Abstract
The objective of this study was the characterisation of the S-layer protein (SlpA) and its functional role in the probiotic activity of Lactobacillus helveticus M92. SlpA was isolated and identified by SDS-PAGE LC-MS/MS analysis. The slpA gene encoding the SlpA from L. helveticus M92 was sequenced and compared with other well characterised slpA genes. Sequence similarity searches revealed high homology with the SlpA of Lactobacillus strains. Purified SlpA showed significantly better immunomodulatory effects in orally immunised mice than L. helveticus M92 cells after SlpA removal. SlpA is involved in the autoaggregation of L. helveticus M92 cells and coaggregation of L. helveticus M92 with S. Typhimurium FP1 as these processes were negatively affected after SlpA removal from the cell surface. Therefore, the influence of oral treatment with L. helveticus M92 on an oral infection of mice by S. Typhimurium FP1 was investigated. Following the oral immunization of mice, with viable L. helveticus M92 and S. Typhimurium FP1 cells, the concentration in the luminal contents of total S-IgA and specific anti-Salmonella S-IgA antibodies, from all immunized mice was significantly higher compared to the control group or a group of mice infected only with S. Typhimurium FP1. These results demonstrate that the observed reduced infection by S. Typhimurium FP1 in mice with L. helveticus M92 is associated with competitive exclusion in the intestinal tract and enhanced immune protection conferred by the L. helveticus M92 and its SlpA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Among lactic acid bacteria many Lactobacillus strains have been characterised as probiotics. These strains were reported to exert health benefits such as protection against infection, e.g. by modulating the immune system. Immunostimulation and the ability to colonize mucosal surfaces have prompted efforts aimed at the use of these strains as vaccine delivery vehicles for oral immunization. Although the molecular basis of these probiotic activities are not well understood, several mechanisms have been proposed: contribution to mucosal barrier function, coaggregation with pathogens, competitive exclusion, modulation of the immune response, decreasing of the luminal pH and secretion of specific compounds such as bacteriocins (Coconnier et al. 2000; Fayol-Messaoudi et al. 2005; Šušković et al. 2010). Still, adhesion of the probiotics to the mucosa is considered a main prerequisite for their survival and establishment in the gastrointestinal tract (GIT) where their health benefits are expected. Ability to temporarily colonize the intestinal epithelia allows probiotics to exert their beneficial effects longer (Servin and Coconnier 2003). Surface-located molecules such as lipoteichoic acid, lectin-like molecules and proteins have been identified as adhesins which specifically interact with different receptor moieties in the intestinal tissue (Martinez et al. 2000; Beganović 2008; Beganović et al. 2010).
Several species of the genus Lactobacillus possess surface S-layer protein (SlpA). Due to their structural regularity and the unique self-assembling properties S-layers have potential for many biotechnological applications (Åvall-Jääskeläinen and Palva 2005; Åvall-Jääskeläinen et al. 2008). Although the functional significance of Lactobacillus SlpA is not completely elucidated, these proteins are assumed to have an important role in bacteria, because a substantial part of the synthetic capacity of the cell is used for their production. The following biological functions have been shown or presumed for S-layers: (i) protective barrier against environmental hazards, (ii) control of the transfer of nutrients and metabolites, (iii) maintenance of cell shape and envelope rigidity, and (iv) promoter for cell adhesion and surface recognition (Vidgrén et al. 1992; Buck et al. 2005).
Strain Lactobacillus helveticus M92 was defined as probiotic according to proposed probiotic selection criteria (Kos et al. 2000; Šušković et al. 2000; Kos et al. 2003; Beganović 2008; Frece et al. 2009; Leboš Pavunc et al. 2010). This strain has the ability to survive simulated GIT conditions, is bile resistant, has antibacterial activity against some enteropathogenic and spore-forming bacteria, adheres to porcine ileal epithelial cells ex vivo, and as such is a potential candidate probiotic (Kos et al. 2000; Šušković et al. 2000; Kos et al. 2003). Furthermore, in vitro studies have shown that L. helveticus M92 assimilated cholesterol in the presence of bile, so it is postulated that this strain might help in lowering serum cholesterol in vivo (Šušković et al. 2000, 2001).
Lactobacillus helveticus M92 possesses an SlpA. A gene coding for the SlpA protein from L. helveticus M92 was detected by Southern blot hybridization (Frece et al. 2005a). Various data suggested that some of the L. helveticus M92 probiotic traits could be mediated by its SlpA, notably data concerning strain adhesion to the host cells. Kos et al. (2003) and Frece et al. (2005a) have shown a protective role of S-layers during transit through the GIT and during freeze-drying of cultures for probiotic applications. The role of the S-layer in the adherence of L. helveticus M92 to mouse and pig intestinal epithelial cells was demonstrated (Kos et al. 2003; Frece et al. 2005a). Adhesion is believed to be a requirement for the realisation of probiotic effects, such as pathogen exclusion and immunomodulation (Buck et al. 2005; Lebeer et al. 2008). Indeed, S-layers of Lactobacillus species have been shown to interact with the receptors on the host epithelial cells, thereby blocking receptor sites on the mucosal surfaces for the adherence of pathogenic species (van der Mei et al. 2003; Liu et al. 2010).
