Abstract
Background
Aberrant expression of SWI/SNF complex subunits is closely associated with tumorigenesis. The clinicopathological and prognostic significance of altered SMARCA2 and SMARCA4 subunits has not been well evaluated in gastric adenocarcinoma.
Methods
We collected 1271 postoperative cases of gastric adenocarcinoma and then constructed tissue microarrays (TMA), from which we obtained the immunohistochemistry expression of SMARCA2 and SMARCA4. Next, we screened the variables related to the loss of SMARCA2 and SMARCA4 by univariate correlation analysis and multivariate logistic regression analysis. Then, we identified the variables related to prognosis by univariate and multivariate Cox regression analysis. Finally, we constructed a nomogram prognostic model and evaluated it.
Results
The loss of SMARCA2 and SMARCA4 occurred in 236 (18.57%) and 86 (6.77%) cases, respectively, including 26 cases of co-loss. After multivariate logistic regression, variables independently associated with SMARCA2 loss were T stage, differentiation status, WHO histological classification, and EBER. Variables independently associated with SMARCA4 loss were differentiation status, WHO histological classification, PD-L1, and MMR. Survival analysis revealed that the SMARCA2 and SMARCA4 lost groups showed worse survival than the corresponding present groups (P = 0.032 and P = 0.0048, respectively). Univariate and multivariate Cox analyses identified independent prognostic factors, including age, T stage, N stage, M stage, SMARCA2, and chemotherapy.
Conclusion
The loss of SMARCA2 and SMARCA4 correlated with poor differentiation, leading to a worse prognosis. SMARCA2, as an independent prognostic factor, combined with other clinicopathological variables, established a novel nomogram prognostic model, which outperformed the AJCC TNM model.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastric cancer (GC) is the fourth cause of cancer-related mortality globally [1], and the median survival time for advanced gastric cancer is less than 12 months despite many treatments [2]. As a highly invasive and heterogeneous malignancy [3], gastric cancer is still a global health problem.
In eukaryotes, DNA wraps around histones to form nucleosomes, which are highly compressed to form chromatin. This structure guarantees genomic stability but impedes genetic information replication and DNA damage repair. Therefore, chromatin remodeling complexes are essential for the dynamic regulation of chromatin. The Switch/Sucrose non-fermentable (SWI/SNF) chromatin remodeling complex is a multiprotein complex consisting of 10 to 15 subunits that utilizes the energy from ATP hydrolysis to disrupt the contact between DNA and histones, achieving nucleosome disassembly and regulation of gene expression [4]. The SWI/SNF complex is involved in various important cellular processes, such as cell proliferation, cell lineage differentiation, and DNA repair, which are frequently significantly altered in carcinoma [5, 6]. Alterations of SWI/SNF complex are found in approximately 20% of human cancers, and have also been proposed as potential drug targets for cancer treatment [7]. SMARCA2 and SMARCA4 are essential ATPase subunits and generate energy by catalyzing the hydrolysis of adenosine triphosphate (ATP) to ensure the proper functioning of the SWI/SNF complex [8]. Although aberrant expression of SMARCA2 and SMARCA4 has been identified in a wide range of human cancers, the significance of the two subunits' alterations in gastric adenocarcinoma is incompletely understood, and only a few studies to date have evaluated the prognostic significance of the two subunits in a large sample of gastric adenocarcinomas [9, 10].
In this study, we evaluated the correlation of immunohistochemical (IHC) expression patterns of SMARCA2 and SMARCA4 with clinicopathological features and prognosis in 1271 patients with gastric adenocarcinoma. Of the studies on SWI/SNF complex in gastric adenocarcinoma to date, the sample size of this study is the largest. And for the first time, we have included SMARCA2 in a clinical prognostic model.
