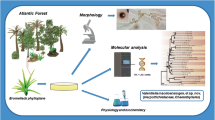
Abstract
Eight strains of a novel yeast species were isolated from rotting wood and wood-boring insects in Atlantic Rain Forest ecosystems in Brazil. Sequences of the D1/D2 domains of the large subunit of the rRNA gene showed that the yeast belongs to the Scheffersomyces clade and that it is related to Candida lignicola and Candida coipomoensis. The new species was isolated from rotting wood of three different localities and a wood-boring insect suggesting that these substrates are its ecological niche. This new yeast species is able to assimilate cellobiose and other compounds related to rotting wood. Strong fermentation of cellobiose in Durham tubes was observed for the strains of this new yeast. The new species produced an intracellular β-glucosidase responsible for cellobiose hydrolysis. The novel species, Candida queiroziae sp. nov., is proposed to accommodate these isolates. The type strain of C. queiroziae is UFMG-CLM 5.1T (=CBS 11853T = NRRL Y-48722T).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Discarded cellulosic biomass derived from forestry, agriculture, and municipal sources are potential feed stocks for the production of bio-fuels, in particularly for the synthesis of fuel ethanol (Hahn-Hägerdal et al. 2006). Although cellulose is the most abundant biopolymer in the world, its chemical structure makes it difficult to hydrolyze (Zhu et al. 2010) and current strategies to produce fuel ethanol from cellulose should favour the simultaneous saccharification and fermentation of the substrate (Lynd et al. 2002, 2005). The process requires pre-treatment of the cellulosic feedstock by stem-explosion or acid treatment, followed by addition of exogenously produced cellulolytic enzymes to hydrolyze cellulose chains and release the monomers required for fermentation (Carere et al. 2008).
During the simultaneous saccharification and fermentation process, cellulose is hydrolyzed by the cellulase enzyme complex (cellobiohydrolases, endoglucanases and β-glucosidases) working in synergy. Endoglucanases (EC 3.2.1.4) randomly cleave the β-1,4 glycosidic linkages of cellulose, cellobiohydrolases (EC 3.2.1.91) attack cellulose chain ends to produce the constitutive unit of cellulose, cellobiose, a disaccharide of glucose linked by a β-1,4 glycosidic bond, and finally β-glucosidases (EC 3.2.1 21) hydrolyze cellobiose into two glucose moieties (Kumar et al. 2008). The use of microorganisms able to ferment cellobiose constitutes a desirable alternative for bio-ethanol production because cellobiose is a competitive inhibitor of cellulases (Bezerra and Dias 2005). These strains also ferment the monomers produced during cellulose hydrolysis, and cellulases are thus not inhibited (Lynd et al. 2005).
In a study on yeast communities associated with rotting wood and wood-boring insects in Atlantic Rain Forest ecosystems in Brazil, we have isolated eight strains of a new asexual cellobiose-fermenting yeast species. Analysis of the sequence of the D1/D2 domains of the large subunit rRNA gene showed that these strains represented a single species closely related to Candida lignicola and C. coipomoensis. In this work, we describe this new species as Candida queiroziae sp.nov.
Materials and methods
Yeast isolation and identification
Five strains were isolated from rotting wood and one from a wood-boring insect collected in the Private Natural Heritage Reserve of the Sanctuary of the Caraça. This is an ecological reserve with 11,233 ha of Atlantic Rain Forest located in the Serra do Espinhaço, state of Minas Gerais, Southeastern Brazil (Cadete et al. 2009). Two other yeast strains were isolated from rotting wood samples collected at the Private Natural Heritage Reserves of Woodstock and Bello & Kerida, respectively, two areas of Atlantic Rain Forest ecosystems located in the city of Nova Friburgo, Rio de Janeiro state, Brazil. The local climate in these ecological reserves is altitudinal tropical, with cold and dry winter and fresh and rainy summers, with annual mean temperatures around 16°C.