In the present study, the main objective was to characterise the SlpA and its functional role in the probiotic activity of L. helveticus M92. Previous research in our laboratory showed that oral administration of L. helveticus M92 can enhance immune functions in mice by increasing the concentrations of serum IgA, IgG, and IgM antibodies (Frece et al. 2005b). Hence, the possibility of inducing an immunogenic response by using purified SlpA in mice was investigated. In addition, the role of the L. helveticus M92 in enhanced protection of mice against oral challenge infection by Salmonella enterica serovar Thyphimurium FP1 was studied.
Materials and methods
Bacterial strains and growth conditions
Strains L. helveticus M92, Lactobacillus fermentum A8 and S. enterica serovar Typhimurium FP1 were obtained from the culture collection of the Department of Biochemical Engineering, Laboratory for Antibiotic, Enzyme, Probiotic and Starter cultures Technology, Faculty of Food Technology and Biotechnology, University of Zagreb. L. helveticus M92 and L. fermentum A8 were stored at −80°C in MRS broth (Difco, Detroit, MI, USA) containing 30% (v/v) glycerol. In order to distinguish and monitor the survival of L. helveticus M92 in the GIT of the mice, rifampicin marking of the strain was performed according to Frece et al. (2005b). This was performed just for the purpose of the present research because a rifampicin-resistant variant of L. helveticus M92 is not applicable in food. S. Typhimurium FP1 was stored at −80°C in the nutrient broth (Biolife, Milano, Italy) with 30% (v/v) glycerol.
Extraction of the L. helveticus M92 SlpA
An overnight culture of L. helveticus M92 grown in MRS broth was used to inoculate 400 ml of MRS broth, to an optical density of 0.05 at 600 nm (UVIKON 931 spectrophotometer, KONTRON Instruments) and then cultivated at 37°C until the exponential phase of growth (OD600 nm of 0.7). The cells were washed twice with an equal volume of ice-cold water and resuspended in 10 ml of 5 M LiCl and incubated for 30 min at room temperature. SlpA from L. helveticus M92 was LiCl extracted and extensively purified by dialysis using a method described by Frece et al. (2005a). After the freeze-drying of the dialysed S-layer (CHRIST Alpha 1-2 LDplus freeze-dryer, SciQuip, Shropshire, UK), protein concentration was determined by the Bradford method (Bradford, 1976) and the purity of the preparation was analysed by denaturing SDS-PAGE on 4–12% polyacrylamide minigels in MES buffer (200 V, 110 mA for 45 min). The gel was stained with Blue safe stain (Invitrogen, Carlsbad, CA) while shaking on an orbital shaker for 60 min after which the gel was washed twice with 100 ml of Milli-Q water.
Protein identification by mass spectrometry
In- gel digestion with sequencing grade modified trypsin (Promega, Madison, WI, USA) was performed as previously described by Beganović et al. (2010). For liquid chromatography tandem mass spectrometry (LC-MS/MS) analysis, tryptic protein digests were resuspended in 25 μl of precolumn loading buffer (0.08% TFA and 2% ACN in water). Tandem mass spectrometry analysis (LC/ESI-MS/MS) was performed on an Ultimate 3,000 LC system (Dionex, Voisins le Bretonneux, France) connected to a linear ion trap mass spectrometer (LTQ, Thermo Fisher, USA) by a nanoelectrospray interface. Peptide samples (4 μl) were loaded at a flow rate of 20 μl/min at precolumn (Pepmap C18; 0.3 × 5 mm, 100 Å, 5 μm; Dionex). After 4 min, the precolumn was connected to the separating nanocolumn Pepmap C18 (0.075 × 15 cm, 100 Å, 3 μm) and the gradient was started at 300 nl/min. All peptides were separated on the nanocolumn using a linear gradient from 2 to 36% of buffer B for 18 min (buffer A: 0.1% formic acid, 2% acetonitrile and eluting buffer B: 0.1% formic acid, 80% acetonitrile). Including the regeneration step, the run length was 50 min. Ionization was performed on the liquid junction with a spray voltage of 1.3 kV applied to a non-coated capillary probe (PicoTip EMITER 10 μm ID; New Objective, USA). Peptides ions were analysed by the Nth-dependent method as follows: (i) full Ms scan (m/z 300–2,000), (ii) ZoomScan (scan of the three major ions), (iii) MS/MS on these three ions with classical peptides fragmentation parameters: Qz = 0.25, activation time = 30 ms, collision energy = 40%. The time during which the same ion cannot be reanalyzed was set to 30 s.
