). The model was highly significant (P < 0.0001) with an F-value of 134.44 and the ‘lack-of-fit’ of 1.60 which was insignificant. In addition, the model possessed a determination coefficient of 0.9918 (R 2), confirming the adequacy of fit of the model. Significance of individual coefficients of the model varied. While all linear and quadratic coefficients were highly significant (P < 0.0001), the interactive coefficients CP (P < 0.0001) and NP (P = 0.0085) but not CN (P = 0.1925) were significant.
Response surface analysis and validation
The response surface plots and their corresponding contour plots of PDD were deduced from the Eq. (1) (Fig. 4). In these contour plots, the steepest and gentlest ascents passing through the optimal conditions represented maximum and minimum decline directions of the response with respect to the variable values, respectively. A round contour suggests that the steepest and gentlest ascents are equal, indicating that the interaction between two variables is insignificant. As described above, interactive coefficient CN was insignificant, matching nearly round contours in these plots (Fig. 4b). The interaction between nitrogen and phosphorus sources appeared to be highly significant, evidenced by an elliptical contour plot (Fig. 4f). The steepest ascents were generally along with the axle of the carbon source, suggesting that PDD was more sensitive to changes in C source concentration than N source. As shown in (Fig. 4a–d), the response surface showed a peak with a carbon source concentration of ≈1.30% and a small deviation from this concentration resulted in a sharp decrease in the dimethoate degradation. As shown in (Fig. 4e), the slopes were steep compared to those in the C:N and C:P plots, suggesting that changes in concentrations of nitrogen and phosphorus sources had a significant impact on the dimethoate degradation. Moreover, it is worth noting that the concentration changing scale of carbon source (0.15%) is much larger than those of nitrogen (0.01%) and phosphorus sources (0.025%), indicating that the dimethoate degradation by X1 was highly sensitive to changes in nitrogen and phosphorus amounts.
To validate the model, we examined the growth and PDD of X1 under optimum conditions of the media predicted by the model. According to the model, the optimum concentrations of the tested variables for the highest PDD 74.79% were: Na3C6H5O7 1.30, NaNO3 0.06, and KH2PO4 0.12%. The experimental value for PDD was 74.23%, which was very close to the predicted maximum rate (Fig. 5). More importantly, this value was higher than all other PDDs that we have obtained in this study (Table 2). This result thus validated the adequacy and feasibility of our model.
Discussion
Degradation of some bio-refractory compounds is measured traditionally by whether they can be used as sole carbon and energy sources by microbes. While this is a common scenario, especially among bacteria isolated from contaminated sites with one or multiple of those compounds, many microorganisms are able to co-metabolize the non-growth substrates concerned only when a growth-supporting substrate is supplemented (Garnier et al. 2000; Liu et al. 2001a). Strain X1 investigated in this study is a good example. From a heavily contaminated site with dimethoate, we failed to isolate any bacteria capable of maintaining a healthy growth on dimethoate as only carbon source.
The isolate showing the strongest resistance to dimethoate was characterized and named as Raoultella sp. X1 based on its 16S rDNA sequence. Raoultella, a new genus defined recently, contains strains showing a potential as biocatalysts for bioremediation of aromatic hydrocarbons (ie. 2, 4, 6-trinitrotoluene (TNT), 2, 4, 5-trichlorophenoxyacetic acid, and benzoic acid) and even opportunistic pathogens found in the environment (Drancourt et al. 2001; Zharikova et al. 2006; Claus et al. 2007). Although dimethoate catabolism has been reported in various microorganisms, there is no example in Raoultella. To our knowledge, it is the first strain of Raoultella found to be able to degrade OP pesticides. Like Aspergillus niger ZHY256 and Klebsiella sp. F51-1-2, the strain X1 reduced the P–S linkage of dimethoate, implicating that this feature may be common in dimethoate degradation enzymes (Liu et al. 2001a; Jiang et al. 2007).
An obvious prerequisite for biological degradation of toxic compounds via co-metabolism is a suitable growth substrate, which serves as a source of carbon and energy. This is particularly important to degradation of highly toxic compounds which may prevent cells from growing to a high level of cell density. Substantial degradation of toxic compounds may only occur when a minimum level of cell density is established. This strategy has been applied to not only pure bacterial isolates but also microbial consortia (Tharakan and Gordon 1999). Growth substrates can be optimally chosen from a wide range of carbon sources, including nontoxic, readily degradable, and relatively cheap organic compounds (Frascari et al. 2005, 2006; Azizian et al. 2007). Dimethoate degradation by Raoultella sp. X1 was greatly promoted when cells were grown on tested TCA intermediates but not on amino acids or carbohydrates in glycolysis. These observations indicate that metabolic pathways differ in their roles of dimethoate degradation via co-metabolism, rendering the exploitation of the process far more difficult than finding a growth-supporting substrate.
