Abstract
Circulating tumor cell detection is one important avenue for early cancer diagnosis. A differential solid-state micropore approach is reported that detects a very low number of tumor cells from blood samples. One micropore is functionalized with an aptamer molecule specific to epidermal growth factor receptor (EGFR), known to be overexpressed on cancer cell surfaces, whereas the other micropore is bare. The translocation behavior of tumor cells is seen to be distinctly different from normal counterparts and control when passed through functionalized micropore. The differentiation stems from the selective interactions between the aptamer molecules and overexpressed EGFR on the tumor cells. Normal cells, on the other hand, do not show selective interaction with aptamers. As a result, cancer cells are distinguished from normal cells just by looking at the translocation data from an aptamer-functionalized micropore. The interactions between the cells and the aptamer in a micropore provide a detection modality that interrogates each cell.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Cancer is one of the leading causes of human mortality around the globe. In the United States, it is placed as the second top cause of deaths. Deaths caused by some cancer types have not dropped significantly during the last five decades (Heron 2011). There are six biological capabilities that are gained during the multistep development of tumors and therefore are considered as the hallmarks of cancer. These are sustaining proliferative signaling, evading growth suppressors, resisting cell death, enabling replicative immortality, inducing angiogenesis, and activating invasion and metastasis. The actual causes of cancer are still not very well understood. However, the major hallmarks of cancer are mutation in genes, inability to repair the genes and abnormal expression of genes (Markowitz and Bertagnolli 2009; Kitamura et al. 2010; Hanahan and Weinberg 2011). These traits may be inherited or caused by environmental factors such as carcinogens, tobacco, radiation, etc. (Anand et al. 2008). During the initial stage of metastasis, the number of circulating tumor cells (CTC) in peripheral blood flow is very low. If these can be enumerated at early stages from blood, it can change the effectiveness of cancer treatment tremendously and alter the prognosis (Smith et al. 2009a, b).
Most of the current approaches for cancer cell detection rely on either target tagging or antibody-based capture. In the area of microdevices, factors such as mechanical forces (Mohamed et al. 2004; Huang et al. 2004), dielectrophoresis (Bustin et al. 2000), optical interactions (MacDonald et al. 2004), immunohistochemistry (Nagrath et al. 2007), magnetic sorting (Tibbe et al. 2002), flow cytometry, etc. (He et al. 2007) have been reported to identify and isolate cancer cells. Affinity-based interactions provide higher efficiency and greater specificity compared to mechanical and electrical sorting techniques (Toner and Irimia 2005). The use of antibodies is, however, subject to high levels of off-target cross-reactivity (Nagrath et al. 2007; Baas et al. 1996; Dalle et al. 2002). It is also challenging to retain specific sample conditions to keep antibodies completely functional. Many technical challenges are also encountered when cross-linking antibodies reproducibly onto the surfaces of miniaturized devices due to non-homogeneity of conjugation and denaturation on the surface. Recently, aptamers have been shown to have similar affinities and specificities such as antibodies (Wan et al. 2010). The aptamers are very stable at a variety of salt and ionic conditions and can be reversibly denatured (Bunka and Stockley 2006; Sullenger and Gilboa 2002). In recent works, aptamers have been used to activate cell signaling pathways and also to label cells (Nagel-Wolfrum et al. 2004; Charlton et al. 1997; Chen et al. 2008; Farokhzad et al. 2004). Devices incorporating aptamers to sort, isolate, and detect tumor cells have also been reported (Phillips et al. 2008; Asghar et al. 2012; Wan et al. 2010, 2012a, b). The main advantages of aptamers over antibodies lie in the ease of chemical synthesis, site-specific labeling, and, most importantly, site-specific immobilization.
