Abstract
Invasive species are one of the most significant factors affecting biodiversity. American mink (Neovison vison) was introduced to Hokkaido, Japan, and is known to compete with other medium-sized mammals and prey on freshwater fish. Therefore, it is important to understand their distribution and the types of environments they prefer. We developed an N. vison-specific environmental DNA (eDNA) detection assay to estimate their distribution. Applying this assay to water samples from 48 rivers in the Shiretoko Peninsula, the World Natural Heritage site in Hokkaido, N. vison-specific DNA was identified in 10 rivers. Including seven rivers from a previous study on N. vision distribution in the peninsula, the environmental characteristics of the 17 rivers with the potential establishment of N. vison populations were investigated using a generalized linear model. The evaluated environmental factors included eDNA concentrations of two salmonid species (Salvelinus curilus and Oncorhynchus masou, potential food resources for N. vison), the presence of salmon hatchery and release programs, land uses around the rivers, and river structures. While the estimated N. vison distribution did not show a clear association with the eDNA concentrations of the two salmonid species, it showed positive and significant associations with the salmon release programs (p = 0.031) and with the proportion of farmland (p = 0.034). These findings imply that human activities have the potential not only to cause the introduction of invasive species but also unintentionally to contribute to the establishment of such species in new environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Freshwater ecosystems worldwide have experienced a rapid decline in biodiversity (Dudgeon et al. 2006; Butchart et al. 2010). Human activities have been proposed as the primary causes of this decline, with the disruption caused by invasive species being one of them (Stuart et al. 2004; Su et al. 2021). Invasive species can potentially disrupt native ecosystems through mechanisms such as competition with native species, predation, and hybridization (Lodge et al. 1998). However, the extent to which their establishment is connected to other human activities in and around native ecosystems remains largely unknown. Therefore, gaining an understanding of the current distribution of invasive species and the factors contributing to their establishment is crucial for the control and management of invasive species (Mehta et al. 2007).
Environmental DNA (eDNA) analysis is a biological monitoring method that facilitates simple, rapid, and non-invasive surveys of aquatic environments (Goldberg et al. 2016; Deiner et al. 2017; Minamoto 2022). Environmental DNA refers to DNA present in the environment and is found in environmental media such as water (Minamoto 2022). Analyzing eDNA enables us to understand the distribution of organisms and estimate their relative abundance (Takahara et al. 2012; Danziger et al. 2022; Greenhalgh et al. 2022). The eDNA analyses have been applied across various taxonomic groups (fish, amphibians, mammals, etc.) to investigate wide-area distributions (Ushio et al. 2017), estimate biomass (Benoit et al. 2023), and monitor invasive species (Manfrin et al. 2022; Mizumoto et al. 2022). Moreover, the ability to conduct analyses using metabarcoding assays allows for the simultaneous detection of multiple taxonomic groups from a single eDNA sample. This feature is a significant advantage of this method for studying invasive species, as it provides crucial biological information required for invasive species control, including their wide-area distribution and its associations with native species (Sepulveda et al. 2020).
American mink (Neovison vison) is an invasive species in Japan. In Hokkaido, the northernmost main island of Japan, American mink is known to compete with native medium-sized mammals such as red fox, raccoon dogs, sables, and martens, and have negative impacts on fish populations, which constitute their major food resources (Hokkaido Blue List 2010, https://www.pref.hokkaido.lg.jp/fs/5/4/9/7/6/9/2/_/blulist2010.pdf). This species was introduced to Hokkaido in 1928 for fur farming but subsequently became feral due to escapes or expulsions (Minami et al. 2016; Invasive Species of Japan, https://www.nies.go.jp/biodiversity/invasive/DB/detail/10190e.html).
In this study, we employed eDNA analysis to investigate the distribution of N. vison within the Shiretoko Peninsula and to identify the factors contributing to its establishment. Firstly, we developed a species-specific detection assay for N. vison and utilized it to assess the species distribution in the study area. Secondly, the abundance of naturally distributed Salmonidae species (Southern Asian Dolly Varden Salvelinus curilus and masu salmon Oncorhynchus masou masou), which could potentially serve as food resources for N. vison, were examined using semi-quantitative eDNA metabarcoding. While other salmonid species inhibit in the study area (O. keta and O. gorbuscha), they migrate to the ocean within a few weeks to months after hatching, whereas most of Southern Asian Dolly Varden and a part of masu salmon remain resident in Shiretoko rivers throughout their life cycles. Finally, we investigated the potential effects of both natural and artificial factors, such as hatchery and salmonid release programs, land uses around the rivers, and river structures, on the distribution of N. vison.
