Abstract
During embryonic development, the lymphatic system emerges by transdifferentiation from the cardinal vein. Although lymphatic and blood vasculature share a close molecular and developmental relationship, they display distinct features and functions. However, even after terminal differentiation, transitions between blood endothelial cells (BEC) and lymphatic endothelial cells (LEC) have been reported. Since phenotypic plasticity and cellular differentiation processes frequently involve epigenetic mechanisms, we hypothesized that DNA methylation might play a role in regulating cell type-specific expression in endothelial cells. By analyzing global gene expression and methylation patterns of primary human dermal LEC and BEC, we identified a highly significant set of genes, which were differentially methylated and expressed. Pathway analyses of the differentially methylated and upregulated genes in LEC revealed involvement in developmental and transdifferentiation processes. We further identified a set of novel genes, which might be implicated in regulating BEC-LEC plasticity and could serve as therapeutic targets and/or biomarkers in vascular diseases associated with alterations in the endothelial phenotype.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The human cutaneous vasculature consists of two tubular networks, the blood and the lymphatic system, which display different but interdependent functions. The lymphatic vasculature is responsible for regulating tissue fluid homeostasis, immune surveillance, and absorption, thereby mediating the uptake of protein-rich fluid from the interstitium. Its capillaries are composed of a single layer of overlapping endothelial cells and lack pericytes. Furthermore, lymphatic capillaries are thin-walled, blind-ended, and connected to the extracellular matrix by anchoring filaments. In contrast to the lymphatic system, the blood vasculature represents a closed circulating system, and mediates the exchange of gases (O2, CO2) and nutrients. Blood capillaries display continuous interendothelial junctions, are surrounded by a continuous basement membrane, and are covered by pericytes [1–4].
Although both, blood vascular and lymphatic endothelium, have distinct functions, they share a close molecular and developmental relationship. During embryonic development, endothelial progenitors called angioblasts differentiate from mesodermal cells and form a primitive vascular network [4, 5]. After determination of arteries and veins, the first lymphatic endothelial cells (LEC) are formed via transdifferentiation from the cardinal vein [1, 4, 6, 7]. Lymphatic development occurs in a stepwise process involving the restricted expression of the prospero-related homebox 1 transcription factor (Prox-1) [8], which commits Prox-1 positive cells to the lymphatic lineage [2, 7]. Upon terminal differentiation, LEC express cell type-characteristic markers including Prox-1 [8] and the transmembrane glycoprotein podoplanin (PDPN) [9]. Also vascular endothelial growth factor receptor 3 (VEGFR-3), the tyrosine kinase receptor for vascular endothelial growth factors C and D (VEGF-C and VEGF-D), becomes largely restricted to the lymphatic endothelium in adult tissue [10]. Interestingly, even after embryonic differentiation, endothelial cells keep the capacity to change their endothelial lineage-specific identity. In vivo studies reported that continued expression of Prox-1 is required for lymphatic identity in mice because reduced levels of Prox-1 resulted in lymphatic vascular defects and adoption of blood endothelial-specific markers [11]. Recently, a transdifferentiation between cultured LEC and blood endothelial cells (BEC) has been reported in response to in vitro extracellular matrix environments [12]. In agreement with this observation, other studies showed that treatment of BEC with interleukin 3 (IL-3) results in expression of lymphatic-specific genes [13]. Endothelial plasticity has also been described in the context of pathobiological processes in vivo. During skin inflammation blood vessels change their phenotype by expressing lymphatic-specific proteins [14]. Importantly the lymphatic marker VEGFR-3 could be detected on blood vessels in tumors and it’s blocking decreased angiogenesis [15]. These data show that even terminally differentiated endothelial cells retain a certain degree of phenotypic plasticity, enabling transdifferentiation processes between the blood vascular and lymphatic system under specific conditions.
One important mechanism, which defines cellular physiology and mediates phenotypic plasticity, is the DNA methylation of cytosines [16, 17]. During the differentiation process, cells acquire a specific DNA methylation pattern reflecting the tissue-specific cellular phenotype [18–22]. Epigenetic mechanisms also play a role during development of the vasculature [23]. In this context, key vascular genes, such as endothelial nitric oxide synthase 3 (eNOS), have been shown to be regulated by DNA methylation [24]. Additionally, gene expression of specific vascular endothelial growth factor receptors 1 and 2 (VEGFR-1 and VEGFR-2) is regulated by promoter DNA methylation in various cancer tissues [25] and activation of the VEGF-C gene by DNA demethylation probably facilitates the metastasis of cancer cells by inducing lymphangiogenesis [26]. These observations indicate that DNA methylation is relevant for regulating the expression of well-known genes that play important roles in mediating vascular-specific functions. Therefore we hypothesized that DNA methylation might be functionally involved in maintaining endothelial phenotypic plasticity and might furthermore regulate genes which are directly connected to transdifferentiation processes in several diseases. To address this point, we aimed to identify endothelial-specific genes which are regulated by DNA methylation and could serve as biomarkers in diseases involving pathological vessel conditions. To our knowledge, no genome-wide comparative DNA methylation analysis of LEC and BEC has been conducted, yet. In order to identify genes that regulate endothelial cell type-specific characteristics, we combined state-of-the-art methylation profiling with global gene expression analyses. Our results revealed that DNA methylation is involved in endothelial lineage-specific gene expression, indicating that epigenetic mechanisms are implicated in regulating LEC-BEC plasticity.
Materials and methods
Ethics statement
The recommendations of the current version of the Declaration of Helsinki as well as the international guidelines (f.e. FDA Regulations, GEP and AWB guidelines) were observed. All participants in the contributing study provided written, informed consent. Punch biopsies were obtained through a clinical study approved by the Freiburger Independent Ethics Committee (010/1973). Written, informed patient consents were given from all volunteers.
Tissue specimens
Full-thickness skin samples were obtained either from a clinical study (punch biopsies) or from material derived from plastic surgeries. In the clinical study, punch biopsy samples were taken under local anesthesia from the upper, sun-protected buttock of 10 female volunteers by Bioskin GmbH (Hamburg, Germany). 8 punch biopsies of 6 mm in diameter were isolated from each participant and transferred into ice-cold Microvascular Endothelial Cell Growth Medium-2 (EGM-2 MV, Lonza, Walkersville, USA) immediately after removal. Sun-protected breast skin biopsies were obtained from different female volunteers after plastic surgery. The size of skin samples collected from plastic surgery was comparable to the size of 8 combined punch biopsies. As described above, breast skin biopsies were also transferred into ice-cold EGM-2 MV immediately after removal. All female volunteers were aged between 22 and 73 years.
