Abstract
Malic enzyme (ME) catalyses the oxidative decarboxylation of l-malate to pyruvate and provides NADPH for intracellular metabolism, such as fatty acid synthesis. Here, the mitochondrial ME (mME) gene from Mortierella alpina was homologously over-expressed. Compared with controls, fungal arachidonic acid (ARA; 20:4 n−6) content increased by 60 % without affecting the total fatty acid content. Our results suggest that enhancing mME activity may be an effective mean to increase industrial production of ARA in M. alpina.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Malic enzyme (ME, EC 1.1.1.40), which catalyses the oxidative decarboxylation of l-malate to pyruvate, is a widely distributed NADPH-supplying enzyme and is suggested to be important during fatty acid synthesis in oleaginous fungi (Ratledge 2004, 2014; Zhang et al. 2007). On the basis of their subcellular localisation, MEs can be divided into cytosolic ME (cME) and mitochondrial ME (mME).
Mortierella alpina, an industrial oleaginous fungi used for the production of arachidonic acid (ARA; 20:4 n−6), can accumulate fatty acids up to 50 % of its dry weight. Two ME genes coding for a cME and a mME are present in its genome (Vongsangnak et al. 2013; Wang et al. 2011), and seven distinct isoforms of ME have been reported (Zhang and Ratledge 2008). Homologous over-expression of the cME gene malE1 increased fungal fatty acid content by 30 % without affecting the degree of fatty acid desaturation (Hao et al. 2014). Interestingly, a membrane-bound ME was suggested by Kendrick and Ratledge (1992) to play a role in the desaturation of fatty acids and could be in the form of a mitochondrially-associated ME. It is unclear, however, whether over-expression of mME would increase the level of unsaturated fatty acids.
In this study, a M. alpina, uracil auxotroph strain CCFM 501 generated by gene deletion (Hao et al. 2014), was used as the recipient for transformation. In contrast to malE1, homologous over-expression of M. alpina mME (malE2) significantly increased the amount of unsaturated fatty acid with little change in the total fatty acid content.
Materials and methods
Strains and growth conditions
Potato/dextrose/agar (PDA) medium was used for the wild-type M. alpina culture. M. alpina uracil auxotrophic strain CCFM 501 (Hao et al. 2014) was maintained on a GY medium consisting of 30 g glucose/l, 5 g yeast extract/l, 2 g KNO3/l, 1 g NaH2PO4/l and 0.3 g MgSO4·7H2O/l, supplemented with 0.5 g 5-fluoroorotic acid (5-FOA)/l and 0.05 g uracil/l. The GY medium contained 30 g glucose/l, 5 g yeast extract/l, 2 g KNO3/l, 1 g NaH2PO4/l and 0.3 g MgSO4·7H2O/l. Escherichia coli TOP10 and Agrobacterium tumefaciens C58C1 were used for the plasmid construction and fungal transformation, and were cultivated on YEP medium consisting of 10 g tryptone/l, 10 g yeast extract/l and 5 g NaCl/l. SC medium was used for the positive selection of the transformers. It consisted of 5 g yeast nitrogen base without amino acids or ammonium sulphate (Difco)/l, 1.7 g (NH4)SO4/l, 20 g glucose/l, 20 mg adenine/l, 30 mg tyrosine/l, 2 mg methionine/l, 2 mg arginine/l, 2 mg histidine/l, 4 mg lysine/l, 4 mg tryptophan/l, 6 mg threonine/l, 6 mg isoleucine/l, 6 mg leucine/l and 6 mg phenylalanine/l. The compositions of the minimal medium (MM) and induction medium (IM) have been described previously (Ando et al. 2009; Takeno et al. 2004). For the fatty acid analysis, M. alpina was grown at 28 °C in the Kendrick and Ratledge (1992) medium with glucose and diammonium tartrate as the principal carbon and nitrogen sources. The proliferative phase cultures of M. alpina (800 ml) were inoculated into a 7.5 l fermenter (BioFlo/CelliGen 115, New Brunswick Scientific, Edison, NJ, USA) to form a 4 l Kendrick and Ratledge medium culture. The fermenters were maintained at 28 °C, stirred at 500 rpm with an aeration rate of 0.5 vvm, and the pH was maintained at 6.0 by the automatic addition of 2 M KOH and HCl.
