Abstract
Two isolates of Pythium nunn, newly recorded in Japan, were obtained from soils in Nagano and Fukuoka prefectures, characterized, and tested for antagonistic activity against P. ultimum var. ultimum. The morphology of both isolates corresponded with those of the original description of P. nunn. The rDNA-ITS sequences of the two isolates were identical to each other and had a high similarity with the sequences of the type strain of P. nunn. The two P. nunn isolates were mycoparasitic toward P. ultimum var. ultimum and suppressed damping off of cucumber seedlings caused by the pathogen at an early stage of plant growth.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pythium nunn Lifshitz, Stanghellini & Baker is a potential biocontrol agent first recorded from a grassland soil in Colorado, USA (Lifshitz et al. 1984a). This species, which has never been recorded as a plant pathogen (Lifshitz et al. 1984b, c), efficiently suppresses pre-emergence damping off of cucumber seedlings caused by P. ultimum Trow var. ultimum in greenhouse conditions (Adams 1990; Lifshitz et al. 1984b; Paulitz and Baker 1987a, b, 1988b; Paulitz et al. 1985, 1990). It also suppresses root rots of azalea and sweet orange caused by Phytophthoracinnamomi Rands, P. citrophthora R. E. & E. H. Smith and P. parasitica Dastur (Fang and Tsao 1995). These studies, however, have been limited to a few isolates from soils in the USA.
We recently obtained P. nunn-like isolates from soils in Japan. The objectives of this study were to identify them and to characterize their antagonistic activity against P. ultimum var. ultimum.
Isolation and identification
Two isolates of P. nunn, UZ041 and UZ415, used in this study were recovered from soils by a baiting technique using cucumber seeds as a baiting substrate (Watanabe 1981). Isolate UZ041 was recovered from soil in a vegetable field in Nagano Prefecture in June 2003. Isolate UZ415 was recovered in Fukuoka Prefecture in April 2007 from soil in a deciduous forest dominated by Japanese little leaf box (Buxus microphylla Sieb. et Zucc. var. japonica Rehd. et Wils.).
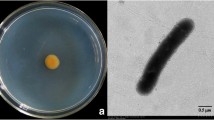
The two isolates were cultured on corn meal agar (CMA) prepared according to Tojo et al. (1998) and grass blade culture prepared according to Martin (1992). Both isolates were maintained on CMA until use. Morphological identification was based on the keys of van der Plaats-Niterink (1981) and the species description of Lifshitz et al. (1984c). The cardinal temperature for hyphal growth of the isolates UZ041 and UZ415 was determined on plates of potato–carrot agar (PCA) (van der Plaats-Niterink 1981) in darkness at temperatures from 4 to 40°C at 3°C intervals and at 42°C. Based on the morphologies and growth temperatures (Table 1; Fig. 1), both isolates were identified as P. nunn. Zoospore production was not reported in the original description of P. nunn (Lifshitz et al. 1984c), but was found in the present isolates (Table 1). The diameter of spherical sporangia differed between isolates UZ041 and UZ415 (Table 1). Isolate UZ041 readily formed sporangia and zoospores, but isolate UZ415 rarely formed them. Zoospore productivity does vary within a species of Pythium spp., especially in species that form spherical sporangia (van der Plaats-Niterink 1981). The present result, with earlier works (Lifshitz et al. 1984c; van der Plaats-Niterink 1981), suggests that zoospore productivity of P. nunn varies among the isolates of the species. The cardinal temperatures for hyphal growth were similar for the two P. nunn isolates, but differed from those in the original description (Table 1). This biological property is also thought to vary intraspecifically in P. nunn as we discuss later. Isolates UZ041 and UZ415 were deposited at the National Institute of Agrobiological Sciences Genebank, Japan as accessions MAFF 241106 and MAFF 241787, respectively.