Materials and methods
Cases collection and tissue microarrays (TMA) construction
With the approval of the Ethics Review Board at Weihai Municipal Hospital (permission code: 2021053), we collected 1347 patients with gastric adenocarcinoma who underwent initial surgical treatment at Weihai Municipal Hospital between January 2014 and December 2020. Two clinical pathologists reviewed the hematoxylin and eosin (H&E)-stained slides in detail and marked representative areas, next took out 2 mm diameter tissue cores from corresponding formalin-fixed and paraffin-embedded (FFPE) donor tissue blocks using a manual tissue sampling gun (jlm-5133, Guangdong, China), then transferred the tissue cores to the hole of the recipient paraffin block (ZSGB-BIO, Beijing, China, 6 × 10 holes). After excluding the tissue spots where there was too little tumor tissue and that detached from the TMA slides during the staining process, 1271 cases were included in this study.
We restaged the enrolled cases according to the current AJCC TNM staging system (8th edition, 2019) and reviewed H&E-stained slides in detail to accurately record pathologic features, including the differentiation status, WHO histological classification, Lauren classification, vascular invasion (VI), and perineural invasion (PNI). We obtained the information about age, sex, tumor location, and tumor size by consulting the electronic medical record. The primary study endpoint was overall survival (OS), defined as the period from the date of diagnosis to death by any cause or the last follow-up. The median follow-up period was 41.6 months (range from 0.03 to 89.2 months).
Immunohistochemistry (IHC) and in situ hybridization (ISH)
We performed IHC staining on 2 µm sections from each TMA block by an automated immunostaining machine (Benchmark ULTRA, Ventana) for SMARCA2, SMARCA4, Her-2, p53, Ki-67, PD-L1, MSH2, MSH6, MLH1, and PSM2.Details of primary antibodies are listed in Table S1. If immunohistochemistry for Her-2 was 2+, we further conducted fluorescence in situ hybridization (FISH) to detect Her-2 amplification status. Following the manufacturer's protocol, we detected EBV infection by EBV-encoded small RNA (EBER) in situ hybridization (EBER-ISH) using EBER assay kits (ZSGB-BIO, ISH-7001).
Assessment criteria
The IHC expression patterns of SMARCA2 and SMARCA4 were categorized as intact (intense nuclear staining in the neoplastic cells was similar to that in control cells), reduced (the nuclear staining was faint but recognizable), lost (the neoplastic cells did not show any nuclear staining), and heterogeneous (lost or reduced expression in only part neoplastic cells). The strong uniform nuclear staining of normal epithelial, inflammatory, and fibroblastic cells was used as the positive control. MMR proteins (MLH1, PMS2, MSH2, and MSH6) were located in cell nuclei and were classified as intact (definite nuclear staining) and lost (complete absence of nuclear staining). Any MMR proteins lost were defined as MMR deficient (dMMR), and all MMR proteins intact were defined as MMR proficient (pMMR). The evaluation criteria of other markers, including p53, Her-2, Ki-67, PD-L1, and TILs, were summarized in Table S2.
Statistical analysis
Univariate correlation analysis between expression status of SMARCA2 and SMARCA4 and clinicopathological features was performed using the Chi-square test or Fisher's exact test. Variables with P < 0.1 were included in the multiple logistic regression model, and the stepwise method was used to identify independent factors. Overall survival (OS) was determined by the Kaplan–Meier method, and the log-rank test was used to determine the difference. Univariate Cox regression analysis was performed to screen out significant variables (P < 0.05) for further multivariate Cox analysis. We identified independent prognostic factors used to construct a nomogram prognostic model, which was assessed by concordance index (C-index), the area under the curve (AUC), net reclassification improvement (NRI), integrated discrimination improvement (IDI), and decision curve analysis (DCA). All statistical analyses were performed using R software (version 4.1.2), and the related R packages were as follows: VennDiagram (V1.7.0), UpSetR (1.4.0), maftools (2.10.0), survival (3.2–13), survminer (0.4.9), epiDisplay (3.5.0.1), MASS (7.3–54), forestplot (2.0.1), rms (6.2–0), pROC (1.18.0), timeROC (0.4), survIDINRI (1.1–1), and nricens (1.6). A two-sided P value of < 0.05 was considered statistically significant.
