Abstract
Non-ribosomal peptide synthetases (NRPS) and type-I polyketide synthases (PKS-I) are multimodular enzymes involved in biosynthesis of oligopeptide and polyketide secondary metabolites produced by microorganisms such as bacteria and fungi. New findings regarding the mechanisms underlying NRPS and PKS-I evolution illustrate how microorganisms expand their metabolic potential. During the last decade rapid development of bioinformatics tools as well as improved sequencing and annotation of microbial genomes led to discovery of novel bioactive compounds synthesized by NRPS and PKS-I through genome-mining. Taking advantage of these technological developments metagenomics is a fast growing research field which directly studies microbial genomes or specific gene groups and their products. Discovery of novel bioactive compounds synthesized by NRPS and PKS-I will certainly be accelerated through metagenomics, allowing the exploitation of so far untapped microbial resources in biotechnology and medicine.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microorganisms produce a large repertoire of bioactive compounds including antibiotics, immunosuppressants, siderophores, antitumor agents and toxins (Walsh 2008). Multidomain modular non-ribosomal peptide synthetases (NRPS) and type-I polyketide synthases (PKS-I) which are detected in bacteria and fungi are often involved in biosynthesis of secondary metabolites (Amoutzias et al. 2008; Clardy et al. 2006; Finking and Marahiel 2004). Gevers et al. (1968) showed that the production of the cyclic decapeptide, gramicidin S, in cell extracts of Bacillus brevis was potent even after the addition of RNase or ribosome inhibitors in the supernatant (Gevers et al. 1968). Additional biochemical analyses demonstrated that gramicidin S synthesis did not include tRNA molecules or aminoacyl-tRNA-synthetases (Gevers et al. 1968). These findings actually led to the discovery of the NRPS.
Intensive research on NRPS and PKS-I gave a great impulse to the identification and application of new secondary metabolites in biomedicine and biotechnology (Amoutzias et al. 2008; Clardy et al. 2006). Studies on NRPS and PKS-I architecture and function revealed a strong analogy (Fig. 1). NRPS are built of catalytic units called modules (Schwarzer and Marahiel 2001; Strieker et al. 2010). Each module consists of 1,000–1,500 amino acids and, according to the co-linearity rule, the number and the order of the modules represents the number and the order of amino acids in the final product (Amoutzias et al. 2008; Schwarzer and Marahiel 2001).
Structure organization of the NRPS and PKS enzymes. a Each module of the non-ribosomal peptide synthetase consists of three main domains: C, A and PCP. The condensation domain (C) is responsible for the formation of the C–N bond between the elongated chain and the activated amino acid. The adenylation (A) domain activates its related amino acid and catalyzes the transfer of the activated substrate to the PCP domain of the same module. The epimerization domain (E) is an auxilliary domain that changes an l-amino acid into a d-amino acid. TE domain releases the final peptide product from the enzyme through cyclization or hydrolysis. b Each module of PKS type I consists of the folwing main domains KS, AT and ACP. The AT domains are responsible for the incorporation of malonyl or methylmalonyl-CoA monomers, while the KS domains form a C–C bond. ACP domains are equivalent to PCP domains of NRPS. The KR domain performs ketoreduction and TE domain releases the final product. NRPS DOMAINS: C condensation, A adenylation, PCP peptidyl-carrier, E epimerization, TE thioesterase. PKS DOMAINS: KS ketosynthase, AT acyl-transferase, ACP acyl-carrier, KR ketoreduction, TE thioesterase
Enzyme modules contain three catalytic domains: in NRPS, the adenylation (A) domain is responsible for recognition and activation of its related amino acid or hydroxyl acid followed by transfer of the activated substrate to the peptidyl-carrier (PCP) domain of the same module. Eventually, the condensation (C) domain of the downstream module catalyzes the formation of the C–N bond between the elongated chain and the activated amino acid (Stachelhaus and Walsh 2000; von Döhren 2004; Strieker et al. 2010). There are also present auxiliary domains such as the epimerization (E) domain which changes an l-amino acid into a d-amino acid as well as the dual/epimerization domains (E/C) which are responsible for both epimerization and condensation (Stachelhaus and Walsh 2000). Cyclization (Cy) domains have been detected in several NRPS gene clusters. These domains can replace C-domains responsible for the incorporation of cysteine, serine or threonine. The oxidation (Ox) domain which is usually located either downstream of the PCP-domain or in the C-terminus of the A-domain (Schwarzer and Marahiel 2001) catalyses the formation of an aromatic thiazol through oxidation of a thiazoline ring (Du et al. 2000). Thioesterase domains (TE) are usually located in the final NRPS module (Trauger et al. 2000). TE domains release the final peptide product from the enzyme through cyclization or hydrolysis (Schwarzer and Marahiel 2001; Strieker et al. 2010; Kopp and Marahiel 2007; Tanovic et al. 2008).
By structural and functional analogy, PKS-I modules contain three main domains: acyl-transferase (AT), ketosynthase (KS) and acyl-carrier (ACP) domains. The AT domains incorporate malonyl or methylmalonyl-CoA, while KS domains form a C–C bond. The ACP domain is equivalent to the PCP domain of NRPS (Amoutzias et al. 2008; Schwarzer and Marahiel 2001). The KR- and DH-domains are auxiliary domains of PKS and they perform ketoreduction and dehydration, respectively (Schwarzer and Marahiel 2001). Enoylreductase (ER) domains have been observed in PKS modules and they perform further reduction of the C–C double bonds (Schwarzer and Marahiel 2001).
