Abstract
Catheter-related bloodstream infection (CRBSI) is one of the most serious complications in hospitalised patients, leading to increased hospitalisation, intensive care admissions, extensive antibiotic treatment and mortality. A greater understanding of these bacterial infections is needed to improve the prevention and the management of CRBSIs. We describe here the systematic culture-independent evaluation of intravascular catheter (IVC) bacteriology. Twelve IVCs (6 central venous catheters and 6 arterial catheters) were collected from 6 patients. By using traditional culture methods, 3 patients were diagnosed with catheter colonisation including 1 patient who also had CRBSI, and 3 had no colonisation. From a total of 839,539 high-quality sequence reads from high-throughput sequencing, 8 microbial phyla and 76 diverse microbial genera were detected. All IVCs examined in this study were colonised with complex microbial communities including “non-colonised IVCs,” as defined using traditional culture methods. Two main community types were observed: Enterobacteriaceae spp., dominant in patients without colonisation or CRBSI; and Staphylococcus spp., dominant in patients with colonisation and CRBSI. More diverse pathogens and a higher microbial diversity were present in patients with IVC colonisation and CRBSI. Community composition did not appear to be affected by patients’ antibiotic treatment or IVC type. Characterisation of these communities is the first step in elucidating roles of these pathogens in disease progression, and to ultimately facilitate the improved prevention, refined diagnosis and management of CRBSI.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Intravascular catheters (IVCs), including central venous catheters (CVCs) and arterial catheters (ACs), are the most frequently used invasive medical devices in hospitals. However, catheter-related bloodstream infection (CRBSI) is one of the most serious nosocomial infections [1, 2]. It is associated with high morbidity, mortality and treatment cost [3]. IVC placement facilitates the access of potential pathogens to the bloodstream, and also provides a ready surface to which they can attach and form biofilms [4, 5].
Many microbial species have been identified from IVC biofilms using traditional culture methods [6]. However, it has been reported that only a small fraction of attendant microorganisms could be detected [7, 8]. Therefore, the identities of microorganisms that participate in catheter biofilm formation and CRBSI may not be fully determined. Next-generation sequencing techniques are widely recognised for their time- and cost-effective descriptions of microbial populations in a high-throughput manner [9]. Importantly, these approaches can also detect and identify microbes without having to culture the microorganisms. In this study, we present the characterisation of biofilms on IVCs recovered from patients with no IVC colonisation, with colonisation and with CRBSI using next-generation sequencing and traditional culture methods.
Materials and methods
The study was carried out in the Intensive Care Unit of the Royal Brisbane and Women’s Hospital (RBWH), Queensland, Australia. Six adult patients with concurrently sited ACs and CVCs were recruited to this study. Ethical approval for the study was granted by the hospital Human Ethics Committee (HREC 2008/024). Written informed consent was provided by participants or their representatives.
The CVCs were ARROWgard Blue® (chlorhexidine acetate and silver sulfadiazine coated; Arrow Int, Reading, PA, USA), and the ACs were of the Vygon Leader Cath brand (Ecouen, France). There was no imposed limitation on dwell time, and the resiting of catheters always occurred at a new body site. Guide-wire exchange was not performed. Dressings and administration sets were maintained by ICU nurses using unit protocols in accordance with guidelines [6].
The IVC samples were taken by qualified registered nurses who were experienced in the preparation of specimens for culture. The distal 2–3 cm of the tip was cut using sterile scissors and deposited in a sterile container. Twelve IVC tips were handled under aseptic conditions and immediately transported to the laboratory for examination, where they were cultured using the roll-plate culture method [10]. Diagnosis of CRBSI was made using conventional methods including a positive IVC tip culture and peripheral venous blood culture with the same microorganism and no other obvious source of BSI [6]. Microorganisms were isolated and identified according to the standard hospital protocol.
