Abstract
New data from sites in Spain, Sicily and North America establish that the spread of the European legume Cytisus scoparius (Scotch broom) across North America involved the concurrent invasion of a European mobile genetic element (symbiosis island [SI]) into North American Bradyrhizobium nodule bacteria. At four SI loci, bacteria from nine C. scoparius populations across the continent were all identical to haplotypes present in European Bradyrhizobium strains. At seven non-symbiotic (housekeeping) loci, these American C. scoparius bacteria grouped into eight diverse lineages related to, or in some cases identical to, symbionts from several native North American legumes. Inoculation experiments showed that American Bradyrhizobium strains with the European SI variant conferred superior growth to C. scoparius plants compared to strains with SI variants found indigenously in North America. Cytisus scoparius invasion in North America thus enabled proliferation of a European SI variant that spread to multiple indigenous American Bradyrhizobium lineages via horizontal gene transfer. Spread of this SI variant, in turn, likely facilitated C. scoparius invasion of North American habitats.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Invasional meltdown has been traditionally viewed as a process of two or more species-level taxa facilitating each other's invasion (e.g., Simberloff and Von Holle 1999; Traveset and Richardson 2014; Braga et al. 2018). Initial work on legumes and nitrogen-fixing root nodule symbionts adopted this perspective and assumed that bacterial co-invaders would be strains with an entire genome derived from the ancestral geographic region (e.g., Parker 2001). However, horizontal gene transfer between indigenous and exotic rhizobia could be important in this process. Rhizobial symbiosis-related genes are typically clustered on mobile genetic elements such as plasmids or symbiosis islands (SI), which experience high rates of horizontal transfer (Sullivan and Ronson 1998; Andrews et al. 2018; Epstein and Tiffin 2021). The mobility of SIs raises the possibility that invasional meltdown could involve co-invasion of individual SI elements as a process distinct from whole-organism invasion. In cases where nodule symbionts from the ancestral range are co-introduced along with a legume into a novel habitat, these rhizobia provide a source of symbiotic genes highly adapted to that exotic legume species, which may then spread to other bacteria. As plants proliferate in the invaded habitat, they may select for the gain of exotic SI variants by indigenous bacteria that are well adapted to other features of the local environment (Le Roux et al. 2017; Andrews et al. 2018). The legume invader represents a potential new niche for indigenous rhizobia, which may be effectively utilized if they acquire a novel SI variant.
In this study, we analyzed invasional meltdown during the colonization of North America by the European legume Cytisus scoparius (Scotch broom). This species has invaded many parts of the world (Peterson and Prasad 1998), and was likely introduced to North America over 160 years ago, with populations established in three Canadian provinces and 29 states in the U.S. (https://www.invasiveplantatlas.org). Cytisus scoparius interacts mostly with nodule symbionts in the genus Bradyrhizobium (Lafay and Burdon 2006; Kalita and Malek 2017; Stepkowski et al. 2018). Bradyrhizobium strains have a chromosomally integrated symbiosis island that comprises about 7% of the genome (Kaneko et al. 2002). Within its native European range, most C. scoparius symbionts belong to a phylogenetic group variously known as the "B. japonicum superclade" (Ormeño-Orrillo and Martínez-Romero 2019) or "B. japonicum supergroup" (Avontuur et al. 2019). However, C. scoparius symbionts from a divergent lineage, the "B. elkanii superclade", have also been detected in one region of Spain (Rodríguez-Echeverría 2010).
A prior survey of 270 Bradyrhizobium isolates from 15 naturalized C. scoparius populations across North America found that 76% of the strains belonged to the B. japonicum superclade, and these were present at every location (Horn et al. 2014). More detailed characterization of 15 of these strains (based on five housekeeping (HK) loci) showed that nearly all were highly similar to C. scoparius nodule bacteria found in Europe.
However, the remaining 24% of North American C. scoparius symbionts belonged to the B. elkanii superclade. Multilocus analysis on seven of these strains suggested that they had mosaic ancestry (Horn et al. 2014). Phylogenetic trees inferred from five HK loci grouped these strains as close relatives of Bradyrhizobium strains associated with indigenous North American legumes. However, at three SI loci, they were highly similar or identical to a C. scoparius nodule symbiont from Spain. Thus, it appeared that bacteria ancestrally associated with other North American legumes may have evolved to become symbionts of C. scoparius by gaining symbiotic genes from a European ancestor.
