Abstract
Understanding ecological niches is essential to comprehend the processes that allow similar species to occur sympatrically. Niche overlap can result in some degree of competition when resources are limited, and therefore, sympatric species must differ to some extent at some niche level in order to co-exist. The two trawling bats that co-occur along the Mediterranean region share their foraging strategy and feeding grounds, potentially consuming similar prey species. However, no research has been conducted to elucidate their dietary niche similarities or differences to test whether these may shape their sympatric foraging occurrence and distribution. We used DNA metabarcoding to study the dietary composition and niche overlap of Myotis capaccinii (an exceptionally endangered species) and M. daubentonii (a relatively common species) during the breeding season in northeastern Iberia. Unlike previous studies, Trichoptera was the most frequently consumed prey order for both bat species, followed by Diptera (mainly Chironomidae). We also report, for the second time, fish consumption by M. capaccinii in the Iberian Peninsula, and provide the fourth report of piscivory for European bats. Although minor differences in diet composition between both trawling bats were found, they presented highly overlapping dietary niches and similar dietary niche breadths, suggesting that they exploit similar trophic resources. Overall, the current results suggest that both species may have found a balance to co-occur in the same foraging niche without interspecific competition being a limiting factor.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A species’ ecological niche comprises all the interactions between the species and the environment, including the resources and required environmental conditions to survive (Soberón 2007). The range of these resources and conditions is defined as the ecological niche breadth, and it has profound implications influencing the species’ vulnerability and resilience in their habitats. In fact, the ecological niches of similar and sympatric species may overlap, resulting in some degree of competition for potentially limited resources. Thus, in order to co-exist, they must differ, at least in some niche dimensions: climatic tolerance, habitat or roost requirements, foraging strategies, or dietary preferences, for instance (Hutchinson 1957; Pianka 1973). However, most differences in niche dimensions between sympatric species remain unravelled, especially for many rare and threatened species, hindering the understanding of their conservation requirements.
Since the ecological niche represents such a broad concept, many authors have focused on the study of the dietary and the trophic niche overlap to understand the biology and dynamics of similar organisms. However, for many decades, the visual analysis of faeces, regurgitates, or gut content was the only method to characterise an organism’s diet (Agosta et al. 2003; Kross et al. 2016; Montoya et al. 2021). The identification limits of small prey items—mainly, the low taxonomic resolution, the reliability of the identifier’s experience, and the lack of hard body parts of some species—have resulted in a generalised underestimation of the diversity of consumed prey items (Pompanon et al. 2012). Nowadays, with the advent of high-throughput sequencing, even severely degraded prey tissue can be identified at the species level, and vast amounts of samples can be processed in a relatively short time (Pompanon et al. 2012; Galan et al. 2018). DNA metabarcoding techniques have allowed a wide variety of studies, including detailed dietary niche analysis for many different taxonomic groups such as fishes (Albaina et al. 2016; Takahashi et al. 2020), birds (da Silva et al. 2020; Cabodevilla et al. 2021), mammalian carnivores and herbivores (Kartzinel et al. 2015; Gebremedhin et al. 2016; Berry et al. 2017; Havmøller et al. 2021), small mammals (Iwanowicz et al. 2016; Biffi et al. 2017), and bats (Edwards et al. 2019; Ingala et al. 2021).
Being the second-largest order of mammals, bats are present in almost every habitat showing a wide range of ecological niches. The 51 European bat species are primarily insectivorous, which makes them especially targeted organisms of diet studies because of the ecosystem services they provide as pest controllers (Puig-Montserrat et al. 2020; Montauban et al. 2021). However, because many of them present similar diets and foraging strategies, potential competition for trophic resources may result (Arlettaz et al. 2000; Burles et al. 2008). In fact, different dietary studies have found high levels of trophic niche overlap between both sympatric and parapatric populations. For instance, Arrizabalaga-Escudero et al. (2018) found similar dietary niches for Rhinolophus euryale and R. mehelyi, which widely shared their foraging habitats. And Ashrafi et al. (2011) found that Plecotus macrobularis and P. austriacus, while foraging in different habitats, also showed high dietary niche overlap.
Trawling bats present a unique behaviour among bats, as they have specialised in catching insects directly from the water surface using their large feet and uropatagium (Aizpurua and Alberdi 2018). This behaviour makes them ideal subject species to assess interspecific competition and niche overlap, since they are strongly related to riparian habitats and present almost identical foraging strategies. Of these aquatic species, the long-fingered bat (Myotis capaccinii, Bonaparte, 1837) is exceptionally threatened all along its distribution area (Paunović 2016). During the last decades, it is facing an extreme population decline, mainly due to disturbance and loss of roosts, foraging riparian habitats, and water body pollution (Hutson et al. 2001; Biscardi et al. 2007). In the northeastern Iberian Peninsula, M. capaccinii shares its foraging habitat with the Daubenton’s bat (Myotis daubentonii, Kuhl, 1817). However, whereas the first one is discontinuously distributed along the Mediterranean basin (Paunović 2016), M. daubentonii is not of conservation concern since it is abundant and widespread throughout Europe to Siberia (Kruskop et al. 2020).
Sharing the same foraging habitats and other behavioural and ecological similarities suggests that trophic niche overlap might be found between both species (Biscardi et al. 2007). In fact, Krüger et al. (2012) already described it for the other pair of sympatric European trawling bats, M. daubentonii and M. dasycneme (the Pond bat, Boie, 1825). Even though M. capaccinii represents a flagship species within the Mediterranean rivers, few studies have accurately described its feeding niche. In the Iberian Peninsula, Almenar et al. (2008) reported Chironomidae as the most preyed arthropod family for M. capaccinii, followed by other Dipteran families, which supported what previous studies showed in Italy and Israel (Levin et al. 2006; Biscardi et al. 2007), all using visual faeces inspections. Similarly, studies conducted in Ireland and Germany and other studies using metabarcoding technique in Finland showed that M. daubentonii had high preferences for Dipterans, mainly Chironomidae and, in less frequency, Trichopterans, and Lepidopterans (Flavin et al. 2001; Nissen et al. 2013; Vesterinen et al. 2013, 2016). Piscivory has been reported on several occasions for M. capaccinii in nature (Aihartza et al. 2003; Levin et al. 2006; Biscardi et al. 2007; Aizpurua et al. 2013), while for M. daubentonii has only been exclusively reported under experimental conditions (Siemers et al. 2001).
Understanding the ecological niches, dietary breadth, and overlap of these species is essential to comprehend the processes that allow M. capaccinii and M. daubentonii to co-exist sympatrically. Studies conducted on their diet in the past suggest a high similarity at a trophic level. However, their niches may differ at some level and to a certain degree in order to co-exist. The present study is aimed at studying the dietary niche of the threatened M. capaccinii and comparing it with the sympatric and abundant M. daubentonii in order to assess mechanisms for their co-existence. Using molecular data, our specific aims are (1) describe the diet of both species in a Mediterranean region; (2) compare prey species richness and diet composition between both trawling bats, and assess potential differences between sexes, ages, reproductive status, months, and regions; (3) evaluate the potential trophic niche overlap between them; and (4) assess and compare the dietary niche breadth of both species.