Results
Clinicopathological features
A total of 1271 gastric adenocarcinoma cases consisting of 947 males (74.51%) and 324 females (25.49%) were finally included in this study. The median age of the cohort was 70 (ranging from 28 to 88) years. The counts of TNM stage I, II, III, and IV were 323 (25.41%), 302 (23.76%), 583 (45.87%), and 63 (4.96%), respectively. Tumors occurred most frequently in antrum (774 cases, 60.9%), followed by body (364 cases, 28.64%) and cardia (133 cases, 10.46%). Other IHC staining results were as follows: Her-2 positive in 79 cases (6.22%), p53 mutation in 531 cases (41.78%), EBER positive in 71 cases (5.59%), PD-L1 positive in 90 cases (7.08%), and dMMR in 187 cases (14.71%).
The expression of SMARCA2/4 and correlation with clinicopathological features
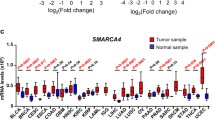
Representative IHC images of SMARCA2 and SMARCA4 were shown in Fig. 1, demonstrating four immunohistochemical expression patterns: intact, reduced, heterogeneous, and lost. Detailed counts were shown in Fig. 2a. Of the 1271 cases, 236 showed SMARCA2 loss (18.57%), and 86 showed SMARCA4 loss (6.77%), including 26 cases with co-loss (Fig. 2b). To gain further insight into the mutational landscape of SMARCA2 and SMARCA4 gene, we downloaded and visualized mutation data of gastric adenocarcinomas from The Cancer Genome Atlas (TCGA) database, which revealed mutation rates of SMARCA2 and SMARCA4 were 5% and 6%, respectively. (Fig. 2c). We integrated the four expression patterns into a dichotomous classification for subsequent statistical analysis. We defined intact, reduced, and heterogeneous patterns as present (i.e., lost vs. present) and defined reduced, heterogeneous, and lost patterns as attenuated (i.e., attenuated vs. intact).
Representative immunohistochemical images of SMARCA2 and SMARCA4 (magnification × 200). a Intact expression pattern of SMARCA4 with intense, uniform nuclear staining; b reduced expression pattern of SMARCA4 with obviously weaker nuclear staining; c heterogeneous expression pattern of SMARCA2 with the coexistence of intense and utterly absent nuclear staining; d lost expression pattern of SMARCA2 with complete absence of nuclear staining
Immunohistochemical expression of SMARCA2 and SMARCA4 and mutations of the corresponding genes. a Distribution of the four immunohistochemical expression patterns of SMARCA2 and SMARCA4. b The Venn diagram showed single-loss and co-loss of SMARCA2 and SMARCA4. c Mutational landscape of SMARCA2 and SMARCA4 gene in gastric adenocarcinoma from TCGA database
Univariate correlation analysis revealed that the variables associated with SMARCA2 loss were T stage, TNM stage, differentiation status, WHO histological classification, EBER, PD-L1, and MMR (Table 1). After multivariate logistic regression, factors independently associated with SMARCA2 loss were T stage, differentiation status, WHO histological classification, and EBER (Fig. 3a). As for SMARCA4, the results of univariate correlation analysis were sex, T stage, differentiation status, WHO histological classification, Lauren classification, Ki-67, and MMR (Table 1). The results of multivariate logistic regression were differentiation status, WHO histological classification, PD-L1, and MMR (Fig. 3b). We further divided the group with SMARCA2 and SMARCA4 loss into single-loss and co-loss subgroups. Univariate correlation analysis found that the co-loss subgroup was more likely to occur lymph node metastasis, poor differentiation, and a dMMR phenotype (Table S3).
Survival analysis
Although there was no survival difference among the four expression patterns of SMARCA2 (Fig. 4a), after integration, the lost group showed worse survival than the present group (P = 0.032) (Fig. 4b). The SMARCA4 lost group also exhibited worse survival than the other three groups (P = 0.043) (Fig. 4d), and this difference was more obvious after integration (P = 0.0048) (Fig. 4e). However, we did not observe the differences between the intact and attenuated groups for either SMARCA2 or SMARCA4 (Fig. 4c, f). We performed survival analysis again after stratification by TNM stage and found that SMARCA2 loss was associated with worse survival in early gastric carcinoma (P = 0.0044) (Fig. 5a), while SMARCA4 loss was related to worse survival in advanced gastric carcinoma (P = 0.0055) (Fig. 5d). In addition, we performed survival analysis in subgroups with single-loss and co-loss of SMARCA2 and SMARCA4; however, we did not observe a difference (Fig. S1).
