Abstract
The tumor suppressor p53 is at the hub of cellular signaling networks that are activated by stress signals including DNA damage. In the present study, we showed that programmed cell death 5 (PDCD5) bound to p53 by glutathione S-transferase (GST)-pulldown, co-immunoprecipitation and co-localization assays. PDCD5 enhanced the stability of p53 by antagonizing Mdm2-induced p53 ubiquitination, nuclear export and proteasomal degradation. We also found that PDCD5 could dissociate the interaction between p53 and Mdm2 and interact with Mdm2 directly to promote its degradation. In cells with or without induction of DNA damage, knockdown of PDCD5 by RNA interference decreased the p53 phosphorylation at Ser9, 20 and 392 residues, as well as the expression of p21 protein. Additionally, chromatin immunoprecipitation assays showed an up-regulated association of PDCD5 at the p53BS2 site of the p21 promoter during DNA damage. Cell cycle analysis also indicated that PDCD5 was required in G1 phase cell arrest during DNA damage. In summary, PDCD5 may contribute to maintain a basal pool of p53 proteins in unstressed conditions, but upon DNA damage it functions as a co-activator of p53 to regulate transcription and cell cycle arrest.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The tumor suppressor p53 is a powerful transcriptional factor and plays a central role in the regulation of cell cycle, DNA repair, apoptosis, senescence and angiogenesis. As a sequence-specific transcription factor, p53 mainly functions by inducing transcription of many different downstream genes, including genes involved in cell cycle arrest, such as p21/WAF1/CIP1 and GADD45, as well as those inducing apoptosis, such as PUMA, Bax and PIG3 [1–3]. However, p53 can also control apoptosis through transcription-independent mechanisms [4].
Under normal conditions, p53 is maintained at a low level by interacting with E3 ubiquitin ligases such as Mdm2 [5], Pirh2 [6], COP1 [7] and ARF-BP1 [8], which mediate p53 degradation by the ubiquitin–proteasome pathway. However, under stressed conditions, such as DNA damage, p53 is stabilized and activated to function primarily as a transcription factor and regulating expression of downstream target genes. This leads to different cellular outcomes such as cell cycle arrest and apoptosis; the former facilitates DNA repair and promotes cell survival, whereas the latter provides an efficient way to remove irreparably damaged cells [9]. These processes are thought to be tightly controlled by its binding partners and post-translational modifications, such as ubiquitination, phosphorylation and acetylation [10]. p53 can be phosphorylated at multiple sites by several different protein kinases in vivo and in vitro [11, 12], such as Ser15 and Ser37 by ATM and ATR [13–15], Ser20 by Chk2 and Chk1 [16, 17], and Ser392 by CKII [18], which modulate its stability and sequence-specific DNA binding activity.
Programmed Cell Death 5 (PDCD5), formerly designated TFAR19 (TF-1 cell apoptosis-related gene 19), was first identified as a gene up-regulated in TF-1 cells undergoing apoptosis [19]. PDCD5 can promote apoptosis in different cell types in response to various stimuli and also enhance TAJ/TROY-induced paraptosis-like cell death [20]. During apoptosis, PDCD5 is rapidly upregulated and translocates from the cytoplasm to nucleus [21]. Conversely, decreased expression of PDCD5 has been detected in various human tumors, such as lung cancer [22], gastric cancer [23], chronic myelogenous leukemia [24], prostate cancer [25], epithelial ovarian carcinomas [26], astrocytic gliomas [27] and chondrosarcoma [28]. At the same time, restoration of PDCD5 with recombinant protein or an adenovirus expression vector can significantly sensitize different cancers to chemotherapies [29–32]. In addition, single-nucleotide polymorphisms in the PDCD5 regulatory region are associated with chronic myelogenous leukemia and lung cancer [22, 33]. Thus, PDCD5 likely plays a critical role in multiple tissues during tumorigenesis. However, the molecular mechanisms underlying PDCD5 functions during cell growth, proliferation and apoptosis remain largely unclear.
We previously demonstrated that PDCD5 interacts with Tip60, enhances the histone acetylation and p53 K120 acetylation, and promotes the expression of Bax, consequently accelerating apoptosis [34]. Here, we offer novel evidence indicating that PDCD5 is a p53 regulator during gene expression and cell cycle. PDCD5 was shown to interact with p53 in vitro and in vivo and increase the stability of p53 by inhibiting its ubiquitination and nuclear export mediated by Mdm2. In DNA damage conditions, PDCD5 co-localized with p53 in the nucleus and regulated gene expression of its downstream genes such as p21 by affecting the phosphorylation and transcriptional activity of p53, thereby affecting the cell cycle progression during the G1/S phase transition.