Materials and methods
Designing Neovison vison specific primer and probe
Species-specific primers and a probe for quantitative real-time PCR (qPCR) were designed to detect N. vison DNA. Mitochondrial DNA (mtDNA) NADH dehydrogenase subunit 2 (ND2) sequences of the target species (N. vison) and closely related species (Mustelidae and medium-sized mammals that potentially inhabiting in Hokkaido) were downloaded from the National Center for Biotechnology Information database (NCBI, https://www.ncbi.nlm.nih.gov) (Table 1). Subsequently, the sequences were aligned using BioEdit (Hall 1999). Based on criteria associated with melting temperature and target-specific bases following a previous study (Mizumoto et al. 2022), species-specific primers and a probe were designed as follows: forward primer (Nvi_NADH_F: 5´-AGGATGAGGAGGACTGA-3´), reverse primer (Nvi_NADH_R: 5´-TAAGAGGTTTAGCAGTGTAA-3´), probe (Nvi_NADH_P: 5´-FAM-CACACATAGGATGAATAATCGCCGTAACA-TAMRA-3´). The resulting amplicon had a length of 124 bp. Potential cross-reactivity of the assay was through an in silico test (i.e., Primer-BLAST was performed on all databases; https://www.ncbi.nlm.nih.gov/tools/primer-blast/index.cgi?LINK_ LOC = BlastHome). Subsequently, species specificity was further confirmed through an in vitro test. The in vitro test involved qPCR with template DNA extracted from tissue samples of the American mink (N. vison), Japanese weasel (Mustela itatsi), Least weasel (Mu. nivalis nivalis), Sable (Martes zibellina brachyura), Japanese marten (Ma. melampus), Common raccoon (Procyon lotor), and feces sample of Hokkaido raccoon dog (Nyctereutes procyonoides albus). The qPCR was conducted using the Stratagene Mx3000P system (Agilent Technologies, Inc.). Each reaction mixture (20 μL final volume) contained 200 nM primers, 150 nM TaqMan probe, 30 nM Reference dye ROX (Agilent Technology, Inc.) in 1 × Brilliant III Ultra-Fast qPCR Master Mix (Agilent Technology, Inc.), and 1 µL DNA template of each species (DNA concentration: > 1 ng/µL). The thermal-cycling profile was set at 95 °C for 3 min, followed by 50 cycles of 10 s at 95 °C and 20 s at 60 °C. To detect false positives due to contamination during the qPCR procedures, ultrapure water was used instead of DNA in one reaction mixture (non-template negative controls).
Field survey
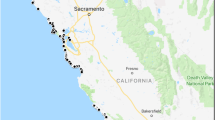
Field surveys were conducted from June 2018 to April 2020 in rivers on the Shiretoko Peninsula. A total of 183 samples were collected from 94 sites in 48 rivers using buckets and moved into new plastic bags (Fig. 1, Table S1). A total of 500 or 250 mL of water was filtered using a Sterivex™-HV filter cartridge with a pore size of 0.45 µm (Merck Millipore, Inc.) and a sterile 50 mL syringe (TERUMO, Inc.), following the procedures outlined in a previous study (Yatsuyanagi et al. 2020). After water filtrations, 2 mL of RNAlater (Thermo Fisher Scientific, Inc.) was injected into the filter cartridge for DNA preservation. Two filtered cartridge samples were collected as field replicates per site. As a negative control (NC), 500 mL of purified water was filtered and preserved, mirroring the field samples, at the end of each sampling day. In addition to these field samples, pool water in an N. vison cage was collected as a positive control at Asahiyama Zoo, Asahikawa, Hokkaido, Japan. Two replicates of 1,000 mL water samples from the pool in the cage were filtered by the same procedure as for field sampling. All field samples were placed in a cooler box with refrigerants, transported back to the laboratory, and stored in a deep freezer at − 80 °C until DNA extraction.