A gene cluster with both NRPS and PKS encoding genes leads to the formation of hybrid NRPS/PKS-derived products. Hybrid systems interact in a head to tail fashion and their order modifies the structure of the bioactive compound (Amoutzias et al. 2008; Schwarzer and Marahiel 2001). Bioactive compounds synthesized by hybrid NRPS–PKS systems can be divided into two main classes. The first one includes natural products synthesized individually by NRPS and PKS and eventually coupled into a hybrid final product. In the second class the NRPS and PKS enzymes are functionally connected thus leading directly to a hybrid peptide-polyketide metabolite (Du et al. 2001).
Recent studies have provided a potential link between ribosomal and non-ribosomal synthesis of peptides. In normal ribosomal biosynthesis aminoacyl-tRNA synthetases catalyze aminoacylation of tRNAs with a specific amino acid. However in 2010, Mocibob et al. suggested that a group of atypical seryl-tRNA synthetase (SerRS)-like proteins discovered in various bacterial genomes may be involved in non-ribosomal peptide synthesis. This assumption is supported by the functional homology between atypical SerRS homologs and adenylation domains in NRPS. Both enzyme types catalyze a two-step reaction which includes the activation of the suitable amino acid as well as the specific aminoacylation of eligible carrier proteins (Mocibob et al. 2010).
Additional evidence regarding the connection of typical ribosomal and non-ribosomal peptide synthesis is offered from transferase PacB which participates in pacidamycin biosynthesis. Specifically, PacB is a tRNA-dependent aminoacyltransferase that catalyzes the incorporation of alanine, derived from an alanyl-tRNA, to the N-terminal residue of a tetrapeptide intermediate. The tetrapeptide intermediate is assembled through a NRPS system and the final yielded product is a pentapeptidyl nucleoside antibiotic (Zhang et al. 2011). The aforementioned examples suggest that non-ribosomal peptide synthesis is not completely divergent from the normal ribosomal logic indicating a possible evolutionary link between these biosynthetic pathways.
Evolution of NRPS and PKS-I
NRPS and PKS-I are involved in secondary metabolite biosynthesis. These megasynthases constitute a great energetic burden for the cell mainly due to their modular architecture (Amoutzias et al. 2008; Jenke-Kodama and Dittmann 2009b). In the case of NRPS a module of three domains that spans roughly 1,000 amino acids is responsible for the incorporation of one monomer in the elongated chain. The most striking example is the cyclosporin synthase, a protein of 15,000 amino acids which produces an undecapeptide (Lawen and Traber 1993). In other words, microbial cells synthesize huge enzymes (megasynthases) which means significant efforts in the level of transcription and translation in order to produce small secondary metabolites. On the other hand cells are capable to synthesize small peptides, comparable to oligopeptides produced by NRPS, ribosomically through normal protein synthesis. Prominent example is bacteriocin synthesis (van Belkum et al. 2011).
The reason why microorganisms use such energy using enzymes for the production of secondary metabolites remains rather elusive. A possible explanation could be that microorganisms take advantage of domain reshuffling in order to produce a vast repertoire of secondary metabolites through NRPS or that secondary metabolites produced by NRPS contain unusual amino acids that might not be easily incorporated to the final product through normal proteinosynthesis. In any case, such an energetic burden must be counterbalanced by the protective or adaptive advantage of the secondary metabolites in a given environment.
It is established by comprehensive studies that NRPS evolve by gene duplications which can be intragenic or intergenic and/or domain deletions. Module skipping, recombination events or point mutations can further contribute to metabolite diversification (Amoutzias et al. 2008; Wenzel et al. 2005; Jenke-Kodama et al. 2005).
Nodularin gene cluster is a representative example of the domain deletion mechanism. Nodularins and microcystins are produced by cyanobacteria and show several similarities regarding their chemical structure and biological activity (Rantala et al. 2004). Nodularin gene cluster has evolved from the microcystin gene cluster through deletion of two modules. Thus nodularin metabolites lack two amino acids in position 3 (Welker and von Döhren 2006).
Diversity of NRPS and PKS products is further increased by the module skipping mechanism leading to metabolites with revised activities. Myxochromides are well established lipopeptides produced from myxobacteria genus. A hybrid NRPS–PKS machinery forms the hexapeptide myxochromide A while a structurally similar assembly line produces the pentapeptide myxochromide S (Wenzel et al. 2005). The fourth amino acid (l-proline) of myxochromide A does not exist in myxochromide S product indicating that one module is skipped during the biosynthesis process of myxochromide S. The proposed mechanism suggests that the biosynthetic intermediate is transferred to the PCP domain of module 4 and the elongation process continues through the incorporation of alanine from the PCP domain of module 5 (Wenzel et al. 2006). Leinamycin (LNM), an antitumor agent, is a typical example of module skipping in polyketides. LNM gene cluster encodes a hybrid NRPS–PKS system which consists of two NRPS and five PKS modules (Tang et al. 2006). However PKS module-6 contains two ACP domains (ACP6-1 and ACP6-2). Using the skipping mechanism, both ACP domains can replace each other during LNM biosynthesis, despite the fact that ACP6-2 seems to be preferred due to its higher loading efficiency (Tang et al. 2006).
