Abstract
Background
As a critical component of exosomes, circular RNAs (circRNAs) have shown great value in cancer diagnosis. This study aimed to identify circRNAs in exosomes for the diagnosis of PTC (papillary thyroid carcinoma).
Methods
We selected hsa_circ_0082002 and hsa_circ_0003863 based on circRNA microarray. The levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 in the sera of healthy control (n = 68), benign thyroid tumors (n = 60), and PTC without and with Hashimoto’s thyroiditis (n = 164) were quantified by qPCR (quantitative polymerase chain reaction). Receiver operating characteristic analyses were conducted to evaluate the diagnostic sensitivity and specificity. Bioinformatics databases were used to predict the microRNAs and proteins binding with hsa_circ_0082002 and hsa_circ_0003863.
Results
The levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 were positively associated and statistically increased in PTC compared to healthy and benign thyroid tumors. Intriguingly, higher levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 were positively correlated with lymph node metastasis and vascular invasion in PTC. Further stability tests show that exosomal hsa_circ_0082002 and hsa_circ_0003863 could exist stably in sera treated by several freeze–thaw cycles at -20 °C and with a storage time shorter than 24 h at 4 °C. Furthermore, hsa_circ_0082002 and hsa_circ_0003863 were predicted to interact with microRNAs and proteins, suggesting that hsa_circ_0082002 and hsa_circ_0003863 might contribute to the occurrence and progression of PTC through interacting with microRNAs and RNA binding proteins.
Conclusion
Collectively, we identified two PTC-related circRNAs incorporated in exosomes and uncovered their potential as tumor markers to diagnose PTC, in particular, more aggressive PTC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Papillary thyroid carcinoma (PTC) is the most common pathological type of thyroid cancer, accounting for 85% to 90% of all cases [1]. Although PTC usually exhibits low malignancy, its recurrence rate remains high. Patients with PTC often develop lymph node metastasis, vascular invasion, and distant metastasis, resulting in a poor prognosis or possibly lifelong complications. Fine needle aspiration biopsy (FNAB) is the most widely used and accurate diagnostic method for thyroid cancer. However, the detection rate of micro-tumor by FNAB is less than satisfactory [2]. And patients generally have low acceptance of the invasive examination method, which limits the application of FNAB in the early diagnosis of thyroid cancer [3]. The standard non-invasive methods for the early diagnosis of thyroid cancer include thyroid globulin assay, positron emission tomography (PET), and computed tomography (CT). However, the sensitivity and specificity of serum thyroglobulin detection remain to be discussed [4,5,6]. PET and CT have specific radiation effects on the human body, and the examination cost is high. Therefore, considering the above situation of clinical diagnosis of thyroid cancer, we urgently need a novel, non-invasive marker to improve the sensitivity and specificity of early diagnosis of thyroid cancer.
Exosomes are extracellular vesicles with a lipid bilayer structure ranging from 30 to 200 nm in diameter [7]. Exosomes mediate cell-to-cell communication by transporting their bioactive contents, including microRNA (miRNA), long non-coding RNA (lncRNA), circular RNA (circRNA), messenger RNA (mRNA), DNA, and proteins to recipient cells, thus affecting the physiological and pathological functions of the recipient cells [7]. Through intercellular communication, exosomes play a vital role in tumorigenesis and development [7, 8]. In recent years, numerous studies have demonstrated that exosomes are involved in multiple phases of tumor occurrence and progression, like tumor growth, cancer cell apoptosis, epithelial-mesenchymal transition, angiogenesis, immunosuppression, and chemotherapy resistance, by releasing their bioactive contents into recipient cells [8, 9].
As a critical cargo of exosomes, circRNA is highly enriched and stably present in exosomes [10]. CircRNA, a non-coding RNA distinguished by a stable closed-loop structure, mainly regulates gene expression during transcription and translation [11, 12]. It is involved in many pathological processes, such as Alzheimer’s disease, atherosclerosis, and cancers [13]. Accumulated documents have shown that circRNA plays a vital role in tumor growth, angiogenesis, metastasis, drug resistance, etc., through different mechanisms [14]. The most crucial function of circRNAs is acting as miRNA sponges to adsorb miRNAs, thereby releasing the inhibitory effect of miRNAs on their target genes [15]. Additionally, circRNA can interact with proteins to regulate gene expression at transcriptional and translational levels and act as translation templates [16]. Recent studies showed many different circRNAs involved in the development of PTC and their altered expression in tissues, blood, and exosomes as a characteristic sign of specific clinicopathological parameters of PTC [17].
Therefore, considering the vital role of circRNAs in PTC and their abundance in exosomes, we aimed to explore PTC-related circRNAs within exosomes, which could be served as potential diagnostic biomarkers of PTC. We first performed circRNA microarray analysis to profile circRNAs within serum exosomes of patients with PTC. Then we selected upregulated hsa_circ_0082002 and hsa_circ_0003863 as the interest circRNAs for further quantification in four subsets (healthy, benign thyroid tumors, PTC plus Hashimoto’s thyroiditis, and PTC). We found that the levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 were significantly elevated in PTC compared to the subpopulations of healthy and benign thyroid tumors, showing higher sensitivity and specificity to diagnose PTC. Intriguingly, the levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 were positively associated, and both of them were related to more aggressive features of PTC, like lymph node metastasis and vascular invasion. The present study identified two exosomal circRNAs as potential biomarkers for PTC diagnosis and further predicted the vital role of exosomal hsa_circ_0082002 and hsa_circ_0003863 in the development of PTC. And the combination of exosomal hsa_circ_0082002 and hsa_circ_0003863 might be a strategy to improve diagnostic sensitivity and specificity for PTC.