Survival analysis performed according to the immunohistochemical expression of SMARCA2 and SMARCA4. There was no difference in survival among the four expression patterns of SMARCA2 (a), and the lost expression pattern of SMARCA4 showed worse survival than the other three expression patterns (d). After binary classification, the lost expression patterns of SMARCA2 (b) and SMARCA4 (e) were all associated with worse survival, while the attenuated expression patterns of SMARCA2 (c) and SMARCA4 (f) were not related to survival. Abbreviations: OS, overall survival; Hete, heterogeneous; Inta, intact; Redu, reduced; Pres, present; Atte, attenuated
Survival analysis performed after stratification of SMARCA2 and SMARCA4 according to TNM stage. SMARCA2 loss resulted in worse survival in early gastric cancer (a) and did not affect survival in advanced gastric cancer (b). SMARCA4 loss led to worse survival in advanced gastric cancer (d) and did not affect survival in early gastric cancer (c)
Univariate and multivariate Cox regression analysis
We performed a univariate Cox regression analysis and identified 15 variables with P < 0.05, including age, T stage, N stage, M stage, size, WHO histological classification, Lauren classification, differentiation status, VI, PNI, p53, MMR, SMARCA2, SMARCA4, and chemotherapy (Table 2). We then adopted these 15 variables into the multivariate Cox regression model and finally screened out six independent prognostic factors, including age, T stage, N stage, M stage, SMARCA2, and chemotherapy (Table 2), which were visualized in the form of a forest plot (Fig. 6a).
Forest plot and nomogram based on multivariate Cox regression analysis. a The forest plot showed that age ≥ 60 years, more advanced T stage, N stage, M stage, and the loss of SMARCA2 were risk factors for prognosis. In contrast, chemotherapy was a favorable factor. b The nomogram showed a good predictor of overall survival at 1, 3, and 5 years
Construction and evaluation of a nomogram prognostic model
Based on the above six independent prognostic factors, we constructed a prognostic nomogram model (Fig. 6b), and we compared the new model with the conventional AJCC TNM model. The C-index and 95% confidence interval (CI) of the new model and AJCC TNM model were 0.786 (0.774, 0.798), and 0.766 (0.754, 0.778), respectively. Subsequently, we evaluated the prediction consistency of the new model, the discrimination and the clinical utility of the two models, then drew calibration curves (Fig. 7a–c), ROC curves (Fig. 7d–f), and clinical decision curves (Fig. 7g–i), showing good prediction consistency, higher discrimination ability and better clinical utility in the new model. Finally, we calculated the NRI and IDI to evaluate the improvement of the predictive ability. Compared with the AJCC TNM model, the NRI and 95% CI of the new model at 1, 3, and 5 years were 0.414 (0.181, 0.667), 0.424 (0.192, 0.551), and 0.298 (0.093, 0.419), respectively. The IDI and 95% CI at 1, 3 and 5 years were 0.020 (0.003, 0.043), P < 0.01; 0.031 (0.016, 0.057), P < 0.001; and 0.012 (0.000, 0.036), P < 0.05, respectively.
Calibration plots, ROC curves, and DCA for the nomogram model and AJCC TNM model. a–c Calibration plots showed good consistency between actual and predicted survival at 1, 3, and 5 years. d–f ROC curves showed a better AUC of the nomogram model than the AJCC TNM model in predicting survival at 1, 3, and 5 years. g–i DCA showed that the nomogram model has better clinical utility than the AJCC TNM model. Abbreviations: DCA, decision curve analysis; ROC, receiver operating characteristic curve; AUC, areas under the ROC curve
Discussion
SMARCA2 and SMARCA4 are ATPase subunits in the SWI/SNF complex and are essential for the activity of the SWI/SNF complex. They all belong to the SWI2/SNF2 family, share approximately 75% structural homology, and have similar ATPase and helicase activities [11]. The two subunits are both mutually exclusive and complementary. Mutual exclusivity is reflected by the fact that SWI/SNF complex utilizes different ATPases or different ratios of ATPases under different conditions [12], whereas complementarity is reflected by the fact that a decrease or absence of one ATPase subunit leads to a compensatory increase in the other [13].