This paper reports label-free detection of cancer cells using aptamer-functionalized micropores at single cell level. A hole with a dimension of the order of a micrometer drilled through a solid-state membrane is essentially called a micropore. Due to the simplicity and sensitivity, micropores have been used to characterize synthetic particles and bioentities such as cells, bacteria, etc. (Strutwolf et al. 2009; Asghar et al. 2012; Chang et al. 2002a; Steinle et al. 2002; Chien et al. 1971; Roberts et al. 2010). These are mechanically robust, stable, and easy to handle. These can be fabricated reliably to exact dimensions. Solid-state micropores can withstand wide range of experimental conditions such as pH, salinity, temperature, etc. (Chang et al. 2002a; Asghar et al. 2012).
The aptamer used was a ribonucleic acid (RNA) molecule known to selectively bind to epidermal growth factor receptor (EGFR) that is overexpressed on primary human glioblastoma (hGBM) cells. The approach detected cancer cells in a simple and highly sensitive cytological modality without any need for cell tagging. Among many other receptor tyrosine kinase oncogenes, EGFR is the most frequently overexpressed in all human malignancies. It is activated when various growth factors bind to it and initiate a signal transduction cascade that promotes cell adhesion, migration, proliferation, invasion, angiogenesis, and antiapoptosis (Mendelsohn 2004). Measurements from silicon dioxide (SiO2) micropores ensured that every cell created a signal. The sensitivity of the device was thus at single cell level. This can have impact on early cancer detection which can dramatically change cancer diagnosis and can subsequently improve mortality rate. It can also provide deeper insights into the interactions between specific cell types and their corresponding ligands.
2 Materials and methods
2.1 Materials
Tetramethylammonium hydroxide (TMAH) was purchased from VWR International. Amino modified DNA molecules were purchased from Integrated DNA Technologies. D/RNAse Free™ decontaminant was purchased from Argos Technologies Inc. Silver wires with 0.375 mm diameter were bought from Warner Instrument. Clorox solution for chloriding of Ag wire was bought from Walmart. The preparation of Ag/AgCl electrodes is detailed in supplementary material. Around 6-month-old rat (Sprague–Dawley) was purchased from Charles River. Blood was collected from the rat’s tail by restraining the rat and then stored in K2-ethylenediaminetetraacetic acid (EDTA) tube to prevent coagulation. The tubes were stored to 4 °C. The hGBM samples were obtained from consenting patient at the University of Texas Southwestern Medical Center (Dallas, TX). The isolation and culturing protocols are described in supplementary material (Wan et al. 2010). All other chemicals used were purchased from Sigma-Aldrich.
2.2 Fabrication of Micropore
The fabrication started with the thermal growth of 1 μm thick silicon dioxide on both sides of a double-side polished (100) p-type silicon wafer. Windows were opened by photolithography to start etching on the front side of the wafer. Wet etching was done using buffered hydrofluoric acid (BHF) to transfer the window from photoresist to SiO2 as shown in Fig. 1a.
Silicon was etched through this oxide window using anisotropic wet etching with 25 % TMAH at 90 °C. The self-limiting etch stopped when silicon etched all the way to SiO2 on the other side, resulting in a square window of 70 μm by 70 μm. The Ga+ focused ion beam (FIB, ZEISS 1540XB) was then used to drill a micropore in the membrane (black square in Fig. 1b). Different acceleration voltages, milling currents and drilling times were used to drill micropores (Fig. 1c) of different diameters in a controllable and reproducible manner (Asghar et al. 2011). Acceleration voltage of 30 kV and 1 nA drilling current typically resulted in 20 μm micropore in a 1 μm thick SiO2 membrane, when drilled for 5 min.
2.3 Preparation of Anti-EGFR Aptamer-Functionalized Micropores
The sequences for the extended anti-EGFR and mutant aptamer were as follows: anti-EGFR aptamer, 5′-GGCGCUCCGACCUUAGUCUCUGUGCCGCUAUAAUGCACGGAUUUAAUCGCCGUAGAAAAGCAUGUCAAAGCCGGAACCGUGUAGCACAGCAGAGAAUUAAAUGCCCGCCAUGACCAG-3′; mutant aptamer, 5′-GGCGCUCCGACCUUAGUCUCUGUUCCCACAUCAUGCACAAGGACAAUUCUGUGCAUCCAAGGAGGAGUUCUCGGAACCGUGUAGCACAGCAGAGAAUUAAAUGCCCGCCAUGACCAG-3′ (extension sequence is underlined). An amino modified capture oligonucleotides, 5′-AmMC6-TTTTTTTTTTTTTTTTTTTTCTGGTCATGGCGGGCATTTAATTC-3′ (active capturing sequence is underlined) was used to capture the aptamer on the surface.