Material and methods
Study area
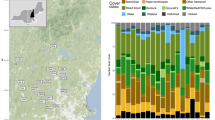
Sample collection was conducted in northeastern Iberia from June to August 2021, during the breeding season of both target species. Sampling locations were selected around known M. capaccinii breeding roosts, mainly located in two different river basins, Segre and Fluvià, where both bat species occur in sympatry. The Segre River originates in the Pyrenees and flows to the central depression, characterised by a more continental climate and arid landscapes. The Fluvià River, while it also rises in the eastern Pyrenees, flows through 90 km to the northern Catalan coastline, surrounded by milder climates and more humid forests. Both rivers have been altered as a result of anthropogenic activities (hydroelectric development, agriculture intensification, and water channelling, among others), causing an impact on river regimes and the associated biodiversity (Vicente-serrano et al. 2017). Bat sampling sessions were undertaken in a total of 27 sites situated along the Segre and Fluvià basins (Fig. 1) and separated by a minimum distance of 1 km.
Bat trapping and sample collection
Trapping sessions were conducted using a minimum of 30 m of mist-nets per night strategically placed over the water surface in pathway areas along the rivers. The sampling effort was standardised to 5 h per night, starting at sunset and checking mist-nets every 15 min. Capture sessions were conducted with the official permission of the State Agency for Environmental Policies of the Government of Catalonia (SF/0137/21). Bats were identified following Dietz and Kiefer (2016), and standard biometric measurements (sex, age, reproductive status, forearm length, and weight) were taken from every specimen. All data is currently included on the online database www.batmonitoring.org (Supplementary Table S1). Myotis capaccinii and M. daubentonii individuals were kept separately in clean cotton bags to avoid cross-contamination of the samples. Guano pellets were collected from the bags and stored in 1.5-ml Eppendorf tubes filled with silica gel granules to keep the samples dry. Before release, bats were marked by cutting off a small patch of fur on the back to avoid replicate sampling. A total of 403 faecal samples were obtained, corresponding to 140 M. capaccinii and 263 M. daubentonii individuals. Samples were stored and refrigerated at 4 °C. All animals were captured and handled under the standards of the American Society of Mammalogists (Sikes et al. 2011).
Laboratory procedures
DNA extraction was performed on one random faecal pellet per bat capture (n = 403) to balance all samples. A single pellet per individual provides a representative record of its dietary composition and reduces costs (Mata et al. 2019). Mata et al. (2021) described the followed protocol with some adjustments. Each pellet was homogenised with 650 µL of lysis buffer (0.1 M Tris–HCl, 0.1 M EDTA, 0.01 M NaCl, 1% N-lauroylsarcosine, pH 7.5–8; Maudet et al. 2002) and incubated at 70 °C for 30 min. Afterwards, samples were short-spinned, and 500 µL was transferred to a new tube containing 25 µL of OB Protease and 200 µL of BL buffer, followed by a second incubation at 70 °C for 10 min. The last steps of DNA precipitation and washing were performed following the instructions of the E.Z.N.A Tissue Kits (Omega Bio-Tek, Norcross, Georgia, USA), except that DNA was eluted twice in 50 µL into different tubes. One negative control was processed for every 23 samples (in a total of 19 extraction blanks). DNA extracts were distributed in 96 well plates, with one well left empty in every plate to serve as PCR blank (in a total of 5 PCR blanks).
Amplification of the DNA was conducted using the primer set Leray-XT (by Wangensteen et al. 2018): forward primer mlCOIintF-XT (5′-GGWACWRGWTGRACWITITAYCCYCC-3′) and reverse primer jgHCO2198 (5′-TAIACYTCIGGRTGICCRAARAAYCA-3′), both modified with Illumina adaptors. Leray-XT is a highly degenerated primer set designed to amplify a COI fragment of about 313 bp. It serves as a universal primer of Metazoa, being able to amplify both arthropods and vertebrates (e.g. bats and fish), while sometimes also co-amplifying non-target groups like fungi and bacteria. This primer has been evaluated and demonstrated to be highly effective for studying insectivorous animals and arthropod communities (Elbrecht et al. 2019). Authors such as Kemp et al. (2019) and Montauban et al. (2021) have already successfully detected insect pest species on bat’s faeces using Leray-XT. To reduce the amplification of bat DNA and maximise the proportion of reads belonging to dietary items, a Myotis daubentonii/capaccinii blocking primer targeting the forward primer mlCOIint-XT (5′-AGTTTATCCTCCCTTAGCAGGAAATCTTGC-C3_spacer-3′) was designed.
The PCR mix contained 5 µL of Qiagen Multiplex Master Mix (Hilden, Germany), 0.3 µL of each 10 nM primer, 0.3 uL of 100 nM blocking primer, 2.1 µL of water, and 2 µL of DNA extract. PCR conditions were as follows: an initial denaturation at 95 °C for 15 min, followed by 35 cycles of 30 s denaturation at 95 °C, annealing at 45 °C for 30 s, and extension at 72 °C for 30 s, and completed with a final extension period of 10 min at 72 °C. The extraction and PCR blanks were amplified along with the samples. Amplification products were diluted 1:4 with water and exposed to a second PCR reaction to incorporate 7-bp long identification tags and the Illumina P5 and P7 adaptors. PCR reactions and cycling conditions were similar to the first PCR except that it used KAPA HiFi HotStart ReadyMix (Rocher, KAPA Biosystems, Basel, Switzerland) and only eight cycles of denaturing, annealing, and extension were performed, with annealing at 55 °C. After PCR, the final products were purified with Agencourt AMPure XP beads (Beckman Coulter, Brea, California, USA), quantified using Epoch Microplate Spectrophotometer (Agilent Technologies, Inc., Santa Clara, California, USA), and then diluted to similar concentrations. The tagged and cleaned PCR products were pooled together into a single library that was quantified using qPCR (KAPA Library Quant Kit qPCR Mix; Rocher) and diluted to 4 nM. Library sequencing was performed using ~ 54% of a MiSeq Kit v3 (600 cycles) for a target of 28 k reads/sample. All PCR conditions described here follow the enzyme guidelines and common practise standards in the literature for the Leray-XT primer set (Wangensteen et al. 2018; Elbrecht et al. 2019).
Bioinformatic analysis
Bioinformatic processing was done using standard metabarcoding pipelines. Paired reads were merged using PEAR (Zhang et al. 2014), followed by the removal of primer sequences and tagging of reads with sample information using the command ‘ngsfilter’ from Obitools (Boyer et al. 2016). Then, reads were collapsed into exact sequence variants (ESVs) using the command ‘obiuniq,’ and singletons were removed per sample with ‘obigrep.’ Next, reads of the different samples were merged into a unique file, and sequence headers were transformed for VSEARCH (Rognes et al. 2016) compatibility. Reads were dereplicated again using ‘–derep_fulllength’ and denoised with ‘–cluster_unoise,’ assuming a minimum sequence length of 300 bp and standard abundance and alpha parameters. Resulting zero-radius operational taxonomic units (zOTUs) were further filtered for chimaeras using ‘–uchime3_denovo’ and clustered at 99% similarity with ‘–cluster_size.’ Reads were then mapped back again to the retained OTUs with ‘–usearch_global’ at an identity level of 99%. Finally, LULU (Frøslev et al. 2017) was used to merge similar OTUs (identity > 84%) with high co-occurrence levels (> 95% of samples), to reduce the number of retained PCR artefacts, sequencing errors, as well as nuclear copies of the mitochondria, that tend to artificially inflate the number of OTUs present in each sample.