PKS-I in bacteria exhibit joint evolution with fatty acid synthases (FAS) of animals (Jenke-Kodama et al. 2005). The mechanisms by which diversification of PKS-I occurs in bacteria include: gene duplications and losses, recombination exchanges and Horizontal Gene Transfer (HGT) (Amoutzias et al. 2008; Jenke-Kodama and Dittmann 2009b; Jenke-Kodama et al. 2006; Rantala et al. 2004). HGT is a common phenomenon between bacteria and fungi as well as among bacteria and fungi leading to acquisition of novel NRPS or PKS-I genes (Amoutzias et al. 2008; Jenke-Kodama et al. 2005; Bode and Müller 2003). Aeruginosins are bioactive compounds produced by cyanobacteria. Aeruginosin gene clusters are present in evolutionary distant taxa, such as Microcystis and Planktothrix and they have probably originated from a common ancestor. However there is evidence indicating that HGT of specific aer genes between Microcystis and Planktothrix lineages took place (Ishida et al. 2009).
Slot and Rokas (2011) observed an unusual HGT phenomenon in Aspergillus nidulans, a filamentous fungus, that synthesizes polyketide-derived secondary metabolites including sterigmatocystin (Yu and Leonard 1995). The sterigmatocystin gene cluster contains 23 genes that form the complete biosynthetic pathway. The whole gene cluster had been transferred horizontally from A. nidulans into Podospora anserine and retained its sequence and functionality indicating that intact transfer of a complete gene cluster between different taxonomic classes can result in production of functional metabolites (Slot and Rokas 2011).
Phylogenomic analysis of PKS-I based on AT domains indicated that PKS-I genes of bacteria were separated in two different clades during evolution. Specifically all AT domains which activate malonyl-CoA, including some domains without any predicted substrates, belong to the first clade while those activating methylmalonyl-CoA or rare substrates are included in the second clade (Jenke-Kodama and Dittmann 2009b).Gene duplications followed by subfunctionalization contributed further to evolution of AT domains in the second clade (Jenke-Kodama et al. 2005).
Over the past two decades, several attempts have been made towards combinatorial biosynthesis of polyketides and non-ribosomal peptides (Condurso and Bruner 2012; Wong and Khosla 2012). Liu et al. (2011) engineered a polyketide product by swapping domains in asperfuranone and sterigmatocystin polyketide synthases of A. nidulans. They replaced the starter unit acyltransferase (SAT) domain in asperfuranone synthase with the SAT domain from sterigmatocystin synthase. The new metabolite produced by the hybrid synthase had the same cyclization and chain length but altered ring organization compared to the natural asperfuranone product (Condurso and Bruner 2012; Liu et al. 2011). These results advance the development of novel genetic engineering protocols and highlight the potential to engineer secondary metabolic pathways.
The first attempt to enlarge a polyketide synthase was described by Rowe et al. (2001). An extension module from the rapamycin PKS was interpolated between the first two extension modules of erythromycin PKS (Rowe et al. 2001). The hybrid multienzyme yielded a novel tetraketide as well as normal triketides produced by normal erythromycin polyketide synthases. However the tetraketide production was not efficient (<5 % of total products) since the major products were the normal molecules of the unextended PKS (Rowe et al. 2001; Thomas et al. 2002). The inserted module was skipped probably due to an ACP-to-ACP chain transfer indicating that skipping mechanism may be dependent on the active ACP of rapamycin module (Thomas et al. 2002). The above results indicate that module skipping can not be avoided in directed evolution taking place in the lab thus ensuring the production of polyketides which have been evolved through normal evolutionary process.
Discovery of novel bioactive compounds through genome-mining
During the last decade rapid development of bioinformatics tools as well as improved sequencing and annotation of microbial genomes led to identification of new NRPS and PKS gene clusters (Jenke-Kodama and Dittmann 2009a; Donadio et al. 2007). Basic Local Alignment Search Tool (BLAST) and the profile hidden Markov model suite of tools (HMMER) are widely used algorithms in NRPS and PKS discovery. BLAST algorithm determines proteins which have in common a short region with known NRPS or PKS but they might be functionally unrelated while HMMER helps to identify proteins that contain a specific domain, for example a C domain, which is characteristic of NRPS, but not of PKS (Amoutzias et al. 2008). GOLD (Genomes Online) database offers access to a plethora of genome and metagenome sequencing projects (Liolios et al. 2010). Presently there are more than 3000 completed projects while approx. 7,700 are ongoing. The completed projects include more than 2,700 bacterial genomes and 150 archaeal genomes (http://www.genomesonline.org). NORINE (http://bioinfo.lifl.fr/norine/) is a specialized database focused on non-ribosomal peptides and contains more than 1,000 entries (Caboche et al. 2008). NORINE provides information about the biological activity, the peptide structure as well as references related to each peptide. In 2010 SBSPKS (http://www.nii.ac.in/~pksdb/sbspks/master.html) which combines three separate tools for sequence analysis of polyketide synthases was published (Anand et al. 2010). Model_3D_PKS has been created to model and analyze 3D structures of individual catalytic and structural PKS domains. Dock_Dom_Anal can be used to identify inter-subunit interactions in modular PKS clusters. The prediction is based on docking domain analysis. NRPS–PKS is an additional tool used to detect and identify novel NRPS and PKS clusters, their catalytic activity, active site residues as well as substrate specificities. NRPS–PKS is based on well-characterized gene clusters with known sequence and structure. A recently published online platform, antiSMASH (http://antismash.secondarymetabolites.org/), extends user’s abilities to analyze secondary metabolite gene clusters (Medema et al. 2011). The antiSMASH is able to identify gene clusters encoding secondary metabolites, including NRPS/PKS compounds, in a plethora of bacterial and fungal genomes and provide further information about their function and chemical structure as well.