In addition to support cell growth, co-metabolites may serve as induces to enzymes and/or cofactors required for degradation (Mai et al. 2001). Enzymes involved in co-metabolic reactions usually act on a series of closely related molecules and are not absolutely specific for a single substrate. It is almost a general phenomenon that competitive inhibition between growth and non-growth substrates occurs since the same key enzymes are responsible for the transformation of non-growth and co-metabolic substrates (Mai et al. 2001). In our case, sodium citrate may induce key enzymes necessary for dimethoate degradation in addition to support growth. Both growth and dimethoate transformation were strongly affected by variation in concentrations of sodium citrate in the culture. Therefore, the optimization of co-metabolism can be obtained by maintaining the proper concentrations of substrates as reported elsewhere (Loh and Wang 1997; Alvarez-Cohen and Speitel 2001; Wang et al. 2003).
On the basis of the strong correlation between growth and degradation, it is not surprising that many factors beyond substrates influencing growth may have an impact on degradation by Raoultella sp. X1. In this study, both environmental (pH, temperature, and osmolarity, etc.) and nutritional (nitrogen and phosphorus) factors were found to be critical to dimethoate degradation. Using RSM, we optimized media for dimethoate degradation by Raoultella sp. X1. The optimal concentrations of the three major components of the medium for dimethoate degradation were sodium citrate 1.30%, sodium nitrate 0.06%, monobasic potassium phosphate 0.12% and the optimum PDD was 74.79%. In this medium, all three variables were significant at its linear and quadratic levels, implicating that they may act as limiting factors and minor variations in its concentration may alter the rate of dimethoate degradation.
In conclusion, the newly isolated Raoultella sp. X1 can be exploited for dimethoate degradation via co-metabolism. The discovery of the isolate reveals that bacteria of the Raoultella genus have a direct usage of biodegradation of OP pesticides. In addition, we found that RSM was particularly useful for optimizing media for dimethoate biodegradation by the microorganism. Further investigation will be focused on deciphering dimethoate degradation pathways by means of molecular biotechnologies. The findings will improve our understandings of the process and the bacterium, which facilitates the application of the bacterium to contaminated sites and/or its degradation genes for construction of engineered strains. In China, this is particularly important and urgent given that a large volume of soils and water bodies have been contaminated with dimethoate resulting from an annual output of 10 kt.
Abbreviations
- OP:
-
Organophosphorus
- PDD:
-
Percentage of degradated dimethoate
- DM:
-
Defined medium
- DM-D:
-
DM supplemented with dimethoate
- DM-DSC:
-
DM-D supplemented with sodium citrate
- DM-Y:
-
DM supplemented with yeast extract
References
Alvarez-Cohen L, Speitel GE (2001) Kinetics of aerobic cometabolism of chlorinated solvents. Biodegradation 12:105–126. doi:10.1023/A:1012075322466
Azizian MF, Istok JD, Semprini L (2007) Evaluation of the in-situ aerobic cometabolism of chlorinated ethenes by toluene-utilizing microorganisms using push-pull tests. J Contam Hydrol 90:105–124. doi:10.1016/j.jconhyd.2006.09.015
Barton JW, Kuritz T, O’Connor LE, Ma CY, Maskarinec MP, Davison BH (2004) Reductive transformation of methyl parathion by the cyanobacterium Anabaena sp. strain PCC7120. Appl Microbiol Biotechnol 65:330–335. doi:10.1007/s00253-004-1557-y
Blair BG, Anderson KL (1998) Comparison of staining techniques for scanning electron microscopic detection of ultrastructural protuberances on cellulolytic bacteria. Biotech Histochem 73:107–113. doi:10.3109/10520299809140514