Construction of the T-DNA binary vector
The primer pair ITF (GCA TGC CAT GGA GARA GCT TGG TAC CGC TAG CTC CCA AGC GARA TTT GTC ATC TCG)/ITR (CGC GGA TCC GAG CTC CCC GGG GGA CTC GAG AGC ATA CGG ARAG TCC ATC AGT TAC G) was used to amplify an intron (IT) DNA sequence from the M. alpina genome. As illustrated in Fig. 1, the IT fragment was double-digested with NcoI and BamHI and introduced into the corresponding endonuclease sites of pET28a-HPHs (Hao et al. 2014) to replace the hpt gene. The resulting plasmid was designated pET28a-ITs. The ITs cassette was gel-purified from XbaI- and SpeI-digested pET28a-ITs, and introduced into XbaI-digested pBIG2-ura5s (Hao et al. 2014) to form pBIG2-ura5s-ITs. The malE2 gene was amplified from M. alpina cDNA with the primer pair malE2F (ATC GGG GTA CCA TGT TGA GGA ATC CTG CTC TCA)/malE2R (TARA TTC CCC CGG GTC AGG GGT GCG ATT CCA G) and sequence-analysed on an ABI PRISM 3730. The malE2 gene was double-digested with KpnI and XmaI, followed by ligation into pBIG2-ura5s-ITs treated with the same restriction enzymes. The resulting malE2 expression plasmid was named pBIG2-ura5s-malE2.
Construction of binary vectors for over-expressing malE2 gene in M. alpina. IT, intron sequence; his 550, his 550 promoter; trpCt, trpC terminator; hpt hygromycin B phosphotransferase gene, ura5 orotate phosphoribosyl transferase gene, RB right border, LB left border, malE2 mitochondrial malic enzyme gene of M. alpina
Agrobacterium tumefaciens-mediated transformation (ATMT)
ATMT was performed essentially as described previously (Hao et al. 2014). The spores of M. alpina CCFM 501 were harvested from 2-week cultures growing on GY agar medium containing 0.05 g uracil/l, centrifuged at 12,000×g for 20 min and washed once with 10 ml fresh liquid IM. The pellet was diluted to 108/ml with fresh liquid IM before use. A. tumefaciens C58C1 was electro-transformed by the binary vector pBIG2-ura5s-malE2. After identification by PCR, a single-bacteria colony was cultured at 28 °C with shaking at 200 rpm for 48 h in 20 ml of MM liquid medium containing 100 μg kanamycin/ml and 100 μg rifampicin/ml. Bacterial cells were harvested by centrifugation at 4,000×g for 5 min, washed once and diluted to an OD600 of 0.3 with fresh IM. The cells were incubated for 8–12 h at 28 °C with shaking at 200 rpm to an OD600 of 1.2. 100 µl of this cell suspension was mixed with an equal volume of the spore suspension and spread onto a cellophane membrane that was placed on solid plates of IM (containing 0.9 g glucose/l). The plates were incubated at 23 °C for 36–48 h in a dark incubator. The membranes were transferred to uracil-free SC plates supplemented with 50 μg cefotaxime/ml and 50 μg spectinomycin/ml and incubated at 28 °C until colonies appeared. The visible mycelia were transferred to uracil-free SC agar plates containing 50 μg cefotaxime/ml and 50 μg spectinomycin/ml and subcultured for three times to obtain stable transformants.
Genomic DNA preparation
After 4 days of cultivation in a GY liquid medium at 28 °C with shaking at 200 rpm, M. alpina mycelium were harvested by filtration and washed twice with sterile water. Genomic DNA was extracted by the method described previously (Chen et al. 2013).
Confirmation of the existence of the T-DNA in the genome
The integration of T-DNA in the genomic DNA was identified by PCR. Two pairs of primers (HisproF1/TrpCR1 and HisproF2/TrpCR2) were used as previously designed (Hao et al. 2014). The PCR cycling conditions were 95 °C for 5 min, followed by 30 cycles of amplification at 95 °C for 30 s, 55 °C for 30 s and 72 °C for 3 min, and a final extension step of 72 °C for 7 min.