The chemistry used to attach aptamers on the SiO2 micropore surfaces was adapted from earlier works (Wan et al. 2012a, b; Iqbal et al. 2007). The micropore silicon chips were 8 × 8 mm2 in size. Those were cleaned in piranha solution (H2O2/H2SO4, 1:3 v/v) for 10 min. After rinsing with deionized (DI) water and drying in nitrogen flow, the substrates were immersed into 2 % v/v (3-aminopropyl)triethoxysilane (APTES) in ethanol for 30 min at room temperature. That step was to create amine groups on surface. The APTES solution was removed and the substrates were sequentially washed with ethanol and DI water. Then, 2 mg of p-phenylene diisothiocyanate (PDITC) was mixed in a solution of dimethyl sulphoxide (DMSO) and pyridine (9:1 v/v) to prepare PDITC solution. The APTES treated substrates were incubated in PDITC solution at 55 °C for 5 h. After 5 h, the PDITC solution was removed and the substrates were washed three times each with isopropyl alcohol (IPA) and diethylpyrocarbonate (DEPC) treated DI water. Then, 10 μM capture DNA solution was added to DMSO to make 1:1 (v/v) DNA:DMSO solution. The substrates were immersed in DNA solution and incubated at 45 °C overnight. The chips were then soaked in ethanol followed by sequential washing with IPA and DEPC treated DI water. Nitrogen was used to dry the chips. RNA aptamer (1 μM) was mixed in hybridization buffer at a ratio of 1:5 (v/v). Then, a hybridization chamber was cleaned with RNase-free DI water thoroughly and a drop of aptamer solution was placed on the chip. The chamber was immersed into a water bath for 1 h at a temperature of 37 °C. The chips were then washed again with IPA and DEPC water three times each. The 6-amino-1-hexanol solution was used to deactivate amine groups that were not bound to the nucleic acids. This solution was made by dissolving 1.1719 g of 6-amino-1-hexanol in 5.2253 ml N,N-diisopropylethylamine and 200 ml of N,N-dimethylformamide. Chips were incubated in the deactivation solution at 45 °C for 5 h. The chips were then washed with ethanol and DEPC treated DI water. Without drying, the chips were put in 1X phosphate-buffered saline (PBS) solution. PBS solution was heated to 45 °C for 10 min and then slowly cooled down to room temperature.
2.4 Experimental Setup
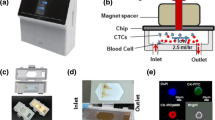
The micropore chip was sandwiched between polydimethylsiloxane (PDMS) gaskets. These gaskets were used to prevent leakage of solution. The chip and gaskets were then held together using two Teflon blocks that had holes as shown in Fig. 2a. Inlet and outlet tubes were connected to the Teflon blocks. Two Ag/AgCl electrodes were connected to the inlet and outlet, as shown in Fig. 2b. A syringe pump (11 PLUS, Harvard Apparatus) was used to maintain a constant flow rate of cell solution as shown in Fig. 2c. The hGBM cells (5,000 cells/ml) were suspended in 1X PBS and pumped through the system while measuring the current across the micropore. The rat blood, collected in tubes with K2-EDTA and stored at 4 °C, was taken out from storage tubes right before the experiments and thawed to room temperature. Then, 1X PBS solution was mixed with rat blood to dilute it one thousand times. Once the blood was homogenously mixed with PBS, the solution was pumped through micropores while measuring ionic current through them. A pair of Ag/AgCl electrodes was used to make contact with the ionic solution. Ionic current was measured using a data acquisition system (National Instrument).