Environmental DNA samples processing
Environmental DNA extractions from Sterivex™-HV filter cartridge were executed using DNeasy Blood and Tissue kits (Qiagen, Hilden, Germany) in accordance with established protocols (Yatsuyanagi et al. 2020). The elution was carried out in 100 µL of AE buffer. To mitigate the risk of DNA contamination, all procedures were carried out while wearing latex gloves in a clean bench. Extracted DNA was then stored at − 20 °C until the quantification process.
Quantification of eDNA concentrations was performed using qPCR, with species-specific primers and probe designed for N. vison in this study. The qPCR conditions were the same as above, with 2 μL of the DNA template included. The qPCR also included a dilution series (200,000, 20,000, 2000, 200, and 20 copies / reaction) of standard DNA synthesized as gBlocks Gene Fragments (Integrated DNA Technologies, Inc., Iowa, USA). All qPCRs for eDNA extracts, standards, and NCs (filtration and extraction) were performed in triplicate. Each river sample underwent six replicates per site per sampling time, consisting of two cartridge samples each with three PCR replicates. The eDNA concentration (copies / L of environmental water sample) was calculated by averaging the results of the six replicates. The minimum detection threshold was set at one copy per reaction, although none of the samples in this study yielded eDNA concentration estimates falling between zero and one copy per reaction.
Evaluation of potential food resources in rivers
To estimate the abundance of S. curilus and O. masou masou, a semi-quantitative eDNA metabarcoding assay with a Salmoninae universal primer set (SalmonU4, Kanbe et al. 2023) was applied to 48 samples (one sample per each river, Table S2). In this analysis, all samples were collected in June or July of 2019, and their concurrent field NCs were analyzed as well. In cases where water samples were collected at multiple sites in the same stream during the same time of the year, downstream samples were prioritized. Amplicon library was prepared using one filtered cartridge sample per site, according to Kanbe et al. (2023).
For quantification, an internal standard with a concentration of 20 copies/reaction was added in the first round PCRs. To mitigate the issues of index hopping and sequencing errors, two types of first PCR primer pairs were employed, differing in the length of N-sequence between the adapter sequence and the specific primer (SalmonU4): one pair featured 6N in both forward and reverse primers (as per the original one in Kanbe et al. 2023), while the other pair had 0N and 12N, respectively. Half of the samples were subjected to amplification with the original primer pair, while the remaining half utilized the alternative primer pair. In the second PCR, purified first PCR products were tagged with unique dual indices within each sample group that was amplified with the same first PCR primer pair. Sequencing was performed using the 2 × 150 bp paired-end protocol on an iSeq100 platform (Illumina, Inc., San Diego CA). Sequence reads were processed following Kanbe et al. (2023) with a minor modification: in the primer sequence removal step, the target primer sequences were defined as the specific primer sequences connected with the N-sequence used in each primer pair. Consequently, although some samples shared one of the indices, misassigned reads were eliminated by the N-sequence tags.
In the Shiretoko Peninsula, there are S. curilus individuals carrying mitochondria from S. leucomaenis due to past introgressive hybridization (Yamamoto et al. 2006). Because S. leucomaenis does not currently inhabit in this peninsula (Yamamoto et al. 2006), and the SalmonU4 primer amplifies a region of mtDNA, the detection of S. leucomaenis eDNA was treated as the detection of introgressed S. curilus eDNA. In addition, although O. masou masou and O. masou ishikawae were indistinguishable in this eDNA assay, we treated the eDNA assigned to any subspecies as O. masou masou because O. masou ishikawae was absent in the study area (Yamamoto et al. 2020). The eDNA concentration of S. curilus and O. masou for each sample was calculated based on the number of read of the internal standard.
Statistical analysis
A generalized linear model (GLM) with the binomial distribution was conducted to assess the relationships between the potential distribution of N. vison and various environmental factors. These factors included the abundance of natural food resources (specifically, S. curilus and O. masou), the proportion of land use area surrounding the river, and aspects of river structure (i.e., river gradient, average elevation of the drainage area, river length, and average slope of the drainage area), as well as artificial fisheries activities such as presence of hatchery operations and salmon release programs (see Table S3). In addition to the influence of natural food resources in rivers, the juvenile salmonids artificially bred in hatcheries and released into rivers were considered as potentially valuable food resources for N. vison. In particular, local salmon hatcheries have undertaken the release of juvenile salmonids, including chum salmon (O. keta) and pink salmon (O. gorbuscha) (National Institute of Fisheries Research and Education, Fisheries Resources Laboratory, Division of Salmon, 2020). Furthermore, abiotic environmental factors, such as land use (forest, farmland) within a 250 m radius around the river, and river structure metrics including river gradient, average elevation of the drainage area (elevation), river length, and average slope of the drainage area (slope), were measured using ArcGIS Pro 3.1.1 (ESRI, Redlands, California, USA). Data on land use and river data were obtained from National Land Information Division (Ministry of Land, Infrastructure, Transport and Tourism, 2023a), while elevation data was sourced from Fundamental Geospatial Information (Ministry of Land, Infrastructure, Transport and Tourism, 2023b). The compilation of these data was performed using ArcGIS Pro 3.1.1.