RT-qPCR analysis
Mortierella alpina total RNA was isolated with Trizol and reverse-transcribed using the PrimeScript RT reagent kit (Takara, Otsu, Shiga, Japan) according to the manufacturer’s instructions. Primers used for the RT-qPCR were summarized in Supplementary Table 1. An ABI-Prism 7900 sequence-detection system (Applied Biosystems, Foster City, CA, USA) was used for the RT-qPCR analysis, together with a Power SYBR Green PCR Master Mix (Applied Biosystems, Foster City, CA, USA) according to the manufacturers’ instructions. The reaction mixtures were composed of 10 µl SYBR Green PCR Master Mix, 0.5 µl of each primer pair, 8 µl distilled water and 1 µl DNA template or distilled water as a no-template control. The PCR cycling conditions were 50 °C for 2 min and 95 °C for 10 min, followed by 40 cycles of amplification at 95 °C for 15 s and 60 °C for 30 s. The expression of the housekeeping gene 18S rRNA was used as the normalisation standard for gene expression; the primers 18SRTF/18SRTR were used as previously described (Hao et al. 2014).
ME activity determination
ME activity was determined as described previously (Zhang et al. 2007). Mycelia were collected by filtration, then frozen and ground in liquid N2 and suspended in an extraction buffer containing 20 % (w/v) glycerol, 100 mM of KH2PO4/KOH, pH 7.5, 1 mM benzamidine and 1 mM DTT. After centrifuging at 10,000×g for 10 min at 4 °C, the supernatant was immediately used to determine the ME activity by continuous spectrophotometric assays at 340 nm and 30 °C. Approx. 0.1 mg protein was added to 1 ml reaction mixture containing 80 mM KH2PO4/KOH at pH 7.5, 3 mM of MgCl2, 0.6 mM NADP+ and 25 mM l-malate.
Preparation of the membrane fraction
The preparation was performed essentially as described previously (Kendrick and Ratledge 1992). Mycelia were harvested and washed twice with 100 ml 50 mM phosphate, pH 7.0, containing 0.25 M sucrose, 2 mM benzamidine and 2 mM DTT. After gently grinding in liquid N2, the cell debris and organelles were removed by centrifuging at 48,000×g for 20 min at 4 °C. The supernatant was transferred and re-centrifuged at 100,000×g for 1 h at 4 °C. The pellet was washed once and resuspended to ~1 mg protein/ml before use.
NADP and NADPH quantification
Mycelia from each sample was rapidly collected and frozed with liquid N2. After lyophilization, dry mycelia of each sample was ground in liquid N2 respectively. NADP and NADPH were determined using the NADP/NADPH Quantification Colorimetric Kit (BioVision, California, USA) according to the manufacturer’s instructions.
FAME analysis
Fatty acids were extracted and methylated from ~20 mg freeze-dried mycelia as described previously (Wang et al. 2011). The fatty acid profiles were analysed as their methyl esters by GC/MS (GC-2010 Plus; GCMS-QP2010 Ultra, Shimadzu Co., Kyoto, Japan) with a 30 m × 0.25 mm Rtx-Waxetr column (film thickness 0.25 µm). The programme was as follows: 40 °C for 5 min, ramp to 120 at 20 °C per min, ramp to 190 at 5 °C per min, hold for 5 min, ramp to 220 at 5 °C per min, hold for 17 min. The carrier gas was helium. Fatty acid quantification was carried out using peak-height area integrals. The pentadecanoic acid was used as the internal standard to quantify the fatty acid methyl esters with aliphatic chain ≤18, and the heneicosanoic acid was used as the internal standard to quantify the fatty acid methyl esters with aliphatic chain >18.
Determination of the cell dry weight, and glucose and ammonium concentrations
The biomass was harvested by filtration and washed twice with distilled water. After being lyophilised, the cell dry weight was determined gravimetrically. The glucose concentration of the medium was determined using a glucose oxidase test kit (Rongsheng Biotech, Shanghai, China) and the ammonium concentration was calculated using the indophenol method, as described previously (Chaney and Marbach 1962).