Materials and methods
Plasmids, siRNA and antibodies
The pcDNA3-PDCD5 and pcDNA3-PDCD5-myc plasmids have been described previously [20]. The pcDNA3-Flag-p53 vector was a gift from Steven B. McMahon (Thomas Jefferson University, Philadelphia, PA, USA). The Mdm2 and His-ubi vectors were kindly provided by Sonia Lain (University of Dundee, Scotland, UK). The luciferase reporter plasmid containing the p21 promoter (p21-Luc) from Koichi Hagiwara (Saitama Medical University, Saitama, Japan). All siRNA including PDCD5 and the control siRNA were synthesized by GeneChem Corporation (Shanghai, China), and their sequences have been reported previously [35]. All shRNA share the same target sequence with siRNA and were constructed by ORIGEN Corporation (Rockville, USA). The anti-Flag, anti-Myc, and anti-actin antibodies were purchased from Sigma Aldrich (St. Louis, MO). The anti-p21, anti-p53 (monoclonal and polyclonal) and anti-Mdm2 were from Santa Cruz Biotechnology (Santa Cruz, CA). The phospho-p53 antibodies were from Cell Signaling Technology (Beverly, MA). The mouse anti-PDCD5 monoclonal antibody (3A3), rabbit anti-PDCD5 polyclonal antibody and FITC-labeled anti-PDCD5 antibody have been described previously [21]. IRDye 800-conjugated secondary antibodies against mouse, rabbit, and goat IgG were purchased from Li-Cor Bioscience (Lincoln, NE). TRITC-labeled rabbit against goat IgG was from Zhongshan Corporation (Beijing, China).
Cell culture, transfection and treatment
U2OS cell line was purchased from ATCC and HeLa cell line was from Cell Bank of Chinese Academy of Science (CAS). These cell lines were cultured in Dulbecco’s modified Eagle’s medium, supplemented with 10 % fetal bovine serum. HeLa cells were transfected by electroporation, as described previously [35]. U2OS cells were transfected using Lipofectamine 2000 (Invitrogen, Carlsbad, CA), according to the manufacturer’s protocol.
Proteasome inhibition was achieved by treating cells with 100 μM of ALLN ((N-acetyl-Leu-Leu-norleucine, Sigma) or 10 μM of MG132 (Sigma). Protein translation inhibition was achieved by treating cells with 100 μg/ml of CHX (Cycloheximide, Sigma) for different times. DNA damage was induced with Actinomycin D (Act. D, 100 nM) or Doxorubicin (Doxo, 0.4 μM). UV irradiation (20 J/m2) was performed using a CX-2000 UV cross-linker (UVP, Inc., Upland, CA).
Immunoprecipitation and Western blot
Cells were collected and disrupted in lysis buffer (300 mM NaCl, 50 mM Tris pH 8.0, 0.4 % NP-40, 10 mM MgCl2, and 2.5 mM CaCl2) supplemented with protease inhibitors (Complete mini EDTA-free; Roche Diagnostics, Mannheim, Germany). After centrifugation, the supernatant was measured using the BCA protein assay reagent (Pierce, Rockford, IL). Subsequently, 1 mg of total cell extracts was diluted in 1 ml of dilution buffer (50 mM Tris, pH 8.0, 0.4 % NP-40). After a pre-clearing step with protein G Sepharose beads, cell lysates were incubated with appropriate antibodies overnight at 4 °C and then with 50 μl of a 50 % slurry of protein G Sepharose for 2 h. Immunoprecipitates were then washed and analyzed by Western blot, as described previously [34]. The protein bands were visualized using an IRDye 800CW-conjugated secondary antibody. The infrared fluorescence image was obtained using an Odyssey infrared imaging system (Li-Cor Bioscience).
GST pull-down assay
GST fusion proteins or GST were expressed in Escherichia coli strain BL21 and purified, then incubated with whole cell lysates extracted from transfected HeLa cells overnight at 4 °C. After five washes, beads were resuspended in 2× SDS loading buffer and analyzed by SDS-PAGE followed by Western blot [34].
Immunofluorescence analysis
U2OS cells were plated on glass coverslips and then treated with or without Doxo for 6 h. Cells were then fixed, permeabilized and blocked as described previously [29]. Then cells were incubated with goat anti-p53 antibody, followed by incubating with TRITC-conjugated goat anti-rabbit IgG or FITC-labeled mouse anti-PDCD5. 4′,6-diamidino-2-phenylindole (DAPI) was used to counterstain the cell nuclei. Coverslips were observed under a Leica SP2 confocal system (Germany).
Quantitative real-time PCR analysis
U2OS cells were transfected with either control or PDCD5-specific siRNA with Lipofectamine 2000. Thirty-six hours later, total RNA were extracted from transfected U2OS cells using TRIzol reagent (Invitrogen), according to the manufacturer’s instructions. First-strand cDNA was generated using the total RNA in a standard reverse transcriptase reaction using a poly(dT) oligonucleotide as a primer and SuperScript II reverse transcriptase (Invitrogen). The cDNA were then analyzed by quantitative real-time PCR. The PDCD5 primers used in the assay were previously reported [34].
Dual-luciferase reporter assay
The luciferase reporter plasmid containing a consensus p21 promoter (p21-Luc) and pRL-TK (Promega) were co-transfected with control or PDCD5-specific shRNA. After 36 h, cells were treated with or without UV or Actinomycin D for 6 h, and then luciferase activity was measured and normalized using the Luciferase Assay System (Vigorous, USA), according to the manufacturer’s protocol.