OTU identification was made with BOLDigger (Buchner and Leese 2020) using the ‘digger_hit’ method, which uses different thresholds to select the assigned taxonomic level (98% similarity to species level, 95% to genus level, 90% to family level, 85% to order level, and < 85% to class level) and find the best fitting hit, while flagging suspicious hits (all sequences are available at the Zenodo repository: https://doi.org/10.5281/zenodo.8036859). Arthropod OTUs were manually checked and curated taking into account species distribution in the Iberian Peninsula and further queried against the NCBI database when no good hits were obtained. Finally, OTUs were classified as ‘prey’ if they belonged to Insecta, Araneae, Collembola, or Actinopterygii and ‘not prey’ in other cases (e.g. fungi, bacteria, and nematodes), except when OTUs were identified only to the class level (e.g. Insecta) or as known external parasites (e.g. bat flies and fleas), in which case they were classified as ‘not prey’ (a total of 4,630,431 reads were assigned as not prey). To remove potential lab contaminations, extraction and PCR blank reads were subtracted from the corresponding samples. Additionally, to further reduce residual or cross-contamination, as well as cross-talk during sequencing, for each sample, all prey taxa with a read count < 1% of the total number of prey reads of that sample were removed. Finally, samples with less than 100 reads belonging to prey items were considered to have failed and were removed from further analysis, resulting in 369 samples (117 corresponding to M. capaccinii and 252 to M. daubentonii).
Statistical analysis
Dietary analyses were conducted with both occurrence (presence/absence data) and relative read abundance (RRA) data considering all the associated biases. While on one side, the RRA is influenced by the differential recovery of markers from prey taxa; on the other, the occurrence data tend to represent rare items at similar weight as common ones (Deagle et al. 2019). Frequency of occurrence (FOO), weighted percentage of occurrence (wPOO) and average RRA were calculated at species, family, and order levels. We defined FOO as the proportion of samples containing the target item, expressed as a percentage; wPOO as the proportion of each prey item within each sample, expressed as a percentage and then rescaled to 100% across all prey items from all the samples; and average RRA rescaled to 100% across all prey items (Deagle et al. 2019). All statistical analyses were conducted in R version 4.1.1 (R Core Team 2021) and RStudio version 1.4.1717 (RStudio Team 2021).
Differences in diet species richness between M. capaccinii and M. daubentonii were assessed using generalised linear models (GLM). Bat species, sex, age, reproductive status, month of capture, and river basin were included as explanatory variables. GLMs were conducted using data from both species (with all explanatory variables) and from each species separately (excluding the bat species variable). In addition, since M. daubentonii was captured in more places than M. capaccinii, these analyses were also conducted only with data from the locations where both species were collected together. All models were run with a negative binomial error distribution to cope with overdispersion (R package aods3; Matthieu and Renaud 2018) and the ‘log’ link function, using the MASS R package (Venables and Ripley 2002). A Tukey post hoc test was used for the categorical predictors, with the R package multcomp (Hothorn et al. 2008) to test the specific effect of each factor level. Variance inflation factors (VIF; R package car; Fox and Weisberg 2019) were calculated for each model to evaluate possible multicollinearity in the explanatory variables (Fox and Monette 1992). Two variables (sex and reproductive status) exceeded the selected threshold (VIF ≤ 2), for the four models. In that case, individual models excluding each of those variables were run, and the one with the lowest Akaike information criterion (AIC) was chosen (Cayuela and de la Cruz 2022). Thus, sex was removed from the first model (including both bat species together) and from the fourth (including only locations where both bats were captured), while reproductive status was excluded in the other two models (using the data of each bat species separately). Then, model selection was performed using the dredge function from the R package MuMIn (Barton 2022), generating a set of models with all possible variable combinations from the saturated model. The final models were selected following an AIC value within Delta 2, including the maximum number of variables to test and with the lowest AIC.
Diet composition was compared using permutational multivariate analysis of variance (PERMANOVA) among both bat species, sex, age, reproductive status, month, and basin explanatory variables. The same analysis was performed for each bat species data separately, and only with the data from those locations where both bat species were captured together. PERMANOVA analyses were performed for presence-absence data (FOO) of each prey per sample (based on Jaccard distance matrix), for weighted occurrence data (wPOO), and for relative read abundance data (RRA; based on Bray Curtis). The three matrices were calculated for prey species, family, and order levels and tested with 9999 permutations using the adonis function from the vegan R package (Oksanen et al. 2020). The betadisper function from vegan R package was used to test for homogeneity of group variance for each significant predictor variable. If the variance was not homogeneous, the PERMANOVA results were excluded. Thus, only predictors with significant PERMANOVA results (p value < 0.05) and non-significant betadisper results (p value > 0.05) were considered to have an effect on diet composition. For the categorical predictors, a similarity percentage analysis (function simper; R package vegan) was conducted to know which prey differed in proportion (FOO, wPOO, and RRA) between groups.
The niche overlap between M. capaccinii and M. daubentonii was calculated using Pianka’s index (Pianka 1973). Pianka values nearing 0 indicate no resource overlap, while values closer to 1 mean that both species present almost identical dietary niche. Pianka’s index was measured from the FOO of prey species with the spaa R package (Zhang 2016).
The trophic niche breadth was assessed with different diversity indexes (Pianka 1973), calculated using the iNEXT R package (Hsieh et al. 2016). It was based on sampling-unit-based incidence data Hill numbers and modulated by the diversity order parameter q (Chao et al. 2014). The calculated indexes were species richness (q = 0), Shannon index (q = 1), and Simpson’s reciprocal index (q = 2). While higher values of the Shannon index express higher diversity with greater evenness, those of Simpson’s reciprocal index express higher diversity with greater evenness and regularity. The niche breadth was calculated using all data from both species and also using only the data collected from the localities where both bats were captured.