Isolation and cultivation of human dermal blood and lymphatic endothelial cells
The eight skin samples of each volunteer were pooled and then disinfected using 1× Hanks’ Balanced Salt Solution (HBSS) with 4 % penicillin/streptomycin (pen/strep) and 0.4 % fungizone (all Invitrogen, Karlsruhe, Germany). After dispase II treatment (2 U/ml, Roche, Mannheim, Germany) with 1 % pen/strep and 1.6 % gentamycine (both Invitrogen, Karlsruhe, Germany) for 4 h at 37 °C, epidermis and dermis were separated. Dermal cells were released by cutting and incubation in 0.25 % trypsin/EDTA (Gibco, Karlsruhe, Germany) at 37 °C. The reaction was stopped with 10 % fetale bovine serum (FBS, PAA, Linz, Austria) in phosphate buffered saline (PBS, Lonza, Walkersville, USA) and the cell suspension was subsequently filtered using a 70 μm cell strainer (BD Biosciences, Heidelberg, Germany). After centrifugation, the pellet (dermis fraction) was resuspended in 1× HBSS with 5 % FBS and cells were incubated with an immunomagnetic bead-conjugated anti-human cluster of differentiation 31 (CD31) antibody (Invitrogen, Dynal, Oslo, Norway) at 4 °C for 30 min using the MACSmix Tube Rotator (Miltenyi Biotec, Bergisch Gladbach, Germany) to isolate primary endothelial cells. The remaining CD31-negative cells were discarded and CD31-positive cells were washed three times prior to resuspension in EGM-2 MV medium (Composition of EGM-2 MV: 500 ml Endothelial Cell Basal Medium-2 (EBM-2) supplemented with 25 ml FBS, 0.2 ml hydrocortisone, 2 ml human fibroblast growth factor, 0.5 ml vascular endothelial growth factor A, 0.5 ml R3-insulin like growth factor 1, 0.5 ml ascorbic acid, 0.5 ml human endothelial growth factor, and 0.5 ml gentamycine; all Lonza, Walkersville, USA). Cells were plated into 25 cm2 tissue culture dishes (Greiner Bio-One, Frickenhausen, Germany) and maintained at 37 °C in a humidified atmosphere containing 5 % CO2. When the cultures reached about 80 % confluence, cells were washed with Hepes Buffered Saline Solution (BSS) (PromoCell, Heidelberg, Germany) and trypsinized with 0.25 % trypsin/EDTA. After 2 min incubation at 37 °C, the reaction was stopped with 0.05 % trypsin inhibitor containing 0.1 % bovine serum albumin (BSA, PromoCell, Heidelberg, Germany). Following blocking in PBS containing 1 % BSA, cells were incubated with primary mouse anti-human D2-40 antibody (Signet, Dedham, USA) which recognizes the lymphatic-specific glycoprotein podoplanin (PDPN). Cells were washed and incubated with a secondary goat anti-mouse antibody, conjugated to magnetic beads (Invitrogen Dynal, Oslo, Norway). Following 30 min incubation at 4 °C in a rotator, PDPN-positive cells (lymphatic endothelial cells, LEC) were washed with PBS and seeded into tissue culture dishes. PDPN-negative cells (blood endothelial cells, BEC) were cleaned up once again using PDPN antibody and magnetic bead-conjugated secondary antibody. For all our studies cells were used up to the fourth passage.
Flow cytometry analysis of blood and lymphatic endothelial cells
Cell surface expression of CD31 and PDPN on endothelial cells isolated from each volunteer was analyzed separately by flow cytometry. Adherent endothelial cells were trypsinized, washed, and after resuspension in cell staining buffer (Biolegend, San Diego, USA), cells were incubated on ice in buffer containing Alexa Fluor 488-labeled anti-human CD31 antibody and Alexa Fluor 647-conjugated anti-human PDPN antibody (both Biolegend, San Diego, USA) for 45 min. Dual staining was performed to obtain the information regarding the expression of both proteins on the specific cell population. Following washing, cells were analyzed by flow cytometry using FACSCanto instrument (BD Biosciences, Heidelberg, Germany) and FlowJo software (TreeStar, Ashland, USA).
DNA and RNA isolation
A total of 300.000 endothelial cells between passage 0 and 4 were trypsinized, pelleted and total cellular RNA or DNA were isolated using the RNeasy Mini Kit and the QIAamp DNA Investigator Kit, respectively (both Qiagen, Hilden, Germany) according to the manufacturer’s instructions. DNA and RNA were quantified using the NanoDrop ND-1000 instrument (Peqlab Biotechnologies, Wilmington, USA). Cell populations isolated from each volunteer were separately analyzed in subsequent experiments and arrays.
Quantitative RT-PCR
The TaqMan Gene Expression Assay (Applied Biosystems, Foster City, USA) was used to further analyze the cell purity of the isolated LEC and BEC as well as for validation of the microarray expression data. RNA of the specific endothelial cell population (BEC or LEC) of each volunteer was reverse-transcribed into cDNA and PCR reactions were carried out in duplicates as recommended by the manufacturer. Quantitative RT-PCR of the genes von Willebrand factor (vWF, Assay ID Hs00169795_m1), podoplanin (PDPN, Assay ID Hs00366766_m1), cluster of differentiation 31 (CD31, Assay ID Hs00169777_m1), prospero-related homebox 1 (Prox-1, Assay ID Hs00896294_m1), homebox A5 (HOXA5, Assay ID Hs00430330_m1), trefoil factor 3 (TFF3, Assay ID Hs00902278_m1), and XIAP associated factor 1 (XAF1, Assay ID Hs01550142_m1) (all Applied Biosystems, Foster City, USA) was performed using the Applied Biosystems 7900HT Fast Real Time PCR System. CT values were calculated by the RQ Manager software 1.2. Target gene expression was normalized on expression of the endogenous control GAPDH (Assay ID Hs99999905_m1) and calculations were performed using the comparative CT method [27].
Array-based DNA methylation profiling
The analysis of the DNA methylation pattern in LEC and BEC of each volunteer was performed using the HumanMethylation450 BeadChip (Illumina, San Diego, USA) [28]. The 450K DNA Methylation array interrogates 485,764 cytosine positions of the human genome whereby 482,421 positions represent CpG (cytosine-phospate-guanine) dinucleotides. 200,339 CpG sites are located in proximal promoters, defined as the sum of CpG sites located within 200 or 1,500 bp upstream of the transcription start site (TSS), the 5′ UTR and first exon. CpGs located in the gene body and 3′ UTR regions were also analyzed. Bisulfite conversion of 500 ng DNA of each sample was carried out according to the manufacturer’s recommendations using the EZ-96 DNA Methylation Kit (Zymo Research Corporation, Irvine, USA). Bisulfite treated DNA was whole-genome amplified, enzymatically fragmented, and hybridized to the HumanMethylation450 BeadChip.
Array-based gene expression analysis
In order to identify differentially expressed genes in LEC versus BEC, samples were analyzed using one-color Whole Human Genome Microarray 4 × 44K chips (G4112F9) from Agilent Technologies which were carried out by Miltenyi Biotech GmbH (Bergisch Gladbach, Germany).