Fifteen rotting wood and 16 wood-boring insect samples were collected from the Caraça Ecological Park in April 2008. Ten samples each were collected from the Ecological Reserves of Woodstock and Bello & Kerida in January 2009. Rotting wood samples were stored in sterile plastic bags and transported under refrigeration to the laboratory over a period of no more than 24 h. Insects were stored in sterile flasks and transported under refrigeration to the laboratory as described by Suh et al. (2003). One gram of each wood sample was placed separately in flasks with 20 ml sterile d-xylose (yeast nitrogen base 0.67%, d-xylose 0.5%, chloramphenicol 0.02%) and carboxymethylcellulose (Yeast Nitrogen Base 0.67%, carboxymethylcellulose 1%, cellobiose 0.05% and chloramphenicol 0.02%) minimal media. The flasks were incubated at 25°C on an incubator shaker (New Brunswick, USA) at 150 rpm for 3–10 days. When growth was detected, 0.5 ml of the cultures was then transferred separately to tubes containing 5 ml sterile carboxymethylcellulose or d-xylose media and the tubes were incubated as described above. For yeast isolation from insects 100 μl of a solution of insect gut macerate was inoculated in the same media and conditions mentioned above. One loopful of culture from each tube was streaked on yeast extract-malt extract agar (YM, glucose 1%, yeast extract 0.3%, malt extract 0.3%, peptone 0.5%, agar 2% and chloramphenicol 0.02%). The plates were incubated at 25°C until yeast colonies developed. The different yeast morphotypes were purified by repeated streaking on YM agar plates and preserved at −80°C or in liquid nitrogen for later identification. The yeasts were characterized by standard methods (Yarrow 1998) and identification followed the taxonomic keys of Kurtzman and Fell (1998).
The ability to ferment cellobiose was tested in Durham tubes containing fermentation basal medium with a final sugar concentration of 2% (w/v) (Yarrow 1998). The tubes were incubated at 25°C on an incubator shaker (New Brunswick) at 100 rpm for 25 days.
DNA sequencing and sequence analysis
The D1/D2 variable domains and internal transcribed spacer (ITS) of the large subunit rRNA gene were amplified by PCR directly from whole cells as described previously (Lachance et al. 1999). The amplified DNA was concentrated, cleaned (Wizard Plus SV Minipreps DNA Purification System, Promega) and sequenced in a Mega-BACETM 1000 automated sequencing system (Amersham Biosciences) in Institute of Biological Sciences of the Federal University of Minas Gerais, Brazil, or sequenced at the Robarts Research Institute, London, Ontario, in Canada. The sequences were edited with the program DNAMAN, version 6 (Lynnon BioSoft, Vaudreuil, QC, Canada). Existing sequences for other yeasts were retrieved from GenBank. The CLUSTAL W software (Thompson et al. 1994) included in DNAMAN was used to align the D1/D2 LSU rRNA gene sequences.The alignment covered 534 positions, of which 63 were polymorphic, 38 were parsimony-informative and 13 were gapped. For tree construction using the built-in Neighbour-Joining algorithm, gapped positions were deleted in pairs and Kimura’s two-parameter distance was applied.
Growth conditions and fermentation assays
Cells were grown on YP medium (1% yeast extract and 2% peptone), adjusted to pH 5.0, supplemented with 2% glucose or cellobiose. Cells were grown with shaking at 28°C (160 rpm) in cotton-plugged Erlenmeyer flasks filled to 1/5 of the volume with medium. The inoculum for growth assays was prepared by transferring a single colony aseptically from a plate into 5 ml of the YP medium containing glucose or cellobiose, and allowing growth to proceed to the stationary phase for 2–3 days before inoculating cells (by a 100 or 1,000× dilution factor) to new media of a similar composition. Culture samples were harvested regularly, centrifuged (5000×g, 1 min) and their supernatants were used for the determination of sugars and ethanol. For batch fermentations, the yeasts were pre-grown on YP-2% sugar to the late exponential phase (~1 g dry yeast l−1), centrifuged (3,500×g, 3 min), washed twice with cold water and inoculated at a high cell density (10 g dry yeast l−1) into YP medium containing the amounts of glucose and/or cellobiose indicated. Batch fermentations were incubated as described above for growth assays, and samples were collected regularly, centrifuged and their supernatants were analyzed as described below.
Determination of β-glucosidase activity
Periplasmic or secreted cellobiose hydrolysis by β-glucosidase was determined in vivo with whole cells previously incubated with 50 mM sodium fluoride to block glycolysis (Silveira et al. 1996), using 67 mM cellobiose in either 67 mM succinate–Tris, pH 5.0, or 67 mM HEPES–NaOH, pH 7.0. Intracellular β-glucosidase activity was determined in situ with permeabilized yeast cells (Stambuk 1999) using 100 mM cellobiose in either 100 mM succinate–Tris, pH 5.0, or 100 mM HEPES–NaOH, pH 7.0. Controls using previously boiled cells were used. Enzyme activity is expressed as U/g dry yeast cells, where one unit corresponds to 1 μmol of glucose produced/min at 28°C.