High-throughput sequencing
Following processing for culture, catheter tips were suspended in 200 μl of lysis buffer, which contained 20 mg/ml lysozyme, 20 mM Tris-HCl (pH 8.0), 2 mM EDTA, 1.2 % Triton, and Proteinase K at 37 °C overnight. Bacterial genomic DNA was extracted from all IVCs using the QIAamp DNA mini kit (Qiagen, Chadstone, VIC, Australia). For each catheter, a control (unused) catheter was taken from the original packaging and rolled back and forth on blood agar plates, with microbial genomic DNA extracted as above. Targeted genes were amplified from purified genomic DNA using the primers F (5’ AAACTYAAAKGAATTGRCGG 3’) and R (5’ ACGGGCGGTGWGTRC3’), which would cover variable regions (V6- V8) of the gene encoding 16S rRNA (rrs gene). For each genomic DNA sample, three replicate polymerase chain reactions (PCRs) were performed. The PCR products were purified with the Qiaquick PCR Purification kit (Qiagen, Australia). Barcoded library construction and paired-end sequencing were performed using the Illumina MiSeq platform, supported and operated by the Australian Centre for Ecogenomics.
Bioinformatics analysis
The raw 16S rRNA sequence data were trimmed from 3’-prime end until their average quality score exceeded 25, and reads containing ambiguous characters were additionally trimmed at the occurrence of the first “N”. Subsequently, low-quality reads were identified and excluded using criteria adapted from Huse et al. [11].
Following these initial sequence quality control assessments, the remaining sequences were analysed using QIIME, and reads grouped into operational taxonomic units (OTUs) using a threshold of 97 % sequence identified. Sequences were chimera checked using ChimeraSlayer. A representative sequence from each OTU was then subjected to BLAST alignment with Greengenes database, as implemented in QIIME, and the OTU table generated through QIIME includes the frequency with which each OTU was observed in each sample. These OTU tables can be represented at phylum, class, order, family and genus/species levels of assignment, and heat maps at the genus level of assignment were constructed in R, using the g-plots package (R package version 2.6.0).
Alpha (within sample) diversity metrics were computed using QIIME. The Shannon diversity and richness values from these analyses were compared using paired t test, as were the differences in individual and relative OTU abundance values. Beta (between-sample) diversity metrics, including hierarchical clustering and weighted and unweighted Unifrac distances, were also computed using QIIME and the FastUnifrac metric. The Jaccard distance was used for principal coordinate analysis, and further statistical comparisons were performed using Calypso software [12]. The two-tailed t test was used to evaluate the difference between variances. A p value of less than 0.05 was considered significant.
Results
Sample characteristics
The 6 recruited patients had a mean age of 47.2 years, and 66.7 % (4/6) were on systemic antimicrobials at the time of catheter removal. The mean duration of catheter placement was 6 days (4–10 days), and there was no difference in dwell time between ACs and CVCs.
Three of the 6 CVCs and 0 of the 6 ACs were colonised (over 15 colony-forming units) on roll-plate cultures. Isolates were all Staphylococcus spp. One of the 6 patients was diagnosed with CRBSI. Both colonisation and CRBSI occurred in patients already receiving multiple antibiotics.
Microbial genomic DNA was extracted and quantified from all the IVC samples, and all these DNA samples produced amplicon libraries for sequencing. Significantly, no microbial DNA was recovered and amplified from sterile, unused IVCs, which indicates that any “contaminating DNA” arising from the manufacture of the IVCs, or the reagents used, was negligible. No negative control DNA samples were used for sequencing and further study.
Bacterial community profiles of IVCs
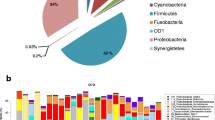
After quality trimming and chimera checking, 839,539 high-quality sequence reads from the 12 IVCs were available (an average of 69,961 reads per sample). The results demonstrated that the microbial communities on IVCs comprised prokaryotes assigned to 7 bacterial and 1 archaeal phyla: Actinobacteria, Bacteroidetes, Chloroflexi, Cyanobacteria, Euryarchaeota, Firmicutes, Proteobacteria and Thermi. However, two of these phyla (Firmicutes and Proteobacteria) represent nearly 90 % of the community’s composition. The OTUs were further assigned to 76 known genera, including Staphylococcus (12.8 % of 16S rRNA reads), Streptococcus (9.7 %), and Enterobacteriaceae (15.9 %).
The microbial communities present on CVCs and ACs were not different from each other in terms of their alpha diversity metrics (Shannon–Weaver, p = 0.28; richness p = 0.72; evenness p = 0.28; and Simpson, p = 0.30). Principal coordinate analyses also suggested that microbial communities present on these catheters could not be differentiated on the basis of catheter type (Fig 1S).