Data deposition
The microarray and methylation data is available in the Gene Expression Omnibus database (http://www.ncbi.nlm.nih.gov/geo/), under accession number GSE34487 (expression array: GSE32709, methylation array GSE34486).
DNA bisulfite sequencing
Deep bisulfite sequencing was carried out using equimolar amplicon pools, as described previously [29]. The following sequence-specific primers were used for amplifying deaminated genomic DNA HOXA5_for, TTATAATGGGTTGTAATTTTAATT; HOXA5_1_rev, AACATATACTTAATTCCCTCCTAC; XAF1_for, TTTTTTTGTAGGGGAGGATTAGAA; XAF1_1_rev, TACAACATAACCAACCCCTACTATC; TFF3_1_for, TAAGGAATTTTTGTGTTTTAGGAGTT; TFF3_1_rev, ATTCAACCCCCACTATTTTAACAA. PCR products were purified using the QIAquick gel extraction Kit (Qiagen, Hilden, Germany). Sequence analysis was conducted by LGC Genomics (Berlin, Germany).
Statistical analyses
For array-based DNA methylation profiling Infinium GenomeStudio software (methylation module v1.8) was used to read the scanner files and to output text files that were suitable for downstream analysis with the R statistical software. Probes with a detection p value >0.01 were excluded from the analysis resulting in detection rates of 99.7–99.9 % for the different samples. To identify genes differentially methylated in human dermal BEC versus LEC the limma package [30] for the R statistical software was used on quantile-normalized data. For statistical analysis and assessing differential methylation, limma uses an empirical Bayes method to moderate the standard errors of the estimated log-fold changes. p values for comparisons between different datasets were statistically adjusted using the Benjamini-Hochberg correction [31]. Averages of methylation values, referred to as average beta values (AVB) were recorded for each sample. One BEC sample was excluded from the analysis due to low-quality data.
For the array-based gene expression analyses Agilent Feature Extraction Software (AFE) was used to read out the microarray image files. The conversion of the data was performed with the R statistical software and its Bioconductor packages Agi4x44PreProcess and limma [30]. After reading the files into R, all data were background-corrected. All arrays were quantile-normalized and transformed to log-2 scale, which enabled comparison of samples loaded on different arrays. After normalization using the AgiPreProcess package a set of quality control steps was performed to filter low-quality probes, as indicated by quality flags set by the AFE. To identify genes differentially expressed in BEC versus LEC, the limma package for the R statistical software was used to compute a moderated t test. Raw values were finally adjusted using the Benjamini-Hochberg procedure [31]. Functional categories and pathway analyses of differentially expressed genes were generated by the use of IPA (Ingenuity® Systems, http://www.ingenuity.com).
For the quantitative RT-PCR analyses of the genes vWF, CD31, Prox-1, and PDPN as well as for HOXA5, TFF3, and XAF1 the GraphPad Software Prism 5 (La Jolla, USA) was used. Normal distribution was checked by means of Shapiro–Wilk’s test and unpaired t test was performed. A significance level of 0.05 (alpha) was chosen for statistical analyses, based on two-sided hypothesis testing.
Results
Isolation and characterization of human dermal endothelial cells
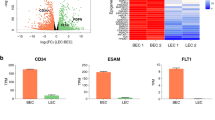
Advanced isolation methods using cell type-specific markers for blood (BEC) and lymphatic endothelial cells (LEC) and the availability of array-based technologies for global methylation and gene expression analyses allow to specifically analyzing lineage-specific features of the vascular system. For comparative array-based analysis, isolated human dermal endothelial cells were separated into LEC and BEC populations by immunomagnetic labeling. To determine whether endothelial cells were contaminated with fibroblasts, visual scoring as well as analyses by flow cytometry were used (see Online Resource 1). Only endothelial cell populations having a purity rate higher than 90 % were used for subsequent analyses. One representative FACS diagram is shown in Fig. 1a. As additional quality control, qRT-PCR was performed for endothelial-specific markers including cluster of differentiation 31 (CD31) and von Willebrand factor (vWF) as well as for the lymphatic-specific markers podoplanin (PDPN) and the prospero-related homeobox 1 transcription factor (Prox-1) (Fig. 1b) [32, 33]. Both cell populations expressed these marker genes in a cell type-specific pattern confirming the distinct vascular origin of both cell populations.
Isolation and characterization of endothelial cells from human skin biopsies a Endothelial cells (EC) were isolated from punch biopsies or skin samples derived from cosmetic surgery. After immunomagnetic purification, the cell surface expression of endothelial-specific marker cluster of differentiation 31 (CD31) and lymphatic-specific marker podoplanin (PDPN) was analyzed using flow cytometry. LEC, lymphatic EC; BEC, blood vascular EC. b Quantitative RT-PCR of von Willebrand factor (vWF), cluster of differentiation 31 (CD31), prospero-related homebox 1 (Prox-1), and podoplanin (PDPN). Target gene expression was normalized to GAPDH expression used as endogenous control. The horizontal black lines denote medians and whiskers the 2.5th and 97.5th percentiles. Significant differences between LEC and BEC are indicated by asterisks. *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001 (unpaired t test). n (EC) = 8, n (LEC) = 10, n (BEC) = 7
LEC and BEC show distinct methylation profiles
The global methylation patterns of 10 primary LEC and 7 primary BEC populations were analyzed using Illumina Infinium Human Methylation 450K arrays. This high-throughput Bead Chip technology allows to determine the genome-wide methylation status of more than 450,000 CpG (cytosine-phospate-guanine) dinucleotides per sample [28]. Prior to detailed data analysis, raw data were preprocessed and quantile-normalized (see “Materials and methods” section). To validate the Infinium 450K methylation data, we used 454 bisulfite sequencing and analyzed the methylation status of four arbitrarily chosen CpG loci which were represented by approximately 0, 50, and 100 % of methylated alleles, respectively (see Online Resource 2). Methylation levels obtained by 454 sequencing were highly similar to the array-predicted methylation value indicating that the array produced reliable results. Hierarchical clustering of the 16 LEC and BEC samples revealed distinct endothelial cell type-specific methylation patterns, as LEC and BEC form separate clusters in the dendrogram (Fig. 2a). Next, we analyzed the distribution of methylation values in LEC and BEC, using Kernel density plotting (Fig. 2b). Both endothelial cell types showed a nearly identical bimodal distribution of averaged methylation values (AVB) with a low methylation peak near 0.1 and a second peak around 0.8–0.9 represented by highly methylated loci. To investigate whether the DNA methylation pattern of single CpG loci differs between LEC and BEC, we subtracted the DNA methylation data of both cell types to generate differential methylation values. A total of 32,660 gene loci represented by 8,138 genes showed a significantly differential methylation (Benjamini-Hochberg adjusted p < 0.01) between both endothelial cell types as indicated in the Volcano plot (Fig. 2c). To illustrate the distribution of average methylation values for these hypo- and hypermethylated CpG loci in LEC and BEC, Kernel density plots are shown in Fig. 2d. The upper panel represents average methylation values of hypo- and hypermethylated loci in LEC versus BEC, corresponding to the green- and blue-colored dots in the scatter plot of Fig. 2c. In contrast to the flattened curve represented by hypomethylated CpGs, a shift towards hypermethylation resulted in a distinct peak composed of highly methylated CpG loci (AVB of approximately 0.8). The same strong gain in methylation was observed for hypermethylated loci in both, LEC (upper panel) and BEC (lower panel), and already suggests a functional relevance of these marks for transcriptional repression of the corresponding genes.