Materials and methods
Serum specimens enrolled in the present study
With the approval of the Ethics Review Board of the Ningbo No.2 Hospital (YJ-NBEY-KY-2021-181-01), we collected serum samples from 68 healthy people, 60 patients with benign thyroid tumors, 44 patients with PTC plus Hashimoto’s thyroiditis, and 120 patients with PTC from the Ningbo No.2 Hospital at the initial diagnosis before surgery between October 2021 and September 2022. The clinical parameters (age, gender, tumor stage, tumor size, distant metastasis, vascular invasion, lymph node metastasis) of each subset are shown in Table 1.
The isolation of serum exosomes
Exosomes were extracted directly from sera with the exosome isolation kit [18]. Briefly, 400 μL of serum without cells and debris were incubated with 80 μL of Total Exosome Isolation Reagent for Serum (Life Technologies, Austin, United States) at 4 °C for 30 min. After incubation, serum exosomes were obtained by centrifugation at 10,000 g for 10 min. The isolated exosomes were resuspended and stored in PBS (Phosphate-Buffered Saline) for further identification and RNA extraction.
The identification of serum exosomes
After serum exosome isolation, we carried out western blot to check three exosome-specific markers (CD9, CD63, and CD81). Additionally, the size and morphology of isolated exosomes were tested by scanning electron microscope (SU-70, HITACHI, Japan) and a potentiometric analyzer (Zetasizer, ZEN3700, Malvern, England), respectively.
Protein extraction and western blot
Exosome proteins were lysed in RIPA buffer (50 mM Tris pH8, 0.1% SDS, 150 mM NaCl, 1% NP40, 0.5% Na-deoxycholate) plus 1 mM PMSF (Phenylmethylsulfonyl Fluoride) and phosphatase inhibitor (Beyotime, China). Then, lysed exosome proteins were firstly separated by SDS-PAGE (Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis) and then transferred to PVDF (Polyvinylidene Fluoride) membranes (Immobilon-P Membrane, Merk Millipore, United States) as previously described[19]. To detect the exosome-specific markers, three primary antibodies specific to the exosomal proteins (CD9, CD63, and CD81), including anti-CD9 (AP1482d-ev, ABGENT, United States), anti-CD63 (AP5333b-ev, ABGENT, United States) and anti-CD81 (bs-6934R, Bioss, China), and the secondary antibodies, peroxidase affinipure goat anti-mouse IgG (115–035-003E, Jackson, United States), and affinipure goat anti-rabbit IgG (111–035-003E, Jackson, United States), were used. Finally, the exosomal protein bands were presented with the super ECL detection reagent (36208ES60, YEASEN, China) and photographed with the infrared imaging system (Li-COR, United States).
RNA isolation from exosomes
To extract total RNAs from exosomes, we resuspended exosomes in PBS to get a 400 µL solution and added 1.2 mL trizol (Invitrogen, Thermo Fisher Scientific, United States). The mixture was incubated at 4 °C for 5 min, then mixed with 240 µL chloroform (ChemMall, China), and continued to incubate at 4 °C for another 5 min. After centrifugation at 12,000 × g for 15 min at 4 °C, the supernatant was transferred to a new tube and mixed with 600 µL isopropanol (ChemMall, China). The RNAs were precipitated by centrifugation at 12,000 × g for 10 min at 4 °C and then purified with 75% ethanol twice. At last, the purified RNAs were dissolved in 10 µL nuclease-free water and then quantified by a NanoDrop spectrophotometer (NanoDrop™ One, Thermo Fisher Scientific, United States).
CircRNA microarray
We isolated RNAs from the serum exosomes of 4 healthy individuals and 4 patients with PTC. The circRNAs in exosomes were profiled with Arraystar Human CircRNA Array (Arraystar, Rockville, MD, USA) following the Arraystar’s Super RNA Labeling protocol. Hierarchical clustering was applied to present the circRNA expression pattern between the healthy and PTC groups. Statistically differentially expressed circRNAs between the healthy and PTC groups were characterized by fold changes > 2.0 and P values < 0.05.
Reverse transcription and quantitative polymerase chain reaction (qPCR)
Following the RNA extraction from exosomes, the reverse transcription kit (ReverTra Ace qPCR RT Master Mix with gDNA Remover, TOYOBO, Japan) was immediately used to synthesize cDNAs as previously described [20]. For qPCR, the divergent primers used to amplify hsa_circ_0082002 and hsa_circ_0003863 were designed by Primer 3 Plus (http://www.primer3plus.com/cgi-bin/dev/primer3plus.cgi), and synthesized by BGI (Beijing Genomics Institution, China). The primer sequences of hsa_circ_0082002, hsa_circ_0003863, and GAPDH were summarized in supplementary table 1. The fluorescent dye SYBR (Yeason, China) was chosen to perform qPCR. The qPCR system was prepared as previously described [19]. Subsequently, the qPCR was run with the procedure of hot-start at 95 °C for 5 min, 40 cycles of 94 °C for 30 s, 60 °C for 30 s, and 72 °C for 30 s, then 98 °C for 10 min, and finally hold at 4 °C on a Mastercycler Gradient (Vaudaux-Eppendorf, Germany). Each reaction was carried out in triplicate. Based on the previous publications[19, 21, 22] and the constant expression levels of GAPDH among different serum subsets, in the present study, we selected GAPDH as the internal reference gene to normalize the qPCR data. The relative expression levels of hsa_circ_0082002 and hsa_circ_0003863 were calculated with the formula 2^ (mean Cq value of GAPDH—mean Cq value of circRNA). The melting curve and agarose gel electrophoresis were used to examine the specificity of qPCR products.