Morphological characteristics of Pythium nunn isolated from Japan. a–g Isolate UZ041, h–k isolate UZ415. a Sporangium with germ tube. b Vesicle with zoospores. c Empty sporangium after dispersion of zoospores. d Globose, intercalary hyphal swelling. e Oogonium with a spine and an antheridium with an inflated stalk. f Terminal oogonium with two antheridia. g Terminal oogonium with an antheridium having crook stalk. h Globose, intercalary hyphal swelling. i Globose, terminal hyphal swelling. j Terminal oogonium and an antheridium that has an inflated stalk. k Oogonium with one spine and a monoclinous antheridium having crook stalk. Bar 10 μm
Sequence analysis
The nucleotide sequences of the ITS region including 5.8S rDNA of isolates UZ041 and UZ415 were determined as described previously (Uzuhashi et al. 2008). Both sequences were 875 bp long, and they were identical each other. The sequences had high similarity (98.6%) with those of P. nunn type isolate CBS 808.98 (AY598709; Lévesque and de Cock 2004; Lifshitz et al. 1984c). The results support the morphological identification of the isolates. The sequence differences between the present isolates and the type isolate were related to difference of growth temperature (Table 1). Although it is difficult to explain why the sequence differences relate to growth temperatures in P. nunn, such intraspecific variation in the ITS sequences relating to growth temperature is widely known for Pythium spp. (Kurokawa and Tojo 2010; Perneel et al. 2006; Uzuhashi et al. 2009). The sequences of isolates UZ041 and UZ415 have been deposited in GenBank as accessions AB468771 and AB537557, respectively.
Hyphal interaction
Pythium nunn isolates UZ041 and UZ415, and P. ultimum var. ultimum isolate OPU774 (Kida et al. 2007) were used. Agar plugs (4 mm diameter) were cut with a cork borer from the margins of actively growing colonies on CMA. The plugs were transferred to opposite sides of a Petri dish (90 mm diameter) with a cellophane film disk (90 mm diameter) placed on water agar. The plates were incubated for 3 days at 25°C, then a strip of the film (15 mm2) was removed from the crossing zone between the mycelia and was examined with a compound light microscope. Hyphae of both P. nunn isolates had penetrated the hyphae of the P. ultimum var. ultimum isolate (Fig. 2). In the penetrated hyphae, cytoplasm had disappeared, and hyphal septa had formed (Fig. 2). These phenomena are consistent with previous observations of hyphal interactions between P. nunn and P. ultimum var. ultimum (Adams 1990; Lifshitz et al. 1984a).
Hyphal interaction between Pythium nunn and P. ultimum var. ultimum. a P. nunn isolate UZ041 versus P. ultimum var. ultimum. b P. nunn isolate UZ415 versus P. ultimum var. ultimum. Arrowheads indicate hyphae of P. nunn (PN), P. ultimum var. ultimum (PU), and septa (S) formed in hyphae of P. ultimum var. ultimum. Bar 10 μm
Disease inhibition
A pot inoculation test was used to test the ability of P. nunn to inhibit disease caused by P. ultimum var. ultimum. Pythium nunn isolates UZ041 and UZ415, P. oligandrum isolate UOP399 (Kinoshita et al. 1994), and P. ultimum var. ultimum isolate OPU774 were used. Pythium oligandrum was used as comparison for biocontrol with P. nunn, because P. oligandrum has been well documented as a biocontrol agent on P. ultimum var. ultimum (Martin and Hancock 1987; Mcquilken et al. 1992) and on many other plant pathogens (e.g., Takenaka et al. 2008). A CMA plug containing mycelia of one of the Pythium isolates was transferred to a 300 ml Erlenmeyer flask containing 3 g of autoclaved seeds of tall fescue (Festuca arundinacea Schreb. cv. Davinchi) and a 12 ml of distilled water. After 10 days of incubation at 25°C in darkness, 15 g of these colonized seeds were mixed with 150 g of commercial nursery soil (Aisai-1, Katakura Chikkarin, Tokyo, Japan) in a mortar. The soil infested with P. ultimum var. ultimum was prepared with mixing 15 g of the inoculum and 3 kg of the nursery soil in a polyethylene bag. A single layer of a water-soluble paper (Scottie toilet tissue, Nippon Paper Crecia, Tokyo, Japan) was cut into a disk (7 cm diameter) and was placed on 90 ml of the P. ultimum var. ultimum infested soil in a ceramic pot (inner diameter 7 cm, inner depth 6.5 cm). Either the P. nunn-infested or the P. oligandrum-infested soil (40 ml) was placed on the paper disk, and seven seeds of cucumber (Cucumis sativus L. cv. Jibai) were placed in the soil. The seeds were covered with the paper disk, then 30 ml of the P. ultimum var. ultimum infested soil was placed on top. Each pot was enclosed in a double layer of polyethylene bag and kept at 25°C under continuous light (73 μmol/m2/s measured at plant levels) in a growth chamber. The soil in the plates was watered every day to keep the moisture near −10 kPa. The percentage emergence was recorded at 4 and 10 days after sowing, and again at 20 days (data not shown). The experiments were repeated eight times with one pot per treatment. Analysis of variance was conducted for the percentage emergence data of different treatments using JMP software (version 8; SAS Institute, Cary, NC, USA). Means of the data were compared by the least significant difference based on a Tukey–Kramer honestly significant different (HSD) test (P < 0.05).