The loss ratio of SMARCA4 in this study (6.77%) is consistent with the mutation ratio in the TCGA database (6%), indicating that mutation is the primary mechanism of SMARCA4 loss [14]. However, the loss ratio of SMARCA2 in this study (18.57%) was much higher than the mutation ratio in the TCGA database (5%) (Fig. 2c). Indeed, mutations in the SMARCA2 gene are uncommon in most SMARCA2 deficient tumors, suggesting that epigenetic regulation plays a more critical role in SMARCA2 inactivation, such as methylation of CpG islands in the promoter region of the SMARCA2 gene [15, 16]. In addition, SMARCA2 gene promoter insertion polymorphisms [17], posttranslational modifications such as acetylation [18], and loss of chromosome 9p (the location where the SMARCA2 gene is located) [15, 19] all result in the loss of SMARCA2. In brief, the mechanism of SMARCA2 loss is more complicated than that of SMARCA4.
The SWI/SNF complex plays an essential role in regulating cell differentiation [5], which was also confirmed by our correlation analysis that the poorly differentiated state was significantly associated with the loss of SMARCA2 or SMARCA4; in other words, SMARCA2 or SMARCA4 loss could lead to poor differentiation of tumors. We can reasonably assume that the tumors partly originate from the differentiation disorder of normal cells due to the loss of SMARCA2 or SMARCA4. Another finding from the correlation analysis of this study was that the loss of SMARCA2 or SMARCA4 could more easily lead to a dMMR phenotype, which could induce a large number of neoantigens and further promote the infiltration of immune cells and ultimately improve the immune microenvironment in tumors [20]. Thus, SMARCA2 and SMARCA4 hold promising potential as immunotherapeutic markers. It was documented that SMARCA4 mutant Small Cell Carcinoma of the Ovary, Hypercalcemic Type (SCCOHT) exhibited active immune microenvironment [21], and SMARCA4 deficient non-small cell lung cancer (NSCLC) responded significantly to immune checkpoint inhibitors [22]. A pan-cancer analysis also confirmed that SMARCA4 was associated with immune infiltration in multiple types of cancers [23]. However, the literature on the relationship between SMARCA2 loss and tumor immune infiltration or immunotherapy has rarely been reported. The correlation between SMARCA2 and the immune microenvironment still needs further investigation.
It has been widely accepted that the SWI/SNF complex is a tumor suppressor and the loss of complex subunits leads to a worse prognosis [7]. This present study also showed that the loss of SMARCA4 or SMARCA2 led to a worse prognosis (Fig. 4b, e). However, a pan-cancer study showed that high SMARCA4 expression is associated with poor prognosis in many types of tumors, including liver hepatocellular carcinoma and kidney renal clear cell carcinoma [24]. Similarly, in pancreatic and ovarian cancer, high expression of SMARCA2 induced chemoresistance and further led to tumor progression, illustrating the tumor-promoting role of SMARCA2 [25, 26]. In brief, SMARCA2 and SMARCA4 acted as tumor suppressors in most cases but tumor promoters in other tumor types or at certain specialized stages, indicating the roles of SMARCA2 and SMARCA4 were context- specific [27].
The majority of current clinical research on SWI/SNF complex subunits mainly focused on the lost pattern, while in this study, we also observed reduced and heterogeneous expression patterns. Survival analyses identified that the reduced and heterogeneous expression of the subunit was insufficient to affect survival (Fig. 4c, f), while when the subunit was completely lost, did it profoundly affect survival (Fig. 4b, e), which is easily explained, the lost pattern resulted in a complete loss of subunit function, whereas the reduced and heterogeneous pattern implied partial retention of the subunit function. The different expression patterns of the subunits implied distinct molecular landscapes and corresponding clinical features, and the underlying molecular mechanisms still required more in-depth investigation.