The aforementioned bioinformatics platforms and software could be valuable tools for discovery of novel NRPS/PKS metabolites through genome mining. Orfamide A is a recently discovered NRPS metabolite using a genome mining approach (Table 1). In Pseudomonas fluorescens Pf-5 genome a cryptic biosynthetic gene cluster was identified and subsequently the substrate specificities and domain organization were predicted using bioinformatics tools. The substrate predictions led to the use of isotopically labeled precursors in P. fluorescens cultures which were incorporated into the peptide natural product. Thus a novel natural product was isolated proving that genome mining approaches allow the discovery of new bioactive compounds avoiding any genetic manipulations (Gross et al. 2007; Challis 2008; Zerikly and Challis 2009).
Salinilactam was discovered in 2007 by mining the genome of the marine actinomycete Salinispora tropica (Table 1) (Udwary et al. 2007). Sequence analysis revealed a cryptic PKS gene cluster. The modular prediction in conjunction with the putative substrates led to the assumption that the product of interest is a polyene macrolactam. These compounds absorb UV light at specific wavelengths and based on this feature salinilactam A was isolated.
Genome mining approaches can also be applied in silent cryptic biosynthetic gene clusters. In the case of aspyridones a silent cryptic gene cluster (apd) encoding a PKS-NRPS system was identified by mining the genome of A. nidulans (Table 1) (Bergmann et al. 2007; Zerikly and Challis 2009). Gene expression of the apd locus had to be induced using a suitable promoter. In this way the metabolic profiles of wild type and mutant strains were compared and the putative product of the cryptic gene cluster was identified and characterized.
Streptomyces genus is known to be a primary bacterial group in terms of producing many antibiotics as well as peptides and polyketides. The first cryptic gene cluster discovered through genome mining in the genome of Streptomyces coelicolor was reported in 2000 (Challis and Ravel 2000). The cch gene cluster contained an NRPS consisting of three modules and it produced a tetrapeptide siderophore named coelichelin (Table 1) (Challis and Ravel 2000; Lautru et al. 2005; Challis 2008).
Discovery of other bioactive compounds produced by Streptomyces spp. using the genome-mining approach continued in recent studies. Holomycin is an N-acylated dithiolopyrrolone antibiotic produced by Streptomyces clavuligerus (Table 1) (Li and Walsh 2010). The putative biosynthetic gene cluster was identified in the genome and in-frame deletions of two biosynthetic genes including one encoding an NRPS module were constructed thus resulting in mutant strains. Wild-type and mutant strains were screened and the putative product was not detected in deletion mutants. Further biochemical analyses led to the identification and purification of holomycin (Li and Walsh 2010).
Laspartomycins are lipopeptide antibiotics produced by Streptomyces viridochromogenes (Table 1). They exhibit significant activity against Gram-positive pathogens including methicillin-resistant Staphylococcus aureus (MRSA). The lpm gene cluster consists of 21 open reading frames (ORFs) (Wang et al. 2011). Disrupting one of the four NRPS genes resulted in complete abolishment of laspartomycins production in mutant strains. Additionally information provided by cloning of the whole lpm gene cluster allowed further understanding of their molecular mechanism and it will contribute in future combinatorial approaches (Wang et al. 2011).
Discovery of novel bioactive compounds through metagenomics
Metagenomics is a fast growing research field used to study microbial genomes or specific gene groups and their products. The most prominent advantage of metagenomics is that it can be applied in the study of unculturable or yet-uncultured microbes which are part of microbial communities present in environmental samples (e.g. soil, sea water) or hosts (e.g. plants, insects, human) (Schmeisser et al. 2007; Meyer et al. 2008; Simon and Daniel 2009). The increasing demand for novel bioactive compounds and the scientific inquiry to study unknown microbial communities led to an explosive growth of metagenomic projects in the last years (Simon and Daniel 2009; Riesenfeld et al. 2004; Sleator et al. 2008).
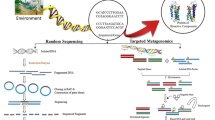
Metagenomic projects adopt in most cases two distinct approaches (Fig. 2): In comparative metagenomics high-throughput sequencing of total DNA isolated from environmental samples leads to identification of gene clusters and metabolic pathways via comparison with genes or pathways of known function found in microbial genomes based solely on DNA sequence similarity (Sleator et al. 2008). Sequence-driven approaches have been applied in the majority of metagenomic projects since they allow the identification of genes without requiring gene expression in a host strain (Simon and Daniel 2009). Random shotgun sequencing of metagenomes involves the construction of small-insert DNA clone libraries which are subsequently analyzed yielding small read lengths (Kennedy et al. 2010). The overlapping contigs are assembled using advanced bioinformatics tools in order to obtain full gene clusters. PCR-DGGE followed by metagenome walking is an alternative approach that allows retrieval of full-lengthed functional genes (Morimoto and Fujii 2009; Simon and Daniel 2009).
Metagenomics strategies. Functional and sequence-based approaches (modified from Simon and Daniel 2009)
On the other hand, in functional metagenomics an environmental sample is collected and total DNA is extracted from it which is then used to construct metagenomic libraries (Kennedy et al. 2010; Chistoserdova 2010; Craig et al. 2010). These libraries are screened in order to identify specific functions such as antibiotic or enzyme production. As soon as the function of interest is identified the clone which contains the responsible gene(s) for that function is sequenced and finally is compared with DNA from other organisms or communities. In this way new gene classes of known or even unknown functions can be directly identified (Kennedy et al. 2010; Chistoserdova 2010). The main limitation of the function-based approach is possibly the poor expression of interesting genes in a heterologous host strain. In most studies E. coli has been the host strain of choice. However additional bacterial strains from Streptomyces, Pseudomonas and Bacillus genera have been proposed (Ekkers et al. 2012). The ability to express the genes of interest in a host strain could be hampered due to a plethora of factors, including differences in codon usage, lack of regulating elements and abnormalities in protein folding (Ekkers et al. 2012; Schmieder and Edwards 2012; Uchiyama and Miyazaki 2009). In addition standard laboratory protocols used for gene expression could be improper since they cannot be applied in all genes and their encoded proteins. Potential toxicity of the gene product as well as absence of gene product secretion can be significant drawbacks of heterologous gene expression (Ekkers et al. 2012; Schmieder and Edwards 2012). Therefore if the aforementioned limitations are overpassed in the future, functional metagenomics could become a powerful tool for isolation of enzymes and compounds appropriate for biotechnological applications (Schmeisser et al. 2007; Simon and Daniel 2009).