Identification of differentially methylated CpGs in blood and lymphatic endothelial cells a Hierarchical clustering of the quantile normalized methylation array data (482,421 CpG dinucleotides) of lymphatic endothelial cells (LEC) versus blood endothelial cells (BEC). Numbers indicate different volunteers. b Kernel density plot representing the averaged methylation values (AVB) distributions of human endothelial cells. Since BEC show a nearly identical distribution as LEC, only one plot is shown. c The Volcano plot represents Benjamini Hochberg adjusted p values versus mean methylation differences between LEC and BEC. Green dots indicate hypomethylated CpG loci, blue dots indicate hypermethylated CpG loci, and grey dots indicate non-significant (Benjamini Hochberg adj. p value ≥0.01) methylation changes in LEC. ΔAVB, methylation changes. d Average methylation distributions of significantly differentially methylated CpG loci in LEC versus BEC are illustrated by Kernel density plots. Green lines represent the distribution of methylation values in 16,834 hypomethylated CpGs in LEC (corresponding to hypermethylated CpGs in BEC), blue lines indicate the distribution of methylation values in 15,826 hypermethylated CpG loci in LEC (corresponding to hypomethylated CpG loci in BEC)
Differential DNA methylation correlates with gene expression changes in endothelial cells
We also determined the mRNA transcription profile of LEC and BEC of each volunteer using Agilent Whole Human Genome Microarrays, which allow the analysis of 45,015 gene probes. The processed data comprised 26,519 highly significant gene probes, which corresponded to 13,406 genes in human endothelial cells. Of the 13,406 genes, 596 genes were differentially expressed between LEC and BEC, as visualized by a Volcano scatter plot (Fig. 3a). These genes corresponded to 303 up- and 293 downregulated genes in LEC compared to BEC (Benjamini Hochberg adjusted p value <0.01). To investigate the correlation between DNA methylation and gene expression, we combined Agilent gene expression data and Infinium methylation data. For initial analyses, we decided to use the complete set of differentially expressed or methylated CpGs (Benjamini-Hochberg adjusted p < 0.01), without specifying any minimum threshold for expression and methylation change. The rationale behind this decision was to derive more general conclusions from our data analysis and to prevent any arbitrary shift of the data set caused by stringent selection criteria at this time point. In detail, we compared the 596 differentially expressed genes with 8,138 genes represented by 32,660 differentially methylated CpGs on the Infinium 450K methylation array. A total of 375 genes showed significant changes in gene expression and DNA methylation, as indicated by the overlap in the Venn diagram (Fig. 3b), suggesting that a great proportion (>60 %) of differentially expressed genes is regulated by epigenetic mechanisms. To investigate the correlation between DNA methylation and gene expression in greater detail, we separated differentially expressed genes in 193 upregulated genes (represented by 1,109 CpG loci) and 182 downregulated genes (represented 711 CpG loci) in LEC versus BEC and analyzed the methylation changes at the corresponding loci. The box plot in Fig. 3c indicates that upregulated genes were generally demethylated whereas the methylation level of downregulated genes increased. To analyze the correlation between methylation status and expression levels in primary human endothelial cells more extensively, the methylation changes were plotted according to their location in different genomic regions (Fig. 3d). In regions close to the transcription start site (TSS) (TSS200, TSS1500 and 5′-UTR) and the first exon of genes, upregulated genes in LEC were less methylated than downregulated genes. CpG loci, which were located in a region covering 0–200 bp upstream of the TSS and the first exon of genes, showed the greatest methylation changes between up- and downregulated genes with upregulated genes being demethylated and downregulated genes being hypermethylated in LEC versus BEC. Similar to that, we observed a tendency for gene body demethylation resulting in higher gene expression. Most of the differentially methylated genes showed significant methylation changes in this region (634 hypo- and 378 hypermethylated CpG loci in LEC versus BEC). In contrast, the 3′-UTR appeared to be hypermethylated in upregulated genes and hypomethylated in downregulated genes. To validate the correlation between methylation and expression we chose three genes which showed a differential methylation in the TSS200 region between LEC and BEC. 454 bisulfite sequencing was performed using equimolar sample pools of 6 BEC and 6 LEC populations (n = 12) and the results confirmed the distinct methylation pattern and a homogenous methylation across the promoter region (Online Resource 3a). Quantitative RT-PCR analyses of 10 LEC and 6 BEC populations validated the differential gene expression levels predicted by microarray analysis (Online Resource 3b). Overall, our validation experiments confirmed the observation that promoter hypermethylation of LEC correlated with a decrease in gene expression whereas hypomethylation caused an increased expression of the corresponding gene.