Agarose gel electrophoresis
We performed agarose gel electrophoresis to detect the qPCR product size of hsa_circ_0082002 and hsa_circ_0003863. And the effect of RNase R digestion on the circular and linear 0082002 and 0003863 amplification fragments was also checked. According to the length of the PCR products, 10% agarose gel was prepared, and the electrophoresis was carried out at 100 V for 40 min. Finally, the bands were visualized in the gel imager (GelDoc XR+, Bio-Rad, Germany).
Sequencing
Sanger sequencing was carried out to verify the circular structure of hsa_circ_0082002 and hsa_circ_0003863 after the amplification by qPCR. We used the divergent primer to detect the specific circRNA identifier BSJ in the qPCR products.
Statistical analyses
SPSS 22.0 Software package (SPSS Inc. Chicago, United States) and GraphPad Prism 8.0 (GraphPad Software, United States) were used for the present study’s statistical analyses. The Ct values of circRNAs between the healthy and PTC subpopulations were compared by student t-test. The levels of circRNAs between tumor sizes of ≤ 1 cm and > 1 cm, Stage I and II-III, lymph node metastasis negative and positive, vascular invasion negative and positive, were compared by Mann–Whitney U test. To achieve normal distribution, the calculated relative levels of circRNAs were transformed by Ln. The levels of circRNAs among the 4 cohorts of healthy, benign thyroid tumors, PTC plus Hashimoto’s thyroiditis, and PTC were compared by ANOVA and Tukey’s HSD test. Receiver operating characteristic (ROC) analyses were carried out to estimate the diagnostic sensitivity and specificity of circRNAs to distinguish healthy and benign, healthy and PTC, benign thyroid tumors and PTC, stage I and II-III, lymph node metastasis negative and positive, as well as vascular invasion negative and positive. The areas under the curves (AUC) of ROC were calculated. Pearson correlation analysis was used to evaluate the correlation of hsa_circ_0082002 and hsa_circ_0003863. The logistic regression model was used to combine hsa_circ_0082002 and hsa_circ_0003863, and the probability of the combination was calculated. The Ct values of exosomal circRNAs in serum samples treated by freeze–thaw cycles and different storage times were compared by student t-test. The possible regulatory mechanisms of hsa_circ_0082002 and hsa_circ_0003863 were predicted by Cancer-Specific CircRNA Database (CSCD, http://gb.whu.edu.cn/CSCD). The interactions of hsa_circ_0082002 and hsa_circ_0003863 with miRNAs and proteins were downloaded from circbank (http://www.circbank.cn/index.html) and were visualized by the software Cytoscape_v3.9.1. A P value < 0.05 was considered statistically significant; all P values are two-sided.
Results
The isolation and the characterization of serum exosomes
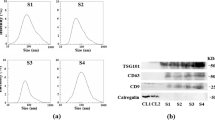
We chose an exosome isolation kit to isolate total exosomes from serum. We first isolated exosomes from the sera of 2 healthy individuals and 2 patients with PTC who were randomly chosen. After the isolation, the specific proteins on exosomes, including CD63, CD81, and CD9, were checked by western blot (Fig. 1a). In addition, the shapes of isolated exosomes from the sera of healthy people (Fig. 1b) and patients with PTC (Fig. 1c) were visualized under the transmission electron microscopy. The typical round or oval structure of exosomes with a diameter range of 40–200 nm could be observed. Also, the serum exosomes isolated from healthy persons (Fig. 1d) and patients with PTC (Fig. 1e) show a size of 146 nm and 137 nm, respectively. The above examination of specific exosomal markers, the shape, and the size indicates the successful isolation of exosomes from sera.
The identification of isolated serum exosomes. The specific exosomal proteins (CD63, CD81, and CD9) were tested by western blot (a). A scanning electron microscope checked the morphology of serum exosomes from healthy donors (b) and patients with PTC (c). The size of serum exosomes from healthy donors (d) and patients with PTC (e) was 146 nm and 137 nm, respectively
The selection of differentially up-regulated circRNAs in the serum exosomes of patients with PTC
To select differentially expressed circRNAs in the serum exosomes of patients with PTC, circRNA microarray was performed with 4 serum samples of healthy control and 4 serum samples of PTC (Fig. 2a). We found 3, 773 differentially expressed circRNAs, of which 1074 were up-regulated, and 2, 699 were down-regulated in PTC compared to healthy control (Fig. 2b). GO annotation and KEGG pathway analyses were carried out to reveal the potential signaling pathways the differentially expressed circRNAs might involve in. GO annotation shows the top 30 enrichment of the differentially expressed circRNAs in biological processes, cellular components, and molecular function (Fig. 2c). KEGG pathway analyses identified the top 20 signaling pathways the differentially expressed circRNAs might involve in, of which “autoimmune thyroid disease”, “thyroid hormone synthesis”, “purine metabolism” and “gastric acid secretion” are the most significant ones (Fig. 2d). Then, we selected 9 circRNAs through the intersection of top 30 circRNAs with most significant fold changes, and top 42 circRNAs with smallest P values (Fig. 2e). We quantified the 9 most significantly dysregulated circRNAs in the serum exosomes of 20 healthy and 20 PTC samples by qPCR. Hsa_circ_0082002 and hsa_circ_0003863 were picked for further study based on their lower Ct values in PTC (Fig. 2f).