Metagenomics approaches which have been successfully applied to discovery and isolation of novel compounds produced by NRPS and PKS are highlighted in the following examples: Pederin is a polyketide produced by an uncultured bacterial symbiont of Paederus fuscipes beetles (Table 1) (Piel 2002). Using sequence-based analysis and genomic library constructions the authors managed to reveal the complete 54 kb gene cluster containing PKS and NRPS genes, responsible for pederin biosynthesis (Piel 2002; Piel et al. 2005).
Hildebrand et al. (2004) identified the bryostatin PKS gene cluster in “Candidatus Endobugula sertula”, a bacterial symbiont of the marine bryozoan Bugula neritina (Table 1). PCR of total DNA using degenerated primers (Davidson et al. 2001) led to amplification of 300 bp products of the β-ketoacyl synthase (KSa) which were used as probes in order to screen genomic libraries. In this way the 65 kb gene cluster of bryostatin was identified and genetic analysis of the bryA gene revealed that it is responsible for the initial steps of bryostatin biosynthesis (Hildebrand et al. 2004).
The discovery of bryostatin led eventually to the identification of a vast amount of polyketide synthases of biotechnological interest. Marine sponges constitute a rich reservoir of bioactive compounds due to their bacterial symbionts. Hence metagenomics strategies in sponges have led to the identification of the sup PKS group The sup PKSs are dominant in marine sponges and encode synthases that are present exclusively in these animals. These synthases produce methyl-branched fatty acids and they are assumed to be of bacterial origin due to their similarity to metabolites that are commonly present in bacteria (Hochmuth and Piel 2009).
Members of the Acidobacteria phylum may also be a rich source of novel polyketide metabolites. A fosmid metagenomic library was constructed using agricultural soil samples. The library was screened using degenerate primers and probes specific for the KS domain of PKS enzymes. The obtained positive clones were further sequenced and sequences were analysed using standard bioinformatics tools (e.g. BLAST). Phylogenetic analysis of the positive clones suggested that they are closely related to PKS of Acidobacteria. (Parsley et al. 2011). Although no polyketide product from Acidobacteria sp. Has been detected so far, nevertheless it seems plausible that they are a rather unexplored source of polyketide compounds.
Application of metagenomic approaches in the tropical marine cyanobacterium Lyngbya bouillonii resulted in the identification of a 58 kb gene cluster encoding a hybrid PKS-NRPS system. Metagenomic DNA libraries were formed using cyanobacterial filaments while simultaneously single cells were isolated and multiple displacement amplification was performed in their whole genome. Combining the two strategies, apr gene cluster was detected and proved to be responsible for the biosynthesis of the antitumor natural product apratoxin A (Table 1) (Grindberg et al. 2011).
Concluding remarks
NRPS and polyketide synthases are pivotal enzymes in secondary metabolism of microorganisms. Many bioactive compounds which are synthesized by these enzymes help microorganisms to adapt and compete under various environmental conditions. New findings regarding the mechanisms underlying NRPS and PKS evolution, illustrate how microorganisms take advantage of stochastic events such as gene duplication or domain deletion in order to expand their metabolic potential. Recent advances in bioinformatics tools and sequencing technologies (e.g. pyrosequencing) allowed the study of whole microbial communities consisting of many unknown species through metagenomics. Metagenomics open new avenues in bioactive compound discovery and it is expected many useful compounds to be discovered and exploited in foreseeable future. Interdisciplinary research on bioactive compounds synthesized by NRPS and PKS will continue to expand in order to exploit the vast repertoire of secondary metabolites produced by microorganisms (including unculturable or yet-uncultured). Advances in bioinformatics and combinatorial chemistry will further facilitate discovery or man-made modification of potentially therapeutic compounds which are much needed.