DNA methylation correlates with gene expression in human endothelial cells a The Volcano scatter plot represents Benjamini Hochberg adjusted p values versus gene expression changes in lymphatic (LEC) versus blood endothelial cells (BEC). Red dots indicate upregulated genes, blue dots indicate downregulated genes in LEC and grey dots indicate non-significant (Benjamini Hochberg adj. p value ≥0.01) expression changes. b The overlap of differentially methylated and expressed genes is shown in the Venn diagram. c Box plots indicate the correlation of methylation and gene expression in LEC versus BEC. Red boxes represent upregulated genes in LEC, blue boxes represent downregulated genes in LEC. The numbers above indicate the numbers of CpG loci used for generation of the respective boxplot. The horizontal black lines denote medians, notches the standard errors, boxes the interquartile range, and whiskers the 2.5th and 97.5th percentiles. d The boxplots show methylation changes for the indicated categories (TSS1500, TSS200, 5′-UTR, 1st Exon, Body, and 3′-UTR). Red boxes represent upregulated genes, blue boxes represent downregulated genes in LEC. The numbers above indicate the numbers of CpG loci used for generation of the respective boxplot. The horizontal black lines denote medians, notches the standard errors, boxes the interquartile range, and whiskers the 2.5th and 97.5th percentiles. TSS, transcription start site
Upregulated and differentially methylated genes in LEC are associated with developmental and transdifferentiation processes
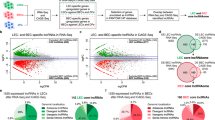
To further characterize our dataset, we selected genes showing the highest differential methylation and gene expression change using previously published thresholds (absolute methylation changes ΔAVB ≥ 0.15, twofold change for gene expression) [34, 35]. We obtained a set of 24,523 differentially methylated CpG loci, represented by 6,663 genes. Out of the 596 significantly differentially expressed genes, 336 genes showed expression changes of at least two-fold. The Venn diagram (Fig. 4a) indicates an overlap of 215 genes. To further analyze the biological relevance of these 107 up- and 108 downregulated genes in LEC, network and functional analysis were generated through the use of IPA (Ingenuity Pathway Analysis, Ingenuity® Systems, http://www.ingenuity.com). Downregulated genes in LEC were enriched in functional categories such as “cancer, genetic disorder, gastrointestinal disease, inflammatory disease, and dermatological diseases and disorders” (Fig. 4b). Interestingly, diseases characterized by vascular malfunctions or abnormal angiogenesis such as psoriasis, coronary artery disease, and inflammatory bowel diseases (i.e. Crohn’s disease) formed part of the category “genetic disorders”. Upregulated genes in LEC were enriched for functional categories associated with developmental processes such as “tissue, embryonic, organ, and organismal development and cellular movement” (Fig. 4b). These functions were further specified by subcategories such as “development of organ, growth of embryonic tissue, development of body axis, cardiogenesis, and lymphangiogenesis” in the category “embryonic development” indicating that DNA methylation is also involved in regulating normal vascular developmental processes. Three of the five most significant functional categories in LEC built the top-score network “tissue development, cellular movement, and embryonic development” (Fig. 4c) which is drawn in Fig. 4d. Interestingly, it includes a number of genes that play important roles in lymphatic vessel development and function such as the vascular endothelial growth factor receptor-3 (VEGFR-3) [10], its co-receptor Neuropilin 2 (Nrp-2), which is also largely limited to the lymphatic endothelium [36, 37], and the LEC-specific marker for differentiation, Prox-1 [8]. Another important transcription factor is ETS-domain protein (SRF accessory protein 2) (abbreviated Elk 3) because mutant mice lacking this protein suffer from dilated lymphatic vessels [38]. Other genes included in the top-score network and known to be overexpressed in LEC versus BEC were basic leucine zipper transcription factor (BATF) [39], cluster of differentiation 36 (CD36) [39], v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) (MAF) [39], T-box 15 (TBX-15) [39], interleukin 7 (IL-7) [40–42], and the extracellular matrix protein reelin (RELN) [41, 42]. In addition to these well-established genes, we also identified novel genes as part of the same network such as the fatty-acid binding protein 5 (FABP-5) that showed differential expression and methylation in LEC compared to BEC. FABP-5 has been implicated to play a role in psoriasis [43, 44] and showed hypomethylation in the TSS1500 island region and upregulation in LEC versus BEC. Another example is the scaffold protein rhotekin (RTKN) which affects cellular apoptosis [45]. The RTKN TSS1500 island region contained significantly hypomethylated CpGs that correlated with an upregulation in LEC. A list of all differentially expressed genes included in our top network and corresponding methylation data are shown in Online Resource 4. To analyze if our selection criteria (twofold change for expression analysis and ΔAVB ≥ 0.15 for methylation analysis) affected the enrichment for functional categories, we also performed pathway analyses using the whole set of 596 genes. The results revealed very similar functions pointing out that the majority of the selected genes are enriched in the same networks (Online Resource 5).
Pathway analysis of differentially methylated and expressed genes in lymphatic compared to blood endothelial cells a The Venn diagram shows the overlap of genes with a differential expression of at least twofold and genes corresponding to CpG loci with methylation changes (ΔAVB) equal or greater than 0.15. LEC lymphatic endothelial cells, BEC blood endothelial cells. b Top five functional categories for either up- or downregulated genes in LEC versus BEC obtained by Ingenuity Pathway Analysis (IPA). c Top three associated networks for genes upregulated in LEC derived from IPA. d Top network built by upregulated and differentially methylated genes in LEC. The more intense the redness, the greater the expression changes between LEC and BEC. AEBP1, AE binding protein 1; BATF, basic leucine zipper transcription factor, ATF-like; CD36, cluster of differentiation 36; ELK3, ETS-domain protein (SRF accessory protein 2); FABP5 fatty acid binding protein 5 (psoriasis-associated), GATA3 GATA binding protein 3, IL-7 Interleukin 7, ITGA10 integrin, alpha 10, MAF v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian), MRC2 mannose receptor C type 2, NRP-2 Neuropilin 2, OTX1 orthodenticle homeobox 1, PCSK6 proprotein convertase subtilisin/kexin type 6, PITX2 paired-like homeodomain 2, Prox-1 prospero-related homeobox 1, RELN reelin, RTKN rhotekin, SLC2A12 solute carrier family 2 (facilitated glucose transporter), member 12, TBX-1 T-box 1, TBX-15 T-box 15, TFPI tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor), VAV3 vav 3 guanine nucleotide exchange factor, VEGFR3 vascular endothelial growth factor receptor 3
In summary, our results indicate that DNA methylation regulates endothelial cell type-specific gene expression and thereby contributes to the characteristics of both endothelial systems. Our data further suggest that genes differentially methylated and expressed in LEC are implicated in developmental and transdifferentiation processes.
Discussion
Lymphatic and blood vessels are involved in many physiological processes during tissue development and regeneration. However, they also play a role in the development of a variety of pathological conditions. For example, lymphatic vessel formation has been shown to promote tumor metastasis whereas impairment of its function leads to reduced lymphatic transport of fluids resulting in lymphedema [42, 46]. In contrast, the formation of new blood vessels is essential for tumor growth and abnormalities in blood vessel function lead to artheriosclerosis as well as skin diseases including psoriasis, ulcers, or allergic dermatitis [46, 47]. Interestingly the adoption of a lymphatic-like phenotype by blood endothelial cells (BEC) has been observed in some diseases [15, 48]. Our results indicate that epigenetic processes play an important role in specifying the vascular lineage of endothelial cells and may further be involved in mediating endothelial plasticity under pathologic conditions.