ChIP analysis
ChIP experiments were carried out essentially as previously described with minor alterations [36]. The chromatin was fragmented to a range of 0.5–1 kb after sonication with a Bioruptor 251. The ChIP DNA samples were purified using the QIAGEN PCR product purification kit, and real-time PCR was used to amplify p53BS2 and Exon 1 within the p21 promoter.
Cell cycle analysis
Treated U2OS cells (5 × 105) were fixed in 70 % (v/v) ethanol overnight at 4 °C. After washing with PBS, cells were incubated with RNase (0.5 mg/mL) at 37 °C for 30 min (Sigma). Finally, after cells were stained with PI (50 μg/mL), DNA content was measured using the CellQuest Pro program on a FACSCalibur flow cytometer, and data were analyzed using ModFit software.
Statistical analysis
Data are presented as the mean ± SD. Differences between groups were analyzed using the Student’s t test for continuous variables. Statistical significance in this study was set at P < 0.05. All reported P values are two-sided. All analyses were performed with GraphPad Prism 5.
Results
PDCD5 interacts and co-localizes with p53
To determine whether PDCD5 interacts with p53 to regulate its function, we performed a series of biochemical experiments. In a GST pull-down assay, GST-PDCD5 but not GST bound specifically to p53 overexpressed in Hela cells (Fig. 1a, lane 3). To confirm the association of PDCD5 and p53 in mammalian cells, lysates of HeLa cells transfected with Myc-PDCD5 and/or Flag-p53 plasmids were immunoprecipitated with an anti-Myc antibody or anti-Flag antibody. Myc-PDCD5 was co-immunoprecipitated (co-IP) by the anti-Flag antibody from the cells expressing Flag-p53, but not from cells without Flag-p53 expression (Fig. 1b, lane 3). In reciprocal co-IP assays, Flag-p53 was detected in the Myc-PDCD5 immunoprecipitates (Fig. 1c, lane 3), demonstrating that the two proteins interacted in a complex in HeLa cells.
PDCD5 interacts with p53. a Pull-down assay with recombinant GST-PDCD5 protein and p53-transfected HeLa cell lysates. GST protein was used as negative control. b Exogenous co-IP of PDCD5 and p53 in HeLa cells transiently transfected with Flag-p53 and/or Myc-PDCD5 as indicated. c Cells were treated as in (b), except that cell lysates were immunoprecipitated with an anti-Myc antibody. d Endogenous co-IP of PDCD5 and p53 in U2OS cells treated with or without UV irradiation (20 J/m2) for 6 h. Non-specific rabbit IgG was used as negative control. e PDCD5 co-localization with p53 in U2OS Cells treated with or without Doxo (0.4 μM, 6 h). p53 was detected with a rabbit anti-p53 antibody, followed by an anti-rabbit TRITC-conjugated secondary antibody. PDCD5 was detected with FITC-anti-PDCD5 and cell nuclei were stained with DAPI and observed using a Leica SP2 confocal microscope. Scale bar 20 μm
We further examined whether the interaction could be detected between endogenous PDCD5 and p53 by performing co-IP using U2OS cell lysates. PDCD5 was co-immunoprecipitated by the p53 antibody (Fig. 1d, lane 2) but not by the normal rabbit IgG (Fig. 1d, lane 1). It is known that p53 and PDCD5 can be induced and up-regulated in DNA damage conditions. Indeed, we found that the amount of p53 bound to PDCD5 increased after UV (20 J/m2) irradiation (Fig. 1d, lane 3).
To further confirm the PDCD5-p53 association, co-localization analysis was performed on U2OS cells treated with or without chemotherapeutic reagent Doxo and then immunostained with purified mouse PDCD5 antibody and rabbit p53 antibody. Cell nuclear morphology was observed by DAPI staining. In DMSO control cells, p53 localized predominantly to the nucleus, although a minor fraction was also observed in the cytoplasm; meanwhile, PDCD5 showed a relatively diffuse distribution in both the cytoplasm and nucleus, and the location of the two proteins overlapped in the nucleus (Fig. 1e upper panel). In Doxo-treated cells, the amount of PDCD5 and p53 were up-regulated as shown by the increased florescence intensity. PDCD5 translocated and accumulated in the nucleus and co-localized with p53 (Fig. 1e, lower panel). Collectively, these results demonstrate that PDCD5 and p53 exist in the same protein complex, and their association is up-regulated during DNA damaging conditions.
PDCD5 enhances stability of the p53 protein
The function of p53 is tightly controlled by its stability. Therefore, we investigated the effect of PDCD5 on the stability of the p53 protein. U2OS cells were transfected for 36 h with either control or PDCD5-specific siRNA that we had previously characterized [35]. As shown by Western blot in Fig. 2a (lane 1 vs. lane 2), the levels of p53 protein decreased after knockdown of PDCD5, whereas the actin levels were unchanged. Since it is known that p53 is degraded through the proteasome, the effect of PDCD5 on proteasome-mediated proteolysis of p53 was tested by incubating the control or PDCD5-specific siRNA-treated U2OS cells with the proteasome inhibitor ALLN (100 μM) for the last 6 h before cell harvest. The reduction of p53 levels by the PDCD5 siRNA was significantly blocked by ALLN (Fig. 2a, lanes 3 vs. lane 1), suggesting that endogenous PDCD5 may be involved in the modulation of p53 level by protecting it from proteasomal degradation. At the same time, knockdown of PDCD5 failed to influence the p53 mRNA levels (Fig. 2b), suggesting that PDCD5 does not regulate p53 at the transcriptional level.