Protein identification was performed using Bioworks 3.2TM software (Thermo scientific). The Bioworks 3.2TM search parameters included: trypsin specificity allowing one missed cleavage site, oxidation variable of methionine. The mass tolerance was fixed to 1.4 Da for precursor ions and 0.5 Da for fragment ions. The search result was filtered using Bioworks 3.2. using following criteria: Xcorrelation score (X corr) > 1.7, 2.5, and 3.0 for mono-, di-, and tricharged peptides, respectively; peptide probabilities lower than 0.01; ΔCn defined by [(X corr1 − X corr2)/X corr1] bigger than 0.1 and only the first match result for each identified peptide. Upon completion of the LC/ESI-MS/MS run, the acquired MS/MS spectrum was analysed on LTQ linear ion trap mass spectrometer by SEQUEST protein search algorithm.
Detection of S-protein genes by PCR
SlpA gene-specific oligonucleotides used for PCR for the detection of slpA genes are listed in Table 1. Chromosomal DNA was isolated from L. helveticus M92 essentially as described by Frece et al. (2005a). A PCR reaction containing 1 μl of diluted template DNA, 0.2 mM deoxynucleoside triphosphate mix, 1 mM MgCl2, 1 pmol/μl each oligonucleotide, and 0.05 U/μl Taq polymerase was prepared and amplified under the following conditions: 94°C for 5 min followed by 25 cycles of 1 min at 94°C, 1 min at the oligonucleotide-specific annealing temperature (Ta) (Table 1), and 2 min of extension at 72°C, and then a hold at 72°C for 8 min. The negative control consisted of 1 μl sterile MilliQ H2O and 1 μl diluted (1/20) DNA from L. fermentum A8 (an SlpA-negative strain). The presence or absence of PCR products and the sizes of the fragments from positive PCR reactions were analyzed using a 1% agarose gel. Nucleotide sequencing was performed with an ABI PRISM 310 Genetic Analyser-Bioscreen (PE Biosystems, USA) and sequence editing was performed with the Sequencher (version 3.0) software (Gene Codes Corporation, Ann Arbor, MI). Homology searches of the databases were done with the BLAST program (http://www.ncbi.nlm.nih.gov/BLAST).
Anti-Salmonella effect of L. helveticus M92 in vitro and in vivo in experimental mice
Autoaggregation assay and coaggregation with S. Typhimurium FP1 were performed according to Kos et al. (2003) to explore the anti-Salmonella effect of L. helveticus M92 in vitro as salmonellosis is among the most common causes of foodborne human gastroenteritis worldwide (Golowczyc et al. 2007). Swiss albino mice have been used as a suitable animal model to study the events following experimental administration of probiotic strains (Frece et al. 2005a, b; Racedo et al. 2006; Frece et al. 2009; Hajduk et al. 2009). In the present study, we used this mouse model to study the dynamics of antibody responses to L. helveticus M92 expressing SlpA. Hence, female Swiss albino mice (four mice per group) for the in vivo experiments were treated as described previously (Frece et al. 2009). Rifampicin-resistant L. helveticus M92 cells and S. Typhimurium FP1 cells were centrifuged at 10,000 g for 2 min, washed 3 times and resuspended in sterile 0.5% NaCl solution to reach final concentration of ca. 1011 CFU/ml for L. helveticus M92, and ca. 103 CFU/ml for S. Typhimurium FP1, respectively, using standard curves for each bacterium. The concentration of probiotic cells corresponds to recommended daily probiotic dose, while the concentration of Salmonella cells represents a possible infective dose. Two groups of mice were orally treated daily with 200 μl of prepared suspension of probiotic cells during seven consecutive days. On the 3rd day, mice in one group (M92+S) were challenged by single oral infection with S. Typhimurium FP1, whereas the second group (M92) were not. A third group of mice (S) was infected with 200 μl of prepared suspension of S. Typhimurium FP1 cells at 3 days. A fourth (control) group of mice was fed only with standard rodent feed and received no bacterial infection. The group of mice treated daily with 200 μl of prepared suspension of L. helveticus M92 cells was used as negative control (M92 group). Each group of experimental animals consisted of four mice. In vivo adhesion test was carried out as described by Frece et al. (2005b).
Oral immunization of mice with purified SlpA, whole L. helveticus M92 cells or L. helveticus M92 without S-layer, as well as treatment with L. helveticus M92 in combination with S. Typhimurium FP1 infection and S. Typhimurium FP1 alone was performed as described previously (Frece et al. 2005b). The total S-IgA and specific S-IgA antibodies against S. Typhimurium FP1 were determined in polystyrene microtiter plates (NUNC) according to the method described by Frece et al. (2005b). All animal studies were performed according to ethical procedures set in the “Guide for the Care and Use of Laboratory Animal’s of the National Research Council’’ (1996).