Analytical methods
Glucose was measured by the glucose oxidase and peroxidase method using a commercial kit (BioDiagnostica-Laborclin, Brazil), and cellobiose was determined as described by Miller (1959). Ethanol was determined with alcohol oxidase (Sigma) and peroxidase (Toyobo do Brasil, Brazil) as described previously (Alves et al. 2007). Growth was followed by turbidity measurements at 570 nm after appropriate dilution, and yeast cell dry weight was determined as described elsewhere (Badotti et al. 2008). Ethanol yield coefficients (Ye/s) were obtained at the end of ethanol production taking into account the amount of sugar utilized.
Results and discussion
The sequences of the D1/D2 domains and ITS region of the large subunit rRNA gene of the eight strains of the new species were identical. This new species belongs to the Scheffersomyces clade and it is closely related to C. lignicola and C. coipomoensis (Fig. 1). The genus Scheffersomyces was proposed by Kurtzman and Suzuki (2010) to include several d-xylose-fermenting yeast species related to Pichia stipitis. This clade contains the species S. stipitis, S. segobiensis, S. spartinae and the anamorphic yeasts C. lignosa, C. insectosa, C. shehatae, C. coipomoensis, C. lignicola and C. ergastensis. The new species differs by 11 substitutions and one gap from C. lignicola, and 13 substitutions from C. coipomoensis in the D1/D2 domains, and by eight indels from C. lignicola and six substitutions from C. coipomoensis in the ITS region of the large subunit rRNA gene. The name C. queiroziae sp. nov. is proposed for the new species.
Neighbour-joining tree of the D1/D2 domains of the large subunit of the rRNA gene showing the placement of Candida queiroziae sp. nov. among related species in the Scheffersomyces clade. Bootstrap values of 50% and above are shown. Candida zeylanoides NRRL Y-1774 is the outgroup species, and all species are analyzed from type strains. Bar 0.01 substitutions per nucleotide position
The strains of the new species were isolated from different sources and localities. The new species is not able to ferment d-xylose, but strong fermentation of cellobiose in Durham tubes was observed after 24 h for strain UFMG-IMX 6.1 isolated from a boring wood insect and after 3–6 days for the yeast strains isolated from wooding root samples (Table 1). The strains of C. queiroziae are physiologically similar to C. lignicola and C. coipomoensis. C. lignicola is able to assimilate l-arabinose and l-rhamnose, which does not occur with the new species. C. queiroziae can be distinguished from C. coipomoensis based on the assimilation of d-Gluconate, which is positive for the new species and negative for C. coipomoensis. In addition, C. queiroziae is able to grow on 10% NaCl plus 5% glucose medium, whereas the other two species did not grow. Isolates of C. queiroziae were examined individually or mixed in pairs on cornmeal, V8, dilute V8, 5% malt extract, yeast carbon base supplemented with 0.01% ammonium sulphate, and Gorodkowa agars incubated at 20 or 28°C until 30 days, but asci or signs of conjugation were not seen.
Candida coipomoensis was isolated from wood in advanced stages of degradation in the evergreen rainy Valdivian forest of Southern Chile (Ramirez and González 1984), and C. lignicola was isolated from insect frass collected from Khao-Yai National Park of Thailand (Jindamorakot et al. 2007). Similar to their closest relatives, C. queiroziae was isolated from rotting wood of three different localities and a wood-boring insect suggesting that these substrates are the ecological niche of this new species.
Growth and fermentation of cellobiose by Candida queiroziae
Candida queiroziae is able to assimilate cellobiose and other compounds related to rotting wood, and tests in Durham tubes indicates that this yeast species can ferment cellobiose within variable periods of time (see Table 1). Figure 2 shows the kinetics of growth on cellobiose by the strain UFMG-CLM 5.1T and UFMG-IMX 6.1. Both strains exhibited a typical growth curve where the sugar is efficiently fermented. After the sugar is exhausted from the medium, the ethanol produced starts to be consumed and used as a carbon source by the yeast. In agreement with the fermentation results in Durham tubes (Table 1), strain UFMG-IMX 6.1 grew and consumed cellobiose faster than strain UFMG-CLM 5.1, although at the end of the fermentation both strains produced practically the same amount of biomass (Fig. 2). An ethanol yield from cellobiose (Ye/s = 0.32 ± 0.03) obtained with these two strains of C. queiroziae is in the same range as those obtained during glucose fermentation (Ye/s = 0.32 ± 0.01) by these yeasts (data not shown). Although during growth strain UFMG-IMX 6.1 apparently utilized cellobiose more efficiently than strain UFMG-CLM 5.1, during batch fermentations at high cell densities (Fig. 3) no significant differences in the fermentation of glucose (Fig. 3a) or cellobiose (Fig. 3b) between these two strains was observed, with cellobiose fermentation by both strains occurring at a slightly slower rate than glucose fermentation.