First, only PBS was pumped through the micropore to establish the baseline. The PBS gave a constant ionic current through the micropores. When the cell solution was pumped through the micropore, each cell blocked it temporarily. Current dips were seen in the ionic current traces at that time. Every downward peak corresponded to a single cell passage. The hGBM cells were passed through a bare micropore and an aptamer-functionalized micropore to observe the differences.
3 Calculation
3.1 Gaussian pulse model
The data of pulses obtained from micropores have a number of parameters that can be used to understand the biophysical interactions occurring within the confinement of a micropore. When a cell passes through the micropore, it replaces or blocks the ionic species or charges from the micropore volume. That charge can then be calculated. The following assumptions were made to calculate the amount of charge physically blocked/replaced by a cell while passing through the micropore.
-
1.
All current pulses were inverted Gaussian in shape.
-
2.
Translocation time of the cell or current pulse width was assumed to be the full width at tenth of maxima (FWTM).
-
3.
The micropore was perfectly circular.
-
4.
Ionic concentration of the PBS was homogenous and uniform. PBS consisted of 137 mM NaCl, 2.7 mM KCl, 4.3 mM Na2HPO4 and 1.47 mM KH2PO4.
-
5.
Cells were perfectly spherical.
The equation of an inverted Gaussian current pulse is (Fig. 3):
where I(t) is current at time t within the pulse, I peak is the peak current, and c is a parameter that controls the width of the pulse (P w).
A simulated pulse used to model the current pulses (one example shown in inset of Fig. 5) obtained from micropore blockage
If the area under a Gaussian pulse is calculated (as shown in Fig. 3), it will be the amount of charge replaced/blocked during the time interval of the pulse width. Now, the area under the pulse, or in other words, the charge blocked, can be calculated by integrating the Gaussian function with respect to time.
and
3.2 Differential Detection
Since every cell corresponds to a current pulse i.e., every cell is essentially blocking charge flow while passing the micropore. This charge is accurately depicted by the current peak (I peak) and pulse width (P w) measured experimentally.
When raw rat blood was run through the micropore under the same experimental conditions, the two distributions of charge blockages were calculated for the bare and aptamer-functionalized micropores using this model. The two distributions were not different as shown in Fig. 4. Both distributions were single-sided Gaussian in nature and showed peaks on the same ranges of blocked charges. The blocked charge, on the other hand, for hGBM cells shows a clear shift in the peak (inset of Fig. 4). The experimental result shown in Fig. 4 can be explained in terms of the interactions between anti-EGFR aptamer and the EGFR present on the surface of hGBM and normal blood cells. As normal blood cells do not have a significant amount of EGFR compared to tumor cells on their surfaces (Normanno et al. 2006), the interactions between the anti-EGFR aptamer and the EGFR on normal rat blood cells are not sufficient to create significant differences between the distributions obtained from aptamer-modified micropore and bare micropore. As a result, distributions for blocked charges of normal blood cells obtained from bare and aptamer-functionalized micropores were quite similar. The statistics of the blocked charge data of hGBM cells, calculated for the bare and functionalized micropores, showed very different Gaussian distributions. The maximum occurrence of events for bare micropore was within 50–100 pC. On the other hand, the maximum occurrence of events for a micropore functionalized with aptamer was within 100–150 pC. The comparison between distributions of the charge blocked shows a clear shift of maxima toward the right for aptamer-modified micropore. It implies that due to the interactions between the EGFR on the cell surface and the anti-EGFR aptamer, the amount of charge blocked by the cells was larger. The overexpression of EGFR on hGBM cells caused significantly higher interactions between the anti-EGFR aptamer on the functionalized pore wall and the tumor cells, resulting in the shift of the histograms. This enhanced aptamer-receptor activity led to spatially separable distributions for bare and functionalized micropores. Despite the overlap, the distribution peaks are clearly located around different maxima. Given the presence of cancer cells in peripheral blood at the outset of cancer (Paterlini-Brechot and Benali 2007), the peaks of the histograms can be used as noninvasive diagnostic metrics.