Statistical analysis
Data were expressed as means of three independent trials ± standard deviation (SD). Data were subjected to a one-way analysis of variance. Statistical analysis was made by Statistica 9.0 software (StatSoft Inc., Tulsa, OK). A P value of <0.05 was considered to indicate a significant difference.
Results
Characterisation of L. helveticus M92 SlpA
A generally employed method for the removal of SlpA from the cell surfaces, LiCl extraction, was applied for the isolation of SlpA from L. helveticus M92. An identification of the L. helveticus M92 S-layer was performed by SDS-PAGE coupled to LC/ESI–MS/MS, which revealed that the molecular mass of the SlpA is 46541.9 Da (Table 2). BLASTP analysis of L. helveticus M92 SlpA sequence, obtained by mass spectrometry analysis, showed that this protein shared a high sequence identity to related SlpA of other L. helveticus and Lactobacillus acidophilus strains (Fig. 1).
PCR amplification with specific primers, designed from the protein sequence obtained by mass spectrometry analysis, was used to amplify the slpA gene from the genome of L. helveticus M92. A single 1.2 kbp PCR product was obtained from L. helveticus M92 chromosomal DNA, while DNA from L. fermentum A8 was used as negative control (Fig. 2). The nucleotide sequence of the slpA gene of L. helveticus M92 revealed an ORF (open reading frame) of 1,439 bp and is deposited in the GenBank database under the accession number HM140425 and needs to be processed for further annotation. A similarity search using the deduced nucleotide sequence of the L. helveticus M92 slpA with the algorithm BLASTN revealed a high level of sequence homology to the other Lactobacillus S-layer genes, showing between notably with L. helveticus (98% sequence identity over >1,330 nucleotides with GenBank accession no. AJ388558, X91199), Lactobacillus crispatus (85% sequence identity over 492 nucleotides with GenBank accession no. AY941197) and L. acidophilus (79% sequence identity over 1,415 nucleotides with GenBank accession no. X71412).
Functional role of SlpA in the probiotic activity of L. helveticus M92
To asses the functional role of the identified SlpA, the influence of S-layer removal from the surface of L. helveticus M92 cells on autoaggregation and coaggregation with the enteropathogen S. Typhimurium FP1 was investigated. These two characteristics were markedly affected by treatment of L. helveticus M92 cells with LiCl (which resulted in ca. 10% lower autoaggregation and ca. 16% lower coaggregation). This suggested that the SlpA is involved in autoaggregation as well as in coaggregation with S. Typhimurium FP1 (Table 3). Coaggregation is a part of competitive exclusion mechanism which, coupled with the antimicrobial activity of the probiotic strain, enables a decrease of the pathogenic load during infections. Therefore, an antagonistic activity of L. helveticus M92 against S. Typhimurium FP1 and its influence on the composition of the intestinal microflora was tested in vivo on Swiss Albino mice. Seven days after the oral administration of L. helveticus M92 in combination with S. Typhimurium FP1 challenge, enterobacterial counts as well as Salmonella sp. counts decreased by ca. 2 log units compared to the enterobacteria and Salmonella sp. counts determined in the group of mice infected only with S. Typhimurium FP1 (Fig. 3). Additionally, the LAB and rifampicin-resistant LAB (representing the probiotic strain administered) counts in the small and large intestines of mice were significantly increased compared to the control group by ca. 1.5–2 log units, respectively (Fig. 3).
Bacterial viable cell count determined in the small intestine (a) and large intestine (b) of mice, 7 days after the oral treatment with L. helveticus M92, L. helveticus M92 in combination with S. Typhimurium FP1, or after the challenge with S. Typhimurium FP1. Total LAB (A) and rifampicin-resistant LAB (B) on MRS-agar; Enterobacteriaceae (C) on Violet red bile glucose agar, Salmonella sp. (D) on Brilliant green violet agar. Error bars represent standard deviations of the mean values of results from three replicates
Furthermore, the oral immunization of mice with purified L. helveticus M92 slpA protein and with L. helveticus M92 cells without SlpA stimulated the immune response in mice (Figs. 4a–c, 5b). After the oral immunisation of mice with purified SlpA, the levels of serum IgA, IgG, and IgM antibodies were significantly higher in comparison to the levels of these antibodies in the control group of mice and in the group of mice immunized with L. helveticus M92 cells either with or without SlpA (Fig. 4a–c). The highest luminal content of anti-S. Typhimurium S-IgA antibody was detected in the group of mice infected with S. Typhimurium FP1 in combination with L. helveticus M92 (Fig. 5a). Total secretory S-IgA antibody levels were the highest after the oral administration of mice with L. helveticus M92 alone and were higher in combination with S. Typhimurium FP1, than in the control group of mice (Fig. 5b).