The presence/absence data for N. vison incorporated results from both the eDNA assay in this study and a prior investigation (Murakami et al. 2011). In the previous study, the presence of N. vison was confirmed by direct observation, photographs, and visual inspection. In our modeling approach, the estimated presence/absence of N. vison served as a response variable, while environmental factors were designated as explanatory variables. To avoid multicollinearity among explanatory variables, Variance Inflation Factor (VIF) was employed in this study. Explanatory variables with the highest VIF were sequentially excluded until the VIF for each remaining variable fell below two. Finally, explanatory variables included stream gradient, slope, channel length, proportion of farmland (%), O. masou eDNA concentration, S. curilus eDNA concentration, and the presence of salmon release program. All statistical analyses were conducted using R software version 3.6.3. (R Core Team 2019).
Results
Specificity and detectability of the assay
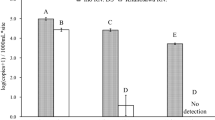
The result of in silico test demonstrated potential amplification of the N. vison DNA exclusively. The in vitro test, using tissue DNA, confirmed the amplification of N. vison-specific DNA and no amplification of six closely related species (Fig. S1). Furthermore, the eDNA in water samples collected from a water pool within N. vison cages at a zoo was also amplified as anticipated.
The current distribution of Neovison vison
As a result of the eDNA distribution survey, N. vison DNA was detected in at least one PCR replicate at 16 sites across 10 rivers (Fig. 1, Table 2). The average eDNA concentration ranged from 37.67 to 2,675.83 copies per liter per site. The highest eDNA concentration was found in the Kikiribetsu River. All negative controls (field NC, extraction NC, and PCR NC) yielded no detection of N. vison DNA.
The potential environmental factors preferred by N. vison
In the eDNA metabarcoding analysis conducted on 48 rivers, eDNA from S. curilus and O. masou were detected from 47 and 25 rivers, respectively (Table S3). Out of the 48 rivers surveyed for N. vison distribution in this study and the previous study (Murakami et al. 2011), N. vison eDNA was detected or previously confirmed in 17 rivers. The results of the GLM analysis revealed that salmon release program and the proportion of farmland around the river had positive associations with the estimated presence of N. vison (p = 0.031 and 0.034, respectively) (Fig. 2, Fig S2, Table 3). No significant association was found between the eDNA concentrations of two salmonid species (S. curilus and O. masou) and the presence of N. vison (p = 0.223 and 0.483, respectively) (Table 3).
The relationship between N. vison and environmental factors.a The relationship between N. vison and the presence or absence of salmon release program. The blue line indicates regression line. The gray shaded area represents 95% confidence interval. b The relationship between N. vison and the proportion of farmland around the river. The blue line indicates regression line. The gray shaded area represents 95% confidence interval
Discussion
Through the application of eDNA analysis, we were able to estimate the distribution of the invasive American mink within the World Natural Heritage site. The newly developed eDNA detection assay proved effective in specifically identifying eDNA from the target species, making it suitable for a variety of surveys concerning this species. Although the initial development of an eDNA detection assay incurs costs, the utilization of eDNA analysis significantly enhances wide-area surveys. This is one of the primary advantages of the eDNA analysis. Given that N. vison is solitary species that does not form colonies, the population density at each site is considered to be low (Ihara et al. 2013). Consequently, the successful eDNA detection of such terrestrial mammals from a water sample represents a noteworthy milestone in expanding the range of applications of eDNA analysis (Ishige et al. 2017; Ushio et al. 2017).