The selection of PTC-related exosomal circRNAs. CircRNA microarray was used to profile the expression patterns of exosomal circRNAs in 4 serum samples from healthy donors and 4 patients with PTC (a). Three thousand seven hundred and seventy-three statistically dysregulated circRNAs were found in the serum exosomes of PTC (b). The GO annotation shows the top 30 enrichment of the differentially expressed circRNAs (c). KEGG pathway analyses identified the top 20 signaling pathways in the differentially expressed circRNAs might involve in (d). The nine most significantly dysregulated circRNAs were selected through the intersection of the top 30 circRNAs with the most significant fold changes and the top 42 circRNAs with the smallest P values (e). The Ct values of the 9 selected circRNAs in the serum exosomes of 20 healthy and 20 PTC samples were evaluated by qPCR (f)
Identifying the circular structure of hsa_circ_0082002 and hsa_circ_0003863
To verify the circular structure of hsa_circ_0082002 and hsa_circ_0003863, we used RNase R to digest the linear RNAs in exosomes. We synthesized the convergent primers of 0082002 and 0003863 to amplify the corresponding linear RNAs of hsa_circ_0082002 and hsa_circ_0003863, respectively. As shown in Fig. 3a, with the digestion of RNase R, the divergent primers could amplify hsa_circ_0082002 and hsa_circ_0003863 fragments, indicating that the circular structure of hsa_circ_0082002 and hsa_circ_0003863 is stable to the digestion of RNase R. Still, the convergent primers failed to amplify the linear RNAs. Also, the qPCR products of hsa_circ_0082002 and hsa_circ_0003863 were sequenced. The sequencing results showed the right junction sites, confirming the successful amplification of hsa_circ_0082002 and hsa_circ_0003863 (Fig. 3b). Additionally, the amplification plots and the melt curve plots of hsa_circ_0082002 (Fig. 3c) and hsa_circ_0003863 (Fig. 3d) show their specific amplification by qPCR. The results suggest that we successfully amplified hsa_circ_0082002 and hsa_circ_0003863 in serum exosomes.
The confirmation of the circular structure of hsa_circ_0082002 and hsa_circ_0003863. The effect of RNase R on the hsa_circ_0082002, hsa_circ_0003863, and their corresponding linear RNAs (a). The sequencing of qPCR products of hsa_circ_0082002 and hsa_circ_0003863 (b). The plots show the amplification and melt curves of hsa_circ_0082002 (c) and hsa_circ_0003863 (d)
The serum levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 in different cohorts
To evaluate the diagnostic potential of exosomal hsa_circ_0082002 and hsa_circ_0003863 for PTC, we quantified their levels by qPCR in the sample populations of healthy, benign thyroid tumors, PTC plus Hashimoto’s thyroiditis and PTC. We analyzed and compared their levels in different cohorts. Surprisingly, we found a similar expression pattern of exosomal hsa_circ_0082002 to hsa_circ_0003863 (Fig. 4a, e). Specifically, the serum levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 increased gradually from healthy, benign thyroid tumors, PTC plus Hashimoto’s thyroiditis samples to PTC. They showed no difference between PTC with and without Hashimoto’s thyroiditis (Fig. 4a, e). ROC analyses were conducted to estimate the diagnostic power of exosomal hsa_circ_0082002 and hsa_circ_0003863 to discriminate different subsets. It was found that exosomal hsa_circ_0082002 had AUC values of 0.740 (Fig. 4b), 0.992 (Fig. 4c), and 0.848 (Fig. 4d) to differentiate healthy and benign thyroid tumors, healthy and all PTC (PTC with and without Hashimoto’s thyroiditis in the present study), and benign thyroid tumors and all PTC, respectively. For exosomal hsa_circ_0003863, the AUC values of 0.682 (Fig. 4f), 0.856 (Fig. 4g), and 0.712 (Fig. 4h) were obtained to distinguish healthy and benign thyroid tumors, healthy and all PTC, and benign thyroid tumors and all PTC, respectively. To further evaluate the diagnostic value of exosomal hsa_circ_0082002 and hsa_circ_0003863, we performed statistical analyses with the healthy samples (n = 68) and PTC samples with T1N0M0 (n = 100). We found that the levels of exosomal hsa_circ_0082002 (4i) and hsa_circ_0003863 (4 k) were statistically elevated in the PTC samples with T1N0M0 compared to those in healthy control, with the AUC values of 0.987 (4j) and 0.814 (4 l), respectively, to discriminate PTC with T1N0M0 and healthy, indicating their promising value as a screening tool for patients with PTC. Altogether, the serum levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 were significantly up-regulated in the patients with PTC, showing apparent diagnostic abilities to discriminate different subpopulations, remarkably benign and malignant thyroid tumors.
The levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 in different sample cohorts. The box plots show the levels of exosomal hsa_circ_0082002 in the subpopulations of healthy (n = 68), benign thyroid tumors (n = 60), PTC plus Hashimoto’s thyroiditis (n = 44), and PTC (n = 120) (a). ROC analyses were used to estimate the diagnostic sensitivity and specificity of exosomal hsa_circ_0082002 to discriminate healthy and benign thyroid tumors (b), healthy and all PTC samples (c), and benign and malignant thyroid tumors (d). The box plots show the levels of exosomal hsa_circ_0003863 in the subsets of healthy (n = 68), benign thyroid tumors (n = 60), PTC plus Hashimoto’s thyroiditis (n = 44), and PTC (n = 120) (e). ROC analyses were used to estimate the diagnostic sensitivity and specificity of exosomal hsa_circ_0003863 to differentiate healthy and benign thyroid tumors (f), healthy and all PTC samples (g), and benign and malignant thyroid tumors (h). The box plots show the levels of exosomal hsa_circ_0082002 (i) and hsa_circ_0003863 (k) in the subpopulations of healthy (n = 68) and PTC with T1N0M0 (n = 100). ROC analyses were used to estimate the diagnostic sensitivity and specificity of exosomal hsa_circ_0082002 (j) and hsa_circ_0003863 (l) to differentiate healthy and PTC with T1N0M0
The correlation of exosomal hsa_circ_0082002 and hsa_circ_0003863 with clinical pathological parameters of PTC
Since exosomal hsa_circ_0082002 and hsa_circ_0003863 were remarkably up-regulated in PTC, we were inspired to investigate their association with clinical factors of PTC, including age, gender, tumor size, tumor stage, lymph node metastasis, and vascular invasion. Regarding age and gender, no correlation could be found for exosomal hsa_circ_0082002 and hsa_circ_0003863 (data not shown). However, regarding the pathological factors, as shown in Table 2, the clinically-used tumor marker calcitonin showed no relationship with tumor size, tumor stage, lymph node metastasis, and vascular invasion. Exosomal hsa_circ_0082002 and hsa_circ_0003863 were associated with tumor stage, lymph node metastasis, and vascular invasion but not tumor size (Table 2). In detail, the serum levels of exosomal hsa_circ_0082002 were up-regulated in PTC with higher tumor stages II-III (Fig. 5a), positive lymph node metastasis (Fig. 5c), and positive vascular invasion (Fig. 5e), having AUC values of 0.747 (Fig. 5b), 0.736 (Fig. 5d), and 0.686 (Fig. 5f) to discriminate subsets of lower and higher tumor stages, positive and negative lymph node metastases, and positive and negative vascular invasion, respectively. For exosomal hsa_circ_0003863, no correlation could be found with tumor stages (Table 2). Still, it is up-regulated in subgroups of positive lymph node metastasis (Fig. 5g) and positive vascular invasion (Fig. 5i), showing AUC values of 0.732 (Fig. 5h) and 0.685 (Fig. 5j) to distinguish the subgroups of positive and negative lymph node metastases, and positive and negative vascular invasion, respectively. Therefore, considering the similar relationship of exosomal hsa_circ_0082002 and hsa_circ_0003863 with the status of lymph node metastasis and vascular invasion, we performed the correlation analyses of the two exosomal circRNAs and found a positive correlation of them in PTC samples (Fig. 5k). Subsequently, we combined exosomal hsa_circ_0082002 and hsa_circ_0003863 to estimate the diagnostic power. As shown in Fig. 5l, compared to single exosomal hsa_circ_0082002 (AUC = 0.736) and hsa_circ_0003863 (AUC = 0.732), the combination of them (AUC = 0.802) showed more excellent diagnostic value to differentiate positive and negative lymph node metastases. Similarly, the AUC values of single exosomal hsa_circ_0082002, hsa_circ_0003863, the combination of them were 0.686, 0.685, 0.726, respectively, to discriminate positive and negative vascular invasion (Fig. 5m). Collectively, higher serum levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 are correlated with more aggressive characteristics of PTC, and the combination of exosomal hsa_circ_0082002 and hsa_circ_0003863 shows better abilities to diagnose lymph node metastasis and vascular invasion of PTC.
The correlation of exosomal hsa_circ_0082002 and hsa_circ_0003863 with clinical pathological parameters of PTC. The plots show the levels of exosomal hsa_circ_0082002 in different subgroups of stage I (n = 152) and stage II-III (n = 10) (a), N0 (n = 100) and N1 (n = 64) (c), negative vascular invasion (n = 146) and positive vascular invasion (n = 18) (e) of PTC. ROC analyses were used to evaluate the diagnostic power of exosomal hsa_circ_0082002 to distinguish stage I and stage II-III (b), negative and positive lymph node metastases (d), and negative and positive vascular invasion (f). The plots show the levels of exosomal hsa_circ_0003863 in different subgroups of N0 (n = 100) and N1 (n = 64) (g), negative vascular invasion (n = 146) and positive vascular invasion (n = 18) (i) of PTC. ROC analyses were used to evaluate the diagnostic power of exosomal hsa_circ_0003863 to distinguish N0 and N1 (h) and negative and positive vascular invasion (j). The correlation of exosomal hsa_circ_0082002 and hsa_circ_0003863 in PTC (n = 164) (k). ROC analyses were performed to examine the diagnostic value of the combination of exosomal hsa_circ_0082002 and hsa_circ_0003863 to discriminate N0 and N1 (l) and negative and positive vascular invasion (m)
The stability of exosomal hsa_circ_0082002 and hsa_circ_0003863 in different circumstances
As a tumor marker, it should be stable for clinical detection. Therefore, we further examined the stability of exosomal hsa_circ_0082002 and hsa_circ_0003863 in different circumstances, including the freeze–thaw cycles at -20 °C and the storage time at 4 °C and room temperature. For exosomal hsa_circ_0082002, 12 serum samples from healthy individuals were randomly selected and divided into 3 groups (4 samples/group). The 3 groups were used to study the effect of the freeze–thaw cycles at -20 °C, the storage time at 4 °C, and room temperature on the levels of exosomal hsa_circ_0082002. We found that the freeze–thaw cycles at -20 °C (Fig. 6a) and the storage time at 4 °C (Fig. 6b) of 4 serum samples had no apparent influence on the Ct values of exosomal hsa_circ_0082002, but the storage time at room temperature might have an effect on the Ct values of exosomal hsa_circ_0082002 (Fig. 6c). For exosomal hsa_circ_0003863, another 12 serum samples from healthy individuals were randomly selected. Similar evaluation methods were applied to exosomal hsa_circ_0082002. We found that the freeze–thaw cycles at -20 °C (Fig. 6d) and the storage time at 4 °C (Fig. 6e) of 4 serum samples had no significant influence on the Ct values of exosomal hsa_circ_0003863, but the longer storage time at room temperature increased the Ct values of exosomal hsa_circ_0003863 in one sample (Fig. 6f). Taken together, these results indicate that exosomal hsa_circ_0082002 and hsa_circ_0003863 are stable with several freeze–thaw cycles at -20 °C, and with the storage time no longer than 24 h at 4 °C.