Total a IgA, b IgG, and c IgM antibodies in sera, determined during and after oral immunisation of mice with purified S-protein, with viable L. helveticus M92 cells and with viable L. helveticus M92 cells after SlpA removal, by ELISA method (C–control). Sampling started on 5th day after the first oral immunization that was performed during seven consecutive days
Determination of a secretory-IgA (S-IgA) specific against S. Typhimurium FP1 and b total secretory IgA (S-IgA) by ELISA method in the intestinal fluid after the oral challenge of mice with viable cells of S. Typhimurium FP1 alone (S) or in combination with L. helveticus M92 (M92+S), or with L. helveticus M92 alone (M92), (C–control). Error bars represent standard deviations of the mean values
Discussion
Probiotics exert several beneficial effects on human health, including interaction with the immune system, production of antimicrobial substances, enhancement of the mucosal barrier function and competition with enteropathogens for adhesion sites (Boesten and de Vos 2008). Although the molecular mechanisms by which probiotic bacteria exert health benefits to the host are largely unknown, it has been accepted that surface molecules, mostly proteins, are involved in their adhesion and colonization in intestinal tract, which are correlated with their probiotic activity (Åvall-Jääskeläinen and Palva 2005). This research is aimed to elucidate if there is a relationship between some important probiotic traits of L. helveticus M92, such as adhesion ability, Salmonella exclusion and immunomodulation in mice intestinal tract, and its SlpA. The slpA gene of L. helveticus M92 was originally identified by Southern blot hybridization (Frece et al. 2005a). Here, the slpA gene was sequenced and sequence similarity searches revealed high homology with the other SlpAs of Lactobacillus strains. The identification of the L. helveticus M92 surface paracrystalline SlpA, encoded by slpA gene, was also achieved by means of mass spectrometry analysis, SDS-PAGE coupled to LC-MS/MS. Previously Kos et al. (2003) suggested that this poorly soluble SlpA could be responsible for the hydrophobicity of L. helveticus M92 cells. Kos et al. (2003) and Frece et al. (2005a) demonstrated, in ex vivo experiments, that SlpA was involved in L. helveticus M92 adhesion to the intestinal epithelial cells of a pig and a mouse. Hence, it was postulated that the SlpA could be responsible for the interactions with intestinal epithelial cells in vivo and for the autoaggregation ability of this strain. The results of the present study demonstrated that the autoaggregation percentage determined for L. helveticus M92 was significantly lower after the removal of S-layer from the bacterial surface. These results support the hypothesis that the S-layer from L. helveticus M92, through autoaggregation, could be involved in adhesion, which is known to be a prerequisite for the colonization of the GIT by probiotic strains in high viable cell count (Kos et al. 2003). This is in agreement with Mobili et al. (2009) who showed a correlation between the structure of SlpAs from different L. kefir strains and aggregation properties of whole bacterial cells.
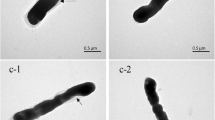
In addition to the above, coaggregation of probiotic strains with pathogens, as well as their ability to displace pathogens through antimicrobial activity, is of importance for the therapeutic manipulation of an aberrant intestinal microbiota (Servin and Coconnier 2003). Interestingly, coaggregation was significantly reduced when L. helveticus M92 cells were lacking SlpA compared to the results obtained with whole L. helveticus M92 cells, again implicating the importance of the S-layer in this process. Coaggregation, which is thought to facilitate the clearance of pathogens during mucus flushing, is described as an additional mechanism to decrease the pathogenic load during infections. Moreover, adhesion to epithelial cells and mucus mediates colonisation of the GIT by lactobacilli and may be prerequisite for competitive exclusion of enteropathogenic bacteria and immunomodulation of the host (Perdigón et al. 2003). Johnson-Henry et al. (2007) reported that SlpA extracts from L. helveticus had inhibited enterohaemorrhagic Escherichia coli adhesion to host epithelial cells, while Buck et al. (2005) and Frece et al. (2005a) demonstrated a decrease of L. crispatus and L. helveticus M92 ability to bind to intestinal epithelial cells in vitro after the removal or disruption of SlpAs. A complementary approach using transmission electron microscopy could be useful for the confirmation of both the presence of the paracrystalline SlpA and the functional role of SlpAs or any other cell surface structures important for probiotic activity, such as adhesion to intestinal epithelial cells (Johnson-Henry et al. 2007).