Pythium nunn isolates UZ041 and UZ415 significantly (P < 0.05) supported the stand of the cucumber seedlings planted in the soil infested with P. ultimum var. ultimum at 4 days after sowing (Table 2; Fig. 3). They did not support the stand after 10 days of the sowing, resulting in post-emergence damping off of all seedlings (Table 2). Pythium oligandrum, on the other hand, inhibited damping off through 10 days after the sowing. The result demonstrated that the present isolates of P. nunn are potential biocontrol agents for P. ultimum var. ultimum. The lower effectiveness of P. nunn than P. oligandrum may be related to a unique mechanism for the disease inhibition by P. nunn. Lifshitz et al. (1984b) reported that the antagonistic activity of P. nunn against P. ultimum var. ultimum was dependent on organic substrates that contained high levels of labile carbon substrates, such as green leaves. Paulitz and Baker (1987a, 1988a) also reported that disease suppression against P. ultimum var. ultimum on cucumber operated only if organic amendments such as bean leaves or rolled oats were also added to the soil, along with a low inoculum density of P. nunn. Moreover, P. nunn is a relatively slow-acting mycoparasite compared with P. oligandrum (Laing and Deacon 1991). These previous reports suggest that P. nunn is a slow-acting antagonist that requires organic amendments for activity. The effect of organic amendments on the antagonistic activity has not yet been determined for the present isolates of P. nunn and will be examined in a future study.
Effects of Pythium nunn or P. oligandrum on cucumber seedlings planted in a commercial nursery soil infested with P. ultimum var. ultimum at 4 days after planting. a P. nunn isolate UZ041 + P. ultimum var. ultimum. b P. nunn isolate UZ415 + P. ultimum var. ultimum. c P. oligandrum + P. ultimum var. ultimum. d Noninoculated soil + P. ultimum var. ultimum (arrowheads indicate the diseased plants covered by mycelia of P. ultimum var. ultimum). e Noninoculated
References
Adams PB (1990) The potential of mycoparasites for biological control of plant diseases. Annu Rev Phytophthol 28:59–72
Fang JG, Tsao PH (1995) Evaluation of Pythium nunn as a potential biocontrol agent against Phytophthora root rots of azalea and sweet orange. Phytopathology 85:29–36
Kida K, Tojo M, Yano K, Kotani S (2007) First report of Pythium ultimum var. ultimum causing damping-off on okra in Japan. Plant Pathol 56:1042
Kinoshita T, Ichitani T, Okumura T (1994) Materials for Pythium flora of Japan VIII. Two species of Pythium: P. pyrilobum and P. oligandrum. Mycoscience 35:191–198
Kurokawa K, Tojo M (2010) First record of Pythium grandisporangium in Japan. Mycoscience. doi:10.1007/s10267-010-0041-z
Laing SAK, Deacon JW (1991) Video microscopical comparison of mycoparasitism by Pythium oligandrum, P. nunn and an unnamed Pythium species. Mycol Res 95:469–479
Lévesque CA, de Cock AWAM (2004) Molecular phylogeny and taxonomy of the genus Pythium. Mycol Res 108:1363–1383
Lifshitz R, Dupler M, Elad Y, Baker R (1984a) Hyphal interactions between a mycoparasite, Pythium nunn and several soil fungi. Can J Microbiol 30:1482–1487