Typical aerobic batch growth on cellobiose by Candida queiroziae UFMG-CLM-5.1T (open symbols), or strain UFMG-IMX-6.1 (black symbols). a Cell density (diamonds), b cellobiose (squares) consumption, and c production of ethanol (triangles), were determined during growth in rich YP medium containing 20 g l−1 of cellobiose. In b are also shown glucose (circles) concentrations in the medium during cellobiose utilization
Sugar batch fermentations by 10 g l−1 (dry weight) of Candida queiroziae UFMG-CLM-5.1T (open symbols) or strain UFMG-IMX-6.1 (black symbols) yeast cells in rich YP medium. Biomass (diamonds) and ethanol (triangles) production was determined during the consumption of 20 g l−1 of a glucose (circles) or b cellobiose (squares). In b are also shown glucose (circles) concentrations in the medium during cellobiose utilization
The lack of glucose production during growth on cellobiose (Fig. 2b), or even during batch fermentation of this sugar (Fig. 3b), prompted us to characterize in more detail the subcellular localization (periplasmic or intracellular) of the β-glucosidase responsible for cellobiose hydrolysis by C. queiroziae. Our results indicate that both strains UFMG-IMX 6.1 and UFMG-CLM 5.1 lack (or have very low) periplasmic β-glucosidase activity (Fig. 4), with rates of cellobiose hydrolysis of less than ~5 U/g dry yeast cells at either pH 5.0 or 7.0. When the yeast cells were permeabilized, a significant β-glucosidase activity could be seen both at pH 5.0 (29–47 U/g dry yeast cells), and especially at pH 7.0 (167–230 U/g dry yeast cells) by both strains, which is consistent with an intracellular β-glucosidase as being responsible for cellobiose hydrolysis by C. queiroziae. While yeast species (e.g. Saccharomycopsis fibuligera, Wickerhamomyces anomalus and C. wickerhamii) ferment cellobiose by expressing β-glucosidases at the cell surface, very few yeast are reported as capable of fermenting this sugar through an intracellular enzyme, a process which requires the uptake of the disaccharide into the cell. Unfortunately, when compared to our current knowledge of other disaccharide uptake system in several yeast species, cellobiose transport systems have been poorly characterized (Freer and Greene 1990; Spencer-Martins 1994). In view of its efficient cellobiose fermentation C. queiroziae may provide a new source of genes, enzymes and/or sugar transporters to engineer industrial strains for efficient ethanol production from renewable biomass (Galazca et al. 2010; Li et al. 2010).
Latin diagnosis of Candida queiroziae Santos, Cadete, Badotti, Mouro, Wallheim, Gomes, Stambuk, Lachance & Rosa sp. nov.
In medio liquido post dies tres (25°C) cellulae singulae aut binae; cellulae ovoidae aut ellipsoideae (2–3 × 2–4 μm). Post unum mensem sedimentum formatur. Cultura in agaro malti post dies 2 (17°C) candida, convexa, rugosa et opalescens. In agaro farinae Zea mays post dies 14 pseudomycelium formantur. Glucosum, galactosum et cellobiosum fermentantur, sucrosum variable at non maltosum at d-xylosum. Glucosum, galactosum, l-sorbosum, maltosum, sucrosum, cellobiosum, trehalosum, d-xylosum, d-arabinosum, ethanolum, glycerolum, erythritolum, ribitolum (variable), d-mannitolum, glucitolum, salicinum, acidum succinicum, acidum citricum, d-ribosum (exigue), hexadecanum (exigue), xylitolum, acidum lacticum (exigue), d-gluconate et N-acetylglucosaminum assimilantur, at non lactosum, melibiosum, raffinosum, melezitosum, inulinum, amylum solubile, l-arabinosum, l-rhamnosum, galactitolum, meso-inositolum, methanolum, d-glucosaminum, acetonum, ethyl acetatum nec isopropanolum. Ethylaminum, lysinum et cadaverinum assimilantur at non kalium nitricum et natrium nitrosum. Augmentum (exigue) in 37°C. Habitat materiam in Brazil. Typus UFMG–CLM 5.1. In collectione zymotica Centraalbureau voor Schimmelcultures, Trajectum ad Rhenum, sub no. CBS 11853 typus stirps deposita est.