Results
Generation of the malE2 over-expressing strains
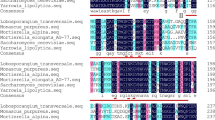
The malE2-encoding DNA fragment (1857 bp) was amplified from M. alpina cDNA with the primer pair malE2F/malE2R. Subcellular localization analysis, using the subCELlular LOcalization predictor (CELLO) version 2.5 (Yu et al. 2006), indicating that the ME coding by malE2 gene (named ME2) is located at mitochondria. Analysis of the malE2 gene (Supplementary Fig. 1) showed a 67.3 % identity with three mME-encoding genes in Mucor circinelloides CBS 277.49 (Vongsangnak et al. 2012). The protein coding by malE2 exhibited an identity of 32.9 % with that coding by malE1. The binary vector pBIG2-ura5s-malE2 (Fig. 1) was constructed for the transformation of the malE2 gene into the M. alpina uracil auxotrophic strain CCFM 501. The presence of T-DNA fragments in the genomic DNA was identified by PCR. In addition to the 818 and 861 bp fragments (representing the ura5 gene expression cassette), the presence of 2021 and 2064 bp products (representing the malE2 gene over-expression cassette) identified the presence of the malE2 over-expression cassette in the genome (Supplementary Fig. 2). There was no evidence in any of the transformants of multiple-copy insertion, which may be due to a high single-copy insertion rate (94 %) of ATMT (Ando et al. 2009).
Over-expression of the malE2 gene increased fatty acid desaturation
As illustrated in Fig. 2, three stable malE2-over-expressing strains (MA-malE2-1, MA-malE2-2 and MA-malE2-3) and control strains (wild type M. alpina, CCFM 501 and the ura5 gene reintroduction strain MAUC1) (Hao et al. 2014) were analysed after culture in a fully-controlled 7.5 l fermenter for 144 h. RT-qPCR analysis showed that the expression levels of the malE2 gene in all of the malE2 over-expressing strains were significantly higher than in the controls (Fig. 2a). The expression levels of the desaturase in MA-malE2-1 were not significantly changed compared to the wild type M. alpina (Supplementary Fig. 3).The ME activity was ~35 ± 4.2 nmol/(min mg) in all three controls, compared with ~55 ± 2.9 nmol/(min mg) in MA-malE2-1 (Fig. 2b). Analysis of the membrane ME activity revealed an ~300 % elevation in the malE2 over-expressing strains compare with the other strains (Fig. 2c). NADPH levels in MA-malE2-1 and MA-malE2-2 (ranged around 1.7 nmol/g dry weight) were also modestly higher than in the wild type M. alpina and MAUC1 (Fig. 2d). In the malE2 over-expressing strains, the cell fatty acid content was not significantly increased but arachidonic acid (ARA; 20:4 n−6) content increased by ~60 % (from 25 to 40 % of total fatty acids; Fig. 2f) compare with the controls, and the cell ω6 PUFA content increased from 42 to 62 % (Table 1). These results indicate that the over-expression of malE2 gene increases cell fatty acid desaturation.
MalE2 expression level (a), ME activity (b), membrane-bound ME activity (c), NADPH level (d), fatty acid content (e) and ARA content (f) of the control strains (M. alpina, CCFM 501 and MAUC1) and malE2-over-expressing strains (MA-malE2-1, MA-malE2-2 and MA-malE2-3). Samples were taken from cultures of each strain growing in Kendrick and Ratledge (1992) medium at a pH of 6.0 in a stirred 7.5 l fermenter with direct aeration for 144 h at 28 °C. The open bars represent the non-over-expressing strains and the black bars represent the malE2-over-expressing strains. Three independent experiments were performed and the error bars represent the standard deviations. Data with different superscript letters are significantly different (p < 0.05)
Fatty acid accumulation in the malE1 and malE2 over-expressing strains
The fatty acid production in the malE1 and malE2 over-expressing strains was analysed. As shown in Fig. 3, the malE1 over-expressing strain MA-malE1-1, the malE2 over-expressing strain MA-malE2-1 and the wild type M. alpina control were cultured in a fully controlled 7.5 l fermenter. The cell growth of the three strains was compared. In each case, ammonia (from ammonium tartrate) had been exhausted from the medium by 24 h. Strain MA-malE1-1 showed a relatively higher cell dry biomass (Fig. 3a) than the other two strains. In all three strains, ME activity showed a steadily decreasing trend (Fig. 3b), which was also described in M. circinelloides (Zhang et al. 2007). The fatty acid content in MA-malE1-1 improved to ~47 % cell dry weight at 144 h in comparison to the MA-malE2-1 and control cultures that had a fatty acid content of ~35 % in the same time period (Fig. 3c), along with a modest increase in PUFA content (Table 1). A higher yield of glucose to fatty acids at 144 than at 24 h in all three strains was observed (Fig. 3c; Table 2). The newly-accumulated fatty acids from 24 to 144 h in MA-malE1-1, MA-malE2-1 and the control were 4.9 ± 0.3, 3.3 ± 0.2 and 3 ± 0.2 g/l, respectively. The ME activity in MA-malE2-1 was significantly concentrated in the membrane during the entire culture period (Fig. 3d).