References
Amoutzias GD, Van de Peer Y, Mossialos D (2008) Evolution and taxonomic distribution of non-ribosomal peptide and polyketide synthases. Future Microbiol 3(3):361–370
Anand S, Prasad MV, Yadav G, Kumar N, Shehara J, Ansari MZ, Mohanty D (2010) SBSPKS: structure based sequence analysis of polyketide synthases. Nucleic Acids Res 38(Web Server issue):W487–W496
Bergmann S, Schümann J, Scherlach K, Lange C, Brakhage AA, Hertweck C (2007) Genomics-driven discovery of PKS-NRPS hybrid metabolites from Aspergillus nidulans. Nat Chem Biol 3(4):213–217
Bode HB, Müller R (2003) Possibility of bacterial recruitment of plant genes associated with the biosynthesis of secondary metabolites. Plant Physiol 132(3):1153–1161
Caboche S, Pupin M, Leclère V, Fontaine A, Jacques P, Kucherov G (2008) NORINE: a database of non-ribosomal peptides. Nucleic Acids Res 36(Database issue):D326–D331
Challis GL (2008) Genome mining for novel natural product discovery. J Med Chem 51(9):2618–2628
Challis GL, Ravel J (2000) Coelichelin, a new peptide siderophore encoded by the Streptomyces coelicolor genome: structure prediction from the sequence of its non-ribosomal peptide synthetase. FEMS Microbiol Lett 187(2):111–114
Chiang YM, Szewczyk E, Nayak T, Davidson AD, Sanchez JF, Lo HC, Ho WY, Simityan H, Kuo E, Praseuth A, Watanabe K, Oakley BR, Wang CC (2008) Molecular genetic mining of the Aspergillus secondary metabolome: discovery of the emericellamide biosynthetic pathway. Chem Biol 15(6):527–532
Chistoserdova L (2010) Recent progress and new challenges in metagenomics for biotechnology. Biotechnol Lett 32(10):1351–1359
Clardy J, Fischbach MA, Walsh CT (2006) New antibiotics from bacterial natural products. Nat Biotechnol 24(12):1541–1550
Condurso HL, Bruner SD (2012) Structure guided approaches toward exploiting and manipulating non-ribosomal peptide and polyketide biosynthetic pathways. Curr Opin Chem Biol, in press
Craig JW, Chang FY, Kim JH, Obiajulu SC, Brady SF (2010) Expanding small-molecule functional metagenomics through parallel screening of broad-host-range cosmid environmental DNA libraries in diverse proteobacteria. Appl Environ Microbiol 76(5):1633–1641
Davidson SK, Allen SW, Lim GE, Anderson CM, Haygood MG (2001) Evidence for the biosynthesis of bryostatins by the bacterial symbiont “Candidatus Endobugula sertula” of the bryozoan Bugula neritina. Appl Environ Microbiol 67(10):4531–4537
Donadio S, Monciardini P, Sosio M (2007) Polyketide synthases and non-ribosomal peptide synthatases: the emerging view from bacterial genomics. Nat Prod Rep 24(5):1073–1109
Du L, Chen M, Sánchez C, Shen B (2000) An oxidation domain in the BlmIII non-ribosomal peptide synthetase probably catalyzing thiazole formation in the biosynthesis of the anti-tumor drug bleomycin in Streptomyces verticillus ATCC15003. FEMS Microbiol Lett 189(2):171–175
Du L, Sánchez C, Shen B (2001) Hybrid peptide-polyketide natural products: biosynthesis and prospects toward engineering novel molecules. Metab Eng 3(1):78–95
Ekkers DM, Cretoiu MS, Kielak AM, Elsas JD (2012) The great screen anomaly—a new frontier in product discovery through functional metagenomics. Appl Microbiol Biotechnol 93(3):1005–1020
Eustáquio AS, Nam SJ, Penn K, Lechner A, Wilson MC, Fenical W, Jensen PR, Moore BS (2011) The discovery of salinosporamide K from the marine bacterium “Salinispora pacifica” by genome mining gives insight into pathway evolution. ChemBioChem 12(1):61–64
Finking R, Marahiel MA (2004) Biosynthesis of non-ribosomal peptides. Annu Rev Microbiol 58:453–488
Gevers W, Kleinkauf H, Lipmann F (1968) The activation of amino acids for biosynthesis of gramicidin S. Proc Natl Acad Sci USA 60(1):269–276
Grindberg RV, Ishoey T, Brinza D, Esquenazi E, Coates RC, Liu WT, Gerwick L, Dorrestein PC, Pevzner P, Lasken R, Gerwick WH (2011) Single cell genome amplification accelerates identification of the apratoxin biosynthetic pathway from a complex microbial assemblage. PLoS ONE 6(4):e18565
Gross H, Stockwell VO, Henkels MD, Nowak-Thompson B, Loper JE, Gerwick WH (2007) The genomisotopic approach: a systematic method to isolate products of orphan biosynthetic gene clusters. Chem Biol 14(1):53–63
Hildebrand M, Waggoner LE, Liu H, Sudek S, Allen S, Anderson C, Sherman DH, Haygood M (2004) bryA: an unusual modular polyketide synthase gene from the uncultivated bacterial symbiont of the marine bryozoan Bugula neritina. Chem Biol 11(11):1543–1552
Hochmuth T, Piel J (2009) Polyketide synthases of bacterial symbionts in sponges—evolution-based applications in natural products research. Phytochemistry 70(15–16):1841–1849
Huang T, Wang Y, Yin J, Du Y, Tao M, Xu J, Chen W, Lin S, Deng Z (2011) Identification and characterization of the pyridomycin biosynthetic gene cluster of Streptomyces pyridomyceticus NRRL B-2517. J Biol Chem 286(23):20648–20657
Ishida K, Christiansen G, Yoshida WY, Kurmayer R, Welker M, Valls N, Bonjoch J, Hertweck C, Börner T, Hemscheidt T, Dittmann E (2007) Biosynthesis and structure of aeruginoside 126A and 126B, cyanobacterial peptide glycosides bearing a 2-carboxy-6-hydroxyoctahydroindole moiety. Chem Biol 14(5):565–576