The possible role of L. helveticus M92 in the competitive exclusion of S. Typhimurium FP1 was investigated because Salmonella infections are one of the primary causes of gastroenteritis in humans. The use of antibiotics in the treatment of Salmonella often becomes less efficient due to the spread of antibiotic resistance (Casey et al. 2004). Therefore, the application of probiotic LAB bearing activity against Salmonella sp. could be effective as an alternative strategy (Coconnier et al. 2000; Casey et al. 2004; Golowczyc et al. 2007). Previously, Kos et al. (2008) found, by in vitro competition tests, that the growth of S. Typhimurium was completely inhibited after 10 h of incubation with L. helveticus M92 (Kos et al. 2008). The SlpAs are involved in coaggregation, but do not possess antimicrobial activity (data not shown). The intact probiotic cells, capable of producing antimicrobials, were necessary for the competitive exclusion of Salmonella. Therefore, in this study, involvement of the L. helveticus M92 in the reduction of gastrointestinal Salmonella infection in vivo was studied. The increased viable cell counts of LAB in small and large intestine of mice were detected 7 days after L. helveticus M92 administration. The viable cell counts of enterobacteria and Salmonella sp., in small and large intestine of mice, were lower compared with those obtained in the group of mice infected only with S. Typhimurium FP1. According to Golowczyc et al. (2007) SlpAs could interact with specific sites on Salmonella surface involved in the first step of mucosal infection or could modify or mask Salmonella structures necessary for the invasion of intestinal epithelial cells. Surface layer extracts from L. helveticus R0052 were recently shown to inhibit the adhesion of E. coli O157:H7 to epithelial cells (Johnson-Henry et al. 2007). Recently Liu et al. (2010) demonstrated that Lactobacillus plantarum surface layer adhesive protein decreased E. coli adhesion to Caco-2 cells and rescued E. coli-induced alterations in tight junction structures and permeability of Caco-2 cell monolayers. This process seems to be partly mediated by high hydrophobicity of the S-layers, and it is not yet known whether it involves interactions with specific receptors. Similar results were obtained for the S-layers of L. crispatus ZJ001, which were shown to play a role in the competitive exclusion against enterohemorrhagic E. coli and S. enterica serovar Typhimurium (Chen et al. 2007). Furthermore, Horie et al. (2002) reported that SlpAs of L. crispatus JCM 5810 inhibited the adhesion of E. coli to Matrigel, and this effect was ascribed to the competition with E. coli for the same binding sites in the extracellular matrix.
The possible competitive exclusion mechanisms of probiotic include ability of their cells to produce antibacterial substances and to compete for nutrients and receptors on the gut enterocytes, but also immune stimulation of the specific and non-specific immune system. Hence, the possible functional role of the orally administered, purified SlpA in the imunomodulation conferred by L. helveticus M92 in mice was studied. Here it must be emphasised that L. helveticus M92 SlpA evoked higher total serum IgA, IgG, and IgM than L. helveticus M92 cells without SlpA, but the S-layer did not evoke a specific humoral immune response after oral application and as such is suitable for probiotic application as an immunomodulator. In addition, the concentrations of the serum IgA, IgG, and IgM antibodies were lower when mice were orally immunised by L. helveticus M92 cells without S-protein compared to the levels of antibodies determined in the samples from the group of mice orally immunised with whole L. helveticus M92 cells, but were still higher compared to the control. SlpA, as the outer shell of proteins in lactobacilli (Delcour et al. 1999), may have the highest probability of the intimate interaction with the immune cells associated with the gut. Previously, between different probiotic strains assessed, L. helveticus M92, showed the highest capacity of activation of the immune system (Frece et al. 2005b). The immunomodulation capacity of the S-layer could be one of its functions, as was reported for the S-layer of the Bacteroides (Yoneda et al. 2003) and Campylobacter species (Grogono-Thomas et al. 2003). It seems that the S-layer, besides its involvement in the adhesive capacity and certain cell surface traits such as hydrophobicity and autoaggregation of L. helveticus M92, contributes to the immunostimulatory activity of this probiotic bacterium. Whereas bacterial interactions are the most accepted mechanism for the reduction of Salmonella count observed by L. helveticus M92 application, stimulation of an effective innate immune response by the probiotic strain is more likely due to the rapidity of this response. Therefore, the effect of probiotic strain L. helveticus M92 on the total and specific mucosal antibody response levels in mice after challenge with S. Typhimurium FP1 was investigated. The significant increase in intestinal secretory IgA (S-IgA) antibody after L. helveticus M92 application is an important result. IgA is the predominant mucosal antibody and plays a key role in protection against dietary antigens and intestinal pathogens. This could be assigned to the surface SlpA of L. helveticus M92. Recently, Konstatinov et al. (2008) found that 45 kDa SlpA from the surface of L. acidophilus NCFM was involved in the regulation of immature dendritic cells (DC) as well as cytokine production. The cellular contacts of DCs and L. acidophilus NCFM involve interactions between dendritic cell-specific intercellular adhesion molecule (ICAM)-3-grabbing nonintegrin (DC-SIGN), a DC-specific receptor DC-SIGN, and SlpA, the dominant protein expressed by L. acidophilus NCFM.
The enhanced immunity and reduced disease severity conferred by L. helveticus M92 in this study against S. Typhimurium FP1, with the evidences from the previous studies of immunity-enhancing and antimicrobial effect of L. helveticus M92 against pathogens (Frece et al. 2005a, b; Kos et al. 2008) suggest that dietary supplementation with this defined probiotic strain may represent an effective biotherapeutic means of countering gastrointestinal infections in humans.