Results
In total, 350 prey items were identified (Supplementary Table S2), with an average of 4.28 ± 2.91 prey items per sample. A total of 7,010,034 reads were obtained, with an average of 18,997 ± 15,871 prey reads per sample (Supplementary Table S3). Myotis capaccinii samples contained 174 different prey items (69.0% identified to species level) corresponding to 65 families and 16 orders, while M. daubentonii samples contained 274 prey items (66.8% identified to species level) representing 95 families and 13 orders. The most detected prey order in both bat species was Trichoptera (80.3% and 82.9% of samples of M. capaccinii and M. daubentonii, respectively), which included the most detected family and species, Hydropsychidae (59.0% and 57.5% FOO), and Cheumatopsyche lepida (40.2% and 47.6% FOO) (Fig. 2). Results from both occurrence data and read counts agreed, revealing the following main orders—Diptera and Ephemeroptera, families—Chironomidae and Psychomyiidae, and species—Hydropsyche exocellata and Psychomyia pusilla (Fig. 2, Supplementary Figs. S1 and S2). Overall, only 25 prey items from the total 350 appeared at a frequency higher than 5% (24 for M. capaccinii and 15 for M. daubentonii; Fig. 2C), while the other prey items were only occasionally eaten. Additionally, being represented just in 0.8% of all samples, the only vertebrate detected in the samples was Gambusia affinis/holbrooki, in adult females of M. capaccinii captured in the Fluvià basin.
Frequency of occurrence (FOO) of prey items identified in the faeces of Myotis capaccinii (n = 117) and Myotis daubentonii (n = 252) during the summer 2021 in the Northeastern Iberian Peninsula, presented at different levels: A all identified orders, B the 2% more frequent families identified for both bat species, and C the 5% more frequent species identified for both bat species. Families and species showing significative differences in FOO between Myotis capaccinii and Myotis daubentonii (p value < 0.05) are also represented with an asterisk symbol
Prey richness
Significant differences in prey richness between bat species were not found. However, prey richness was affected by the month (p value < 0.05) for models including both bat species together and when only M. capaccinii was considered (Table 1). Individuals of M. capaccinii fed on a larger number of prey items in June than during July and August, with an average of 5.89 ± 3.41 (Fig. 3). M. daubentonii prey richness did not present differences within any variable (Table 1). Moreover, when only data from the locations where both bat species were collected together was considered, significant differences between months and reproductive status were found.
Prey richness variation in the Myotis capaccinii diet along the three sampled summer months, June (n = 53), July (n = 37), and August (n = 27), in the Northeastern Iberian Peninsula. Bars represent the 95% confidence intervals for each month. Tukey post hoc test results confirmed that June significantly differed from the other months (p value > 0.05)
Diet composition
Results on diet composition were similar whether presence-absence (FOO), weighted occurrence (wPOO), or relative read abundance (RRA) data were used (Supplementary Tables S4, S5, and S6). Diet composition significantly differed between both bats at prey species (Fig. 4A) and family levels. The diet composition differences between bat species resulted from a significantly higher FOO and wPOO of Hydropsyche exocellata, Chironomus annularius, Polycentropus flavomaculatus, and another 84 species in M. capaccinii compared to the M. daubentonii. At family level, 11 out of 112 families were found with a significantly higher FOO, 26 for wPOO, and 7 for RRA in M. capaccinii than in M. daubentonii, with Chironomidae being the most relevant family. Dietary composition diverged among sexes at the species level for FOO and wPOO. These results were consistent in the models including both bat species, only M. capaccinii and only locations with both bats. Differences in diet composition were also detected for FOO and wPOO between age and reproductive status at species, family, and order levels (Supplementary Tables S4, S5, and S6). Moreover, while monthly variation was found in M. capaccinii diet with FOO (Fig. 4B), wPOO, and RRA data, for M. daubentonii, differences in diet composition could not be attributed to any variable due to heterogeneity in dispersion between groups. The diet composition differences of M. capaccinii between months resulted from 15 prey species, seven families, and three orders that show significantly different FOO among months (60 species, 21 families, and nine orders for wPOO and 11 families and six orders for RRA). Altogether, Cheumatopsyche lepida was more dominant in July than in June and August, Hydropsyche exocellata and Psychomyia pusilla more in August than in June and July, and Diptera (Chironomidae) more in June than in August. Finally, diet composition based on RRA data showed differences between adults and juveniles of M. capaccinii.
Principal coordinates analysis (PCoA) graphic representation, using only the first two axes. The ordinations are based on: A prey species in the diet of M. capaccinii (n = 117) and M. daubentonii (n = 252) in the Northeastern Iberian Peninsula; B prey species in the diet of M. capaccinii faeces in the three sampled months, June (n = 53), July (n = 37), and August (n = 27), and use the Jaccard distance matrix for presence-absence data of each prey per sample. Each dot corresponds to a single sample
Niche overlap and niche breadth
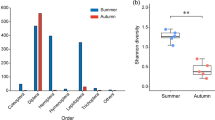
Pianka’s index showed an overlap of the dietary niche between both bat species of 89.26%, suggesting highly similar diets. In fact, no differences in dietary niche breadth were found between M. capaccinii and M. daubentonii with any of the analyses (species richness, Fig. 5; Shannon index, Supplementary Fig. S3; and reciprocal Simpson’s index, Supplementary Fig. S4), as the 95% confidence intervals highly overlapped. Similar results were also observed using only the data from the locations where both bats were collected.
Myotis capaccinii (Myocap, n = 117) and Myotis daubentonii (Myodau, n = 252) dietary niche breadth in the Northeastern Iberian Peninsula, during the summer 2021, based on the Hill number species richness (q = 0). Sample coverage rarefaction (solid line) and extrapolation (dashed line) are represented with the 95% confidence intervals (shaded regions) obtained from a bootstrap method based on 100 replications. The solid dot and triangle refer to the reference sample of each bat species
Discussion
This is the first study using molecular data to characterise the dietary niche of two Mediterranean trawling bat populations, Myotis capaccinii and M. daubentonii, and compare them in order to assess its implication for their co-existence. The present study achieves a great taxonomic resolution to species level and shows Trichoptera as the most consumed prey order, contrary to what previous authors found. For M. capaccinii, we report fish consumption for the second time in the Iberian Peninsula (corresponding to the fourth report of piscivory in this species across Europe) and seasonal influence during the breeding season. Finally, our results reveal similar trophic niches between both bat species, but subtle dissimilarities in prey composition, which, together with the fact that both species were feeding on abundant prey species, suggests a balance between both species that allows co-existence in the same foraging niche without interspecific strong competition for feeding resources.
The most frequently consumed prey items by both M. capaccinii and M. daubentonii in the present study differ from what has been found in previous dietary research on these species. Our results show Trichoptera (Hydropsychidae) and Diptera (Chironomidae) as the dominant prey orders for both bat species, followed by Ephemeroptera, Lepidoptera, and other less occurred orders. However, previous studies on M. capaccinii and M. daubentonii showed chironomids as the most frequently consumed prey (e.g. Almenar et al. 2008; Biscardi et al. 2007; Krüger et al. 2012; Vesterinen et al. 2013). While the results presented here indicate that Trichoptera constitutes the major part of the diet of both trawling bat species, studies using visual analysis (e.g. Biscardi et al. 2007; Almenar et al. 2008) reported Trichopterans in a lower frequency of occurrence, especially for M. capaccinii. For M. daubentonii, Nissen et al. (2013) detected Trichopterans (23%) only slightly more frequently than chironomids (22.5%). Ephemeropterans were detected in a few samples by Almenar et al. (2008), who explained the lack of this prey item in M. capaccinii faeces due to the soft and easily digestible bodies of ephemeropterans (Rabinowitz and Tuttle 1982).