The stability of exosomal hsa_circ_0082002 and hsa_circ_0003863 in different circumstances. The Ct values of exosomal hsa_circ_0082002 in 4 samples treated with different freeze–thaw cycles at -20 °C (a) and different storage times at 4 °C (b) and room temperature (c). The Ct values of exosomal hsa_circ_0003863 in 4 samples treated with different freeze–thaw cycles at -20 °C (d) and different storage times at 4 °C (e) and room temperature (f)
The potential functional mechanisms of hsa_circ_0082002 and hsa_circ_0003863
Considering the up-regulation of exosomal hsa_circ_0082002 and hsa_circ_0003863 and their correlation with lymph node metastasis and vascular invasion in PTC, we were interested in exploring their potential role in PTC and underlying mechanisms. We first predicted the possible function of hsa_circ_0082002 and hsa_circ_0003863 through the CSCD database (http://gb.whu.edu.cn/CSCD/). As shown in Fig. 7a, b, the structure diagrams of hsa_circ_0082002 and hsa_circ_0003863 uncovered the microRNA response elements (MREs), the sites of RNA binding proteins, and the open reading frame, suggesting that they might bind with miRNAs, RNA binding proteins, and even encode proteins in physiological and pathological conditions. Furthermore, the bioinformatic database circbank (http://www.circbank.cn/searchCirc.html) was used to predict potential interacting miRNAs and proteins of hsa_circ_0082002 and hsa_circ_0003863. We found 98 miRNAs and 42 miRNAs interacting with hsa_circ_0082002 and hsa_circ_0003863, respectively, of which miR-6757-3p, miR-146a-3p, miR-149-5p, miR-216b-5p, and miR-5580-5p could bind with both of hsa_circ_0082002 and hsa_circ_0003863 (Fig. 7c). There were 11 proteins and 7 proteins binding with hsa_circ_0082002 and hsa_circ_0003863, respectively, and U2AF65, HuR, AGO2, PTB, eIF4AIII, and UPF1 might bind with both of hsa_circ_0082002 and hsa_circ_0003863 (Fig. 7d). Besides, we performed bioinformatic analyses of the exosomal hsa_circ_0082002 and hsa_circ_0003863 data derived from exoRBase2.0 (Home-exoRBase2.0) and found that the levels of plasma exosomal hsa_circ_0082002 (Supplementary Fig. 1a) were lower in a majority of cancers than those in healthy control. Whereas, the levels of plasma exosomal hsa_circ_0003863 (Supplementary Fig. 1b) were higher in a majority of cancers than those in healthy control. Furthermore, we carried out GO annotation analyses with the data from circNET (CircNet (cuhk.edu.cn) and revealed that hsa_circ_0082002 were mainly involved in nuclear division and mitotic nuclear division (Supplementary Fig. 2a), and people’s characteristics of Byzanthine arch palate, and congenital epicanthus (Supplementary Fig. 2b). About hsa_circ_0003863, it is mainly involved in catabolic process (Supplementary Fig. 2c), and fibrosarcoma, and meningioma (Supplementary Fig. 2d). These results indicate that hsa_circ_0082002 and hsa_circ_0003863, in particular hsa_circ_0003863 might play vital roles in cancers. Since one circRNA plays its role mainly through interacting with miRNAs and proteins, or even coding functional peptides or proteins, these findings provide clues for further elucidating the contribution of hsa_circ_0082002 and hsa_circ_0003863 to PTC and their possible mechanisms.
The potential functional mechanisms of hsa_circ_0082002 and hsa_circ_0003863 in PTC. The structure diagrams show the possible function of hsa_circ_0082002 (a) and hsa_circ_0003863 (b) by the CSCD database. The potential interacting miRNAs (c) and proteins (d) of hsa_circ_0082002 and hsa_circ_0003863 were predicted with the bioinformatic database circbank
Discussion
Recently, exosomes have emerged as a new frontier of cancer liquid biopsy. Their dysregulated contents, including miRNAs, lncRNAs, circRNAs, DNAs, and proteins, are frequently reported as promising tumor markers for cancer detection, prognosis prediction, and monitoring treatment response[23]. The present study aimed to identify PTC-related circRNAs in exosomes to diagnose PTC. We selected hsa_circ_0082002 and hsa_circ_0003863 based on circRNA microarray and preliminary testing in healthy and PTC serum samples (Fig. 2). We further quantified the levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 in the serum samples of healthy, benign thyroid tumors and PTC without and with Hashimoto’s thyroiditis by qPCR. It was found that the levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 were increased in PTC samples compared to the samples of healthy and benign thyroid tumors, showing diagnostic abilities to distinguish PTC from healthy and benign thyroid tumors (Fig. 4). Intriguingly, in PTC samples, the levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 are positively associated. Moreover, higher levels of exosomal hsa_circ_0082002 and hsa_circ_0003863 were positively correlated with the characteristics of lymph node metastasis and vascular invasion in PTC (Fig. 5), indicating their involvement in PTC progression. Further stability tests show that exosomal hsa_circ_0082002 and hsa_circ_0003863 could exist stably in sera treated by several freeze–thaw cycles at -20 °C and with a storage time shorter than 24 h at 4 °C (Fig. 6), indicating the feasibility to quantify exosomal hsa_circ_0082002 and hsa_circ_0003863 in the clinic. Based on the potential of exosomal hsa_circ_0082002 and hsa_circ_0003863 in diagnosing PTC, particularly more malignant PTC, we