Another finding for survival analysis in our present study was that SMARCA2 loss caused a worse prognosis in early gastric adenocarcinoma (Fig. 5a), whereas SMARCA4 loss was associated with a worse prognosis in advanced gastric adenocarcinoma (Fig. 5d), suggesting that the two subunits played different roles in different stages. A study of undifferentiated gastric carcinomas revealed that SMARCA4 loss, rather than SMARCA2 loss, led to an unfavorable prognosis [28]. In contrast, our large-scale cases of gastric adenocarcinoma showed that SMARCA2 was an independent factor of poor prognosis, indicating that the prognostic significance of these two subunits also varied between groups of different differentiation states.
Loss of SMARCA2 and SMARCA4 was found to be mutually exclusive in one study of undifferentiated gastrointestinal carcinoma [29]; however, the small sample size led to a decline in persuasion. Co-loss of SMARCA2 and SMARCA4 has been described in SCCOHT, NSCLC, and endometrial carcinoma [30,31,32]. We also found that immunohistochemical loss of SMARCA2 and SMARCA4 can occur concomitantly or independently in this present study (Fig. 2b). Further investigation revealed that the co-loss group was more likely to occur lymph node metastasis and poor differentiation than the single-loss group; however, it did not show a worse prognosis (Fig. S1). The mechanism underlying the co-loss of SMARCA2 and SMARCA4 has not yet been fully understood.
There are many studies concerned SMARCA4 in tumors, particularly in undifferentiated carcinomas [14, 33], whereas the role of SMARCA2 has been neglected. SMARCA2 and SMARCA4 were all associated with poor prognosis in our study; however, after multivariate Cox regression analysis, only SMARCA2 was an independent prognostic factor. The independent prognostic role of SMARCA2 was also confirmed in our previous lung cancer study [34]. Given the histological and prognostic significance, we consider that SMARCA2 lost gastric adenocarcinoma may represent a unique molecular subgroup that merits further treatment targeting SMARCA2. Ideas for utilizing SMARCA2 in anticancer therapy are emerging, such as histone deacetylase (HDAC) inhibitors [35], Enhancer of zeste homolog 2 (EZH2) inhibitors [36] and synthetic lethality approach targeted against SMARCA2 ATPase domain or bromodomain [37].
Currently, the AJCC TNM staging system is essential for the prognosis of patients with gastric carcinoma [38]. However, some patients with the same TNM stage showed significantly different prognoses. Therefore, a more scientific predictive system is urgently needed. Here, we constructed a novel nomogram prognostic model that included T stage, N stage, M stage, age, chemotherapy information, and expression status of SMARCA2. The calibration was evaluated by the calibration curve, which reflects the consistency between predicted and actual survival probabilities. The discrimination ability was assessed by AUC, or C-index, reflecting the model's accuracy in discriminating individuals. Clinical utility was evaluated by DCA, reflecting whether the model could benefit patients by influencing the clinical decision. In addition, to increase sensitivity when comparing the predictive ability of two models, we applied NRI and IDI, which reflect the extent to which the prediction performance can be improved [39]. This study showed that the nomogram prognostic model outperformed the conventional AJCC TNM model in predictive consistency, discrimination, and clinical utility.
Conclusion
In gastric adenocarcinoma, the loss of SMARCA2 and SMARCA4 correlated with poor differentiation and led to a worse prognosis. We combined SMARCA2 with other well-established prognostic factors to develop a novel nomogram prognostic model and found that the new model outperformed the conventional AJCC TNM model in concordance, discrimination, and clinical utility. SMARCA2 is not only an independent prognostic factor but also an emerging therapeutic target, and testing for SMARCA2 is recommended for patients with gastric adenocarcinoma.
Data availability
The datasets for this study are available from the corresponding author.