Histogram showing distribution of the amount of charge blocked by normal rat blood cells while passing through bare and aptamer-functionalized micropores. The inset shows histogram of distribution of amount of charge blocked by hGBM cells while passing through bare and aptamer-functionalized micropores
4 Results
4.1 Translocation behavior
Figure 5 shows scatter plots of the pulse widths and peaks obtained from hGBM cells passing through bare and aptamer-functionalized micropores. Although the data points are distributed over a wide region, there is a clear clustering of bare micropore data points within 50–450 µS and 0.6–2.5 µA (within the dashed boundary).
Scatter plots of hGBM cells passage through a bare micropore (red dot) and same sized micropore that was functionalized with anti-EGFR aptamer (blue diamonds). The inset shows example of an actual ionic current trace of a typical pulse from an aptamer-functionalized micropore of 20 μm diameter when single hGBM cell passes at a flow rate of 20 μl/min. The electrical data acquisition is done at 200 kHz (color figure online)
The data represented by blue diamonds in Fig. 5 shows scatter plot of the pulse widths and current peaks for hGBM cells passing through an aptamer grafted micropore. The data points are fairly confined within 50–230 μs and 2.5 and 3.8 µA.
Even though there is a small overlap but the clustering of data points is spatially separate. The pulses from the aptamer-modified micropore clearly have different center of clustering than that for the bare micropore. The current pulses are much deeper when the micropore was functionalized with an aptamer. When an hGBM cell passed through a functionalized micropore, the interactions between EGFRs on tumor cell and the aptamers attached to the micropore’s wall had two effects. One, the ions were blocked, and second, a few ions that could make it through had reduced mobility. These factors led to an effective increase in resistance of the pore. As for the nature of interactions, the most fundamental mechanism here is the binding between the aptamer and EGFR. This binding is still not completely understood as this is neither duplex formation that happens in nucleic acids nor it is irreversible. The understanding on aptamer-receptor binding hints at formation of secondary structures such as hairpins, G-quartets, and loops that show selective affinity to certain target molecules.
In case of EGFR binding, we have reported before that this interaction is due to the hairpin structure formation in the aptamer molecule, which could be reversed to release EGFR (and the cells) from the surface of the chip (Wan et al. 2012a).
The current blocked by an hGBM cell was higher than that of normal cells due to higher resistance, even if cell sizes were the same (Fig. 6). As a result, the aptamer-functionalized micropore clearly differentiated between cells with more EGFR on their surfaces and cells with less/no EGFR due to its selective interaction with the EGFR on the cell surfaces.
Schematic showing interactions of hGBM cells with aptamer-functionalized and bare micropores (not to scale). a The tumor cells have significantly higher concentration of EGFR on the cells walls. The anti-EGFR aptamers are in hairpin form. b The cells interact with the aptamer. Left pane shows a normal cell that has significantly less binding with the aptamer-grafted micropore than that for the tumor cell shown in the right pane. The black cloud depicts the binding event. c The schematic of the ionic current pulses resulting from respective cell passages as in b
On the other hand, Fig. 7 shows scatter plot of the pulse widths and current peaks obtained from rat blood cells passing through the bare and aptamer-functionalized micropores. The data points obtained from both experiments are distributed over a wide region. The pulse width ranges from 50 to 3,500 μs and current peak ranges from 800 to 15,000 nA for the bare pore. Similarly, the pulse width varies between 50 and 4,500 μs and current peak varies between 800 and 55,000 nA for the aptamer-functionalized micropore. Contrary to the data for tumor cells, the two distributions here completely overlap. The normal expression of EGFR on normal cells results in similar interactions with the bare micropore and the aptamer-functionalized micropore. This makes the two distributions spatially inseparable. The experimental setup, thus, had the unique ability to differentiate between cells which had a significantly different number of EFGR on their surface. As tumor cells express a sufficiently larger concentration of EGFR compared to their normal counterparts, this unique differentiability can be extended to detect cancer cells from peripheral blood, and in many more applications. There is, however, a caveat regarding the overlap of the data range from cancer cells that might be enveloped within the data points stemming from the various types of cells in blood. This is discussed later. In a nutshell, normal blood cells do not have overexpressed EGFR on their surfaces, thus these do not interact with the aptamer-functionalized micropores.