PDCD5 enhances stability of the p53 protein. a U2OS cells were transfected with control or PDCD5-specific siRNA for 36 h. ALLN (100 μM) was added where indicated 6 h before cell harvest. The total cell lysates were analyzed for the presence of p53, PDCD5 and actin by Western blot. b U2OS cells were treated as in (a), and the mRNA levels of both PDCD5 and p53 were analyzed by quantitative real-time PCR. Data are presented as mean ± SD. c U2OS cells were transiently transfected as in (a), then cells were treated with 100 μg/ml of CHX for different lengths of time as indicated. p53 was detected in total cell lysates by Western blot. d Relative p53 protein levels were quantified using a Li-Cor Odyssey scanner (*p < 0.05, **p < 0.001)
To test the idea that PDCD5 regulates proteasomal degradation of p53, we next determined the influence of PDCD5 on the half-life of p53. U2OS cells were transiently transfected with either control or PDCD5-specific siRNA for 36 h. Cells were then treated with the translation inhibitor CHX (100 μg/ml) for different periods up to 60 min before harvesting. As shown in Fig. 2c, d knockdown of endogenous PDCD5 was associated with an acceleration in p53 decay compared with control siRNA (*p < 0.05, **p < 0.001, Fig. 2d), suggesting that the decrease of p53 protein levels after PDCD5 knockdown was due to destabilization of the p53 protein.
PDCD5 impairs the Mdm2-mediated degradation, ubiquitination and nuclear export of p53
The degradation of p53 is known to be regulated by the ubiquitin–proteasome system. In this process, Mdm2 functions as an ubiquitin ligase that promotes p53 ubiquitination and degradation. Therefore, we next tested the influence of PDCD5 on the Mdm2-dependent degradation of p53. U2OS cells were transfected with plasmids to express Mdm2 with or without PDCD5 for 24 h. As shown in Fig. 3a, the p53 level decreased in cells overexpressing Mdm2 (lane 2 vs. lane 1), while p53 accumulation was restored in cells overexpressing both with Mdm2 and PDCD5 (lane 3 vs. lane 2).
PDCD5 blocks the Mdm2-mediated degradation, ubiquitination and nuclear export of p53. a U2OS cells were transfected with Mdm2 and/or PDCD5 as indicated for 24 h. Mdm2, p53, PDCD5 and actin were detected in total cell lysates. b U2OS cells were transiently transfected with different combinations of mammalian expression vectors for His-ubiquitin, Mdm2 and PDCD5. After 24 h, cells were treated with 10 μM MG132 for 4 h. The cells lysates were then immunoprecipitated with an anti-p53 antibody and probed with an anti-ubiquitin antibody. c U2OS cells were transfected as indicated for 24 h. The location of p53 and PDCD5 were detected by immunofluorescence analysis. Cell nuclei were stained with DAPI. d Statistical analysis of (c). Two hundred cells were chosen randomly from each group. The intensities of cytoplasmic and nuclear p53 were determined and the ratio of cytoplasmic to nuclear p53 calculated. One representative experiment of three is shown. Data are presented as mean ± SD *P < 0.05
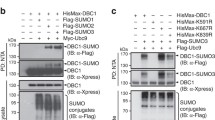
To investigate whether PDCD5 affects the ubiquitination of p53, U2OS cells were transiently transfected with different combinations of mammalian expression vectors for His-ubiquitin, Mdm2 and PDCD5 for 24 h, followed by treatment with the proteasome inhibitor MG132 (10 μM) for 4 h. The cell lysates were then immunoprecipitated using an anti-p53 antibody, and the samples were probed in a Western blot with an anti-ubiquitin antibody to detect ubiquitinated p53. As shown in Fig. 3b, the level of ubiquitinated p53 was increased in the cells overexpressing His-ubiquitin and Mdm2 (lane 2), whereas PDCD5 reduced the ubiquitination of p53 (lane 3 vs. lane 2). At the same time, we observed that overexpression of PDCD5 could decrease the accumulation of Mdm2 protein (Fig 3b, lane 3).This result was consisted with Fig. 4c (see below).