Here, we re-examine this possible case of SI invasion via horizontal transfer in North American B. elkanii superclade symbionts of C. scoparius. We analyzed additional strains from a larger set of locations in both North America and Europe, and also expanded the set of genetic markers (7 HK loci and 4 SI loci), to permit more precise phylogenetic conclusions. We focused on three specific issues. First, do all B. elkanii superclade symbionts in North American C. scoparius populations have a SI derived from Europe? Second, how many independent acquisitions of the European SI variant have taken place in distinct Bradyrhizobium lineages? This will provide insights as to whether SI transfer is a rare event, or whether it has happened recurrently during legume invasion. Finally, is host legume growth differentially affected by symbionts of different phylogeographic origin?
Materials and methods
Bacterial strains
Four groups of Bradyrhizobium strains were analyzed (Supplementary Table S1): (1) 28 strains sampled from nine naturalized North American C. scoparius populations (five in the eastern U.S. [NY, RI, MA], and four in the western U.S. [WA, CA]; Horn et al. 2014). (2) nine B. elkanii-lineage strains from Tribe Genisteae legumes in the native European range of C. scoparius (Cardinale et al. 2008; Rodríguez-Echeverría 2010; Rodríguez-Echeverría et al. 2014). (3) 65 B. elkanii-lineage isolates from diverse indigenous legumes in the U.S. and Mexico. (4) 17 strains isolated from nodules developing on C. scoparius plants that were exposed to soil from eight legume habitats in New York where C. scoparius has never been present ("naïve soil"), representing novel symbionts potentially similar to those that C. scoparius encountered when it first arrived in North America (see Supplementary Methods).
DNA Sequencing
DNA was purified by heating cells in a lysis buffer followed by chloroform extraction (Parker et al. 2012). Portions of 11 gene loci were amplified and sequenced (see Boudehouche et al. 2020 for protocols). The seven HK loci were 23S rRNA (468 bp), gyrB (781 bp); rpoB (626 bp), dnaK (603 bp), rplC (471 bp); glnII (608 bp), and recA (513 bp). The 4 SI loci analyzed were nifD (822 bp), nifH (631 bp), nodA (633 bp), and nodC (552 bp).
A NeighborNet analysis on the seven concatenated HK loci and on the four concatenated SI loci was run using SplitsTree v. 4.16.2 (Huson and Bryant 2006). In both analyses (Supplementary Figs. S1 and S2), reticulation was low around the terminal branches of the network, so strain relationships can be reasonably approximated by a bifurcating tree. Therefore, trees were inferred as described (Boudehouche et al. 2020) for concatenated HK loci and for concatenated SI loci using MrBayes v. 3.2.7a (Ronquist et al. 2012).
Inoculation experiment
Cytisus scoparius growth rates were compared using one European strain (H11), four North American C. scoparius strains with the European SI variant, and four strains originating from nodules developing on C. scoparius plants exposed to naïve soil (from non-invaded habitats). Seeds were sterilized with sulfuric acid, then planted in a rhizobia-free 1:3 mixture of sand and potting soil. Plants were grown in a greenhouse with n = 14 seedlings randomly assigned to each treatment (9 bacterial strains and a set of uninoculated control plants). Six days after planting, seedlings in the nine inoculation treatments were given 5 mL of broth culture containing approximately 108 bacterial cells mL−1. Bacterial cross-contamination was avoided by using standard precautions involving watering by subirrigation and placing a perlite-gravel barrier on top of the soil (Wilkinson et al. 1996). These precautions appeared to be effective because no nodules developed on uninoculated control plants. Every 7–10 days, plant positions in the greenhouse were re-randomized. Total dry plant biomass, nodule number, and the diameter of the two largest nodules per plant were measured after 6 weeks.