Terrestrial mammals typically visit rivers only temporarily, engaging in activities such exploration, foraging, and excretion (Williams et al. 2018). While N. vison is known to have dependency on water, they do not consistently remain in the water, limiting their opportunities for direct contact with water bodies. This behavioral characteristic of the target species increases the likelihood of false-negative detections when collecting water samples, and the eDNA concentrations may not provide an accurate indicator of their population size. In fact, none of our samples showed 100% detections among the six replicates, indicating the stochastic nature of capturing eDNA from terrestrial mammals. That was the reason why we used eDNA detection solely as a presence indicator for N. vison in our analyses. One potential solution to this issue is to increase the sampling and detecting effort by expanding the number of eDNA samples and PCR replicates per river. Given that eDNA analysis requires minimal time for sampling per site, it is worthwhile to consider augmenting the spatio-temporal collection of water samples. This approach could help reduce the risk of false negatives and provide a more reliable indicator of local biomass of the terrestrial organisms at the same time (Mizumoto et al. 2022). Another potential solution would involve increasing the water volume per sample (Schultz and Lance 2015) or collecting environmental media from terrestrial sources such as soil and stemflow (Leempoel et al. 2020; Sakata et al. 2023). These adjustments, where feasible, would be especially effective when the eDNA concentration of a target species is low. If the application of such methods allows for high-resolution and extensive monitoring, it will be possible to investigate distribution patterns at a higher resolution by using data from each of those detection sites. It is expected to provide more detailed information on N. vison.
The presence of salmon release programs exhibited a significant association with potential N. vison distribution (Table 3). While salmon juveniles are small, the quantities of fish released into rivers through hatchery propagation programs are often substantial. Hence, these releases present a suitable food resource for N. vison, particularly in the spring when other food resources remain scarce in the Shiretoko Peninsula. It is also noteworthy that mink individuals have frequently been observed around salmon hatcheries (personal communications), suggesting their interest in salmon juveniles in hatchery ponds. Since the population density of N. vison fluctuates in response to the food availability, it is reasonable to assume that a higher abundance of individuals can be found in areas with abundant food resources (Ihara et al. 2013). Furthermore, the proportion of farmland use around the river was positively associated with the potential habitat of N. vison. This species primarily relies on fish and small mammals such as mice and rats (Uraguchi et al. 1987), and small mammals are known to be abundant in farmland areas (Tattersall et al. 2002). Given that farmland, as an artificial land use, provides a suitable feeding ground for N. vison, it may attract the establishment of N. vison populations. Consequently, it is likely that human activities related to fisheries and agriculture have the potential to contribute to the reproduction and establishment of N. vison populations in the study area. Nevertheless, the precise mechanisms underlying the relationships between N. vison eDNA detections (serving as a proxy for local N. vison establishment in this study) and the associated environmental factors warrant careful investigation in future research.
The estimations of the abundance of natural food resources, based on eDNA concentrations of two salmonid species (S. curilus and O. masou), did not reveal a significant association with the estimated distribution of N. vison in the Shiretoko Peninsula (Table 3). These two fish species were selected as potential food resources because they are regularly found in the Shiretoko rivers. However, biomass estimates derived from eDNA analysis may not necessarily reflect the number of individuals and their sizes. For instance, if large individuals are sparsely distributed or very small individuals are dispersed throughout the area, it may be challenging for N. vison to utilize them as a primary food resource. In addition, the timing of the sampling in this study differs between sampling for salmonids (2019.6 and 7) and for N. vison (2018.6 and 7, 2019. 6, 7, 10, 2020. 4). In general, summer should be suitable for eDNA sampling for fish (Hayami et al. 2020). The wide range of sampling period for N. vison encompasses and covers the sampling period for salmonids, and the summer sampling should have been ideal for this study because salmonids are abundant in study rivers in summer and so as N. vison in and around the rivers for foraging these fishes. Thus, to estimate the relative biomass among the rivers, representative values were obtained from samples from the same season of the same year. Moreover, it would be better to sample multiple times over a long period of time to increase the frequency of detection because N. vison is not always present at water bodies. Therefore, although the analysis did not show a significant relationship with the abundance of natural food resources, the difference in eDNA sampling timing would not have affected the results.