Description of Candida queiroziae Santos, Cadete, Badotti, Mouro, Wallheim, Gomes, Stambuk, Lachance & Rosa sp. nov.
In yeast extract (0.5%), glucose (2%) broth after 3 days at 25°C, the cells are ovoid to ellipsoidal (2–3 × 2–4 μm). Budding is multilateral (Fig. 5). A sediment is formed after a month, but a pellicle is not observed. On YM agar after 2 days at 17°C, colonies are white, convex, and opalescent. In Dalmau plates after 2 weeks on cornmeal agar, pseudomycelia are formed. Ascospores are not formed. Fermentation of glucose, galactose and cellobiose is positive. Sucrose fermentation is variable. Maltose and d-xylose are not fermented. Assimilation of carbon compounds: glucose, galactose, l-sorbose, maltose, sucrose, cellobiose, trehalose, d-xylose, d-arabinose, ethanol, glycerol, erythritol, ribitol (variable), d-mannitol, glucitol, salicin, succinic acid, citric acid, d-ribose (very weak), hexadecane (very weak), xylitol, lactic acid (very weak), d-gluconate and N-acetyl-glucosamine. No growth occurs on lactose, melibiose, raffinose, melezitose, inulin, soluble starch, l-arabinose, l-rhamnose, galactitol, meso-inositol, methanol, d-glucosamine, acetone, ethyl acetate, isopropanol. Assimilation of nitrogen compounds: positive for lysine, ethylamine–HCl, cadaverine, and negative for nitrate and nitrite. Growth in amino-acid-free medium is positive. Growth at 37°C is weak. Growth on YM agar with 10% sodium chloride is positive. Growth in 50% glucose/yeast extract (0.5%) is negative. Starch-like compounds are not produced. In 100 μg cycloheximide ml−1 the growth is positive. Urease activity is negative. Diazonium Blue B reaction is negative. The habitat is rotting wood in Atlantic Rain Forest ecosystem, in the states of Minas Gerais and Rio de Janeiro, Brazil. The type strain accession number of C. queiroziae is UFMG-CLM 5.1. It was isolated from rotting wood in Brazil. It has been deposited in the collection of the Yeast Division of the Centraalbureau voor Schimmelcultures, Utrecht, The Netherlands, as strain CBS 11853T (=NRRL Y-48722T). The Mycobank number is MB 519120.The epithet queiroziae (que′i′ro′zi′ae) L. gen. n. queiroziae, of Queiroz, in honor to Dr Luzinete Aciole de Queiroz for her contribution to the mycology in Brazil.
References
Alves SL Jr, Herberts RA, Hollatz C, Miletti LC, Stambuk BU (2007) Maltose and maltotriose active transport and fermentation by Saccharomyces cerevisiae. J Am Soc Brew Chem 65:99–104. doi:10.1094/ASBCJ-2007-0411-01
Badotti F, Dário MG, Alves SL Jr, Cordioli ML, Miletti LC, Araujo PS, Stambuk BU (2008) Switching the mode of sucrose utilization by Saccharomyces cerevisiae. Microb Cell Fact 7:1–4. doi:10.1186/1475-2859-7-4
Bezerra RM, Dias AA (2005) Enzymatic kinetic of cellulose hydrolysis: inhibition by ethanol and cellobiose. Appl Biochem Biotechnol 126:49–59. doi:10.1007/s12010-005-0005-5
Cadete RM, Santos RO, Melo MA, Mouro A, Gonçalves DL, Stambuk BU, Gomes FCO, Lachance MA, Rosa CA (2009) Spathaspora arboriae sp. nov., a d-xylose-fermenting yeast species isolated from rotting wood in Brazil. FEMS Yeast Res 9:1338–1342. doi:10.1111/j.1567-1364.2009.00582.x
Carere CR, Sparling R, Cicek N, Lewin DB (2008) Third generation biofuels via direct cellulose fermentation. Int J Mol Sci 9:1342–1360. doi:10.3390/ijms9071342
Freer SN, Greene RV (1990) Transport of glucose and cellobiose by Candida wickerhamii and Clavispora lusitaniae. J Biol Chem 265:12864–12868
Galazca JM, Tian C, Beeson WT, Martinez B, Glass NL, Cate JHD (2010) Cellodextrin transport in yeast for improved biofuel production. Science 330:84–86. doi:10.1126/science.1192838