Results
Relationships for symbiosis island loci
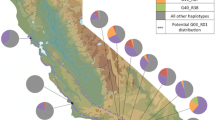
Bayesian analysis of four concatenated SI loci showed that all 28 North American C. scoparius strains clustered within a European clade (Fig. 1). Additionally, every individual locus for all North American C. scoparius strains had an identical haplotype to particular European haplotypes, except for nodA in one strain (csma27). All North American C. scoparius strains shared a nifH haplotype with strain Sj15c, and shared a nifD haplotype with six European strains (H11, AbruD10, AlisF2, CollC19, CollC20, ValeG5). The North American C. scoparius strain nodC haplotype was identical to that of seven European strains (Sj15c, H11, AbruD10, AlisF2, CollC19, CollC20, ValeG5). All but one North American C. scoparius strain shared a nodA haplotype with six European strains (H11, AbruD10, AlisF2, CollC19, CollC20, ValeG5). The nodA haplotype of strain csma27 from California had a single nucleotide difference from that haplotype, and was not present in any of the European strains analyzed. The csma27 nodA locus was amplified and sequenced again yielding the same result, confirming that the SNP was not a sequencing artifact.
Relationships for housekeeping loci
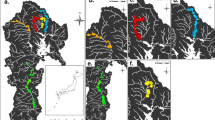
Major phylogenetic incongruity was evident between the SI tree (Fig. 1) and the tree inferred for housekeeping loci (Fig. 2). In contrast to SI loci, where North American C. scoparius strains were nearly all identical and matched European strains, their HK genes were diverse, and grouped into eight lineages (A-H) interspersed among native North American legume strains (Fig. 2). The largest lineage (G) contained 13 isolates from five populations in CA, MA, NY, and RI. These strains were identical at all seven housekeeping loci to strain drlt6, isolated from the native North American legume Desmodium rotundifolium in the same habitat (Lake Tiorati, NY) as five of the lineage G isolates. Although these strains were identical at all of the HK loci, SI loci were vastly different between strain drlt6 and the C. scoparius strains (Fig. 1). Thus, the lineage G C. scoparius strains could have evolved from a strain like drlt6 by acquiring a European SI variant.
Lineage B contained the next largest cluster of North American C. scoparius strains, with five strains from three populations in Washington State. The closest relative of these strains was cscv3, a strain that developed on C. scoparius plants exposed to naïve soil from New York.
Lineage F contained four strains from one population in Rhode Island. These strains had housekeeping genes nearly identical to strain aai25, a native North American legume strain from North Carolina (aai25 differed from lineage F at just one nucleotide site in one locus [rplC]).
Affinities of novel C. scoparius symbionts
Among 17 strains isolated from nodules developing on plants exposed to naïve soil from habitats in New York State where this European legume has never been present, none had the European SI variant (Fig. 1). Instead, their SI sequences fell into a variety of lineages with affinities to native North American legume strains. The HK tree also indicated affinities to North American legume strains (Fig. 2).
Symbiotic effectiveness
Plants inoculated with North American C. scoparius strains carrying the European SI variant exhibited an average biomass 78% higher than uninoculated plants (Supplementary Fig. S3). Strains originating from nodules on C. scoparius plants exposed to naïve soil (with other SI variants) had significantly poorer growth (F(1,6 df) = 52.46; P = 0.0004; see Fig. S3 and ANOVA details in Supplementary Methods). Strains with the European SI variant would likely have higher reproductive success in habitats dominated by C. scoparius, because such strains formed more abundant and larger nodules (Supplementary Table S3). In two further experiments examining additional strains, North American C. scoparius strains with the European SI variant were also found to confer significantly higher growth than strains with SI variants found indigenously in North America (Supplementary Table S4; ANOVA P = 0.0052 and P = 0.00083).
Discussion
Most native legumes in eastern North America associate primarily with B. elkanii superclade strains (Koppell and Parker 2012; Parker 2012). Thus, as C. scoparius colonized North American habitats, it would have encountered numerous Bradyrhizobium strains in the B. elkanii superclade that were adapted to a variety of indigenous American legumes. The strains isolated from nodules that developed on C. scoparius plants exposed to naïve soil confirmed that B. elkanii superclade strains readily colonize this legume host (Figs. 1, 2). However, these B. elkanii superclade strains had diverse symbiotic gene variants (Fig. 1) that apparently made them poor mutualist partners of C. scoparius (Fig. S3, Table S4). If nodule symbionts from Europe had been co-introduced along with C. scoparius into North America, these rhizobia would have provided a source of symbiotic genes highly adapted to this legume species, potentially available for other bacteria to acquire via horizontal transfer. The gain of a C. scoparius-adapted SI variant by indigenous North American Bradyrhizobium strains could have given them a competitive advantage for interacting successfully with this legume, compared to related strains carrying SI variants adapted to other legumes. It is clear that multiple independent gains of the European SI variant occurred in North America, because symbionts with this SI variant were distributed across several diverse clades in the tree for housekeeping loci (Fig. 2).