Furthermore, the microbial communities on IVCs from patients treated with antibiotics (4 out of 6) were indistinguishable from those patients who did not receive antibiotics; both in terms of principal coordinate analysis or their alpha diversity metrics (Shannon–Weaver, p = 0.28; richness p = 0.52; evenness p = 0.32; and Simpson, p = 0.35; Fig. 2S).
Catheter colonisation, CRBSI impact on biofilm community composition
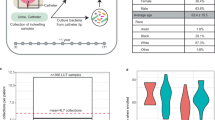
The biofilm communities present upon patients with colonisation and CRBSI were significantly different from those without colonisation and CRBSI, as shown by OTU abundance (Fig. 1a), the principal coordinate analysis (Fig 1b) and alpha diversity metrics (Shannon–Weaver, p = 0.04; richness p = 0.12; evenness p = 0.08; and Simpson, p = 0.03). The microbial communities on IVCs from patients who were diagnosed with IVC colonisation were predominant by Staphylococcus. The abundance of Staphylococcus increased on IVCs from patients with CRBSI, while Enterobacteriaceae dominated in patients without colonisation or CRBSI (Fig. 1c). The microbial communities recovered from patients with colonisation and CRBSI possessed a high abundance of Acinetobacter, Staphylococcus, Stenotrophomonas and Aerococcus, while a greater abundance of Enterobacteriaceae, Pseudomonas and Propionibacterium was found from patients without colonisation or CRBSI (Fig. 1a). Greater microbial diversity and a higher proportion of pathogens were present on patients with colonisation and CRBSI, which indicate that they may trigger the process of clinically significant microbial colonisation and CRBSI.
Comparing microbial communities on intravenous catheters (IVCs) from patients with and without colonisation by a a relative abundance on IVCs of most abundant operational taxonomic units (OTU) across samples; b a significantly different microbial group (p < 0.05, ANOVA). Significant differences between groups (t test) are annotated as *p < 0.05, **p < 0.01. c Principal coordinate analysis of phylogenetic distance among communities according to the weighted Unifrac metric
Discussion
This study provides the first culture-independent examination of microbial biofilms on IVCs from patients with no infection, catheter colonisation and CRBSI (detected by culture). Two main community types were observed: Enterobacteriaceae spp., dominant in patients without colonisation or CRBSI; and Staphylococcus spp., dominant in patients with colonisation and CRBSI. Microbial diversity was lower in patients without colonisation or CRBSIs. In our previous studies, we used 454 pyrosequencing to characterise microbial communities on IVCs and found that Gram-negative bacteria predominated on IVCs when only non-colonised IVC samples were examined from patients without CRBSI [7, 13]. Such results suggest that Gram-negative bacteria might pre-colonise IVCs without causing infection and might be merely “normal” colonisers on IVCs. In this study, complex microbial biofilms were present on patients with CRBSI instead of single bacterial species, as described using culture methods, with Gram-positive Staphylococcus spp. dominant on microbial biofilms. Community composition did not appear to be affected by patients’ antibiotic treatment or IVC type.
Moreover, only CVCs, not ACs, were associated with IVC colonisation or CRBSI using culture methods. Yet, on molecular study, ACs had similar bacterial communities to CVCs. The possible explanations for this might include:
-
1.
DNA-based molecular methods might detect DNA on IVCs from dead bacteria.
-
2.
ACs have only a single infusion fluid, which might reduce bacterial growth (such as lipid-rich propofol or parenteral nutrition administered via CVCs.
-
3.
Traditional culture methods only examine microbial biofilms on external IVC surfaces and leave the bacteria on inner surfaces under evaluation and ACs may have had inner surface colonisation, related to multiple access for blood sampling.
It has been reported that ACs are an under-recognised cause of CRBSI and are associated with infection rates similar to those of short-term CVCs [14]. This study supports the assertion that ACs should be considered an equally likely site of catheter colonisation in critically ill patients as CVCs.
Characterisation of microbial biofilms is the first step in elucidating the roles of these pathogens in disease progression and in ultimately facilitating the prevention, diagnosis and management of CRBSI. Limitations of this study included the small sample size and the lack of IVCs from patients with Gram-negative CRBSI. Future study with larger cohorts of patients with and without CRBSI is required to verify the results.