Cell dry weight (a), ME activity (b), fatty acid content (c), membrane-bound ME activity (d) and ARA content (e) of the control strain (M. alpina) and over-expressing strains (MA-malE1-1 and MA-malE2-1). Samples were taken at the indicated times from cultures of both strains growing in Kendrick and Ratledge (1992) medium at a pH of 6.0 in a stirred 7.5 l fermenter with direct aeration for up to 144 h at 28 °C. The closed squares represent the control strain, the closed circles represent the malE1-over-expressing strain MA-malE1-1 and the closed triangles represent the malE2-over-expressing strain MA-malE2-1. Three independent experiments were performed, and the error bars represent standard deviations
The difference in ARA content between MA-malE2-1 and the control became increasingly significant during the fatty acid accumulation phase (after nitrogen exhaustion), increasing from ~16–60 % (Fig. 3e), indicating that the over-expression of malE2 caused a relatively higher ARA synthesis rate. The newly-accumulated ARA from 24 to 144 h in MA-malE1-1, MA-malE2-1 and the control were 1.5 ± 0.1, 1.4 ± 0.1 and 0.8 ± 0.1 g/l, respectively. The results indicate an increased productivity of total fatty acids in the malE1 over-expressing strain and improved ARA production in both the malE1 and the malE2 over-expressing strains.
Discussion
PUFAs are essential for human health and must be acquired via the diet (Chen et al. 2007; Ratledge 2004; Simopoulos 1999). ARA is industrially produced by M. alpina (Ratledge 2013). However, the mechanism of fatty acid synthesis and desaturation in oleaginous fungi are still not fully understood. During fatty acid synthesis, ME play a role as an NADPH supplier (Zhang et al. 2007; Ratledge 2014). In this study, by homologously over-expressing the malE2 gene, the M. alpina cell ARA content improved by 60 % compared with controls but the total fatty acid content remained at its original level. The membrane ME activity was significantly higher in the malE2 over-expressing strain than in the other strains. These results indicate that the activity of ME coding by the malE2 gene is associated with cell fatty acid desaturation catalysed by membrane desaturases in M. alpina (Chen et al. 2013; Wang et al. 2011). Similar membrane-bound ME activity was originally suggested in another industrial oleaginous fungi, M. circinelloides (Kendrick and Ratledge 1992). This could be partly explained by the regulation of lipid synthesis via the organelle junctions of mitochondria and the endoplasmic reticulum (Rowland and Voeltz 2012).
The membrane-bound ME activity increased by more than 250 % in the malE2 over-expressing strains compared with the controls, but the ARA and ω-6 PUFA content improved only ~60 and 48 %, respectively. These results suggest that fatty acid desaturation in M. alpina is also restricted by other factors. A more significant improvement in ARA content rather than the total PUFA content may indicate a preference of this membrane-bound ME for delta-5 desaturase, which catalyses the desaturation of dihomo-gammalinolenic acid (DGLA) to ARA. ARA, the major fatty acid product of M. alpina, might be further improved by the over-expression of malE1 and malE2 to increase fatty acid content as well as desaturation.
Conclusion
Our results suggest that fatty acid desaturation in M. alpina may be regulated by multiple steps, including the supplementing of NADPH by membrane-bound ME.