PDCD5 dissociates the p53-Mdm2 interaction and promotes Mdm2 degradation. a GST pull-down assay was performed with p53-GST and the whole lysates of cells overexpressing Mdm2 with or without recombinant PDCD5. b GST pull-down assay was performed with GST-PDCD5 and the whole lysates of cells overexpressing Mdm2. c U2OS cells were transfected to overexpress Mdm2 and increasing amounts (0, 2, 4, 6 μg) of PDCD5 for 24 h. Mdm2, PDCD5 and actin were detected in the cell lysates. d U2OS cells were transfected as indicated for 24 h. After another 4 h of treatment with 10 μM MG132, His-ubiquitin proteins were purified with Ni beads and probed with an anti-Mdm2 antibody. Mdm2, PDCD5 and actin were detected in the cell lysates
We then examined the possibility that PDCD5 blocks the nuclear export of p53 mediated by Mdm2. U2OS cells were transfected with expression vectors for Mdm2, and/or PDCD5 for 24 h, and then the location of p53 was observed by confocal microscopy. In empty-vector transfected cells, p53 was mainly located in the nucleus (Fig. 3c, upper panel). Partial p53 translocation from nucleus to cytoplasm was observed in U2OS cells overexpressing Mdm2 (Fig. 3c, middle panel). However, in cells overexpressing both Mdm2 and PDCD5, the Mdm2-mediated exportation of p53 was partially blocked (Fig. 3c, lower panel). The calculated ratios of cytoplasmic to nuclear p53 indicated statistically differences between the Vector and Mdm2-overexpressing U2OS cells, as well as between those overexpressing Mdm2 alone and the combination of Mdm2 and PDCD5 (Fig. 3d, *P < 0.05).
PDCD5 disrupts the p53-Mdm2 interaction and promotes Mdm2 degradation
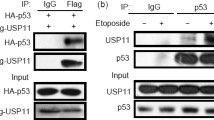
Disruption of the p53-Mdm2 interaction by multiple regulators is a pivotal event for p53 stabilization and subsequent biological responses. To further investigate the mechanisms underlying PDCD5 stabilization of p53, we examined whether PDCD5 could affect the interaction between p53 and Mdm2. GST-p53 or GST was incubated with the whole lysates of Hela cells overexpressing Mdm2 with or without recombinant human PDCD5. As shown in Fig. 4a, p53 could strongly bind to Mdm2 (lane 2), but this interaction between p53 and Mdm2 decreased significantly in the presence of recombinant PDCD5 (lane 3), implying that PDCD5 may disrupt the p53-Mdm2 interaction. Simultaneously, we found that PDCD5 could be pulled down with the GST-p53 fusion protein (Fig. 4a, lane 3), demonstrating the direct interaction between p53 and PDCD5 in vitro.
The relationship between PDCD5 and Mdm2 was also investigated by the GST pull-down assay. As shown in Fig. 4b, GST-PDCD5, but not GST interacted with Mdm2 from overexpressing Hela cells. Interestingly, we found that PDCD5 could decrease the protein level of Mdm2 in a dose-dependent manner (Fig. 4c). Furthermore, knockdown of endogenous PDCD5 could increase the accumulation of Mdm2 and decrease the ubiquitination level of Mdm2 (Fig. 4d, lane 2).
Knockdown of PDCD5 decreases the phosphorylation of p53
We next investigated if PDCD5 could influence the post-translational modifications of p53 required for its function. U2OS cells were transfected for 36 h with either control or PDCD5-specific shRNA and then treated with or without UV or Act. D for 12 h. Equal quantity of DMSO was as a negative control. As shown in Fig. 5a, b knockdown of endogenous PDCD5 caused the reduction of total p53 compared with control shRNA (lane1), while p53 phosphorylation was decreased at Ser9, Ser20 and Ser392 in UV- and Actinomycin D- treated cells (lane 3 and lane 5).
Knockdown of endogenous PDCD5 decreases the phosphorylation. a U2OS cells were transfected for 36 h with either control or PDCD5-specific shRNA. After treatment of DMSO, 20 J/m2 UV or 100 nM Actinomycin D for 12 h, cell lysates were analyzed for the phosphorylated and total p53, PDCD5 and actin by Western blot. b Statistical analysis of (a), relative levels of phosphorylated and total p53, PDCD5 protein were quantified using a Li-Cor Odyssey scanner
Knockdown of PDCD5 attenuates expression and transcription of p21
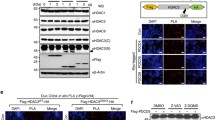
We have previously shown that PDCD5 can promote the p53-reponsive gene Bax [34], which plays a critical role in this apoptotic pathway. Here we examined whether PDCD5 could affect the induction of another p53 target gene, p21 involved in cell cycle arrest. U2OS cells were transfected with control or PDCD5-specific siRNA for 36 h and then treated with or without Actinomycin D for 12 h. As shown in Fig. 6a, the cells were treated with Actinomycin D or not, the knockdown of PDCD5 led to the decrease of p21 protein levels (lane 1 vs. 2 and lane 3 vs. 4). To examine the transcriptional activity of the p21 promoter after PDCD5 knockdown, U2OS cells were transfected with control or PDCD5-specific shRNA together with a p21 promoter-driven luciferase reporter plasmid and the pRL-TK internal control. After 36 h, cells were treated with or without UV or Actinomycin D for 6 h before measuring p21-luc luciferase activity. In both DMSO control and stressed cells, the knockdown of PDCD5 significantly impaired the transcriptional activity of p53 on the p21 promoter compared with control shRNA-transfected cells (Fig. 6b, *P < 0.05).