Scatter plot of widths versus peaks of the pulses from rat blood cells passage through a bare (blue X), and same sized micropore that is functionalized with anti-EGFR aptamer (red O). Diameter of micropore is 20 μm, and flow rate is 20 μl/min. The electrical data acquisition is done at 200 kHz. Inset shows a magnified version of the densely populated region near origin. The functionalized pore data is shown in front of bare micropore data of the main figure. The two data are dense and lie on top of each other (color figure online)
5 Discussion
This paper shows that the data points obtained from the passage of hGBM cells through bare and aptamer-modified micropores have different centers of clustering and these are spatially separable. Also, the distributions of blocked charges show a clear right shift of peaks for pores functionalized with the aptamer. In contrast, normal blood cells do not show any significant difference in the distributions when passed through bare and aptamer-modified micropores. For normal blood cells, both distributions are one-sided Gaussian in nature and there is no relative peak shift observed. These findings are attributed to the interactions between the anti-EGFR aptamer and the EGFR expression levels on the surface of hGBM and normal blood cells. Unlike hGBM cells, normal blood cells do not have overexpressed EGFR on their surface. So, their distributions are not spatially separable as can be done for hGBM cells. This unique capability for analysis can alleviate the challenge of cancer cell data enveloped within normal cell data in a scatter plot (Figs. 5, 7).
Variation in sizes, amount of charges, receptor-ligand interactions (Wan et al. 2012a), and differences between stiffness of different cell types may induce differences in the translocation behavior (Chang et al. 2002b). White blood cells (WBCs) have been reported to be in the range of 11–22.5 μm while hGBM cells have been known to be around 20 μm (Wan et al. 2010). We used 20 μm diameter micropores. Given the sizes, heterogeneity, and stiffnesses of cells are the same for all types of the cells in normal rat blood, their behavior while passing through the two micropores (bare and functional) is not much different (and hence similar histograms). The inset of Fig. 4 (hGBM cells) shows the distribution peak is clearly shifted to the right (toward higher blocked charges) for the aptamer-functionalized micropore. It implies that with the hGBM cells, with their own size variations, heterogeneity, and stiffness range, the shift in the distribution peak for functionalized micropore came only from the aptamer binding effect. The variations in sizes and other factors can thus be discounted as major contributors in the discrimination effect between normal and cancer cells.
In short, the approach has the potential to detect and quantify tumor cells from a blood sample. The same operating principle might be applicable to detect other biological entities such as genes, pathogens, proteins, etc. if their corresponding aptamers are employed. The only challenges will then be to fabricate micropores comparable to the sizes of the target entities and to functionalize these with the specific ligands.
6 Conclusions
A differential detection modality of cancer cells has been demonstrated with a micropore functionalized with an aptamer specific to epidermal growth factor receptor. The differentiation was due to differences in tumor cell interactions with aptamer-functionalized micropore than those with the bare micropore. Normal cells did not exhibit selective interactions with aptamers. The sensitivity of detection was down to the single cell level. The presented method provides a detection modality that interrogates each cell without the requirement of tagging them.