further study the possible function of hsa_circ_0082002 and hsa_circ_0003863 in PTC. We utilized bioinformatic software to predict the functional mechanism of hsa_circ_0082002 and hsa_circ_0003863. It was found that both hsa_circ_0082002 and hsa_circ_0003863 could interact with miRNAs and proteins. However, hsa_circ_0082002 but not hsa_circ_0003863 has an open reading frame (Fig. 7), indicating its potential to encode proteins. These results suggest that hsa_circ_0082002 and hsa_circ_0003863 might contribute to the occurrence and progression of PTC through interacting with miRNAs and RNA binding proteins. Of course, deeper investigations designed with in vitro and in vivo studies should be performed to further understand their functional mechanism in PTC. Hsa_circ_0082002 is 1, 214 nt in length, deriving from the proto-oncogene MET. Recently, it has been identified as an oncogenic circRNA in renal cell carcinoma (RCC) and hepatocellular carcinoma (HCC). In RCC, hsa_circ_0082002 was found to promote tumor proliferation by enhancing the mRNA decay of CDKN2 and competitively absorbing miR-1197 to upregulate SMAD3 mRNA[24]. In HCC, hsa_circ_0082002 drives an epithelial-to-mesenchymal transition and improves the immunosuppressive tumor microenvironment via the Snail/DPP4/CXCL10 axis[25]. Besides, hsa_circ_0082002 was reported to be involved in the angiogenesis of neovascular diseases[26], tamoxifen resistance of breast cancer cells[27], and tumor cell growth of non-small cell lung cancer[28] and colorectal cancer[29]. However, regarding its role in PTC, particularly its levels in exosomes, no related publications can be found until now. About hsa_circ_0003863, it is 489 nt in length, and its host gene is RAB27A (a GTPase required for sEV secretion). There are no publications about the function of hsa_circ_0003863 in physiological and pathological conditions. Therefore, it is the first time for us to uncover the dysregulated levels of hsa_circ_0082002 and hsa_circ_0003863 in serum exosomes of PTC and their diagnostic potential as tumor markers for PTC. Larger patient cohorts with more comprehensive follow-up data should be collected to further verify the diagnostic sensitivity and specificity of exosomal hsa_circ_0082002 and hsa_circ_0003863 before their clinical usage as tumor markers for PTC. In the present study, only the sera from patients with benign thyroid tumors or PTC at initial diagnosis were obtained, therefore it is impossible to examine the level changes of exosomal hsa_circ_0082002, and hsa_circ_0003863 before and after surgery, before and after systemic therapy to study their kinetics. These problems should be figured out in a future study to further determine the diagnostic and prognostic value of exosomal hsa_circ_0082002, and hsa_circ_0003863. Besides, more studies should be carried out to explore the role and the underlying functional mechanisms of hsa_circ_0082002 and hsa_circ_0003863 in PTC.
Collectively, we identified two PTC-related circRNAs (hsa_circ_0082002 and hsa_circ_0003863) incorporated in exosomes and uncovered their potential as tumor markers to diagnose PTC, in particular, more aggressive PTC.
Availability of data and material
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- PTC:
-
Papillary thyroid carcinoma
- circRNA:
-
Circular RNA
- ROC:
-
Receiver operator characteristic
- AUC:
-
Area under the ROC curve
- qPCR:
-
Quantitative polymerase chain reaction
- FNAB:
-
Fine needle aspiration biopsy
- PET:
-
Positron emission tomography
- CT:
-
Computed tomography
- miRNA:
-
MicroRNA
- lncRNA:
-
Long non-coding RNA
References
Ramia de Cap M (2021) Multifocal papillary thyroid carcinoma. Am J Clin Pathol 155(6):913. https://doi.org/10.1093/ajcp/aqab005
Feldkamp J, Führer D, Luster M et al (2016) Fine needle aspiration in the investigation of thyroid nodules. Dtsch Arztebl Int 113(20):353–359. https://doi.org/10.3238/arztebl.2016.0353
Liu T, Tilak M, Awad S et al (2022) A literature review of factors associated with pain from fine needle aspiration biopsy of thyroid nodules. Endocr Pract 28(6):628–636. https://doi.org/10.1016/j.eprac.2022.03.007
Torréns JI, Burch HB (2001) Serum thyroglobulin measurement. Utility in clinical practice. Endocrinol Metab Clin North Am 30 (2):429–467. doi:https://doi.org/10.1016/s0889-8529(05)70194-8
Algeciras-Schimnich A (2018) Thyroglobulin measurement in the management of patients with differentiated thyroid cancer. Crit Rev Clin Lab Sci 55(3):205–218. https://doi.org/10.1080/10408363.2018.1450830
Cho JS, Kim HK (2022) Thyroglobulin Levels as a Predictor of Papillary Cancer Recurrence After Thyroid Lobectomy. Anticancer Res 42(11):5619–5627. https://doi.org/10.21873/anticanres.16070
Kalluri R, LeBleu VS (2020) The biology, function, and biomedical applications of exosomes. Science. https://doi.org/10.1126/science.aau6977
Mashouri L, Yousefi H, Aref AR et al (2019) Exosomes: composition, biogenesis, and mechanisms in cancer metastasis and drug resistance. Mol Cancer 18(1):75. https://doi.org/10.1186/s12943-019-0991-5
Dai J, Su Y, Zhong S et al (2020) Exosomes: key players in cancer and potential therapeutic strategy. Signal Transduct Target Ther 5(1):145. https://doi.org/10.1038/s41392-020-00261-0
Wang Y, Liu J, Ma J et al (2019) Exosomal circRNAs: biogenesis, effect and application in human diseases. Mol Cancer 18(1):116. https://doi.org/10.1186/s12943-019-1041-z