References
Sung H, Ferlay J, Siegel RL et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249
Zhang XY, Zhang PY (2017) Gastric cancer: somatic genetics as a guide to therapy. J Med Genet 54(5):305–312
Gao JP, Xu W, Liu WT et al (2018) Tumor heterogeneity of gastric cancer: from the perspective of tumor-initiating cell. World J Gastroenterol 24(24):2567–2581
Narlikar GJ, Sundaramoorthy R, Owen-Hughes T (2013) Mechanisms and functions of ATP-dependent chromatin-remodeling enzymes. Cell 154(3):490–503
Clapier CR, Iwasa J, Cairns BR et al (2017) Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat Rev Mol Cell Biol 18(7):407–422
Alfert A, Moreno N, Kerl K (2019) The BAF complex in development and disease. Epigenetics Chromatin 12(1):19
Mittal P, Roberts CWM (2020) The SWI/SNF complex in cancer - biology, biomarkers and therapy. Nat Rev Clin Oncol 17(7):435–448
Centore RC, Sandoval GJ, Soares LMM et al (2020) Mammalian SWI/SNF chromatin remodeling complexes: emerging mechanisms and therapeutic strategies. Trends Genet 36(12):936–950
Huang SC, Ng KF, Chang IY et al (2021) The clinicopathological significance of SWI/SNF alterations in gastric cancer is associated with the molecular subtypes. PLoS ONE 16(1):e0245356
Gluckstein MI, Dintner S, Arndt TT et al (2021) Comprehensive immunohistochemical study of the SWI/SNF complex expression status in gastric cancer reveals an adverse prognosis of SWI/SNF deficiency in genomically stable gastric carcinomas. Cancers (Basel) 13 (15)
Chiba H, Muramatsu M, Nomoto A et al (1994) Two human homologues of Saccharomyces cerevisiae SWI2/SNF2 and Drosophila brahma are transcriptional coactivators cooperating with the estrogen receptor and the retinoic acid receptor. Nucleic Acids Res 22(10):1815–1820
Reisman DN, Sciarrotta J, Bouldin TW et al (2005) The expression of the SWI/SNF ATPase subunits BRG1 and BRM in normal human tissues. Appl Immunohistochem Mol Morphol 13(1):66–74
Wang X, Sansam CG, Thom CS et al (2009) Oncogenesis caused by loss of the SNF5 tumor suppressor is dependent on activity of BRG1, the ATPase of the SWI/SNF chromatin remodeling complex. Cancer Res 69(20):8094–8101
Mardinian K, Adashek JJ, Botta GP et al (2021) SMARCA4: implications of an altered chromatin-remodeling gene for cancer development and therapy. Mol Cancer Ther 20(12):2341–2351
Xia QY, Zhan XM, Fan XS et al (2017) BRM/SMARCA2-negative clear cell renal cell carcinoma is associated with a high percentage of BRM somatic mutations, deletions and promoter methylation. Histopathology 70(5):711–721
Jancewicz I, Siedlecki JA, Sarnowski TJ et al (2019) BRM: the core ATPase subunit of SWI/SNF chromatin-remodelling complex-a tumour suppressor or tumour-promoting factor? Epigenetics Chromatin 12(1):68
Ouyang X, Ye XL, Wei HB (2017) BRM promoter insertion polymorphisms increase the risk of cancer: a meta-analysis. Gene 626:420–425
Bourachot B, Yaniv M, Muchardt C (2003) Growth inhibition by the mammalian SWI-SNF subunit Brm is regulated by acetylation. EMBO J 22(24):6505–6515
La Rochelle J, Klatte T, Dastane A et al (2010) Chromosome 9p deletions identify an aggressive phenotype of clear cell renal cell carcinoma. Cancer 116(20):4696–4702
Puliga E, Corso S, Pietrantonio F et al (2021) Microsatellite instability in gastric cancer: between lights and shadows. Cancer Treat Rev 95:102175
Jelinic P, Ricca J, Van Oudenhove E et al (2018) Immune-active microenvironment in small cell carcinoma of the ovary, hypercalcemic type: rationale for immune checkpoint blockade. J Natl Cancer Inst 110(7):787–790
Naito T, Umemura S, Nakamura H et al (2019) Successful treatment with nivolumab for SMARCA4-deficient non-small cell lung carcinoma with a high tumor mutation burden: a case report. Thorac Cancer 10(5):1285–1288
Peng L, Li J, Wu J et al (2021) A pan-cancer analysis of SMARCA4 alterations in human cancers. Front Immunol 12:762598