Chambers HW, Boone JS, Carr RL, Chambers JE (2001) Chemistry of organophosphorus insecticides. In: Krieger R (ed) Handbook of pesticide toxicology, 2nd edn. Academic Press, San Diego, pp 913–917
Claus H, Bausinger T, Lehmler I, Perret N, Fels G, Dehner U, Preuss J, Konig H (2007) Transformation of 2, 4, 6-trinitrotoluene (TNT) by Raoultella terrigena. Biodegradation 18:27–35. doi:10.1007/s10532-005-9033-7
Cochran WG, Cox GM (1992) Experimental designs. Wiley, New York
Costa LG (2006) Current issues in organophosphate toxicology. Clin Chim Acta 366:1–13. doi:10.1016/j.cca.2005.10.008
De Silva HJ, Samarawickrema NA, Wickremasinghe AR (2006) Toxicity due to organophosphorus compounds: what about chronic exposure? Trans R Soc Trop Med Hyg 100:803–806. doi:10.1016/j.trstmh.2006.05.001
DebMandal M, Mandal S, Pal NK, Aich A (2008) Potential metabolites of dimethoate produced by bacterial degradation. World J Microbiol Biotechnol 24:69–72. doi:10.1007/s11274-007-9440-5
Deshpande NM, Dhakephalkar PK, Kanekar PP (2001) Plasmid-mediated dimethoate degradation in Pseudomonas aeruginosa MCMB-427. Lett Appl Microbiol 33:275–279. doi:10.1046/j.1472-765X.2001.00995.x
Drancourt M, Bollet C, Carta A, Rousselier P (2001) Phylogenetic analyses of Klebsiella species delineate Klebsiella and Raoultella gen. nov., with description of Raoultella ornithinolytica comb, nov., Raoultella terrigena comb. nov and Raoultella planticola comb. nov. Int J Syst Evol Microbiol 51:925–932
Frascari D, Zannoni A, Fedi S, Pii Y, Zannoni D, Pinelli D, Nocentini M (2005) Aerobic cometabolism of chloroform by butane-grown microorganisms: long-term monitoring of depletion rates and isolation of a high-performing strain. Biodegradation 16:147–158. doi:10.1007/s10532-004-4877-9
Frascari D, Pinelli D, Nocentini M, Fedi S, Pii Y, Zannoni D (2006) Chloroform degradation by butane-grown cells of Rhodococcus aetherovorans BCP1. Appl Microbiol Biotechnol 73:421–428. doi:10.1007/s00253-006-0433-3
Garnier PM, Auria R, Augur C, Revah S (2000) Cometabolic biodegradation of methyl tert-butyl ether by a soil consortium: effect of components present in gasoline. J Gen Appl Microbiol 46:79–84. doi:10.2323/jgam.46.79
Girbal L, Rols JL, Lindley ND (2000) Growth rate influences reductive biodegradation of the organophosphorus pesticide demeton by Corynebacterium glutamicum. Biodegradation 11:371–376. doi:10.1023/A:1011682714928
Horne I, Sutherland TD, Harcourt RL, Russell RJ, Oakeshott JG (2002) Identification of an opd (organophosphate degradation) gene in an Agrobacterium isolate. Appl Environ Microbiol 68:3371–3376. doi:10.1128/AEM.68.7.3371-3376.2002
Jiang H, Yang C, Qu H, Liu Z, Fu QS, Qiao CF (2007) Cloning of a novel aldo-keto reductase gene from Klebsiella sp. strain F51-1-2 and its functional expression in Escherichia coli. Appl Environ Microbiol 73:4959–4965. doi:10.1128/AEM.02993-06
Jung IG, Park OH (2005) Enhancement of cometabolic biodegradation of trichloroethylene (TCE) gas in biofiltration. J Biosci Bioeng 100:657–661. doi:10.1263/jbb.100.657
Kanagawa T, Dazai M, Takahara Y (1980) Degradation of O,O-dimethyl phosphorothioate by activated sludge. Agric Biol Chem 44:2631–2635
Kan ES, Deshusses MA (2006) Cometabolic degradation of TCE vapors in a foamed emulsion bioreactor. Environ Sci Technol 40:1022–1028. doi:10.1021/es0510055
Kim Y, Arp DJ, Semprini L (2000) Chlorinated solvent co-metabolism by butane-grown mixed culture. J Environ Eng 126:934–942. doi:10.1061/(ASCE)0733-9372(2000)126:10(934)
Kim Y, Istok JD, Semprini L (2004) Push-pull tests for assessing in situ aerobic cometabolism. Ground Water 42:329–337. doi:10.1111/j.1745-6584.2004.tb02681.x
Kim Y, Istok JD, Semprini L (2006) Push-pull tests evaluating in situ aerobic cometabolism of ethylene, propylene, and cis-1, 2-dichloroethylene. J Contam Hydrol 82:165–181. doi:10.1016/j.jconhyd.2005.10.003
Liu CY, Speitel GE, Georgiou G (2001a) Kinetics of methyl t-butyl ether co-metabolism at low concentrations by pure cultures of butane-degrading bacteria. Appl Environ Microbiol 67:2197–2201. doi:10.1128/AEM.67.5.2197-2201.2001