Chen L, Wang C, Sun H et al (2021) The bioinformatics toolbox for circRNA discovery and analysis. Brief Bioinform 22(2):1706–1728. https://doi.org/10.1093/bib/bbaa001
Kristensen LS, Andersen MS, Stagsted LVW et al (2019) The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet 20(11):675–691. https://doi.org/10.1038/s41576-019-0158-7
Chen Y, Wang J, Wang C et al (2022) Deep learning models for disease-associated circRNA prediction: a review. Brief Bioinform. https://doi.org/10.1093/bib/bbac364
Chen L, Shan G (2021) CircRNA in cancer: fundamental mechanism and clinical potential. Cancer Lett 505:49–57. https://doi.org/10.1016/j.canlet.2021.02.004
Panda AC (2018) Circular RNAs Act as miRNA Sponges. Adv Exp Med Biol 1087:67–79. https://doi.org/10.1007/978-981-13-1426-1_6
Huang A, Zheng H, Wu Z et al (2020) Circular RNA-protein interactions: functions, mechanisms, and identification. Theranostics 10(8):3503–3517. https://doi.org/10.7150/thno.42174
Zhu G, Chang X, Kang Y, Zhao X, Tang X, Ma C, Fu S (2021) CircRNA: A novel potential strategy to treat thyroid cancer (Review). Int J Mol Med. https://doi.org/10.3892/ijmm.2021.5034
Chen Y, Ma X, Lou C et al (2022) PLA2G10 incorporated in exosomes could be diagnostic and prognostic biomarker for non-small cell lung cancer. Clin Chim Acta 530:55–65. https://doi.org/10.1016/j.cca.2022.02.016
Chen Y, Lou C, Ma X et al (2022) Serum exosomal hsa_circ_0069313 has a potential to diagnose more aggressive non-small cell lung cancer. Clin Biochem 102:56–64. https://doi.org/10.1016/j.clinbiochem.2022.01.005
Shen X, Yang Y, Chen Y et al (2022) Evaluation of EpCAM-specific exosomal lncRNAs as potential diagnostic biomarkers for lung cancer using droplet digital PCR. J Mol Med (Berl) 100(1):87–100. https://doi.org/10.1007/s00109-021-02145-4
Jiang B, Zhang J, Sun X et al (2022) Circulating exosomal hsa_circRNA_0039480 is highly expressed in gestational diabetes mellitus and may be served as a biomarker for early diagnosis of GDM. J Transl Med 20(1):5. https://doi.org/10.1186/s12967-021-03195-5
Zhi F, Ding Y, Wang R et al (2021) Exosomal hsa_circ_0006859 is a potential biomarker for postmenopausal osteoporosis and enhances adipogenic versus osteogenic differentiation in human bone marrow mesenchymal stem cells by sponging miR-431-5p. Stem Cell Res Ther 12(1):157. https://doi.org/10.1186/s13287-021-02214-y
Yu D, Li Y, Wang M et al (2022) Exosomes as a new frontier of cancer liquid biopsy. Mol Cancer 21(1):56. https://doi.org/10.1186/s12943-022-01509-9
Yang L, Chen Y, Liu N et al (2022) CircMET promotes tumor proliferation by enhancing CDKN2A mRNA decay and upregulating SMAD3. Mol Cancer 21(1):23. https://doi.org/10.1186/s12943-022-01497-w
Huang XY, Zhang PF, Wei CY et al (2020) Circular RNA circMET drives immunosuppression and anti-PD1 therapy resistance in hepatocellular carcinoma via the miR-30-5p/snail/DPP4 axis. Mol Cancer 19(1):92. https://doi.org/10.1186/s12943-020-01213-6
Yao MD, Jiang Q, Ma Y et al (2022) Targeting circular RNA-MET for anti-angiogenesis treatment via inhibiting endothelial tip cell specialization. Mol Ther 30(3):1252–1264. https://doi.org/10.1016/j.ymthe.2022.01.012
Liu J, Dai Z, Li M et al (2022) Circular RNA circMET contributes to tamoxifen resistance of breast cancer cells by targeting miR-204/AHR signaling. Biochem Biophys Res Commun 627:200–206. https://doi.org/10.1016/j.bbrc.2022.07.097
Pei X, Chen SW, Long X et al (2020) circMET promotes NSCLC cell proliferation, metastasis, and immune evasion by regulating the miR-145-5p/CXCL3 axis. Aging (Albany NY) 12(13):13038–13058. https://doi.org/10.18632/aging.103392
Liu Y, Chen L, Liu T et al (2022) Genome-wide circular RNA (circRNA) and mRNA profiling identify a circMET-miR-410-3p regulatory motif for cell growth in colorectal cancer. Genomics 114(1):351–360. https://doi.org/10.1016/j.ygeno.2021.11.038
Acknowledgements
We thank Prof. Junming Guo for providing us with the qPCR instrument.
Funding
This study is supported by the Natural Science Foundation of Ningbo (2023J322, 2023J077, and 2021J313); the Research Foundation of Ningbo No.2 Hospital (2022HMK40); the Young Technical Backbone Talent of Health in Ningbo; the Zhu Xiushan Talent Award Fund of Ningbo NO.2 Hospital (2021HMYQ06); the Project of NINGBO Leading Medical & Health Discipline (2022-F18).
Author information
Authors and Affiliations
Contributions
Study design and supervision: LD, XW, XM. Development of methodology: LD, WH, HJ. Acquisition of data (acquired and managed patients): LD, YW, QL. Analysis and interpretation of data (e.g., statistical analysis, biostatistics): LD. Writing, review, and/or revision of the manuscript: LD, XW, XM.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interest.
Ethics approval and consent to participate
This study was performed following the Declaration of Helsinki. Written informed consent was obtained from all patients, and the study was approved by the ethics committee (the Clinical Research Ethics Committee of Ningbo No.2 Hospital) with the number of. YJ-NBEY-KY-2021-181-01.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
About this article
Cite this article
Dai, L., Hu, W., Jiang, H. et al. The diagnostic potential of two exosome-derived circRNAs for papillary thyroid cancer. Int J Clin Oncol 28, 1461–1474 (2023). https://doi.org/10.1007/s10147-023-02400-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10147-023-02400-3