In the tree for housekeeping loci, no North American strains identical to European B. elkanii superclade strains were detected (Fig. 2). This suggests that while the European SI variant spread successfully across North America (Fig. 1), the original European B. elkanii superclade symbionts that introduced this SI variant have not persisted over time.
When native rhizobia first encounter exotic legumes during the early stages of invasion, symbiotic effectiveness may be impaired (Rodríguez-Echeverría et al. 2012). Spread of the European SI variant in North American Bradyrhizobium populations likely facilitated C. scoparius invasion, because strains with the European SI variant conferred superior plant growth relative to strains with other SI variants found indigenously in North America (Fig. S3, Table S4). To better understand the invasion of C. scoparius in other regions, as well as invasion ecology for other exotic legumes worldwide, horizontal transfer of SI variants merits further investigation.
Availability of data and material
All DNA sequences obtained for the project have been made available in GenBank (Supplementary Table S2).
References
Andrews M, De Meyer S, James EK, Stępkowski T, Hodge S, Simon MF, Young JPW (2018) Horizontal transfer of symbiosis genes within and between rhizobial genera: occurrence and importance. Genes 9:321. https://doi.org/10.3390/genes9070321
Avontuur JR, Palmer M, Beukes CW, Chan WY, Coetzee MPA, Blom J, Stepkowski T, Kyrpides NC, Woyke T, Shapiro N, Whitman WB, Venter SN, Steenkamp ET (2019) Genome-informed Bradyrhizobium taxonomy: where to from here? Syst Appl Microbiol 42:427–439. https://doi.org/10.1016/j.syapm.2019.03.006
Boudehouche W, Parker MA, Boulila F (2020) Relationships of Bradyrhizobium strains nodulating three Algerian Genista species. Syst Appl Microbiol 43:126074. https://doi.org/10.1016/j.syapm.2020.126074
Braga RR, Gómez-Aparicio L, Heger T et al (2018) Structuring evidence for invasional meltdown: broad support but with biases and gaps. Biol Invasions 20:923–936. https://doi.org/10.1007/s10530-017-1582-2
Cardinale M, Lanza A, Bonnì ML, Marsala S, Puglia AM, Quatrini P (2008) Diversity of rhizobia nodulating wild shrubs of Sicily and some neighbouring islands. Arch Microbiol 190:461–470. https://doi.org/10.1007/s00203-008-0394-2
Epstein B, Tiffin P (2021) Comparative genomics reveals high rates of horizontal transfer and strong purifying selection on rhizobial symbiosis genes. Proc Roy Soc Lond B 288:20201804. https://doi.org/10.1098/rspb.2020.1804
Horn K, Parker IM, Malek W, Rodríguez-Echeverría S, Parker MA (2014) Disparate origins of Bradyrhizobium symbionts for invasive populations of Cytisus scoparius (Leguminosae) in North America. FEMS Microbiol Ecol 89:89–98. https://doi.org/10.1111/1574-6941.12335
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267. https://doi.org/10.1093/molbev/msj030
Kalita M, Małek W (2017) Molecular phylogeny of Bradyrhizobium bacteria isolated from root nodules of tribe Genisteae plants growing in southeast Poland. Syst Appl Microbiol 40:482–491. https://doi.org/10.1016/j.syapm.2017.09.001
Kaneko T, Nakamura Y, Sato S et al (2002) Complete genomic sequence of nitrogen-fixing symbiotic bacterium Bradyrhizobium japonicum USDA110. DNA Res 9:189–197
Koppell JH, Parker MA (2012) Phylogenetic clustering of Bradyrhizobium symbionts on legumes indigenous to North America. Microbiol 158:2050–2059. https://doi.org/10.1099/mic.0.059238-0
Lafay B, Burdon JJ (2006) Molecular diversity of rhizobia nodulating the invasive legume Cytisus scoparius in Australia. J Appl Microbiol 100:1228–1238. https://doi.org/10.1111/j.1365-2672.2006.02902.x
Le Roux JJ, Hui C, Keet JH, Ellis AG (2017) Co-introduction vs ecological fitting as pathways to the establishment of effective mutualisms during biological invasions. New Phytol 215:1354–1360. https://doi.org/10.1111/nph.14593