PDCD5 is involved in expression of p21, a key mediator of p53-induced cell cycle arrest. a U2OS cells were transfected for 24 h with either control or PDCD5-specific siRNA. After treatment of 100 nM Actinomycin D for 12 h, cell lysates were analyzed for p21, PDCD5 and actin by Western blot. b U2OS cells were transfected with control or PDCD5-specific shRNA together with the p21 promoter-driven luciferase reporter plasmid (p21-luc) and pRL-TK. After 36 h, cells were treated with or without 20 J/m2 UV or 100 nM Actinomycin D for 6 h, and then p21-luc luciferase activity was measured and normalized. *P < 0.05. c Illustration of the p21 gene promoter, including the two p53 binding sites, p53BS1 and p53BS2. d U2OS cells were treated with or without Doxo (0.4 μM) for 6 h, crosslinked and subjected to ChIP with an anti-PDCD5 antibody and irrelative rabbit IgG. The precipitated DNA was analyzed by real-time PCR
The changes in p21 expression after PDCD5 knockdown prompted us to investigate whether PDCD5 interacts with the p21 promoter. ChIP assay was performed in U2OS cells before and after treatment with Doxo (0.4 μM), followed by quantitative PCR analysis of the ChIP DNA samples to analyze protein-promoter interactions. The p21 promoter is known to have two p53 binding sites, p53BS1 and p53BS2 (Fig. 6c). Our ChIP results showed that PDCD5 was preferentially associated at the p53BS2 site of the p21 promoter after Doxo treatment, but not the exon 1 region of p21 (Fig. 6d), suggesting that PDCD5 may associate with the p21 promoter to facilitate transcriptional activation after DNA damage.
PDCD5 is required in G1 phase cell arrest during DNA damage
The involvement of PDCD5 in p21 regulation prompted us to analyze whether the knockdown of PDCD5 impacts the cell cycle progression, U2OS cells were transfected for 36 h with either control or PDCD5-specific siRNA and then treated with or without Actinomycin D for 12 h. Propidium iodide (PI) staining and flow cytometry were used to analyze the DNA content and the cell cycle. In DMSO control, PDCD5 did not affect the cell cycle progression (Fig. 7, upper panel). After treatment with Actinomycin D, however, the G1 phase cells were decreased by ~20 %, with a concomitant increase in the G2/M phase cells after knockdown of PDCD5 (Fig. 7, lower panel), indicating that PDCD5 was required for G1 phase cell arrest during DNA damage.
PDCD5 is required in G1 phase cell arrest during DNA damage. U2OS cells were transfected for 36 h with either control or PDCD5-specific siRNA and then were treated 100 nM Actinomycin D for 12 h. DMSO was used as negative control. PI staining and flow cytometry were used to show the DNA content and cell cycle. One representative experiment of three is shown
Discussion
In this study, we offer novel evidence for PDCD5 as a positive regulator and co-activator of p53 during gene expression and cell cycle arrest. PDCD5 was shown to interact with p53 in vitro and in vivo and increase the stability of p53 by inhibiting its ubiquitination, nuclear export and degradation mediated by Mdm2. Furthermore, in stressed conditions, PDCD5 was demonstrated to translocate to the nucleus, affect the phosphorylation of p53, regulate gene expression at the p21 promoter and facilitate p53 induction of cell cycle arrest.
In cells without DNA damage, p53 is a short-lived protein, and its stability is regulated by ubiquitination and subsequent protein degradation [5]. Many p53-binding molecules have been proven to stabilize p53 through blocking the Mdm2-p53 feedback loop, such as p14ARF, SOX4 and the ribosomal proteins S7, L5, L23 and L11 [37–40]. Our results indicate that PDCD5 also stabilized p53 and regulated its functions during apoptosis and cell cycle. As PDCD5 could interact with p53 and Mdm2 to form a complex, it might dissociate the interaction of p53 and Mdm2 by competitive binding and impairing the Mdm2-mediated ubiquitination and nuclear export of p53. Recently, our collaborator’s studies obtained from Nuclear Magnetic Resonance showed that the PDCD5 could bind with the N-terminal domain of p53(p5315 − 61) [41], which overlapped with the binding site of p53 (p5315 − 29) and Mdm2 [42]. So it is possible that PDCD5 could compete with MDM2 to bind with p53. Additionally, PDCD5 promoted the ubiquitination and degradation of Mdm2 through an unknown mechanism which will require more study to clarify.
Decreased p53 levels have been observed in many different cancers, and at least 5–10 % of all human tumors possess inappropriate overexpression of Mdm2. Interestingly, PDCD5 is also significantly decreased in many cancer tissues at both the mRNA and protein levels. Therefore, we propose that a decreased PDCD5 level may influence the normal autoregulatory loop between Mdm2 and p53, thereby promoting tumor progression. Because the interaction between Mdm2 and p53 is a primary mechanism for inhibition of the p53 function in cancers retaining wild-type p53, targeting this interaction by different approaches to reactivate p53 has emerged as a promising new cancer therapeutic strategy. Based on our studies, PDCD5 is proposed to disassociate the intracellular Mdm2-p53 interaction and induce the accumulation of p53 and the activation of the p53 pathway in tumor cells. Hence, PDCD5 may represent a potential cancer therapeutic agent.