Other studies using molecular techniques detected dipterans and lepidopterans as the most occurred families in both bat species around Europe (Alberdi et al. 2020), while another study conducted in Finland using DNA metabarcoding to assess the diet composition of M. daubentonii concluded that chironomids were its main prey item even though other prey species were also highly available (Vesterinen et al. 2016). Vesterinen et al. (2016) findings suggested a specialist feeding behaviour for M. daubentonii, which contradicts the claims of other authors who link M. capaccinii and M. daubentonii to opportunistic diets according to the available trophic resources (Almenar et al. 2008; Nissen et al. 2013). In our case study, chironomids, Trichopterans, and Ephemeropterans are abundant insects in the northeastern Iberian rivers with mass emerging periods and swarming behaviour above the freshwater surfaces (Puig 1999). The most frequently preyed species—Cheumatopsyche lepida, Psychomyia pusilla, and Hydropsyche exocellata—are also known to occur in the study area regularly (Bonada et al. 2008). Thus, the diet described in the current study seems to reflect the local prey availability, which could be related to opportunistic behaviour like Almenar et al. (2008) and Nissen et al. (2013) suggested.
However, primer biases must be seriously considered when comparing our results to other metabarcoding studies since, for example, the ZBJ primer (Zeale et al. 2011) used by Vesterinen et al. (2016) and Alberdi et al. (2020) has been reported to be particularly sensitive to Diptera and Lepidoptera (Alberdi et al. 2018). As mentioned before, Leray-XT (Wangensteen et al. 2018) is an ideal primer to recover several of terrestrial arthropod taxa (Elbrecht et al. 2019). Hence, the disparity observed in the most consumed prey compared to previous studies could also be attributed to the utilisation of different primer sets.
This is the second report of fish consumption in the Iberian Peninsula for M. capaccinii. One lactating and two adult passive females preyed on Gambusia affinis/holbrooki in the Fluvià basin during July and August. The first record was registered in the Southern Iberian Peninsula (Aihartza et al. 2003), where otoliths belonging to Gambusia holbrooki were found in M. capaccinii faeces (Aizpurua et al. 2013). Gambusia holbrooki was introduced in the early twentieth century at the Northeastern Iberia, and it is now distributed along this region, especially in the coastline rivers like the Fluvià (Aparicio 2016). The introduction of exotic species may severely influence the balance in our ecosystems by, for example, adding a new element to the trophic network of native species. Piscivory in M. capaccinii individuals has also been detected in other locations within its Mediterranean distribution, namely, Italy (Biscardi et al. 2007) and Israel (Levin et al. 2006). The low frequency of samples in the present study containing fish suggests that it has been a sporadic event, probably fostered by the high abundance of this fish species in the area.
Myotis capaccinii presented significant temporal variation in diet richness and composition through the breeding season. Higher prey richness at the beginning of the summer season may be related to higher energy requirements of early and mid-lactating females (Racey and Entwistle 2000). The early lactation period is an extremely vulnerable time for females, especially when they forage while holding their young on their bodies, therefore, a relatively high diversity of prey species may be needed to compensate for this energy demand (McLean and Speakman 1999; Dietz et al. 2006). In fact, Kunz et al. (1995) suggested that pregnant and lactating females preferred prey with higher fat content. While it is to be expected that differences in prey richness also occur at different reproductive status levels, the present study could not prove it, probably due to the relatively low number of samples for some categories.
Although the diet composition fluctuates over time for M. capaccinii, Hydropsychidae and Chironomidae remain the dominant prey families throughout the study period. Only 88 prey species were found to vary in their FOO between months. These changes may be influenced by local climate and the phenology of available prey, being more abundant in specific periods. Cheumatopsyche lepida (Trichoptera), for example, was much more frequently consumed during July than the rest of the months, which agrees with the peak of Trichoptera observed in July by Raitif et al. (2018) in France. Temporal variation in prey composition has also been found in other bat species like Eptesicus serotinus in Germany, also being associated with the phenology of available prey (Tiede et al. 2020). Nevertheless, the present study did not reveal differences between months in M. daubentonii diet, in line with Vesterinen et al. (2016) who did not find changes along the autumn season in Finland.
M. capaccinii and M. daubentonii exploited similar trophic resources as their dietary niche highly overlapped, and both dietary niche breadths were very similar for all three diversity indexes. The high niche overlap would indicate interspecific competition at the dietary level, like Biscardi et al. (2007) suggested. This might explain the slight partitioning of dietary resources found between both bat species, perhaps favoured by subtle variations in their behaviour. The few prey species that significantly differed among both bats occurred in higher frequency within M. capaccinii faeces. This variation might be due to the specificity of M. capaccinii to feed on aquatic species and the morphological and behavioural differences that make it more adapted to eat a higher range of those species (Almenar et al. 2009). Especially, a hairy uropatagium and tibia, a wing attached to the tibia and larger feet (Dietz and Kiefer 2016) might facilitate capturing larger aquatic prey found at deeper water levels (Aizpurua et al. 2013), such as Hydropsyche exocellata or Polycentropus flavomaculatus (Barata et al. 2005), which seem to be important food sources for M. capaccinii (Fig. 2C). An additional distinction in feeding behaviour between M. capaccinii and M. daubentonii may arise from differences in their consumption of prey at various life cycle stages. Unfortunately, due to the limitations of metabarcoding techniques in distinguishing between different life cycle stages, it was not feasible to investigate distinction in both bats’ preferences regarding this aspect.
The results of the present study suggest that there was no direct competition between M. capaccinii and M. daubentonii. The fact that both bats fed mainly on abundant prey species probably reduced the level of competition (Abrams 1980). Similar results were observed for M. daubentonii and the northern European trawling bat, M. dasycneme (Krüger et al. 2012, 2014), with high trophic niche overlap and similar niche breadth, but small differences in prey composition and prey types. They observed that while M. dasycneme fed on more aquatic prey, M. daubentonii also fed on terrestrial prey, suggesting differences in foraging habitats. This pattern was also found for the sympatric Rhinolophus euryale and R. mehelyi, as they highly overlapped their dietary niche with minor differences in species composition (Arrizabalaga-Escudero et al. 2018). The authors suggested that the foraging habitat segregation described by Salsamendi et al. (2012) might explain the subtle dietary dissimilarities.
Thus, for species with a high niche overlap, consuming abundant prey species with slight differences in the dietary niche may be enough to allow them to co-exist without interspecific trophic competition being a limiting factor. Yet, a rapid change in prey dynamics and abundance could lead to strong competition between them in the future. Understanding the mechanisms that allow species with similar niches to occur in sympatry can help to project future population dynamics according to different environmental and ecological change scenarios, especially for endangered species.
Data availability
All DNA sequences are available in the Zenodo repository https://doi.org/10.5281/zenodo.8036859.