Ishida K, Welker M, Christiansen G, Cadel-Six S, Bouchier C, Dittmann E, Hertweck C, Tandeau de Marsac N (2009) Plasticity and evolution of aeruginosin biosynthesis in cyanobacteria. Appl Environ Microbiol 75(7):2017–2026
Jenke-Kodama H, Dittmann E (2009a) Bioinformatic perspectives on NRPS/PKS megasynthases: advances and challenges. Nat Prod Rep 26(7):874–883
Jenke-Kodama H, Dittmann E (2009b) Evolution of metabolic diversity: Insights from microbial polyketide synthases. Phytochemistry 70(15–16):1858–1866
Jenke-Kodama H, Sandmann A, Müller R, Dittmann E (2005) Evolutionary implications of bacterial polyketide synthases. Mol Biol Evol 22(10):2027–2039
Jenke-Kodama H, Börner T, Dittmann E (2006) Natural biocombinatorics in the polyketide synthase enes of the actinobacterium Streptomyces avermitilis. PLoS Comput Biol 2(10):e132
Kennedy J, Flemer B, Jackson SA, Lejon DP, Morrissey JP, O’Gara F et al (2010) Marine metagenomics: new tools for the study and exploitation of marine microbial metabolism. Mar Drugs 8(3):608–628
Kopp F, Marahiel MA (2007) Macrocyclization strategies in polyketide and non-ribosomal peptide biosynthesis. Nat Prod Rep 24(4):735–749
Lautru S, Deeth RJ, Bailey LM, Challis GL (2005) Discovery of a new peptide natural product by Streptomyces coelicolor genome mining. Nat Chem Biol 1(5):265–269
Lawen A, Traber R (1993) Lawen A, Traber R (1993) Substrate specificities of cyclosporin synthetase and peptolide SDZ 214-103 synthetase. Comparison of the substrate specificities of the related multifunctional polypeptides. J Biol Chem 268(27):20452-65. J Biol Chem 268(27):20452–20465
Li B, Walsh CT (2010) Identification of the gene cluster for the dithiolopyrrolone antibiotic holomycin in Streptomyces clavuligerus. Proc Natl Acad Sci USA 107(46):19731–19735
Liolios K, Chen IM, Mavromatis K, Tavernarakis N, Hugenholtz P, Markowitz VM, Kyrpides NC (2010) The Genomes On Line Database (GOLD) in 2009: status of genomic and metagenomic projects and their associated metadata. Nucleic Acids Res 38(Database issue):D346–D354
Liu T, Chiang YM, Somoza AD, Oakley BR, Wang CC (2011) Engineering of an “unnatural” natural product by swapping polyketide synthase domains in Aspergillus nidulans. J Am Chem Soc 133(34):13314–13316
Loper JE, Henkels MD, Shaffer BT, Valeriote FA, Gross H (2008) Isolation and identification of rhizoxin analogs from Pseudomonas fluorescens Pf-5 by using a genomic mining strategy. Appl Environ Microbiol 74(10):3085–3093
Medema MH, Blin K, Cimermancic P, de Jager V, Zakrzewski P, Fischbach MA, Weber T, Takano E, Breitling R (2011) antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res 39(Web Server issue):W339–W346
Meyer F, Paarmann D, D’Souza M, Olson R, Glass EM, Kubal M et al (2008) The metagenomics RAST server—a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinformatics 9:386
Mocibob M, Ivic N, Bilokapic S, Maier T, Luic M, Ban N, Weygand-Durasevic I (2010) Homologs of aminoacyl-tRNA synthetases acylate carrier proteins and provide a link between ribosomal and non-ribosomal peptide synthesis. Proc Natl Acad Sci USA 107(33):14585–14590
Morimoto S, Fujii T (2009) A new approach to retrieve full lengths of functional genes from soil by PCR-DGGE and metagenome walking. Appl Microbiol Biotechnol 83(2):389–396
Parsley LC, Linneman J, Goode AM, Becklund K, George I, Goodman RM, Lopanik NB, Liles MR (2011) Polyketide synthase pathways identified from a metagenomic library are derived from soil Acidobacteria. FEMS Microbiol Ecol 78(1):176–187. doi:10.1111/j.1574-6941.2011.01122.x
Piel J (2002) A polyketide synthase-peptide synthetase gene cluster from an uncultured bacterial symbiont of Paederus beetles. Proc Natl Acad Sci USA 99(22):14002–14007
Piel J, Hui D, Wen G, Butzke D, Platzer M, Fusetani N, Matsunaga S (2004) Antitumor polyketide biosynthesis by an uncultivated bacterial symbiont of the marine sponge Theonella swinhoei. Proc Natl Acad Sci USA 101(46):16222–16227
Piel J, Butzke D, Fusetani N, Hui D, Platzer M, Wen G, Matsunaga S (2005) Exploring the chemistry of uncultivated bacterial symbionts: antitumor polyketides of the pederin family. J Nat Prod 68(3):472–479
Rantala A, Fewer DP, Hisbergues M, Rouhiainen L, Vaitomaa J, Börner T, Sivonen K (2004) Phylogenetic evidence for the early evolution of microcystin synthesis. Proc Natl Acad Sci USA 101(2):568–573
Rateb ME, Houssen WE, Arnold M, Abdelrahman MH, Deng H, Harrison WT, Okoro CK, Asenjo JA, Andrews BA, Ferguson G, Bull AT, Goodfellow M, Ebel R, Jaspars M (2011) Chaxamycins A–D, bioactive ansamycins from a hyper-arid desert Streptomyces sp. J Nat Prod 74(6):1491–1499
Riesenfeld CS, Goodman RM, Handelsman J (2004) Uncultured soil bacteria are a reservoir of new antibiotic resistance genes. Environ Microbiol 6(9):981–989
Rowe CJ, Böhm IU, Thomas IP, Wilkinson B, Rudd BA, Foster G, Blackaby AP, Sidebottom PJ, Roddis Y, Buss AD, Staunton J, Leadlay PF (2001) Engineering a polyketide with a longer chain by insertion of an extra module into the erythromycin-producing polyketide synthase. Chem Biol 8(5):475–485