References
Ando A, Sumida Y, Negoro H, Suroto DA, Ogawa J, Sakuradani E, Shimizu S (2009) Establishment of Agrobacterium tumefaciens-mediated transformation of an oleaginous fungus, Mortierella alpina 1s-4, and its application for eicosapentaenoic acid producer breeding. Appl Environ Microbiol 75:5529–5535
Chaney AL, Marbach EP (1962) Modified reagents for determination of urea and ammonia. Clin Chem 8:130–132
Chen YQ, Edwards IJ, Kridel SJ, Thornburg T, Berquin IM (2007) Dietary fat-gene interactions in cancer. Cancer Metast Rev 26:535–551
Chen H, Gu Z, Zhang H, Wang M, Chen W, Lowther WT, Chen YQ (2013) Expression and purification of integral membrane fatty acid desaturases. PLoS One 8:e58139
Hao G, Chen H, Wang L, Gu Z, Song Y, Zhang H, Chen W, Chen YQ (2014) Role of malic enzyme during fatty acid synthesis in oleaginous fungus Mortierella alpina. Appl Environ Microbiol 80:2672. doi:10.1128/AEM.00140-14
Kendrick A, Ratledge C (1992) Desaturation of polyunsaturated fatty acids in Mucor circinelloides and the involvement of a novel membrane-bound malic enzyme. Eur J Biochem 209:667–673
Ratledge C (2004) Fatty acid biosynthesis in microorganisms being used for single cell oil production. Biochimie 86:807–815
Ratledge C (2013) Microbial production of polyunsaturated fatty acids as nutraceuticals. In: McNeil B, Archer D, Giavasis I, Harvey L (eds) Microbial production of food ingredients, enzyme and nutraceuticals. Woodhead Publishing Ltd, Oxford, pp 531–558
Ratledge C (2014) The role of malic enzyme as the provider of NADPH in oleaginous microorganisms: a reappraisal and unsolved problems. Biotechnol Lett 36. doi:10.1007/s10529-014-1532-3
Rowland ARA, Voeltz GK (2012) Endoplasmic reticulum–mitochondria contacts: function of the junction. Nat Rev Mol Cell Biol 13:607–625
Simopoulos AP (1999) Essential fatty acids in health and chronic disease. Am J Clin Nutr 70:560S–569S
Takeno S, Sakuradani E, Murata S, Inohara-Ochiai M, Kawashima H, Ashikari T, Shimizu S (2004) Cloning and sequencing of the ura3 and ura5 genes, and isolation and characterization of uracil auxotrophs of the fungus Mortierella alpina 1s-4. Biosci Biotechnol Biochem 68:277–285
Vongsangnak W, Zhang Y, Chen W, Ratledge C, Song Y (2012) Annotation and analysis of malic enzyme genes encoding for multiple isoforms in the fungus Mucor circinelloides CBS 277.49. Biotechnol Lett 34:941–947
Vongsangnak W, Ruenwai R, Tang X, Hu X, Zhang H, Shen B, Song Y, Laoteng K (2013) Genome-scale analysis of the metabolic networks of oleaginous zygomycete fungi. Gene 521:180–190
Wang L, Chen W, Feng Y, Ren Y, Gu Z, Chen H, Wang H, Thomas MJ, Zhang B, Berquin IM, Li Y, Wu J, Zhang H, Song Y, Liu X, Norris JS, Wang S, Du P, Shen J, Wang N, Yang Y, Wang W, Feng L, Ratledge C, Chen YQ (2011) Genome characterization of the oleaginous fungus Mortierella alpina. PLoS One 6:e28319
Yu CS, Chen YC, Lu CH, Hwang JK (2006) Prediction of protein subcellular localization. Proteins 64:643–651
Zhang Y, Ratledge C (2008) Multiple isoforms of malic enzyme in the oleaginous fungus, Mortierella alpina. Mycol Res 112:725–730
Zhang Y, Adams IP, Ratledge C (2007) Malic enzyme: the controlling activity for lipid production? Overexpression of malic enzyme in Mucor circinelloides leads to a 2.5-fold increase in lipid accumulation. Microbiology 153:2013–2025
Acknowledgments
This work was supported by the National Natural Science Foundation of China (No. 21276108, 31271812), the Program for New Century Excellent Talents (NCET-13-0831), the National High Technology Research and Development Program of China (2011ARA100905, 2012ARA022105C), the National Science Fund for Distinguished Young Scholars (31125021), the National Basic Research Program 973 of China (2012CB720802), the 111 project B07029, and the Fundamental Research Funds for the Central Universities (No. JUSRP51320B).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hao, G., Chen, H., Du, K. et al. Increased fatty acid unsaturation and production of arachidonic acid by homologous over-expression of the mitochondrial malic enzyme in Mortierella alpina . Biotechnol Lett 36, 1827–1834 (2014). https://doi.org/10.1007/s10529-014-1546-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-014-1546-x