Hahn-Hägerdal B, Galbe M, Gorwa-Grauslund MF, Lidén G, Zacchi G (2006) Bio-ethanol—the fuel of tomorrow from the residues of today. Trends Biotechnol 24:549–556. doi:10.1016/j.tibtech.2006.10.004
Jindamorakot S, Limtong S, Yongmanitechai W, Tuntirungkij M, Potacharoen W, Kawasaki H, Nakase T (2007) Two new anamorphic yeasts. Candida thailandica sp.nov. and Candida lignicola sp.nov., isolated from insect frass in Thailand. FEMS Yeast Res 7:1409–1414. doi:10.1111/j.1567-1364.2007.00305.x
Kumar R, Singh S, Singh OV (2008) Bioconversion of lignocellulosic biomass: biochemical and molecular perspectives. J Ind Microbiol Biotechnol 35:377–391. doi:10.1007/s10295-008-0327-8
Kurtzman CP, Fell JW (1998) The Yeasts, a taxonomic study, 4th edn. Elsevier, Amsterdam
Lachance MA, Bowles JM, Starmer WT, Barker JSF (1999) Kodamaea kakaduensis and Candida tolerans, two new ascomycetous yeast species from Australian Hibiscus flowers. Can J Microbiol 45:172–177. doi:10.1139/cjm-45-2-172
Li S, Du J, Sun J, Galazca JM, Glass NL, Cate JHD, Yang X, Zhao H (2010) Overcoming glucose repression in mixed sugar fermentation by co-expressing a cellobiose transporter and a β-glucosidase in Saccharomyces cerevisiae. Mol Biosyst 6:2129–2132. doi:10.1039/C0MB00063A
Lynd LR, Weimer PJ, van Zyl WH, Pretorius IS (2002) Microbial cellulose utilization: fundamentals and biotechnology. Microbiol Mol Biol Rev 66:506–577. doi:10.1128/MMBR.66.3.506-577.2002
Lynd LR, van Zyl WH, McBride LE, Laser M (2005) Consolidated bioprocessing of cellulosic biomass: an update. Curr Opin Biotechnol 16:577–583. doi:10.1016/j.copbio.2005.08.009
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–429
Ramirez C, González A (1984) Five new filamentous glucose-fermenting, Candida isolated from decayed wood in the evergreen rainy from Valdivian forest of southern chile. Mycopathologia 88:83–92
Silveira MCF, Carvajal E, Bom EPS (1996) Assay for in vivo yeast invertase activity using NaF. Anal Biochem 238:26–28. doi:10.1006/abio.1996.0244
Spencer-Martins I (1994) Transport of sugars in yeasts: implications in the fermentation of lignocellulosic materials. Bioresour Technol 50:51–57. doi:10.1016/0960-8524(94)90220-8
Stambuk BU (1999) A simple experiment illustrating metabolic regulation: induction versus repression of yeast α-glucosidase. Biochem Educ 27:177–180. doi:10.1016/S0307-4412(98)00302-1
Suh SO, Marshall CJ, Hugh JVM, Blackwell M (2003) Wood ingestion by passalid beetles in the presence of xylose-fermenting gut yeasts. Mol Ecol 12:3137–3145. doi:10.1046/j.1365-294X.2003.01973.x
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Yarrow D (1998) Methods for the isolation and identification of yeasts. In: Kurtzman CP, Fell JW (eds) The Yeasts, a taxonomic study, 4th edn. Elsevier, Amsterdam, pp 77–100
Zhu JY, Pan X, Zalesny RS Jr (2010) Pretreatment of woody biomass for biofuel production: energy efficiency, technologies, and recalcitrance. Appl Microbiol Biotechnol 87:847–857. doi:10.1007/s00253-010-2654-8
Acknowledgments
This work was funded by Conselho Nacional de Desenvolvimento Cientifico e Tecnológico (CNPq), Fundação do Amparo a Pesquisa do Estado de Minas Gerais (FAPEMIG), and the Natural Science and Engineering Research Council of Canada (M.A.L.). The authors are grateful to Acir Moreno for assistance during the field collection in the Private Natural Heritage Reserves of Woodstock and Bello & Kerida.
Conflict of interest
None
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Santos, R.O., Cadete, R.M., Badotti, F. et al. Candida queiroziae sp. nov., a cellobiose-fermenting yeast species isolated from rotting wood in Atlantic Rain Forest. Antonie van Leeuwenhoek 99, 635–642 (2011). https://doi.org/10.1007/s10482-010-9536-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-010-9536-z