References
Tomlinson D, Mermel LA, Ethier MC, Matlow A, Gillmeister B, Sung L (2011) Defining bloodstream infections related to central venous catheters in patients with cancer: a systematic review. Clin Infect Dis 53(7):697–710
Mermel LA, Farr BM, Sherertz RJ, Raad II, O’Grady N, Harris JS, Craven DE (2001) Guidelines for the management of intravascular catheter-related infections. Infect Control Hosp Epidemiol 22(4):222–242
Cecinati V, Brescia L, Tagliaferri L, Giordano P, Esposito S (2012) Catheter-related infections in pediatric patients with cancer. Eur J Clin Microbiol Infect Dis 31(11):2869–2877
Zhang L, Gowardman J, Rickard CM (2011) Impact of microbial attachment on intravascular catheter-related infections. Int J Antimicrob Agents 38(1):9–15
Lindsay D, von Holy A (2006) Bacterial biofilms within the clinical setting: what healthcare professionals should know. J Hosp Infect 64(4):313–325
O’Grady NP, Alexander M, Burns LA, Dellinger EP, Garland J, Heard SO, Lipsett PA, Masur H, Mermel LA, Pearson ML et al (2011) Guidelines for the prevention of intravascular catheter-related infections. Clin Infect Dis 52(9):e162–e193
Zhang L, Morrison M, Nimmo GR, Sriprakash KS, Mondot S, Gowardman JR, George N, Marsh N, Rickard CM (2013) Molecular investigation of bacterial communities on the inner and outer surfaces of peripheral venous catheters. Eur J Clin Microbiol Infect Dis 32(8):1083–1090
Perez E, Williams M, Jacob JT, Reyes MD, Chernetsky Tejedor S, Steinberg JP, Rowe L, Ganakammal SR, Changayil S, Weil MR et al (2014) Microbial biofilms on needleless connectors for central venous catheters: comparison of standard and silver-coated devices collected from patients in an acute care hospital. J Clin Microbiol 52(3):823–831
Grice EA, Kong HH, Renaud G, Young AC, Bouffard GG, Blakesley RW, Wolfsberg TG, Turner ML, Segre JA (2008) A diversity profile of the human skin microbiota. Genome Res 18(7):1043–1050
Maki DG, Weise CE, Sarafin HW (1977) A semiquantitative culture method for identifying intravenous catheter-related infections. N Engl J Med 296:1305–1309
Huse SM, Huber JA, Morrison HG, Sogin ML, Welch DM (2007) Accuracy and quality of massively parallel DNA pyrosequencing. Genome Biol 8(7):R143
Cantacessi C, Giacomin P, Croese J, Zakrzewski M, Sotillo J, McCann L, Nolan MJ, Mitreva M, Krause L, Loukas A (2014) Impact of experimental hookworm infection on the human gut microbiota. J Infect Dis 210(9):1431–1434
Zhang L, Gowardman J, Morrison M, Krause L, Playford EG, Rickard CM (2014) Molecular investigation of bacterial communities on intravascular catheters: no longer just Staphylococcus. Eur J Clin Microbiol Infect Dis 33(7):1189–1198
O’Horo JC, Maki DG, Krupp AE, Safdar N (2014) Arterial catheters as a source of bloodstream infection: a systematic review and meta-analysis. Crit Care Med 42(6):1334–1339
Funding
LZ is supported by a Griffith University Bridging Fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflicts of interest.
Ethical approval
Received.
Supplemental material
Below is the link to the electronic supplementary material.
Fig. S1
Comparing microbial communities on ACs and CVCs by a the Shannon index; b principal coordinate analysis (DOCX 84 kb)
Fig. S2
Comparing microbial communities on IVCs from patients with and without antibiotic treatment by a the Shannon index; b principal coordinate analysis (DOCX 82 kb)
Rights and permissions
About this article
Cite this article
Zhang, L., Gowardman, J., Morrison, M. et al. Microbial biofilms associated with intravascular catheter-related bloodstream infections in adult intensive care patients. Eur J Clin Microbiol Infect Dis 35, 201–205 (2016). https://doi.org/10.1007/s10096-015-2530-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-015-2530-7