The effect of DNA methylation on a gene depends on the position of methylated CpGs (cytosine-phospate-guanine) relative to the transcription start site (TSS) [49]. Hypermethylation of promoter regions correlates with gene expression whereas hypomethylation generally allows active transcription of the corresponding gene [50]. However, the function of intragenic DNA methylation is still controversially discussed and may be associated with active transcription or repression of the corresponding gene [22, 51–54]. Therefore we comprehensively analyzed whether the position of differentially methylated CpGs affects the expression of the corresponding gene. Using genome-wide methylation and gene expression analyses, we identified a strong correlation between hypermethylation of promoter regions (i.e. localized 1,500 bp around the TSS) and transcriptional repression which is in agreement with previous findings [50, 55, 56]. Interestingly, we also observed a substantial number of differentially methylated CpGs in gene bodies. For the first exon, we found a strong correlation between hypomethylation and active gene expression, which was slightly reduced for the entire gene body region. Several groups have provided evidence linking gene body methylation and transcription by observing positive as well as negative correlations between gene body methylation and active gene expression [22, 51–53, 57]. Our data indicate that in our model system gene body methylation negatively correlates with gene expression levels. The functional relevance of intragenic DNA methylation in human endothelial cells might be the induction of a more closed chromatin state [52] or the blocking of transcriptional elongation [52, 58, 59]. In contrast to all other genomic regions analyzed, methylation of CpG dinucleotides in the 3′UTR correlated with repression of the corresponding genes in endothelial cells. It has been proposed that methylation in the 3′UTR may play a role in suppression of antisense transcripts, regulation of polyadenylation, and termination of transcription, respectively [51]. However, further experiments will be required to clarify the precise function of DNA methylation beyond promoter regions in endothelial cells.
Combined pathway analysis of differentially methylated and upregulated genes in primary lymphatic endothelial cells (LEC) revealed functional categories associated with physiological system development. These findings suggest that epigenetic mechanisms are involved in regulating developmental processes of the vascular system. These might include the formation of new vessels during development or regeneration as well as the regulation of transdifferentiation of LEC from the venous blood vascular system during embryogenesis. Additionally, our top-score network derived from epigenetically regulated genes included lymphatic-specific genes as well as genes that have been shown to be involved in transdifferentiation processes during pathological conditions. One of these genes, Prox-1 (prospero-related homebox 1), is a well-known homebox transcription factor involved in lymphatic lineage commitment [8]. Mice lacking Prox-1 fail to develop a lymphatic vasculature during embryonic development [8] and Prox-1 expression in BEC leads to reprogramming and adoption of a lymphatic-like phenotype [40]. It has also been reported that stimulation of cultured BEC with the lipid growth factor lysophosphatidic acid [60] or the Kaposi sarcoma-associated herpesvirus [48] induces expression of Prox-1. The latter stimulation leads to a comprehensive reprogramming of BEC to a lymphatic phenotype resulting in induction of about 70 % of lymphatic lineage-specific genes including the upregulation of interleukin 7 (IL-7) [40, 41] [42, 48], v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) (MAF) [39, 48], reelin (RELN) [48] [42], and the cluster of differentiation 36 (CD36) [39, 48]. Using genome-wide methylation and expression arrays, we confirmed that all of these markers were upregulated in LEC compared to BEC. In addition, we showed that all of these genes are differentially methylated indicating that epigenetic mechanisms are involved in their regulation. Especially IL-7 showed a significant hypomethylation of all analyzed CpGs in the TSS1500 region. IL-7 is known as a lymphangiogenic growth factor, which increases the expression of the lymphatic markers lymphatic vessel endothelial hyaluronan receptor 1 (LYVE-1), podoplanin (PDPN), and Prox-1 in human endothelial cells [61]. Even in breast cancer cells IL-7 induces lymphangiogenic properties which might be involved in lymphatic spread of tumors [62]. The vascular endothelial growth factor receptor 3 (VEGFR-3) is another important regulator of lymphatic function included in our top-network. The corresponding gene was hypomethylated and upregulated in LEC gene bodies. During early stages of development, VEGFR-3 is expressed in all endothelial cells and becomes largely restricted to the lymphatic endothelium after final specification of blood and lymphatic lineages [10]. Notably, VEGFR-3 expression has also been observed in tumor neovessels and its inhibition suppressed angiogenic sprouting, branching, and vessel density [15, 63]. Knockout of its co-receptor Neuropilin 2 (Nrp-2) in mice also causes abnormal lymphatic development [37]. Similarly, we also found the Nrp-2 gene to be differentially methylated and expressed in the lymphatic and blood vascular system (i.e. hypomethylated and upregulated in LEC).
These results provide novel insights into endothelial plasticity [64] because our observations indicate that many well-known lymphatic marker genes involved in the regulation of endothelial plasticity during development and/or progression of diseases are likely regulated by DNA methylation. Additionally, we also identified novel genes, which may play a role in the same context since they had been integrated to the same top-network derived from Ingenuity analysis and showed differential methylation and expression comparing both vascular systems. Two interesting candidates are the fatty-acid binding protein 5 (FABP-5) and the apoptose-associated protein rhotekin. FABP-5 increases VEGF-A (vascular endothelial growth factor A) expression [65] and has been shown to be upregulated in psoriasis [43, 44] and tumors [66–68]. Since VEGF-A stimulates LEC proliferation [69] and its overexpression in transgenic mice results in chronic skin inflammation with enlargement of lymphatic vessels and lymphatic hyperplasia [70], the maintenance of DNA hypermethylation of the FABP-5 gene in BEC could prevent inflammatory diseases. However, additional analyses are required to further specify the mechanism of epigenetic regulation for these candidate genes and to functionally characterize their potential involvement in vascular development, regeneration or diseases.
In summary, our data provide evidence that endothelial cell type-specific gene expression in LEC and BEC is regulated by DNA methylation. We further identified differentially methylated and expressed genes, which have been shown to be involved in BEC-LEC plasticity. As some of the genes are directly connected to transdifferentiation processes in several diseases, these and other genes identified in our study could serve as candidate biomarkers to indicate the status of pathogenesis. Additionally, linking epigenetic mechanisms to blood and lymphatic vessel plasticity, our study suggests new approaches for the treatment of diseases involving pathological vessel conditions: silencing of genes modulating endothelial plasticity by aberrant DNA hypermethylation may be reversed by epigenetic therapy approaches in the future.