References
Anand P, Kunnumakara AB, Sundaram C, Harikumar KB, Tharakan ST, Lai OS, Sung B, Aggarwal BB (2008) Cancer is a preventable disease that requires major lifestyle changes. Pharm Res 25(9):2097–2116
Asghar W, Ilyas A, Deshmukh RR, Sumitsawan S, Timmons RB, Iqbal SM (2011) Pulsed plasma polymerization for controlling shrinkage and surface composition of nanopores. Nanotechnol 22(28):285304–285311
Asghar W, Wan Y, Ilyas A, Bachoo R, Kim Y-t, Iqbal SM (2012) Electrical fingerprinting, 3D profiling and detection of tumor cells with solid-state micropores. Lab Chip 12(13):2345–2352
Baas IO, Berg FMVD, Mulder J-WR, Clement MJ, Slebos RJC, Hamilton SR, Offerhaus GJA (1996) Potential false-positive results with antigen enhancement for immunohistochemisty of the p53 gene product in colorectal neoplasms. J Pathol 178(3):264–267
Bunka DHJ, Stockley PG (2006) Aptamers come of age—at last. Nat Rev Microbiol 4(8):588–596
Bustin SA, Gyselman VG, Siddiqi S, Dorudi S (2000) Cytokeratin 20 is not a tissue-specific marker for the detection of malignant epithelial cells in the blood of colorectal cancer patients. Int J Surg Investig 2(1):49–57
Chang H, Ikram A, Kosari F, Vasmatzis G, Bhunia A, Bashir R (2002a) Electrical characterization of micro-organisms using microfabricated devices. J Vac Sci Tech B Microelectronl Nanom Struct 20(5):2058–2064
Chang WC, Liepmann D, Lee LP (2002b) A biomimetic method for extracting leukocytes from blood in microfluidic devices. Microtechnol Med Biol 2nd Annu Int IEEE-EMB Spec Top Conf, pp 184–188
Charlton J, Sennello J, Smith D (1997) In vivo imaging of inflammation using an aptamer inhibitor of human neutrophil elastase. Chem Biol 4(11):809–816
Chen HW, Medley CD, Sefah K, Shangguan D, Tang Z, Meng L, Smith JE, Tan W (2008) Molecular recognition of small-cell lung cancer cells using aptamers. ChemMedChem 3(6):991–1001
Chien S, Luse SA, Bryant CA (1971) Hemolysis during filtration through micropores: a scanning electron microscopic and hemorheologic correlation. Microvasc Res 3(2):183–203
Dalle F, Lopez J, Caillot D, Cuisenier B, Laubriet AE, Dumont L, Bonnin A (2002) False-positive results caused by cotton swabs in commercial Aspergillus antigen latex agglutination test. Eur J Clin Microbiol Infect Dis 21(2):130–132
Farokhzad OC, Jon S, Khademhosseini A, Tran T-NT, LaVan DA, Langer R (2004) Nanoparticle-aptamer bioconjugates a new approach for targeting prostate cancer cells. Cancer Res 64(21):7668–7672
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674
He W, Wang H, Hartmann LC, Cheng J-X, Low PS (2007) In vivo quantitation of rare circulating tumor cells by multiphoton intravital flow cytometry. Proc Natl Acad Sci 104(28):11760–11765
Heron M (2011) Deaths: leading Causes for 2007. Natl Vital Stat Rep 59(8):1–95
Huang LR, Cox EC, Austin RH, Sturm JC (2004) Continuous particle separation through deterministic lateral displacement. Sci 304(5673):987–990
Iqbal SM, Akin D, Bashir R (2007) Solid-state nanopore channels with DNA selectivity. Nat Nanotechnol 2(4):243–248
Kitamura A, Hosoda W, Sasaki E, Mitsudomi T, Yatabe Y (2010) Immunohistochemical detection of EGFR mutation using mutation-specific antibodies in lung cancer. Clin Cancer Res 16(13):3349–3355
MacDonald MP, Neale S, Paterson L, Richies A, Dholakia K, Spalding GC (2004) Cell cytometry with a light touch: sorting microscopic matter with an optical lattice. J Biol Regul Homeost Agents 18(2):200–205
Markowitz SD, Bertagnolli MM (2009) Molecular basis of colorectal cancer. N Engl J Med 361(25):2449–2460
Mendelsohn J (2004) EGF receptors as a target for cancer therapy. Trans Am Clin Climatol Assoc 115:249–254
Mohamed H, McCurdy LD, Szarowski DH, Duva S, Turner JN, Caggana M (2004) Development of a rare cell fractionation device: application for cancer detection. NanoBiosci IEEE Trans 3(4):251–256