Ormeño-Orrillo E, Martínez-Romero E (2019) A genomotaxonomy view of the Bradyrhizobium genus. Front Microbiol 10:1–13. https://doi.org/10.3389/fmicb.2019.01334
Parker MA (2001) Mutualism as a constraint on invasion success for legumes and rhizobia. Divers Distrib 7:125–136. https://doi.org/10.1046/j.1472-4642.2001.00103.x
Parker MA (2012) Legumes select symbiosis island sequence variants in Bradyrhizobium. Mol Ecol 21:1769–1778. https://doi.org/10.1111/j.1365-294X.2012.05497.x
Peterson DJ, Prasad R (1998) The biology of Canadian weeds. 109. Cytisus scoparius (L.) Link. Can J Plant Sci 78:497–504. https://doi.org/10.4141/P97-079
Rodríguez-Echeverría S (2010) Rhizobial hitchhikers from Down Under: Invasional meltdown in a plant-bacteria mutualism? J Biogeogr 37:1611–1622. https://doi.org/10.1111/j.1365-2699.2010.02284.x
Rodríguez-Echeverría S, Fajardo S, Ruiz-Díez B, Fernáne-Pascual M (2012) Differential effectiveness of novel and old legume-rhizobia mutualisms: implications for invasion by exotic legumes. Oecologia 170:253–261. https://doi.org/10.1007/s00442-012-2299-7
Rodríguez-Echeverría S, Moreno S, Bedmar EJ (2014) Genetic diversity of root nodulating bacteria associated with Retama sphaerocarpa in sites with different soil and environmental conditions. Syst Appl Microbiol 37:305–310. https://doi.org/10.1016/j.syapm.2013.11.003
Ronquist F, Teslenko M, Van Der Mark P et al (2012) MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Simberloff D, Von Holle B (1999) Positive interactions of nonindigenous species: invasional meltdown? Biol Invasions 1:21–32. https://doi.org/10.1007/s10530-017-1582-2
Stępkowski T, Banasiewicz J, Granada CE, Andrews M, Passaglia LMP (2018) Phylogeny and phylogeography of rhizobial symbionts nodulating legumes of the Tribe Genisteae. Genes 9:163. https://doi.org/10.3390/genes9030163
Sullivan JT, Ronson CW (1998) Evolution of rhizobia by acquisition of a 500-kb symbiosis island that integrates into a phe-tRNA gene. Proc Natl Acad Sci USA 95:5145–5149. https://doi.org/10.1073/pnas.95.9.5145
Traveset A, Richardson DM (2014) Mutualistic interactions and biological invasions. Annu Rev Ecol Evol Syst 45:89–113. https://doi.org/10.1146/annurev-ecolsys-120213-091857
Wilkinson HH, Spoerke JM, Parker MA (1996) Divergence in symbiotic compatibility in a legume-Bradyrhizobium mutualism. Evolution 50:1470–1477. https://doi.org/10.1111/j.1558-5646.1996.tb03920.x
Acknowledgements
We thank T. Powell and K. Prior for helpful suggestions on the manuscript, A. Biro for laboratory assistance, I.M. Parker and W. Malek for providing bacterial strains, and A.R. Parker for help with seed collection. Financial support was provided by NSF Grants DEB-0212369 and MCB-0640246.
Funding
Financial support was provided by NSF Grants DEB-0212369 and MCB-0640246.
Author information
Authors and Affiliations
Contributions
All authors collected strains of nodule bacteria for the research. R.F.K. and M.A.P. performed the DNA sequencing and data analysis. R.F.K. wrote the manuscript with support from S.R.-E., P.Q., and M.A.P.
Corresponding author
Ethics declarations
Conflicts of interest
All of the authors declare that they have no conflicts of interest or competing financial, professional, or personal interests that might have influenced the work described in this manuscript.
Consent for publication
All authors give the Publisher permission to publish the work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kruger, R.F., Rodríguez-Echeverría, S., Quatrini, P. et al. Invasional meltdown via horizontal gene transfer of a European symbiosis island variant in North American nodule symbionts of Cytisus scoparius. Biol Invasions 24, 2113–2120 (2022). https://doi.org/10.1007/s10530-022-02776-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-022-02776-9