References
Tammela T, Alitalo K (2010) Lymphangiogenesis: molecular mechanisms and future promise. Cell 140(4):460–476
Hong YK, Shin JW, Detmar M (2004) Development of the lymphatic vascular system: a mystery unravels. Dev Dyn 231(3):462–473
Karpanen T, Alitalo K (2008) Molecular biology and pathology of lymphangiogenesis. Annu Rev Pathol 3:367–397
Adams RH, Alitalo K (2007) Molecular regulation of angiogenesis and lymphangiogenesis. Nat Rev Mol Cell Biol 8(6):464–478
Risau W (1997) Mechanisms of angiogenesis. Nature 386(6626):671–674
Srinivasan RS et al (2007) Lineage tracing demonstrates the venous origin of the mammalian lymphatic vasculature. Genes Dev 21(19):2422–2432
Oliver G, Harvey N (2002) A stepwise model of the development of lymphatic vasculature. Ann N Y Acad Sci 979:159–165 (discussion 188–196)
Wigle JT, Oliver G (1999) Prox1 function is required for the development of the murine lymphatic system. Cell 98(6):769–778
Breiteneder-Geleff S et al (1999) Angiosarcomas express mixed endothelial phenotypes of blood and lymphatic capillaries: podoplanin as a specific marker for lymphatic endothelium. Am J Pathol 154(2):385–394
Kaipainen A et al (1995) Expression of the fms-like tyrosine kinase 4 gene becomes restricted to lymphatic endothelium during development. Proc Natl Acad Sci USA 92(8):3566–3570
Bixel MG, Adams RH (2008) Master and commander: continued expression of Prox1 prevents the dedifferentiation of lymphatic endothelial cells. Genes Dev 22(23):3232–3235
Cooley LS et al (2010) Reversible transdifferentiation of blood vascular endothelial cells to a lymphatic-like phenotype in vitro. J Cell Sci 123(Pt 21):3808–3816
Groger M et al (2004) IL-3 induces expression of lymphatic markers Prox-1 and podoplanin in human endothelial cells. J Immunol 173(12):7161–7169
Groger M et al (2007) A previously unknown dermal blood vessel phenotype in skin inflammation. J Invest Dermatol 127(12):2893–2900
Tammela T et al (2008) Blocking VEGFR-3 suppresses angiogenic sprouting and vascular network formation. Nature 454(7204):656–660
Feinberg AP (2007) Phenotypic plasticity and the epigenetics of human disease. Nature 447(7143):433–440
Bird A (2007) Perceptions of epigenetics. Nature 447(7143):396–398
Reik W (2007) Stability and flexibility of epigenetic gene regulation in mammalian development. Nature 447(7143):425–432
Suzuki MM, Bird A (2008) DNA methylation landscapes: provocative insights from epigenomics. Nat Rev Genet 9(6):465–476
Laurent L et al (2010) Dynamic changes in the human methylome during differentiation. Genome Res 20(3):320–331
Li E (2002) Chromatin modification and epigenetic reprogramming in mammalian development. Nat Rev Genet 3(9):662–673
Lister R et al (2009) Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462(7271):315–322
Ribatti D, Nico B, Crivellato E (2009) Morphological and molecular aspects of physiological vascular morphogenesis. Angiogenesis 12(2):101–111
Chan Y et al (2004) The cell-specific expression of endothelial nitric-oxide synthase: a role for DNA methylation. J Biol Chem 279(33):35087–35100
Kim JY et al (2009) The expression of VEGF receptor genes is concurrently influenced by epigenetic gene silencing of the genes and VEGF activation. Epigenetics 4(5):313–321
Matsumura S et al (2007) DNA demethylation of vascular endothelial growth factor-C is associated with gene expression and its possible involvement of lymphangiogenesis in gastric cancer. Int J Cancer 120(8):1689–1695
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3(6):1101–1108
Sandoval J et al (2011) Validation of a DNA methylation microarray for 450,000 CpG sites in the human genome. Epigenetics 6(6):692–702
Gronniger E et al (2010) Aging and chronic sun exposure cause distinct epigenetic changes in human skin. PLoS Genet 6(5):e1000971
Smyth GK (2004) Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3:Article3
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Series B 57(1):289–300
Baluk P, McDonald DM (2008) Markers for microscopic imaging of lymphangiogenesis and angiogenesis. Ann N Y Acad Sci 1131:1–12
Kriehuber E et al (2001) Isolation and characterization of dermal lymphatic and blood endothelial cells reveal stable and functionally specialized cell lineages. J Exp Med 194(6):797–808
Sozzani R et al (2010) Spatiotemporal regulation of cell-cycle genes by SHORTROOT links patterning and growth. Nature 466(7302):128–132
Bibikova M, Fan JB (2009) Genome-wide DNA methylation profiling. Wiley Interdiscip Rev Syst Biol Med 2(2):210–223
Yuan L et al (2002) Abnormal lymphatic vessel development in neuropilin 2 mutant mice. Development 129(20):4797–4806
Karpanen T et al (2006) Functional interaction of VEGF-C and VEGF-D with neuropilin receptors. FASEB J 20(9):1462–1472
Ayadi A et al (2001) Net-targeted mutant mice develop a vascular phenotype and up-regulate egr-1. EMBO J 20(18):5139–5152
Nelson GM et al (2007) Differential gene expression of primary cultured lymphatic and blood vascular endothelial cells. Neoplasia 9(12):1038–1045
Petrova TV et al (2002) Lymphatic endothelial reprogramming of vascular endothelial cells by the Prox-1 homeobox transcription factor. EMBO J 21(17):4593–4599
Podgrabinska S et al (2002) Molecular characterization of lymphatic endothelial cells. Proc Natl Acad Sci USA 99(25):16069–16074
Saharinen P et al (2004) Lymphatic vasculature: development, molecular regulation and role in tumor metastasis and inflammation. Trends Immunol 25(7):387–395
Romanowska M et al (2008) PPARdelta enhances keratinocyte proliferation in psoriasis and induces heparin-binding EGF-like growth factor. J Invest Dermatol 128(1):110–124
Madsen P et al (1992) Molecular cloning and expression of a novel keratinocyte protein (psoriasis-associated fatty acid-binding protein [PA-FABP]) that is highly up-regulated in psoriatic skin and that shares similarity to fatty acid-binding proteins. J Invest Dermatol 99(3):299–305
Liu CA et al (2004) Rho/Rhotekin-mediated NF-kappaB activation confers resistance to apoptosis. Oncogene 23(54):8731–8742
Carmeliet P, Jain RK (2000) Angiogenesis in cancer and other diseases. Nature 407(6801):249–257
Carmeliet P (2003) Angiogenesis in health and disease. Nat Med 9(6):653–660
Hong YK et al (2004) Lymphatic reprogramming of blood vascular endothelium by Kaposi sarcoma-associated herpesvirus. Nat Genet 36(7):683–685
Portela A, Esteller M (2010) Epigenetic modifications and human disease. Nat Biotechnol 28(10):1057–1068
Goll MG, Bestor TH (2005) Eukaryotic cytosine methyltransferases. Annu Rev Biochem 74:481–514
Rauch TA et al (2009) A human B cell methylome at 100-base pair resolution. Proc Natl Acad Sci USA 106(3):671–678
Lorincz MC et al (2004) Intragenic DNA methylation alters chromatin structure and elongation efficiency in mammalian cells. Nat Struct Mol Biol 11(11):1068–1075
Zilberman D et al (2007) Genome-wide analysis of Arabidopsis thaliana DNA methylation uncovers an interdependence between methylation and transcription. Nat Genet 39(1):61–69