Guerrero-Martinez JA, Reyes JC (2018) High expression of SMARCA4 or SMARCA2 is frequently associated with an opposite prognosis in cancer. Sci Rep 8(1):2043
Zhang Z, Wang F, Du C et al (2017) BRM/SMARCA2 promotes the proliferation and chemoresistance of pancreatic cancer cells by targeting JAK2/STAT3 signaling. Cancer Lett 402:213–224
Xu X, Zheng Z, Jia L et al (2018) Overexpression of SMARCA2 or CAMK2D is associated with cisplatin resistance in human epithelial ovarian cancer. Oncol Lett 16(3):3796–3804
Wu Q, Lian JB, Stein JL et al (2017) The BRG1 ATPase of human SWI/SNF chromatin remodeling enzymes as a driver of cancer. Epigenomics 9(6):919–931
Chang B, Sheng W, Wang L et al (2021) SWI/SNF complex-deficient undifferentiated carcinoma of the gastrointestinal tract: clinicopathologic study of 30 cases with an emphasis on variable morphology, immune features, and the prognostic significance of different SMARCA4 and SMARCA2 subunit deficiencies. Am J Surg Pathol
Agaimy A, Daum O, Markl B et al (2016) SWI/SNF Complex-deficient undifferentiated/rhabdoid carcinomas of the gastrointestinal tract: a series of 13 cases highlighting mutually exclusive loss of SMARCA4 and SMARCA2 and frequent co-inactivation of SMARCB1 and SMARCA2. Am J Surg Pathol 40(4):544–553
Karnezis AN, Wang Y, Ramos P et al (2016) Dual loss of the SWI/SNF complex ATPases SMARCA4/BRG1 and SMARCA2/BRM is highly sensitive and specific for small cell carcinoma of the ovary, hypercalcaemic type. J Pathol 238(3):389–400
Herpel E, Rieker RJ, Dienemann H et al (2017) SMARCA4 and SMARCA2 deficiency in non-small cell lung cancer: immunohistochemical survey of 316 consecutive specimens. Ann Diagn Pathol 26:47–51
Ramalingam P, Croce S, McCluggage WG (2017) Loss of expression of SMARCA4 (BRG1), SMARCA2 (BRM) and SMARCB1 (INI1) in undifferentiated carcinoma of the endometrium is not uncommon and is not always associated with rhabdoid morphology. Histopathology 70(3):359–366
Chetty R, Serra S (2020) SMARCA family of genes. J Clin Pathol 73(5):257–260
Sun S, Li Q, Zhang Z et al (2022) SMARCA2 deficiency in NSCLC: a clinicopathologic and immunohistochemical analysis of a large series from a single institution. Environ Health Prev Med 27:3
Kahali B, Gramling SJ, Marquez SB et al (2014) Identifying targets for the restoration and reactivation of BRM. Oncogene 33(5):653–664
Chan-Penebre E, Armstrong K, Drew A et al (2017) Selective killing of SMARCA2- and SMARCA4-deficient small cell carcinoma of the ovary hypercalcemic type cells by inhibition of EZH2: in vitro and in vivo preclinical models. Mol Cancer Ther 16 (5):850–860
O’Neil NJ, Bailey ML, Hieter P (2017) Synthetic lethality and cancer. Nat Rev Genet 18(10):613–623
Shiraishi N, Sato K, Yasuda K et al (2007) Multivariate prognostic study on large gastric cancer. J Surg Oncol 96(1):14–18
Zhou ZR, Wang WW, Li Y et al (2019) In-depth mining of clinical data: the construction of clinical prediction model with R. Ann Transl Med 7(23):796
Acknowledgements
Not applicable.
Funding
None.
Author information
Authors and Affiliations
Contributions
Study design: ZZ, SZ. Patient tissue collection: ZZ, SS, ZL, SX. Pathology review: SZ, ZZ, QJL. Supply of reagents: QL, YZ. Data analysis: ZZ, SS. Manuscript: ZZ. Manuscript review: SZ, ZGC.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This study was approved by the Ethics Review Board of the Weihai Municipal Hospital (permission code: 2,021,053).
Informed consent
Informed consent was obtained from patients before enrollment in this study.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
About this article
Cite this article
Zhang, Z., Li, Q., Sun, S. et al. Expression of SMARCA2 and SMARCA4 in gastric adenocarcinoma and construction of a nomogram prognostic model. Int J Clin Oncol 28, 1487–1500 (2023). https://doi.org/10.1007/s10147-023-02403-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10147-023-02403-0