While small terrestrial mammals could also serve as a significant food resource for N. vison, this study did not conduct biomass estimations for terrestrial mammals. It is noteworthy that previous reports have indicated the presence of numerous small mammals, including rodents, in the study area (Minami et al. 2016). Therefore, including biomass estimations of such organisms in future studies is crucial for gaining a comprehensive understanding of the preferred environment of N. vison. Advances in eDNA and other state-of-art technologies in ecology will undoubtedly contribute to this endeavor.
The important point in applying eDNA to the management of N. vison would be that N. vison is a mammal that lives close to water bodies, that eDNA remains for some time (days to weeks) (Barnes and Turner 2015), and that eDNA flows down the river (Jo and Yamanaka 2022). Based on these characteristics, regular water sampling and eDNA analysis would make it possible to quickly detect an invasion into that river or area. Early detection before establishment is an important factor in invasive species management (Mehta et al. 2007). In addition, the effectiveness of monitoring sites at risk of invasion will be enhanced by understanding the environmental factors that helps the invasion and establishment of the target species and by evaluating the invasive potential at each monitoring site.
The number of eDNA studies on aquatic invasive species is increasing for the purpose of their management (Sieber et al. 2022; Wang et al. 2022). This study has demonstrated the applicability of eDNA monitoring through river water sampling to terrestrial mammals as well. The examination of N. vison establishment factors revealed no association with the proxy of biomass of naturally distributed fish species. However, it did identify significant associations with human activities such as the release of salmon juveniles and agricultural farming. This suggests that human activities can play a crucial role in the establishment of invasive mammals by providing food resources. Consequently, human beings not only contribute to the introduction of invasive species but also play a part of their establishment in new environments, potentially leading to the out competition of native species like weasels, sables, and martens. Given the difficulty of restricting human activities, it becomes essential to control invasive species by understanding their environmental preferences. For instance, if highly preferred food resources are identified, it may be possible to increase the efficiency of extermination by using them as attractants. Therefore, insights into establishment factors of invasive species have the potential to enhance the control of invasive species. To achieve this, it is important to efficiently understand species distribution and conditions necessary for establishment through eDNA and ecological monitoring, and to promote appropriate human activities for invasive species management based on this information.
References
Barnes MA, Turner CR (2015) The ecology of environmental DNA and implications for conservation genetics. Conserv Genet 17(1):1–17. https://doi.org/10.1007/s10592-015-0775-4
Benoit NP, Robinson KM, Kellogg CT, Lemay MA, Hunt BP (2023) Using qPCR of environmental DNA (eDNA) to estimate the biomass of juvenile Pacific salmon (Oncorhynchus spp.). Environmental DNA. 5(4):683–696
Butchart SHM, Walpole M, Collen B et al (2010) Global biodiversity : indicators of recent declines. Science 328:1164–1168. https://doi.org/10.1126/science.1187512
Danziger AM, Olson ZH, Frederich M (2022) Limitations of eDNA analysis for carcinus maenas abundance estimations. BMC Ecol Evol 22:14. https://doi.org/10.1186/s12862-022-01969-z
Deiner K, Bik HM, Mächler E et al (2017) Environmental DNA metabarcoding: transforming how we survey animal and plant communities. Mol Ecol 26:5872–5895. https://doi.org/10.1111/mec.14350
Dudgeon D, Arthington AH, Gessner MO et al (2006) Freshwater biodiversity: importance, threats, status and conservation challenges. Biol Rev Camb Philos Soc 81:163–182. https://doi.org/10.1017/S1464793105006950
Goldberg CS, Turner CR, Deiner K et al (2016) Critical considerations for the application of environmental DNA methods to detect aquatic species. Methods Ecol Evol 7:1299–1307. https://doi.org/10.1111/2041-210X.12595