References
Åvall-Jääskeläinen S, Palva A (2005) Lactobacillus surface layers and their applications. FEMS Microbiol Rev 29:511–529
Åvall-Jääskeläinen S, Hynönen U, Ilk N, Pum D, Sleytr UB, Palva A (2008) Identification and characterization of domains responsible for self-assembly and cell wall binding of the surface layer protein of Lactobacillus brevis ATCC 8287. BMC Microbiol 8(165):1–15
Beganović J (2008) Application of proteomics and other molecular methods in the characterisation of functionality of the probiotic bacteria. Dissertation, University of Zagreb
Beganović J, Guillot A, van de Guchte M, Jouan A, Gitton C, Loux V, Roy K, Huet S, Monod H, Monnet V (2010) Characterization of the insoluble proteome of Lactococcus lactis by SDS-PAGE LC-MS/MS leads to the identification of new markers of adaptation of the bacteria to the mouse digestive tract. J Proteome Res 9:677–688
Boesten RJ, de Vos WM (2008) Interactomics in the human intestine: Lactobacilli and Bifidobacteria make a difference. J Clin Gastroenterol 42(Suppl 3 Pt 2):S163–S167
Bradford M (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Buck BL, Altermann E, Svingerud T, Klaenhammer TR (2005) Functional analysis of putative adhesion factors in Lactobacillus acidophilus NCFM. Appl Environ Microbiol 71(12):8344–8351
Casey PG, Casey GD, Gardiner GE, Tangney M, Stanton C, Ross RP, Hill C, Fitzgerald GF (2004) Isolation and characterisation of anti-Salmonella lactic acid bacteria from porcine gastrointestinal tract. Lett Appl Microbiol 39:431–438
Chen XY, Xu JJ, Shuai JB, Chen JS, Zhang ZF, Fang WH (2007) The S-layer proteins of Lactobacillus crispatus strain ZJ001 is responsible for competitive exclusion against Escherichia coli O157: H7 and Salmonella typhimurium. Int J Food Microbiol 115:307–312
Coconnier MH, Lievin V, Lorrot M, Servin AL (2000) Antagonistic activity of Lactobacillus acidophilus LB against intracellular Salmonella enterica serovar Typhimurium infecting human enterocyte-like Caco-2/TC-7 cells. Appl Environ Microbiol 66:1152–1157
Delcour J, Ferain T, Deghorain M, Palumbo E, Hols P (1999) The biosynthesis and functionality of the cell-wall of lactic acid bacteria. Antonie van Leeuwenhoek 76:159–184
Fayol-Messaoudi D, Berger CN, Coconnier-Polter MH, Liévin-Le Moal V, Servin AL (2005) pH-, lactic acid-, and non-lactic acid-dependent activities of probiotic lactobacilli against Salmonella enterica Serovar Typhimurium. Appl Environ Microbiol 71(10):6008–6013
Frece J, Kos B, Svetec IK, Zgaga Z, Mrsa V, Šušković J (2005a) Importance of S-layer proteins in probiotic activity of Lactobacillus acidophilus M92. J Appl Microbiol 98:285–292
Frece J, Kos B, Beganović J, Vuković S, Šušković J (2005b) In vivo testing of functional properties of three selected probiotic strains. World J Microbiol Biotechnol 21:1401–1408
Frece J, Kos B, Svetec IK, Zgaga Z, Beganović J, Leboš A, Šušković J (2009) Synbiotic effect of Lactobacillus helveticus M92 and prebiotics on the intestinal microflora and immune system of mice. J Dairy Res 76:98–104
Golowczyc MA, Mobili P, Garrote GL, Abraham AG, De Antoni GL (2007) Protective action of Lactobacillus kefir carrying S-layer protein against Salmonella enterica serovar Enteridis. Int J Food Microbiol 118:264–273
Grogono-Thomas R, Blaser MJ, Ahmadi M, Newell DG (2003) Role of S-layer protein antigenic diversity in the immune responses of sheep experimentally challenged with Campylobacter fetus subsp. Fetus. Infect Immun 71:147–154
Hajduk G, Kos B, Šušković J, Frece J, Leboš A, Beganović J (2009) Probiotic properties of Bifidobacterium animalis subsp. lactis BB-12 in baby cereal flakes enriched with inulin. Ital J Food Sci 4(21):473–486
Horie M, Kajikawa HS, Toba T (2002) Identification of Lactobacillus crispatus by polymerase chain reaction targeting S-layer protein gene. Lett Appl Microbiol 35(1):57–61
Johnson-Henry KC, Hagen KE, Gordonpour M, Tompkins TA, Sherman PM (2007) Surface-layer protein extracts from Lactobacillus helveticus inhibit enterohaemorrhagic Escherichia coli O157:H7 adhesion to epithelial cells. Cell Microbiol 9(2):356–367
Kos B, Šušković J, Goreta J, Matošić S (2000) Effect of protectors on the viability of Lactobacillus acidophilus M92 in simulated gastrointestinal conditions. Food Technol Biotechnol 36:121–127