The functions of p53 are also controlled by post-translational modifications, such as phosphorylation, acetylation, ubiquitination and sumoylation [43]. Many proteins such as Mdm2, ATM/ATR, SUSP4 and Tip60 form a complicated network to regulate cellular activities through changing the post-translational modifications of p53. In our previous paper, we showed that PDCD5 together with Tip60 could increase the acetylation of p53 at K120 [34]. In particular, the effects of phosphorylation of p53 on its stability and transactivation have been investigated intensively. The phosphorylation of p53 at N-terminal residues Ser15 and Ser20 is crucial for the interaction between p53 and Mdm2 [17], whereas the phosphorylation at Ser 9, 46 and 392 is related to transcriptional activity. In this paper, we showed PDCD5 could affect the phosphorylation of p53 at multiple sites including Ser9, 20 and 392 after DNA damage. Phosphorylation of p53 was shown to weaken its interaction with Mdm2, thereby stabilizing p53 and promoting the activation of p53. Our results suggest that PDCD5 could be involved in both stabilization and transcriptional activity of p53, consistent with other investigations. The influence of PDCD5 on p53 phosphorylation also partly explains how it could stabilize p53 aside from dissociating p53 and Mdm2. The question of how PDCD5 might affect the phosphorylation of p53 remains to be answered in further investigations.
PDCD5 knockdown by RNA interference failed to block cell cycle arrest at the G1 phase and decrease the accumulation of p21/CIP1/WAF1 after DNA damage, suggesting an important role of PDCD5 on the regulation for cell cycle. The association of PDCD5 with p53BS2 in the p21 promoter suggests that PDCD5 likely plays a direct role in promoting the expression of p21 after DNA damage. The mechanisms governing PDCD5 association with the p21 promoter after DNA damage is yet to be identified.
Taken together, our findings indicate that PDCD5 can affect p53 at least on three levels, as illustrated in Fig. 8. First, in unstressed cells, PDCD5 can dissociate p53 and Mdm2 and maintain the basal level of p53. Second, upon DNA damage, PDCD5, together with unknown kinases or phosphatases, enhances the phosphorylation of p53 and is recruited to the p21 promoter, finally leading to p21 transactivation and G1 phase arrest and allowing cells to initiate productive DNA repair processes. Third, if the above-mentioned processes fail, PDCD5 together with Tip60 can promote p53K120 acetylation and transactivation of Bax expression in order to initiate an irreversible apoptotic response.
References
Hollstein M, Sidransky D, Vogelstein B, Harris CC (1991) p53 mutations in human cancers. Science 253:49–53
Laptenko O, Prives C (2006) Transcriptional regulation by p53: one protein, many possibilities. Cell Death Differ 13:951–961
Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100:57–70
Mihara M, Erster S, Zaika A et al (2003) p53 has a direct apoptogenic role at the mitochondria. Mol Cell 11:577–590
Kubbutat MH, Jones SN, Vousden KH (1997) Regulation of p53 stability by Mdm2. Nature 387:299–303
Leng RP, Lin Y, Ma W et al (2003) Pirh2, a p53-induced ubiquitin-protein ligase, promotes p53 degradation. Cell 112:779–791
Dornan D, Wertz I, Shimizu H et al (2004) The ubiquitin ligase COP1 is a critical negative regulator of p53. Nature 429:86–92
Chen D, Kon N, Li M et al (2005) ARF-BP1/Mule is a critical mediator of the ARF tumor suppressor. Cell 121:1071–1083
Brady CA, Jiang D, Mello SS et al (2011) Distinct p53 transcriptional programs dictate acute DNA-damage responses and tumor suppression. Cell 145:571–583
Bode AM, Dong Z (2004) Post-translational modification of p53 in tumorigenesis. Nat Rev Cancer 4:793–805
Higashimoto Y, Saito S, Tong XH et al (2000) Human p53 is phosphorylated on serines 6 and 9 in response to DNA damage-inducing agents. J Biol Chem 275:23199–23203
Sakaguchi K, Saito S, Higashimoto Y et al (2000) Damage-mediated phosphorylation of human p53 threonine 18 through a cascade mediated by a casein 1-like kinase. Effect on Mdm2 binding. J Biol Chem 275:9278–9283
Abraham RT (2001) Cell cycle checkpoint signaling through the ATM and ATR kinases. Genes Dev 15:2177–2196
Banin S, Moyal L, Shieh S et al (1998) Enhanced phosphorylation of p53 by ATM in response to DNA damage. Science 281:1674–1677
Tibbetts RS, Brumbaugh KM, Williams JM et al (1999) A role for ATR in the DNA damage-induced phosphorylation of p53. Genes Dev 13:152–157
Hirao A, Kong YY, Matsuoka S et al (2000) DNA damage-induced activation of p53 by the checkpoint kinase Chk2. Science 287:1824–1827
Shieh SY, Ikeda M, Taya Y, Prives C (1997) DNA damage-induced phosphorylation of p53 alleviates inhibition by MDM2. Cell 91:325–334