Klose RJ, Bird AP (2006) Genomic DNA methylation: the mark and its mediators. Trends Biochem Sci 31(2):89–97
Deaton AM, Bird A (2011) CpG islands and the regulation of transcription. Genes Dev 25(10):1010–1022
Eckhardt F et al (2006) DNA methylation profiling of human chromosomes 6, 20 and 22. Nat Genet 38(12):1378–1385
Hellman A, Chess A (2007) Gene body-specific methylation on the active X chromosome. Science 315(5815):1141–1143
Rountree MR, Selker EU (1997) DNA methylation inhibits elongation but not initiation of transcription in Neurospora crassa. Genes Dev 11(18):2383–2395
Barry C, Faugeron G, Rossignol JL (1993) Methylation induced premeiotically in Ascobolus: coextension with DNA repeat lengths and effect on transcript elongation. Proc Natl Acad Sci USA 90(10):4557–4561
Lin CI et al (2008) Lysophosphatidic acid up-regulates vascular endothelial growth factor-C and lymphatic marker expressions in human endothelial cells. Cell Mol Life Sci 65(17):2740–2751
Al-Rawi MA et al (2005) The effects of interleukin-7 on the lymphangiogenic properties of human endothelial cells. Int J Oncol 27(3):721–730
Al-Rawi MA et al (2005) Interleukin 7 upregulates vascular endothelial growth factor D in breast cancer cells and induces lymphangiogenesis in vivo. Br J Surg 92(3):305–310
Valtola R et al (1999) VEGFR-3 and its ligand VEGF-C are associated with angiogenesis in breast cancer. Am J Pathol 154(5):1381–1390
Rocha SF, Adams RH (2009) Molecular differentiation and specialization of vascular beds. Angiogenesis 12(2):139–147
Jing C et al (2001) Human cutaneous fatty acid-binding protein induces metastasis by up-regulating the expression of vascular endothelial growth factor gene in rat Rama 37 model cells. Cancer Res 61(11):4357–4364
Hummerich L et al (2006) Identification of novel tumour-associated genes differentially expressed in the process of squamous cell cancer development. Oncogene 25(1):111–121
Munz M, Zeidler R, Gires O (2005) The tumour-associated antigen EpCAM upregulates the fatty acid binding protein E-FABP. Cancer Lett 225(1):151–157
Adamson J et al (2003) High-level expression of cutaneous fatty acid-binding protein in prostatic carcinomas and its effect on tumorigenicity. Oncogene 22(18):2739–2749
Hirakawa S et al (2003) Identification of vascular lineage-specific genes by transcriptional profiling of isolated blood vascular and lymphatic endothelial cells. Am J Pathol 162(2):575–586
Kunstfeld R et al (2004) Induction of cutaneous delayed-type hypersensitivity reactions in VEGF-A transgenic mice results in chronic skin inflammation associated with persistent lymphatic hyperplasia. Blood 104(4):1048–1057
Acknowledgments
We thank the microarray unit of the DKFZ Genomics and Proteomics Core Facility for providing the Illumina Human Methylation arrays and related services. We thank Frank Lyko (Division of Epigenetics, DKFZ Heidelberg, Germany) for helpful discussions and valuable comments on the manuscript and Sylvain Foret (James Cook University, Townsville) for critical reading of the manuscript.
Conflict of interest
The authors have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Sabine Hagemann and Marc Winnefeld contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10456_2012_9264_MOESM1_ESM.tif
Online Resource 1 Determination of contamination of endothelial cells by other cell types. (a) Fibroblasts could be clearly distinguished from endothelial cells based on their different morphology. (b) FACS diagram showing a contamination of blood endothelial cells with fibroblasts (podoplanin (PDPN) and cluster of differentiation 31 (CD31) negative). (TIFF 1420 kb)
10456_2012_9264_MOESM2_ESM.tif
Online Resource 2 Validation of array-predicted methylation levels in blood and lymphatic endothelial cells. PCR amplification was performed using equimolar sample pools and sequencing results are shown for the arbitrarily chosen genes KN motif and ankyrin repeat domains 3 (KANK3), zinc finger, CCHC domain containing 12 (ZCCHC12), zinc finger, DHHC-type containing 11 (ZDHHC11), and kalirin (KALRN). 454 sequencing coverage ranged from 35 to 389 reads, as indicated. Red boxes indicate methylated, blue boxes unmethylated CpG dinucleotides, and white boxes indicate sequence gaps. CpG loci interrogated by the array are represented by black triangles. The tables below show averaged methylation values (AVB) for respective CpG loci calculated from Illumina array data. LEC, lymphatic endothelial cells; BEC, blood endothelial cells. (TIFF 1267 kb)
10456_2012_9264_MOESM3_ESM.tif
Online Resource 3 Validation of correlation of differentially methylated and expressed genes by 454 bisulfite sequencing and qRT-PCR (a) PCR amplification was carried out using equimolar sample pools of 6 blood (BEC) and 6 lymphatic endothelial cell populations (LEC) and sequencing results are shown for homebox A5 (HOXA5), trefoil factor 3 (TFF3), and XIAP associated factor 1 (XAF1). Sequencing coverage ranged from 106 to 262 reads, as indicated. Red boxes indicate methylated, blue boxes indicate unmethylated CpG dinucleotides, and white boxes represent sequence gaps. The horizontal grey bar illustrates the amplified sequence. CpGs interrogated on the Infinium methylation array are indicated by black triangles. (b) To validate distinct expression levels in LEC and BEC, quantitative RT-PCR analysis of the genes HOXA5, TFF3, and XAF1 was performed. Target gene expression was normalized to expression values of the endogenous control GAPDH. The horizontal black lines denote medians and whiskers the 2.5th and 97.5th percentiles. Significant differences are indicated by asterisks. * = p ≤ 0.05; ** = p ≤ 0.01; *** = p ≤ 0.001 (unpaired t-test). n (LEC) = 10, n (BEC) = 6. (TIFF 889 kb)
10456_2012_9264_MOESM4_ESM.tif
Online Resource 4 List of the 24 genes included in the top network “tissue development, cellular movement, and embryonic development”. Green or blue marks indicate that a least one CpG locus with a methylation change greater than 0.15 and Benjamini Hochberg adj. p.val < 0.01 exists in the defined genomic region. Green/blue color gradients indicate the presence of both hypo- and hypermethylated CpG loci. “Promoter” includes the regions TSS1500, TSS200, 5′UTR, and the first exon of genes. TSS, transcription start site. (TIFF 311 kb)
10456_2012_9264_MOESM5_ESM.tif
Online Resource 5 Pathway Analysis of 375 differentially methylated and expressed genes. Ingenuity Pathway Analysis reveals significantly enriched functional categories for upregulated and downregulated genes in lymphatic (LEC) versus blood endothelial cells (BEC). (TIFF 201 kb)
Rights and permissions
About this article
Cite this article
Brönneke, S., Brückner, B., Peters, N. et al. DNA methylation regulates lineage-specifying genes in primary lymphatic and blood endothelial cells. Angiogenesis 15, 317–329 (2012). https://doi.org/10.1007/s10456-012-9264-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10456-012-9264-2