Liu YH, Chung YC, Xiong Y (2001b) Purification and characterization of a dimethoate-degrading enzyme of Aspergillus niger ZHY256, isolated from sewage. Appl Environ Microbiol 67:3746–3749. doi:10.1128/AEM.67.8.3746-3749.2001
Loh KC, Wang SJ (1997) Enhancement of biodegradation of phenol and a nongrowth substrate 4-chlorophenol by medium augmentation with conventional carbon sources. Biodegradation 8:329–338. doi:10.1023/A:1008267607634
Mai P, Jacobsen OS, Aamand J (2001) Mineralization and co-metabolic degradation of phenoxyalkanoic acid herbicides by a pure bacterial culture isolated from an aquifer. Appl Microbiol Biotechnol 56:486–490. doi:10.1007/s002530000589
Munnecke DM, Hsieh DP (1976) Pathways of microbial metabolism of parathion. Appl Environ Microbiol 31:63–69
Richins RD, Mulchandani A, Chen W (2000) Expression, immobilization, and enzymatic characterization of cellulose-binding domain-organophosphorus hydrolase fusion enzymes. Biotechnol Bioeng 69:591–596. doi:10.1002/1097-0290(20000920)69:6<591::AID-BIT2>3.0.CO;2-X
Shelton DR, Somich CJ (1988) Isolation and characterization of coumaphos-metabolizing bacteria from cattle dip. Appl Environ Microbiol 54:2566–2571
Siddavattam D, Khajamohiddin S, Manavathi B, Pakala SB, Merrick M (2003) Transposon-like organization of the plasmid-borne organophosphate degradation (opd) gene cluster found in Flavobacterium sp. Appl Environ Microbiol 69:2533–2539. doi:10.1128/AEM.69.5.2533-2539.2003
Singh BK, Walker A (2006) Microbial degradation of organophosphorus compounds. FEMS Microbiol Rev 30:428–471. doi:10.1111/j.1574-6976.2006.00018.x
Singh BK, Walker A, Morgan JAW, Wright DJ (2003) Role of soil pH in the development of enhanced biodegradation of fenamiphos. Appl Environ Microbiol 69:7035–7043. doi:10.1128/AEM.69.12.7035-7043.2003
Singh BK, Walker A, Morgan JAW, Wright DJ (2004) Biodegradation of chlorpyrifos by Enterobacter strain B-14 and its use in bioremediation of contaminated soils. Appl Environ Microbiol 70:4855–4863. doi:10.1128/AEM.70.8.4855-4863.2004
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt F, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington, pp 607–654
Sui H, Li XG, Huang GQ, Jiang B (2006) A study on cometabolic bioventing for the in situ remediation of trichloroethylene. Environ Geochem Health 28:147–152. doi:10.1007/s10653-005-9025-x
Sun BL, Cole JR, Sanford RA, Tiedje JM (2000) Isolation and characterization of Desulfovibrio dechloracetivorans sp. nov, a marine dechlorinating bacterium growing by coupling the oxidation of acetate to the reductive dechlorination of 2-chlorophenol. Appl Environ Microbiol 66:2408–2413. doi:10.1128/AEM.66.6.2408-2413.2000
Tharakan JP, Gordon JA (1999) Co-metabolic biotransformation of trinitrotoluene (TNT) supported by aromatic and non-aromatic cosubstrates. Chemosphere 38:1323–1330. doi:10.1016/S0045-6535(98)00533-5
Wang SJ, Loh KC, Chua SS (2003) Prediction of critical cell growth behavior of Pseudomonas putida to maximize the co-metabolism of 4-chlorophenol with phenol and sodium glutamate as carbon sources. Enzyme Microb Technol 32:422–430. doi:10.1016/S0141-0229(02)00315-0
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. doi:10.1128/AEM.00062-07
Zharikova NV, Markusheva TV, Galkin EG, Korobov VV, Zhurenko EY, Sitdikova LR, Kolganova TV, Kuznetsov BB, Turova TP (2006) Raoultella planticola, a new strain degrading 2, 4, 5-trichlorophenoxyacetic acid. Appl Biochem Microbiol 42:258–262. doi:10.1134/S0003683806030069
Acknowledgments
This work was supported by Chinese Science Foundation for Distinguished Group (No. 50621063) and Zhejiang University research startup funding for HG.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Liang, Y., Zeng, F., Qiu, G. et al. Co-metabolic degradation of dimethoate by Raoultella sp. X1. Biodegradation 20, 363–373 (2009). https://doi.org/10.1007/s10532-008-9227-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-008-9227-x