Schmeisser C, Steele H, Streit WR (2007) Metagenomics, biotechnology with non-culturable microbes. Appl Microbiol Biotechnol 75(5):955–962
Schmieder R, Edwards R (2012) Insights into antibiotic resistance through metagenomic approaches. Future Microbiol 7(1):73–89
Schwarzer D, Marahiel MA (2001) Multimodular biocatalysts for natural product assembly. Naturwissenschaften 88(3):93–101
Simon C, Daniel R (2009) Achievements and new knowledge unraveled by metagenomic approaches. Appl Microbiol Biotechnol 85(2):265–276
Sleator RD, Shortall C, Hill C (2008) Metagenomics. Lett Appl Microbiol 47(5):361–366
Slot JC, Rokas A (2011) Horizontal transfer of a large and highly toxic secondary metabolic gene cluster between fungi. Curr Biol 21(2):134–139
Stachelhaus T, Walsh CT (2000) Mutational analysis of the epimerization domain in the initiation module PheATE of gramicidin S synthetase. Biochemistry 39(19):5775–5787
Strieker M, Tanovic A, Marahiel MA (2010) Strieker M, Tanovic A, Marahiel MA (2010) non-ribosomal peptide synthetases :Structures and dynamics. Curr Opin Struct Biol 20(2): 234–40. Curr Opin Struct Biol 20(2):234–240
Tang GL, Cheng YQ, Shen B (2006) Polyketide chain skipping mechanism in the biosynthesis of the hybrid non-ribosomal peptide-polyketide antitumor antibiotic leinamycin in Streptomyces atroolivaceus S-140. J Nat Prod 69(3):387–393
Tanovic A, Samel SA, Essen LO, Marahiel MA (2008) Crystal structure of the termination module of a non-ribosomal peptide synthetase. Science 321(5889):659–663
Thomas I, Martin CJ, Wilkinson CJ, Staunton J, Leadlay PF (2002) Skipping in a hybrid polyketide synthase. Evidence for ACP-to-ACP chain transfer. Chem Biol 9(7):781–787
Trauger JW, Kohli RM, Mootz HD, Marahiel MA, Walsh CT (2000) Peptide cyclization catalysed by the thioesterase domain of tyrocidine synthetase. Nature 407(6801):215–218
Uchiyama T, Miyazaki K (2009) Functional metagenomics for enzyme discovery: challenges to efficient screening. Curr Opin Biotechnol 20(6):616–622
Udwary DW, Zeigler L, Asolkar RN, Singan V, Lapidus A, Fenical W, Jensen PR, Moore BS (2007) Genome sequencing reveals complex secondary metabolome in the marine actinomycete Salinispora tropica. Proc Natl Acad Sci USA 104(25):10376–10381
van Belkum MJ, Martin-Visscher LA, Vederas JC (2011) Structure and genetics of circular bacteriocins. Trends Microbiol 19(8):411–418. doi:10.1016/j.tim.2011.04.004
von Döhren H (2004) Biochemistry and general genetics of non-ribosomal peptide synthetases in fungi. Adv Biochem Eng Biotechnol 88:217–264
Walsh CT (2008) The chemical versatility of natural-product assembly lines. Acc Chem Res 41(1):4–10
Wang Y, Chen Y, Shen Q, Yin X (2011) Molecular cloning and identification of the laspartomycin biosynthetic gene cluster from Streptomyces viridochromogenes. Gene 483(1–2):11–21
Welker M, von Döhren H (2006) Cyanobacterial peptides—nature’s own combinatorial biosynthesis. FEMS Microbiol Rev 30(4):530–563
Wenzel SC, Kunze B, Höfle G, Silakowski B, Scharfe M, Blöcker H, Müller R (2005) Structure and biosynthesis of myxochromides S1-3 in Stigmatella aurantiaca: evidence for an iterativebacterial type I polyketide synthase and for module skipping in non-ribosomal peptide biosynthesis. ChemBioChem 6(2):375–385
Wenzel SC, Meiser P, Binz TM, Mahmud T, Müller R (2006) Non-ribosomal peptide biosynthesis: point mutations and module skipping lead to chemical diversity. Angew Chem Int Ed Engl 45(14):2296–2301
Wong FT, Khosla C (2012) Combinatorial biosynthesis of polyketides—a perspective. Curr Opin Chem Biol, in press
Yin WB, Grundmann A, Cheng J, Li SM (2009) Acetylaszonalenin biosynthesis in Neosartorya fischeri. Identification of the biosynthetic gene cluster by genomic mining and functional proof of the genes by biochemical investigation. J Biol Chem 284(1):100–109
Yu JH, Leonard TJ (1995) Sterigmatocystin biosynthesis in Aspergillus nidulans requires a novel type I polyketide synthase. J Bacteriol 177(16):4792–4800
Zerikly M, Challis GL (2009) Strategies for the discovery of new natural products by genome mining. ChemBioChem 10(4):625–633
Zhang W, Ntai I, Kelleher NL, Walsh CT (2011) tRNA-dependent peptide bond formation by the transferase PacB in biosynthesis of the pacidamycin group of pentapeptidyl nucleoside antibiotics. Proc Natl Acad Sci USA 108(30):12249–12253
Acknowledgments
This work was supported by the Postgraduate Programs: “Molecular Biology and Genetics Applications-Diagnostic Markers” and “Biotechnology-Quality assessment in Nutrition and the Environment”, Department of Biochemistry and Biotechnology, University of Thessaly.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nikolouli, K., Mossialos, D. Bioactive compounds synthesized by non-ribosomal peptide synthetases and type-I polyketide synthases discovered through genome-mining and metagenomics. Biotechnol Lett 34, 1393–1403 (2012). https://doi.org/10.1007/s10529-012-0919-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-012-0919-2