Greenhalgh JA, Collins RA, Edgley DE et al (2022) Environmental DNA-based methods detect the invasion front of an advancing signal crayfish population. Environ DNA 4:596–607. https://doi.org/10.1002/edn3.280
Hall TA (1999) Bioedit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hayami K, Sakata MK, Inagawa T et al (2020) Effects of sampling seasons and locations on fish environmental DNA metabarcoding in dam reservoirs. Ecol Evol 10(12):5354–5367. https://doi.org/10.1002/ece3.6279
Ihara S, Inaba O, Fujiwara K, Yoji S (2013) Estimated density of american mink, Neovison vison, in two tributaries of the abukuma rivers. Wildl Conser Jpn 14:9–13
Ishige T, Miya M, Ushio M, Sado T, Ushioda M, Maebashi K, Yonechi R, Lagan P, Matsubayashi H (2017) Tropical-forest mammals as detected by environmental DNA at natural saltlicks in Borneo. In Biol Conserv 210:281–285. https://doi.org/10.1016/j.biocon.2017.04.023
Jo T, Yamanaka H (2022) Meta-analyses of environmental DNA downstream transport and deposition in relation to hydrogeography in riverine environments. Freshw Biol 67(8):1333–1343. https://doi.org/10.1111/fwb.13920
Kanbe T, Mizumoto H, Mitsuzuka T, Nakajima N, Araki H (2023) Co-occurrence patterns of endangered Sakhalin taimen and introduced rainbow trout in Hokkaido, Japan, inferred by environmental DNA metabarcoding. Aquat Conserv Mar Freshwat Ecosyst 33(12):1492–1500. https://doi.org/10.1002/aqc.4022
National Institute of Fisheries Research and Education, Fisheries Resources Laboratory, Division of Salmon, (2020) Artificial hatchery release plan of salmon. http://salmon.fra.affrc.go.jp/zousyoku/plan/R2syuuroku.pdf
Leempoel K, Hebert T, Hadly EA (2020) A comparison of eDNA to camera trapping for assessment of terrestrial mammal diversity. Proc R Soc B 287(1918):20192353. https://doi.org/10.1098/rspb.2019.2353
Lodge DM, Stein RA, Brown KM, Covich AP, Brönmark C, Garvey JE, Klosiewskt SP (1998) Predicting impact of freshwater exotic species on native biodiversity: challenges in spatial scaling. Aust J Ecol 23(1):53–67
Manfrin C, Mirimin L, Zanetti M et al (2022) Highly sensitive environmental DNA detection of topmouth gudgeon, Pseudorasbora parva: a comparison of qPCR and microfluidic qdPCR. Biol Invasions 24:2121–2133. https://doi.org/10.1007/s10530-022-02761-2
Mehta S, Haight R, Homans F, Polasky S, Venette R (2007) Optimal detection and control strategies for invasive species management. Ecol Econ 61:237–245. https://doi.org/10.1016/j.ecolecon.2006.10.024
Minami Y, Tsumita U, Shimoyama S, Yoshikawa T (2016) Impact of the American mink, Neovison vison, on small rodents inhabiting riparian forests along the headstream of Kushiro River in eastern Hokkaido. [In Japanese] Natural Environmental Science Research 29: 1-10 https://doi.org/10.32280/nesr.29.0_1
Minamoto T (2022) Environmental DNA analysis for macro-organisms: species distribution and more. DNA Res 29:1–9. https://doi.org/10.1093/dnares/dsac018
Ministry of Land, Infrastructure, Transport and Tourism (2023a) National regional planning bureau: digital national land information. Available online at: https://nlftp.mlit.go.jp/ksj/ [Accessed 25 April 2023]. [in Japanese]
Ministry of Land, Infrastructure, Transport and Tourism (2023b) Fundamental Geospatial Information. Available online at: https://www.gsi.go.jp/kiban/ [Accessed 25 April 2023]. [in Japanese]
Mizumoto H, Kishida O, Takai K et al (2022) Utilizing environmental DNA for wide-range distributions of reproductive area of an invasive terrestrial toad in Ishikari river basin in Japan. Biol Invasions 24:1199–1211. https://doi.org/10.1007/s10530-021-02709-y
Murakami T, Ikeda T, Shimada K (2011) Current distribution of an invasive alien species, the American Mink Neovison vison, in Shiretoko Peninsula and its adjacent areas. Bull Shiretoko Mus 33:61–67
R Core Team (2019). R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Aus- tria. Retrieved from https://www.R-project.org/
Sakata A, Sado T, Oka SI, Ushio M, Miya M (2023) Collection of environmental DNA from stemflow for monitoring arboreal biodiversity: preliminary validation using lichens. MethodsX. 11:102448