References
Abrams P (1980) Some comments on measuring niche overlap. Ecology 61:44–49. https://doi.org/10.2307/1937153
Agosta SJ, Morton D, Kuhn KM (2003) Feeding ecology of the bat Eptesicus fuscus: “preferred” prey abundance as one factor influencing prey selection and diet breadth. J Zool 260:169–177. https://doi.org/10.1017/S0952836903003601
Aihartza JR, Goiti U, Almenar D, Garin I (2003) Evidences of piscivory by Myotis capaccinii (Bonaparte, 1837) in Southern Iberian Peninsula. Acta Chiropterologica 5:193–198. https://doi.org/10.3161/001.005.0204
Aizpurua O, Alberdi A (2018) Ecology and evolutionary biology of fishing bats. Mamm Rev 48:284–297. https://doi.org/10.1111/mam.12136
Aizpurua O, Garin I, Alberdi A et al (2013) Fishing long-fingered bats (Myotis capaccinii) prey regularly upon exotic fish. PLoS One 8:e80163. https://doi.org/10.1371/journal.pone.0080163
Albaina A, Aguirre M, Abad D et al (2016) 18S rRNA V9 metabarcoding for diet characterization: a critical evaluation with two sympatric zooplanktivorous fish species. Ecol Evol 6:1809–1824. https://doi.org/10.1002/ece3.1986
Alberdi A, Aizpurua O, Gilbert MTP, Bohmann K (2018) Scrutinizing key steps for reliable metabarcoding of environmental samples. Methods Ecol Evol 9:134–147. https://doi.org/10.1111/2041-210X.12849
Alberdi A, Razgour O, Aizpurua O et al (2020) DNA metabarcoding and spatial modelling link diet diversification with distribution homogeneity in European bats. Nat Commun 11:1154. https://doi.org/10.1038/s41467-020-14961-2
Almenar D, Aihartza J, Goiti U et al (2008) Diet and prey selection in the trawling long-fingered bat. J Zool 274:340–348. https://doi.org/10.1111/j.1469-7998.2007.00390.x
Almenar D, Aihartza J, Goiti U et al (2009) Foraging behaviour of the long-fingered bat Myotis capaccinii: implications for conservation and management. Endanger Species Res 8:69–78. https://doi.org/10.3354/esr00183
Aparicio E (2016) Peixos continentals de Catalunya. Ecologia, conservació i guia d’identificació. Lynx Edicions, Barcelona
Arlettaz R, Godat S, Meyer H (2000) Competition for food by expanding pipistrelle bat populations (Pipistrellus pipistrellus) might contribute to the decline of lesser horseshoe bats (Rhinolophus hipposideros). Biol Conserv 93:55–60. https://doi.org/10.1016/S0006-3207(99)00112-3
Arrizabalaga-Escudero A, Clare EL, Salsamendi E et al (2018) Assessing niche partitioning of co-occurring sibling bat species by DNA metabarcoding. Mol Ecol 27:1273–1283. https://doi.org/10.1111/mec.14508
Ashrafi S, Beck A, Rutishauser M et al (2011) Trophic niche partitioning of cryptic species of long-eared bats in Switzerland: implications for conservation. Eur J Wildl Res 57:843–849. https://doi.org/10.1007/s10344-011-0496-z
Barata C, Lekumberri I, Vila-Escalé M et al (2005) Trace metal concentration, antioxidant enzyme activities and susceptibility to oxidative stress in the tricoptera larvae Hydropsyche exocellata from the Llobregat river basin (NE Spain). Aquat Toxicol 74:3–19. https://doi.org/10.1016/j.aquatox.2005.04.002
Barton K (2022) MuMIn: multi-model inference
Berry TE, Osterrieder SK, Murray DC et al (2017) DNA metabarcoding for diet analysis and biodiversity: a case study using the endangered Australian sea lion (Neophoca cinerea). Ecol Evol 7:5435–5453. https://doi.org/10.1002/ece3.3123
Biffi M, Laffaille P, Jabiol J et al (2017) Comparison of diet and prey selectivity of the Pyrenean desman and the Eurasian water shrew using next-generation sequencing methods. Mamm Biol 87:176–184. https://doi.org/10.1016/j.mambio.2017.09.001
Biscardi S, Russo D, Casciani V et al (2007) Foraging requirements of the endangered long-fingered bat: the influence of micro-habitat structure, water quality and prey type. J Zool 273:372–381. https://doi.org/10.1111/j.1469-7998.2007.00337.x
Bonada N, Zamora-Muñoz C, El Alami M et al (2008) New records of Trichoptera in reference Mediterranean-climate rivers of the Iberian Peninsula and north of Africa: taxonomical, faunistical and ecological aspects. Graellsia 64:189–208. https://doi.org/10.3989/graellsia.2008.v64.i2.32
Boyer F, Mercier C, Bonin A et al (2016) obitools: a unix-inspired software package for DNA metabarcoding. Mol Ecol Resour 16:176–182. https://doi.org/10.1111/1755-0998.12428
Buchner D, Leese F (2020) BOLDigger - a Python package to identify and organise sequences with the Barcode of Life Data systems. Metabarcoding Metagenom 4:19–21. https://doi.org/10.3897/mbmg.4.53535
Burles DW, Brigham RM, Ring RA, Reimchen TE (2008) Diet of two insectivorous bats, Myotis lucifugus and Myotis keenii, in relation to arthropod abundance in a temperate Pacific Northwest rainforest environment. Can J Zool 86:1367–1375. https://doi.org/10.1139/Z08-125
Cabodevilla X, Mougeot F, Bota G et al (2021) Metabarcoding insights into the diet and trophic diversity of six declining farmland birds. Sci Rep 11:21131. https://doi.org/10.1038/s41598-021-00519-9
Cayuela L, de la Cruz M (2022) Análisis de datos ecológicos en R. Mundi-Prensa, Madrid
Chao A, Gotelli NJ, Hsieh TC et al (2014) Rarefaction and extrapolation with Hill numbers: a framework for sampling and estimation in species diversity studies. The Harvard community has made this article openly available. Please share how this access benefits you. Your Story Matters Ecol Monogr 84:45–67. https://doi.org/10.1890/13-0133.1
da Silva LP, Mata VA, Lopes PB et al (2020) High-resolution multi-marker DNA metabarcoding reveals sexual dietary differentiation in a bird with minor dimorphism. Ecol Evol 10:10364–10373. https://doi.org/10.1002/ece3.6687
Deagle BE, Thomas AC, McInnes JC et al (2019) Counting with DNA in metabarcoding studies: how should we convert sequence reads to dietary data? Mol Ecol 28:391–406. https://doi.org/10.1111/mec.14734
Dietz C, Kiefer A (2016) Bats of Britain and Europe. Bloomsbury Publishing, London
Dietz M, Encarnação JA, Kalko EKV (2006) Small scale distribution patterns of female and male Daubenton’s bats (Myotis daubentonii). Acta Chiropterologica 8:403–415. https://doi.org/10.3161/1733-5329(2006)8[403:SSDPOF]2.0.CO;2
Edwards CE, Swift JF, Lance RF et al (2019) Evaluating the efficacy of sample collection approaches and DNA metabarcoding for identifying the diversity of plants utilized by nectivorous bats. Genome 62:19–29. https://doi.org/10.1139/gen-2018-0102