Lifshitz R, Sneh B, Baker R (1984b) Soil suppressiveness to a plant pathogenic Pythium species. Phytopathology 74:1054–1061
Lifshitz R, Stanghellini ME, Baker R (1984c) A new species of Pythium isolated from soil in Colorado. Mycotaxon 20:373–379
Martin FN (1992) Pythium. In: Singleton LL, Mihail JD, Rush CM (eds) Methods for research on soilborne phytopathogenic fungi. APS Press, St. Paul, pp 39–49
Martin FN, Hancock JG (1987) The uses of Pythium oligandrum for biological control of preemergence damping-off caused by P. ultimum. Phytopathology 77:1013–1020
McQuilken MP, Whipps JM, Cooke RC (1992) Use of oospore formulations of Pythium oligandrum for biological control of Pythium damping-off in cress. Phytopathology 135:125–134
Paulitz TC, Baker R (1987a) Biological control of Pythium damping-off of cucumbers with Pythium nunn: population dynamics and disease suppression. Phytopathology 77:335–340
Paulitz TC, Baker R (1987b) Biological control of Pythium damping-off of cucumbers with Pythium nunn: influence of soil environment and organic amendments. Phytopathology 77:341–346
Paulitz TC, Baker R (1988a) The formation of secondary sporangia by Pythium ultimum: the influence of organic amendments and Pythium nunn. Soil Biol Biochem 20:151–156
Paulitz TC, Baker R (1988b) Interactions between Pythium nunn and Pythium ultimum on bean leaves. Can J Microbiol 34:947–951
Paulitz TC, Windham M, Baker R (1985) Pythium nunn—a potential biological control agent. Phytopathology 75:1326–1327
Paulitz TC, Ahmad JS, Baker R (1990) Integration of Pythium nunn and Trichoderma harzianum isolate T-95 for the biological control of Pythium damping-off of cucumber. Plant Soil 121:243–250
Perneel M, Tambong JT, Adiobo A, Foloren C, Saborío F, Lévesque A, Höfte M (2006) Intraspecific variability of Pythium myriotylum isolated from cocoyam and other host crops. Mycol Res 110:583–593
Takenaka S, Sekiguchi H, Nakaho K, Tojo M, Masunaka A, Takahashi H (2008) Colonization of Pythium oligandrum in the tomato rhizosphere for biological control of bacterial wilt disease analyzed by real-time PCR and confocal laser-scanning microscopy. Phytopathology 98:187–195
Tojo M, Nakazono E, Tsushima S, Morikawa T, Matsumoto N (1998) Characterization of two morphological groups of isolates of Pythium ultimum var. ultimum in a vegetable field. Mycoscience 39:135–144
Uzuhashi S, Tojo M, Kobayashi S, Tokura K, Kakishima M (2008) First records of Pythium aquatile and P. macrosporum isolated from soils in Japan. Mycoscience 49:276–279
Uzuhashi S, Tojo M, Kobayashi S, Kakishima M (2009) Pythium apinafurcum: its morphology, molecular phylogeny, and infectivity for plants. Mycoscience 50:281–290
van der Plaats-Niterink AJ (1981) Monograph of the genus Pythium. Stud Mycol 21:1–242
Watanabe T (1981) Distribution and populations of Pythium species in the northern and southern parts of Japan. Ann Phytopathol Soc Jpn 47:449–456
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kobayashi, S., Uzuhashi, S., Tojo, M. et al. Characterization of Pythium nunn newly recorded in Japan and its antagonistic activity against P. ultimum var. ultimum . J Gen Plant Pathol 76, 278–283 (2010). https://doi.org/10.1007/s10327-010-0244-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10327-010-0244-3