Kos B, Šušković J, Vuković S, Šimpraga M, Frece J, Matošić S (2003) Adhesion and aggregation ability of probiotic strain Lactobacillus acidophilus M92. J Appl Microbiol 94:981–987
Kos B, Šušković J, Beganović J, Gjuračić K, Frece J, Iannaccone C, Canganella F (2008) Characterization of the three selected probiotic strains for the application in food industry. World J Microbiol Biotechnol 24:699–707
Lebeer S, Vanderleyden J, De Keersmaecker SCJ (2008) Genes and molecules of lactobacilli supporting probiotic action. Microbiol Mol Biol Rev 72(4):728–764
Leboš Pavunc A, Beganović J, Kos B, Buneta A, Beluhan S, Šušković J (2010) Influence of microencapsulation and transglutaminase on viability of probiotic strain Lactobacillus helveticus M92 and consistency of set yoghurt. Int J Dairy Techol 76 (in press). doi:10.1111/j.1471-0307.2010.00647.x
Liu TS, P Zhang, Ma Y, Qin H (2010) Lactobacillus plantarum surface layer adhesive protein protects intestinal epithelial cells against tight junction injury induced by enteropathogenic Escherichia coli. Mol Biol Rep. doi:10.1007/s11033-010-0457-8
Martínez B, Sillanpää J, Smit E, Korhonen TK, Pouwels PH (2000) Expression of cbsA encoding the collagen-binding S-protein of Lactobacillus crispatus JCM5810 in Lactobacillus casei ATCC 393T. J Bacteriol 182(23):6857–6861
Mobili P, Serradell MA, Trejo SA, Puigvert FXA, Abraham GA GL, Antoni De (2009) Heterogeneity of S-layer proteins from aggregating and non-aggregating Lactobacillus kefir strains. Antonie van Leeuwenhoek 95:363–372
National Research Council (1996) Guide for the care and use of laboratory animals. Institute of Laboratory Animal Resources, National Academy Press, Washington, DC
Perdigón G, Locascio M, Medici M, Pesce de Ruiz Holgado A, Oliver G (2003) Interaction of bifidobacteria with the gut and their influence in the immune function. Biocell 27:1–9
Racedo S, Villena J, Medina M, Agüero G, Rodríguez V, Alvarez S (2006) Lactobacillus casei administration reduces lung injuries in a Streptococcus pneumoniae infection in mice. Microbes Infect 8:2359–2366
Servin AL, Coconnier MH (2003) Adhesion of probiotic strains to the intestinal mucosa and interaction with pathogens. Best Pract Res Clin Gastroenterol 17(5):741–754
Šušković J, Kos B, Matošić S, Besendorfer V (2000) The effect of bile salts on survival and morphology of potential probiotic strain Lactobacillus acidophilus M92. World J Microbiol Biotechnol 16:673–678
Šušković J, Kos B, Goreta J, Matošić S (2001) Role of lactic acid bacteria and bifidobacteria in synbiotic effect. Food Technol Biotechnol 39:227–235
Šušković J, Kos B, Beganović J, Leboš Pavunc A, Habjanič K, Matošić S (2010) Antimicrobial activity—the most important property of probiotic and starter lactic acid bacteria. Food Technol Biotechnol 48(3):296–307
Van der Mei HC, van de Belt-Gritter B, Pouwels PH, Martinez B, Busscher HJ (2003) Cell surface hydrophobicity is conveyed by S-layer: a study in recombinant lactobacilli. Colloids Surf B Biointerfaces 28:127–134
Vidgrén G, Palva I, Pakkanen R, Lounatmaa K, Palva A (1992) S-layer protein gene of Lactobacillus brevis: cloning by polymerase chain reaction and determination of the nucleotide sequence. J Bacteriol 174(22):7419–7427
Yoneda M, Hirofuji T, Motooka N, Nozoe K, Shigenaga K, Anan H, Miura M, Kabashima H, Matsumoto A, Maeda K (2003) Humoral immune responses to S-layer-like proteins of Bacteroides forsythus. Clin Diagn Lab Immunol 10:383–387
Acknowledgments
The authors are grateful for the financial support provided by Ministry of Science, Education and Sports of the Republic of Croatia (Project 0581990 “Probiotics, prebiotics and functional starter cultures”). The authors wish also to thank to the staff of PAAPSO, INRA Jouy en Josas, France for mass spectrometry analysis. Jasna Beganović was recipient of a Marie Curie fellowship for Early Stage Research Training, inside LABHEALTH (MEST-CT-2004-514428).
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Beganović, J., Frece, J., Kos, B. et al. Functionality of the S-layer protein from the probiotic strain Lactobacillus helveticus M92. Antonie van Leeuwenhoek 100, 43–53 (2011). https://doi.org/10.1007/s10482-011-9563-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-011-9563-4