Elbrecht V, Braukmann TWA, Ivanova NV et al (2019) Validation of COI metabarcoding primers for terrestrial arthropods. PeerJ 7(e7745):1–23. https://doi.org/10.7717/peerj.7745
Flavin DA, Biggane SS, Shiel CB et al (2001) Analysis of the diet of Daubenton’s bat Myotis daubentonii in Ireland. Acta Theriol (warsz) 46:43–52. https://doi.org/10.1007/BF03192415
Fox J, Monette G (1992) Generalized collinearity diagnostics. J Am Stat Assoc 87:178–183. https://doi.org/10.1080/01621459.1992.10475190
Fox J, Weisberg S (2019) An {R} Companion to applied regression
Frøslev TG, Kjøller R, Bruun HH et al (2017) Algorithm for post-clustering curation of DNA amplicon data yields reliable biodiversity estimates. Nat Commun 8. https://doi.org/10.1038/s41467-017-01312-x
Galan M, Pons J-B, Tournayre O et al (2018) Metabarcoding for the parallel identification of several hundred predators and their prey: application to bat species diet analysis. Mol Ecol Resour 18:474–489. https://doi.org/10.1111/1755-0998.12749
Gebremedhin B, Flagstad Ø, Bekele A et al (2016) DNA metabarcoding reveals diet overlap between the endangered walia ibex and domestic goats - implications for conservation. PLoS One 11:e0159133. https://doi.org/10.1371/journal.pone.0159133
Havmøller RW, Jacobsen NS, Havmøller LW et al (2021) DNA metabarcoding reveals that African leopard diet varies between habitats. Afr J Ecol 59:37–50. https://doi.org/10.1111/aje.12817
Hothorn T, Bretz F, Westfall P (2008) Simultaneous inference in general parametric models. Biometrical J 50:346–363. https://doi.org/10.1002/bimj.200810425
Hsieh TC, Ma KH, Chao A (2016) iNEXT: an R package for rarefaction and extrapolation of species diversity (Hill numbers). Methods Ecol Evol 7:1451–1456. https://doi.org/10.1111/2041-210X.12613
Hutchinson GE (1957) Concluding remarks. In: Cold spring harbor symposia on quantitative biology. pp 415–427
Hutson AM, Mickleburgh SP, Racey PA (2001) Microchiropteran bats: global status survey and conservation action plan. IUCN, Gland, Switzerland and Cambridge, UK
Ingala MR, Simmons NB, Wultsch C et al (2021) Molecular diet analysis of neotropical bats based on fecal DNA metabarcoding. Ecol Evol 11:7474–7491. https://doi.org/10.1002/ece3.7579
Iwanowicz DD, Vandergast AG, Cornman RS et al (2016) Metabarcoding of fecal samples to determine herbivore diets: a case study of the endangered Pacific pocket mouse. PLoS One 11:e0165366. https://doi.org/10.1371/journal.pone.0165366
Kartzinel TR, Chen PA, Coverdale TC et al (2015) DNA metabarcoding illuminates dietary niche partitioning by African large herbivores. Proc Natl Acad Sci USA 112:8019–8024. https://doi.org/10.1073/pnas.1503283112
Kemp J, López-Baucells A, Rocha R et al (2019) Bats as potential suppressors of multiple agricultural pests : a case study from Madagascar. Agric Ecosyst Environ 269:88–96. https://doi.org/10.1016/j.agee.2018.09.027
Kross SM, Bourbour RP, Martinico BL (2016) Agricultural land use, barn owl diet, and vertebrate pest control implications. Agric Ecosyst Environ 223:167–174. https://doi.org/10.1016/j.agee.2016.03.002
Krüger F, Clare EL, Greif S et al (2014) An integrative approach to detect subtle trophic niche differentiation in the sympatric trawling bat species Myotis dasycneme and Myotis daubentonii. Mol Ecol 23:3657–3671. https://doi.org/10.1111/mec.12512
Krüger F, Harms I, Fichtner A et al (2012) High trophic similarity in the sympatric North European trawling bat species Myotis daubentonii and Myotis dasycneme. Acta Chiropterologica 14:347–356. https://doi.org/10.3161/150811012X661666
Kruskop SV, Godlevska L, Bücs S et al (2020) Myotis daubentonii (errata version published in 2021). In: IUCN Red List Threat. Species 2020. https://doi.org/10.2305/IUCN.UK.2020-2.RLTS.T85342710A195858793.en%0ACopyright
Kunz TH, Whitaker JO, Wadanoli MD (1995) Dietary energetics of the insectivorous Mexican free-tailed bat (Tadarida brasiliensis) during pregnancy and lactation. Oecologia 101:407–415. https://doi.org/10.1007/BF00329419
Levin E, Barnea A, Yovel Y, Yom-Tov Y (2006) Have introduced fish initiated piscivory among the long-fingered bat? Mamm Biol 71:139–143. https://doi.org/10.1016/j.mambio.2006.01.002
Mata VA, da Silva LP, Veríssimo J et al (2021) Combining DNA metabarcoding and ecological networks to inform conservation biocontrol by small vertebrate predators. Ecol Appl 31:e02457. https://doi.org/10.1002/eap.2457
Mata VA, Rebelo H, Amorim F et al (2019) How much is enough? Effects of technical and biological replication on metabarcoding dietary analysis. Mol Ecol 28:165–175. https://doi.org/10.1111/mec.14779
Matthieu L, Renaud L (2018) aods3: analysis of overdispersed data using S3 methods
Maudet C, Miller C, Bassano B et al (2002) Microsatellite DNA and recent statistical methods in wildlife conservation management: applications in Alpine ibex [Capra ibex (ibex)]. Mol Ecol 11:421–436. https://doi.org/10.1046/j.0962-1083.2001.01451.x
McLean JA, Speakman JR (1999) Energy budgets of lactating and non-reproductive brown long-eared bats (Plecotus auritus) suggest females use compensation in lactation. Funct Ecol 13:360–372. https://doi.org/10.1046/j.1365-2435.1999.00321.x
Montauban C, Mas M, Wangensteen OS et al (2021) Bats as natural samplers: first record of the invasive pest rice water weevil Lissorhoptrus oryzophilus in the Iberian Peninsula. Crop Prot 141:105427. https://doi.org/10.1016/j.cropro.2020.105427
Montoya A, Cabodevilla X, Fargallo JA et al (2021) Vertebrate diet of the common kestrel (Falco tinnunculus) and barn owl (Tyto alba) in rain-fed crops: implications to the pest control programs. Eur J Wildl Res 67:79. https://doi.org/10.1007/s10344-021-01515-0
Nissen H, Krüger F, Fichtner A, Sommer RS (2013) Local variability in the diet of daubenton’s bat (myotis daubentonii) in a lake landscape of Northern Germany. Folia Zool 62:36–41. https://doi.org/10.25225/fozo.v62.i1.a5.2013
Oksanen J, Blanchet FG, Friendly M et al (2020) vegan: community ecology package
Paunović M (2016) Myotis capaccinii. In: IUCN red list threat. Species 2016. https://doi.org/10.2305/IUCN.UK.2016-2.RLTS.T14126A22054131.en. Accessed 25 Dec 2021