Schultz MT, Lance RF (2015) Modeling the sensitivity of field surveys for detection of environmental DNA (eDNA). PLoS ONE 10:e0141503. https://doi.org/10.1371/journal.pone.0141503
Sepulveda AJ, Nelson NM, Jerde CL, Luikart G (2020) Are environmental DNA methods ready for aquatic invasive species management? Trends Ecol Evol 35(8):668–678. https://doi.org/10.1016/j.tree.2020.03.011
Sieber N, Hartikainen H, Krieg R et al (2022) Parasite DNA detection in water samples enhances crayfish plague monitoring in asymptomatic invasive populations. Biol Invasions 24:281–297. https://doi.org/10.1007/s10530-021-02644-y
Stuart SN, Chanson JS, Cox NA et al (2004) Status and trends of amphibian declines and extinctions worldwide. Science 306:1783–1786. https://doi.org/10.1126/science.1103538
Su G, Logez M, Xu J et al (2021) Human impacts on global freshwater fish biodiversity. Science 371:835–838. https://doi.org/10.1126/science.abd3369
Takahara T, Minamoto T, Yamanaka H et al (2012) Estimation of fish biomass using environmental DNA. PLoS ONE 7:e35868. https://doi.org/10.1371/journal.pone.0035868
Tattersall FH, Macdonald DW, Hart BJ et al (2002) Is habitat linearity important for small mammal communities on farmland? J Appl Ecol 39:643–652. https://doi.org/10.1046/j.1365-2664.2002.00741.x
Uraguchi K, Saitoh T, Kondo N, Abe H (1987) Food habits of the feral mink (Mustela vison Schreber) in Hokkaido. J Mammal Soc Jpn 12:57–67
Ushio M, Fukuda H, Inoue T et al (2017) Environmental DNA enables detection of terrestrial mammals from forest pond water. Mol Ecol Resour 17:e63–e75. https://doi.org/10.1111/1755-0998.12690
Wang X, Zhang H, Lu G, Gao T (2022) Detection of an invasive species through an environmental DNA approach: the example of the red drum Sciaenops ocellatus in the East China sea. Sci Total Environ 815:152865. https://doi.org/10.1016/j.scitotenv.2021.152865
Williams KE, Huyvaert KP, Vercauteren KC et al (2018) Detection and persistence of environmental DNA from an invasive, terrestrial mammal. Ecol Evol 8:688–695. https://doi.org/10.1002/ece3.3698
Yamamoto S, Kitano S, Maekawa K, Koizumi I, Morita K (2006) Introgressive hybridization between Dolly Varden Salvelinus malma and white-spotted charr Salvelinus leucomaenis on Hokkaido island. Jpn. J Fish Biol 68:68–85
Yamamoto S, Morita K, Kikko T, Kawamura K, Sato S, Gwo J (2020) Phylogeography of a salmonid fish, masu salmon Oncorhynchus masou subspecies-complex, with disjunct distributions across the temperate northern Pacific. Freshw Biol 65(4):698–715. https://doi.org/10.1111/fwb.13460
Yatsuyanagi T, Ishida R, Sakata MK et al (2020) Environmental DNA monitoring for short-term reproductive migration of endemic anadromous species, Shishamo smelt (Spirinchus lanceolatus). Environ DNA 2:130–139. https://doi.org/10.1002/edn3.50
Acknowledgements
We would like to express our deep gratitude to Shuji Mitani from Forest Realize Co., Ltd, and Tetsu Yatsuyanagi for cooperating with the field survey, Takahiro Murakami from Shari Municipal Shiretoko Museum for providing the tissue samples and meaningful information, and Gen Bando from Asahiyama Zoo for cooperating with the water sampling in the zoo.
Funding
This study was funded by the Environment Research and Technology Development Fund (JPMEERF20204004) of the Environmental Restoration and Conservation Agency provided by the Ministry of Environment of Japan, and JSPS KAKENHI grant no. 20H03005, 20K06086, and 23H00329 (HA).
Author information
Authors and Affiliations
Contributions
TT, HM, and HA conceived and designed the study. TT and TN collected field data and samples. TT, TK, and SI performed laboratory experiments and environmental DNA analysis. MKS, TT, and TM performed the statistical analysis. MKS and TT wrote the first draft of the manuscript. All authors discussed the results and contributed to the development of the manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors declare that they have no conflict of interest.
Data availability
All raw data are included in the Supporting Information.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Takaba, T., Sakata, M.K., Kanbe, T. et al. Human activity-associated establishment of invasive mink population estimated using environmental DNA. Biol Invasions (2024). https://doi.org/10.1007/s10530-024-03407-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10530-024-03407-1