Nagel-Wolfrum K, Buerger C, Wittig I, Butz K, Hoppe-Seyler F, Groner B (2004) The interaction of specific peptide aptamers with the DNA binding domain and the dimerization domain of the transcription factor Stat3 inhibits transactivation and induces apoptosis in tumor cells. Mol Cancer Res 2(3):170–182
Nagrath S, Sequist LV, Maheswaran S, Bell DW, Irimia D, Ulkus L, Smith MR, Kwak EL, Digumarthy S, Muzikansky A, Ryan P, Balis UJ, Tompkins RG, Haber DA, Toner M (2007) Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nat 450(7173):1235–1239
Normanno N, De Luca A, Bianco C, Strizzi L, Mancino M, Maiello MR, Carotenuto A, De Feo G, Caponigro F, Salomon DS (2006) Epidermal growth factor receptor (EGFR) signaling in cancer. Gene 366(1):2–16
Paterlini-Brechot P, Benali NL (2007) Circulating tumor cells (CTC) detection: clinical impact and future directions. Cancer Lett 253(2):180–204
Phillips JA, Xu Y, Xia Z, Fan ZH, Tan W (2008) Enrichment of cancer cells using aptamers immobilized on a microfluidic channel. Anal Chem 81(3):1033–1039
Roberts GS, Kozak D, Anderson W, Broom MF, Vogel R, Trau M (2010) Tunable nano/micropores for particle detection and discrimination: scanning ion occlusion spectroscopy. Small 6(23):2653–2658
Strutwolf J, Scanlon MlD, Arrigan DWM (2009) Electrochemical ion transfer across liquid/liquid interfaces confined within solid-state micropore arrays-simulations and experiments. Analyst 134(1):148–158
Smith RA, Cokkinides V, Brawley OW (2009a) Cancer screening in the United States, 2009: a review of current American Cancer Society guidelines and issues in cancer screening. CA Cancer J Clin 59(1):27–41
Smith RA, Cokkinides V, von Eschenbach AC, Levin B, Cohen C, Runowicz CD, Sener S, Saslow D, Eyre HJ (2009b) American cancer society guidelines for the early detection of cancer. CA Cancer J Clin 52(1):8–22
Steinle ED, Mitchell DT, Wirtz M, Lee SB, Young VY, Martin CR (2002) Ion channel mimetic micropore and nanotube membrane sensors. Anal Chem 74(10):2416–2422
Sullenger BA, Gilboa E (2002) Emerging clinical applications of RNA. Nat 418:252–258
Tibbe AGJ, de Grooth BG, Greve J, Dolan GJ, Rao C, Terstappen LWMM (2002) Magnetic field design for selecting and aligning immunomagnetic labeled cells. Cytom 47(3):163–172
Toner M, Irimia D (2005) Blood-on-a-chip. Annu Rev Biomed Eng 7:77–103
Wan Y, Kim Y-t, Li N, Cho SK, Bachoo R, Ellington AD, Iqbal SM (2010) Surface-immobilized aptamers for cancer cell isolation and microscopic cytology. Cancer Res 70(22):9371–9380
Wan Y, Liu Y, Allen PB, Asghar W, Mahmood MAI, Tan J, Duhon H, Kim Y-t, Ellington A, Iqbal SM (2012a) Capture, isolation and release of cancer cells with aptamer-functionalized glass beads. Lab Chip 22:4693–4701
Wan Y, Mahmood M, Li N, Allen PB, Kim Y-t, Bachoo R, Ellington AD, Iqbal SM (2012b) Nanotextured substrates with immobilized aptamers for cancer cell isolation and cytology. Cancer 118(4):1145–1154
Acknowledgments
We thank Azhar Ilyas and Muhymin Islam for help with the manuscript preparation and for useful discussions. The aptamers were provided by Dr. Andrew D. Ellington from the University of Texas at Austin, and tumor samples were provided by Dr. Robert Bachoo from the University of Texas Southwestern Medical Center at Dallas. This work was supported with National Science Foundation CAREER grant ECCS-1201878 and an award to SMI by the Cancer Research Foundation of North Texas, Arlington, Texas.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bellah, M.M., Iqbal, S.M. & Kim, YT. Differential behavior of EGFR-overexpressing cancer cells through aptamer-functionalized micropores. Microfluid Nanofluid 17, 983–992 (2014). https://doi.org/10.1007/s10404-014-1402-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10404-014-1402-2