Pianka ER (1973) The structure of lizard communities. Annu Rev Ecol Syst 4:53–74. https://doi.org/10.1146/annurev.es.04.110173.000413
Pompanon F, Deagle BE, Symondson WOC et al (2012) Who is eating what: diet assessment using next generation sequencing. Mol Ecol 21:1931–1950. https://doi.org/10.1111/j.1365-294X.2011.05403.x
Puig-Montserrat X, Flaquer C, Gómez-Aguilera N et al (2020) Bats actively prey on mosquitoes and other deleterious insects in rice paddies: potential impact on human health and agriculture. Pest Manag Sci 76:3759–3769. https://doi.org/10.1002/ps.5925
Puig M (1999) Els macroinvertebrats dels rius catalans, Primera. Edigraf S.A
R Core Team (2021) R: a language and environment for statistical computing
Rabinowitz AR, Tuttle MD (1982) A test of the validity of two currently used methods of determining bat prey preferences. Acta Theriol (warsz) 27:283–293. https://doi.org/10.4098/at.arch.82-25
Racey PA, Entwistle AC (2000) Life-history and reproductive strategies of bats. London, UK
Raitif J, Plantegenest M, Agator O et al (2018) Seasonal and spatial variations of stream insect emergence in an intensive agricultural landscape. Sci Total Environ 644:594–601. https://doi.org/10.1016/j.scitotenv.2018.07.021
Rognes T, Flouri T, Nichols B et al (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584: https://doi.org/10.7717/peerj.2584
RStudio Team (2021) RStudio: integrated development environment for R
Salsamendi E, Garin I, Arostegui I et al (2012) What mechanism of niche segregation allows the coexistence of sympatric sibling rhinolophid bats? Front Zool 9:1–12. https://doi.org/10.1186/1742-9994-9-30
Siemers BM, Dietz C, Nill D, Schnitzler HU (2001) Myotis daubentonii is able to catch small fish. Acta Chiropterologica 3:71–75
Sikes RS, Gannon WL, Mammalogists the AC and UC of the AS of (2011) Guidelines of the American Society of Mammalogists for the use of wild mammals in research. J Mammal 92:235–253. https://doi.org/10.1644/10-MAMM-F-355.1
Soberón J (2007) Grinnellian and Eltonian niches and geographic distributions of species. Ecol Lett 10:1115–1123. https://doi.org/10.1111/j.1461-0248.2007.01107.x
Takahashi M, DiBattista JD, Jarman S et al (2020) Partitioning of diet between species and life history stages of sympatric and cryptic snappers (Lutjanidae) based on DNA metabarcoding. Sci Rep 10:4319. https://doi.org/10.1038/s41598-020-60779-9
Tiede J, Diepenbruck M, Gadau J et al (2020) Seasonal variation in the diet of the serotine bat (Eptesicus serotinus): a high-resolution analysis using DNA metabarcoding. Basic Appl Ecol 49:1–12. https://doi.org/10.1016/j.baae.2020.09.004
Venables WN, Ripley BD (2002) Modern applied statistics with S, Fourth. Springer, New York
Vesterinen EJ, Lilley T, Laine VN, Wahlberg N (2013) Next generation sequencing of fecal DNA reveals the dietary diversity of the widespread insectivorous predator Daubenton’s bat (Myotis daubentonii) in southwestern Finland. PLoS One 8:e82168. https://doi.org/10.1371/journal.pone.0082168
Vesterinen EJ, Ruokolainen L, Wahlberg N et al (2016) What you need is what you eat? Prey selection by the bat Myotis daubentonii. Mol Ecol 25:1581–1594. https://doi.org/10.1111/mec.13564
Vicente-serrano SM, Zabalza-martínez J, Borràs G et al (2017) Effect of reservoirs on streamflow and river regimes in a heavily regulated river basin of Northeast Spain. Catena 149:727–741. https://doi.org/10.1016/j.catena.2016.03.042
Wangensteen OS, Palacín C, Guardiola M, Turon X (2018) DNA metabarcoding of littoral hardbottom communities: high diversity and database gaps revealed by two molecular markers. PeerJ 6:e4705. https://doi.org/10.7717/peerj.4705
Zeale MRK, Butlin RK, Barker GLA et al (2011) Taxon-specific PCR for DNA barcoding arthropod prey in bat faeces. Mol Ecol Resour 11:236–244. https://doi.org/10.1111/j.1755-0998.2010.02920.x
Zhang J (2016) spaa: species association analysis
Zhang J, Kobert K, Flouri T, Stamatakis A (2014) PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30:614–620. https://doi.org/10.1093/bioinformatics/btt593
Acknowledgements
We are especially grateful to Toni Arrizabalaga, Pau Sainz de la Maza and Pep Xarles for all the logistic support and Ferran Páramo for his contribution to data management. We thank the Barcelona Zoo Foundation and the Catalan Government for supporting and funding the project. We thank Carme Tuneu-Corral and Xavier Puig-Montserrat, for their fieldwork assistance and Benjamí Francesc Vallmanya Subirada for his valuable advice, motivation, and knowledge of the study area. We also appreciate the support of the forest rangers, especially the GSM group, and the local councils of Camarassa, Balaguer, Artesa de Segre, Alfarràs, and Besalú. We thank Maria Mas Navarro, Owen S. Wangensteen, Francisco Amorim, Luis P. da Silva, Javier Dieguez-Uribeondo, Jesús Muñoz Fuente, Jose Manuel Serrano Talavera, Leopold Füreder, and Luis Cayuela Delgado for their constructive comments during the project.
Funding
This research was funded by the scholarship program for research projects or in situ conservation of species and habitats of the Barcelona Zoo Foundation, call 2020. This project also counted with the support by the Catalan Government (registration number DB201804) and the Granollers council. D.L.-B. was funded by AGAUR (grant number 2020DI113). V.A.M. research contract is funded by Fundação para a Ciência e Tecnologia (FCT; 2020.02547.CEECIND).
Author information
Authors and Affiliations
Contributions
A.L.-B., D.L.-B., C.F., and E.B. conceived the idea; D.L.-B. and C.F. acquired the financial support; C.F. did project administration; A.L.-B. and D.L.-B. supervised and led the research planning and execution; V.A.M. led the metabarcoding analysis; D.L.-B. and E.B. prepared the material and collected the samples; V.A.M. and E.B. conducted the bioinformatic analysis and statistical analysis; E.B. analysed the data and wrote the manuscript. All authors discussed the results, commented, and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Blanch, E., López-Baucells, A., Mata, V.A. et al. To share or not to share: DNA metabarcoding reveals trophic niche overlap between sympatric trawling bats. Eur J Wildl Res 69, 90 (2023). https://doi.org/10.1007/s10344-023-01712-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10344-023-01712-z