Lu H, Fisher RP, Bailey P, Levine AJ (1997) The CDK7-cycH-p36 complex of transcription factor IIH phosphorylates p53, enhancing its sequence-specific DNA binding activity in vitro. Mol Cell Biol 17:5923–5934
Liu H, Wang Y, Zhang Y et al (1999) TFAR19, a novel apoptosis-related gene cloned from human leukemia cell line TF-1, could enhance apoptosis of some tumor cells induced by growth factor withdrawal. Biochem Biophys Res Commun 254:203–210
Wang Y, Li X, Wang L et al (2004) An alternative form of paraptosis-like cell death, triggered by TAJ/TROY and enhanced by PDCD5 overexpression. J Cell Sci 117:1525–1532
Chen Y, Sun R, Han W et al (2001) Nuclear translocation of PDCD5 (TFAR19): an early signal for apoptosis? FEBS Lett 509:191–196
Spinola M, Meyer P, Kammerer S et al (2006) Association of the PDCD5 locus with lung cancer risk and prognosis in smokers. J Clin Oncol 24:1672–1678
Yang Y, Zhao M, Li W et al (2006) Expression of programmed cell death 5 gene involves in regulation of apoptosis in gastric tumor cells. Apoptosis 11:993–1001
Ruan G, Qin Y, Chen S et al (2006) Abnormal expression of the programmed cell death 5 gene in acute and chronic myeloid leukemia. Leuk Res 30:1159–1165
Du Y, Xiong L, Lou Y, Tan W, Zheng S (2009) Reduced expression of programmed cell death 5 protein in tissue of human prostate cancer. Chin Med Sci J 24:241–245
Zhang X, Wang X, Song X et al (2011) Clinical and prognostic significance of lost or decreased PDCD5 expression in human epithelial ovarian carcinomas. Oncol Rep 25:353–358
Li H, Wang Q, Gao F et al (2008) Reduced expression of PDCD5 is associated with high-grade astrocytic gliomas. Oncol Rep 20:573–579
Chen C, Zhou H, Xu L et al (2010) Prognostic significance of downregulated expression of programmed cell death 5 in chondrosarcoma. J Surg Oncol 102:838–843
Ruan G, Zhao H, Chang Y et al (2008) Adenovirus-mediated PDCD5 gene transfer sensitizes K562 cells to apoptosis induced by idarubicin in vitro and in vivo. Apoptosis 13:641–648
Chen C, Zhou H, Xu L et al (2010) Recombinant human PDCD5 sensitizes chondrosarcomas to cisplatin chemotherapy in vitro and in vivo. Apoptosis 15:805–813
Shi L, Song Q, Zhang Y et al (2010) Potent antitumor activities of recombinant human PDCD5 protein in combination with chemotherapy drugs in K562 cells. Biochem Biophys Res Commun 396:224–230
Wang Y, Song Q, Zhang Y et al (2009) Recombinant human PDCD5 protein enhances chemosensitivities of hematologic malignancies. Chin Sci Bull 54:3981–3989
Ma X, Ruan G, Wang Y et al (2005) Two single-nucleotide polymorphisms with linkage disequilibrium in the human programmed cell death 5 gene 5′ regulatory region affect promoter activity and the susceptibility of chronic myelogenous leukemia in Chinese population. Clin Cancer Res 11:8592–8599
Xu L, Chen Y, Song Q et al (2009) PDCD5 interacts with Tip60 and functions as a cooperator in acetyltransferase activity and DNA damage-induced apoptosis. Neoplasia 11:345–354
Chen L, Wang Y, Ma D, Chen Y (2006) Short interfering RNA against the PDCD5 attenuates cell apoptosis and caspase-3 activity induced by Bax overexpression. Apoptosis 11:101–111
Li P, Wang D, Yao H et al (2010) Coordination of PAD4 and HDAC2 in the regulation of p53-target gene expression. Oncogene 29:3153–3162
Chen D, Zhang Z, Li M et al (2007) Ribosomal protein S7 as a novel modulator of p53-MDM2 interaction: binding to MDM2, stabilization of p53 protein, and activation of p53 function. Oncogene 26:5029–5037
Dai MS, Lu H (2004) Inhibition of MDM2-mediated p53 ubiquitination and degradation by ribosomal protein L5. J Biol Chem 279:44475–44482
Jin A, Itahana K, O’Keefe K, Zhang Y (2004) Inhibition of HDM2 and activation of p53 by ribosomal protein L23. Mol Cell Biol 24:7669–7680
Lohrum MA, Ludwig RL, Kubbutat MH, Hanlon M, Vousden KH (2003) Regulation of HDM2 activity by the ribosomal protein L11. Cancer Cell 3:577–587
Yao H, Feng Y, Zhou T, Wang J, Wang Z (2012) NMR studies of the interaction between human programmed cell death 5 and human p53. Biochemistry 51:2684–2693
Schon O, Friedler A, Freund S, Fersht AR (2004) Binding of p53-derived ligands to MDM2 induces a variety of long range conformational changes. J Mol Biol 336:197–202
Meek DW (1994) Post-translational modification of p53. Semin Cancer Biol 5:203–210
Acknowledgments
This work was supported by Grants from the National Key Project for Basic Research of China (973, 2011CB910103) and the National Natural Science Foundation of China (30871263).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xu, L., Hu, J., Zhao, Y. et al. PDCD5 interacts with p53 and functions as a positive regulator in the p53 pathway. Apoptosis 17, 1235–1245 (2012). https://doi.org/10.1007/s10495-012-0754-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10495-012-0754-x